Abstract
The gram-negative bacterium Pseudomonas aeruginosa is an opportunistic pathogen associated with drug resistance complications and, as such, an important object for drug discovery efforts. One attractive target for development of therapeutics is the ADP-ribosyltransferase Exotoxin-S (ExoS), an early effector of the type III secretion system that is delivered into host cells to affect their transcription pattern and cytoskeletal dynamics. The purpose of this study was to formulate a real-time assay of purified recombinant ExoS activity for high-throughput application. We characterized the turnover kinetics of the fluorescent dinucleotide 1,N6-etheno-NAD+ as co-substrate for ExoS. Further, we found that the toxin relied on any of five tested isoforms of human 14-3-3 to modify vH-Ras and the Rho-family GTPases Rac1, -2, and -3 and RhoC. We then used 14-3-3β–stimulated ExoS modification of vH-Ras to screen a collection of low-molecular-weight compounds selected to target the poly-ADP ribose polymerase family and identified 3-(4-oxo-3,5,6,7-tetrahydro-4H-cyclopenta[4,5]thieno[2,3-d]pyrimidin-2-yl)propanoic acid as an ExoS inhibitor with micromolar potency. Thus, we present an optimized method to screen for inhibitors of ExoS activity that is amenable to high-throughput format and an intermediate affinity inhibitor that can serve both as assay control and as a starting point for further development.
Introduction
Pseudomonas aeruginosa is a gram-negative opportunistic pathogen that is best known as a common cause of hospital infections. Pseudomonas species feature both intrinsic and acquired mechanisms of antibiotic resistance and thereby pose a serious threat to human and livestock health. At an early stage in infection, the pathogen makes contact with a cell surface and delivers a small set of effectors into the cytosol using a type III secretion system. These effectors include four exotoxins: the bimodular ExoS and ExoT, the lipase ExoU, and the adenylate cyclase ExoY.1,2 ExoS and ExoT share 76% amino acid homology and are bifunctional enzymes containing an N-terminal GTPase activating protein (GAP) domain and a C-terminal ADP-ribosyltransferase (ART) domain. 2 Together, their activities elicit effects that contribute to creating a replication competent niche for the bacteria. The GAP activities of the toxins inactivate Rho GTPases Rho, Rac, and Cdc42, thereby modulating the state of actin polymerization in the host cell. 3 The ART activities are directed toward diverse proteins; ExoS ART domain targets include Ras- and Rho-family GTPases4–6 and ezrin/radixin/moesin proteins. 7 Expression of ExoS ADP-ribosylation activity is dependent on the association with a mammalian cofactor, originally termed factor activating exoenzyme-S, 8 and later identified as the zeta/delta isoform of 14-3-3 (YWHAZ). 9 The interaction with 14-3-3 proteins is mediated by a C-terminal segment of ExoS.10–12
The prominent early effects of ExoS ART activity, combined with the fact that this activity is required for inhibition of phagocytosis in pneumonia, 13 make the ExoS ART domain a promising target for intervention strategies. A chemical probe directed at this domain would be useful both for studying cellular pathology and for development of a therapeutic entity to overcome antibiotic resistance.
Methods to detect toxin-mediated ADP-ribosylation include the use of either radioisotope-labeled or biotinylated NAD+ for detection of ADP-ribosyl groups on target proteins and the use of fluorescent NAD analog, 1,N6-etheno-NAD+ (εNAD+) for detection of εNAD+ hydrolysis. The latter has been used to study a variety of enzymes with NAD-glycohydrolase activity, including phosphodiesterase, 14 Diphtheria toxin, 15 Pertussis toxin, 16 Vibrio splendidus Vis toxin, 17 and ecto-ARTs. 18 A real-time assay using εNAD+ has been developed for mono-ARTs 19 and for the Diphtheria-class Pseudomonas toxin ExoA. 20 Here, our aim was to formulate an assay of ExoS activity that was amenable to compound screening in a high-throughput format. We verified functional cleavage of εNAD+ by recombinant ExoS ART domain and quantified the stimulating effects of several 14-3-3 isoforms on the modification of several small soluble GTPases. Then, we performed a pilot screen and identified a commercially available compound that mimics nicotinamide and that inhibits ExoS activity with micromolar potency in vitro.
Materials and Methods
Materials
Fine chemicals and media were purchased from SigmaAldrich. 1,N6-etheno-NAD+ was obtained from SigmaAldrich and 1,N6-etheno-AMP from Jena Bioscience. Commercial sources of the poly-ADP ribose polymerase (PARP) inhibitor-like compounds are given in
Molecular Cloning
The cDNA fragment encoding ExoS residues 233–453, subcloned in the NdeI and EcoRI sites of pET28a, was contributed by Yngve Östberg (Umeå University). The cDNA fragments encoding ExoS residues 199–453, 215–453, 233–416, 242–416, 242–435, 242–453, 266–416, 266–435, and 266–453 were PCR-amplified from extracts of P. aeruginosa PAK cells and inserted into pNIC28-Bsa4 by ligase independent cloning. An expression plasmid (pGEX-2TR) for human vH-Ras was provided by Anne Ridley (King’s College, London). The cDNA fragments encoding all other small GTPases were contributed by Pontus Aspenström (Karolinska Institutet, Stockholm), and these were subcloned into pNIC28-Bsa4. Expression plasmids containing the 14-3-3 encoding genes YWHAB (pTvHR21-SGC), YWHAE (pLIC-SGC1), YWHAG (pTvHR21-SGC), YWHAH (pNIC28-Bsa4), and YWHAQ (pNIC28-Bsa4) were contributed by the Structural Genomics Consortium (Oxford University) and obtained from Addgene.
Protein Expression and Purification
All proteins except vH-Ras were expressed as hexahistidine fusion proteins in Escherichia coli strains BL21(DE3) or C41 in rich medium, by induction during logarithmic growth phase using 0.5 mM isopropyl β-D-1-thiogalactopyranoside and overnight bacterial growth at 18 °C. Purification of ExoS constructs, 14-3-3 isoforms, and GTPases was done using immobilized metal affinity chromatography followed by size exclusion chromatography (SEC). vH-Ras was expressed as a glutathione S-transferase fusion protein in E. coli strain BL21(DE3) and purified using glutathione sepharose 4B (GE Healthcare) followed by SEC. Protein containing fractions was pooled after analysis by sodium dodecyl sulfate polyacrylamide gel electrophoresis, and target protein integrity was verified by time-of-flight mass spectrometry.
Enzymatic Assay
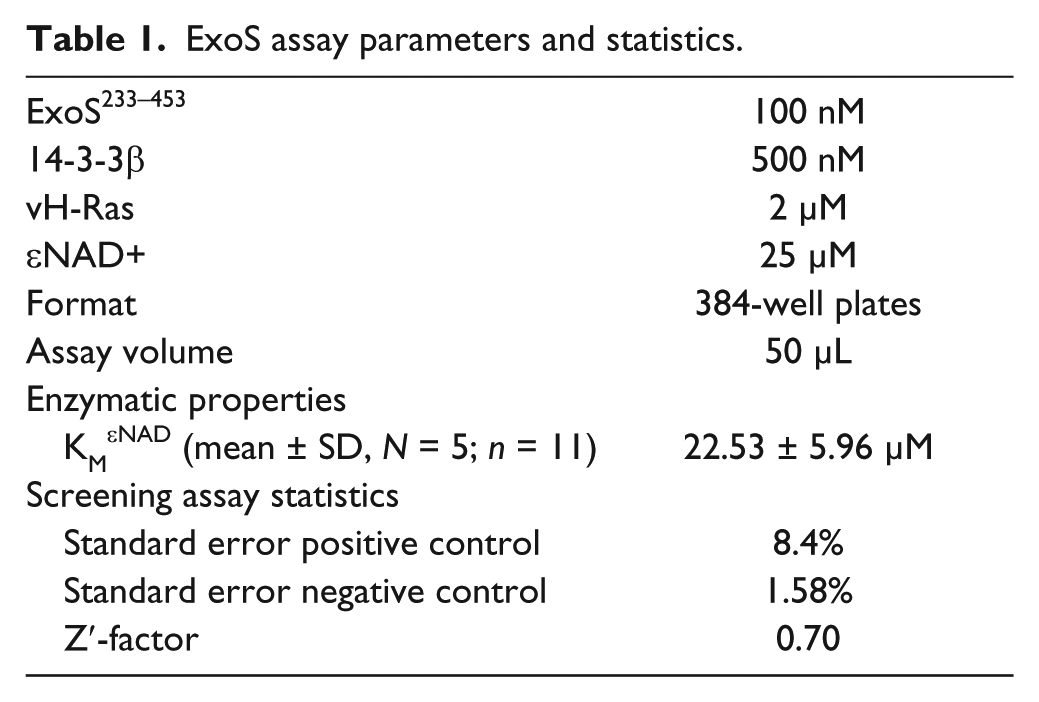
Assays were carried out in black flat-bottom 384-well plates (Corning 3573 or Greiner 781900) with a final assay volume of 50 µL. The optimized conditions for compound screening assays contained 100 nM ExoS, 500 nM 14-3-3β, 2 µM vH-Ras, and 25 µM εNAD+ in 20 mM HEPES pH 7.5, 50 mM NaCl, 4 mM MgCl2, and 0.5 mM TCEP. For positive inhibition assay controls, ExoS233-453 was substituted for ExoS242-416 or 14-3-3 protein was omitted. The DMSO concentration was kept at 1% when testing compounds. Enzymatic reactions, at ambient temperature (22 °C), were started by addition of εNAD+, and fluorescence was followed over time in a CLARIOstar multiplate reader (BMG Labtech) using λex = 302/10 nm (filter) and λem = 410/10 nm (monochromator). Fluorescence increase was linear over 30 min. Where applicable, fluorescence was related to concentrations of fluorescent product using dilution series of 1,N6-etheno-AMP. Fluorescence data were analyzed and kinetic parameters calculated using Graph Pad (Prism Software). KM values for ExoS determined based on either fluorescence increase in the linear time range or using endpoint assays at 30 min were identical within experimental error.
Screening Library
A screening set of compounds with molecular weights up to 300 Da was chosen from a previously assembled library of PARP inhibitors and chemically related compounds
21
(see
Results and Discussion
To formulate a real-time fluorescence-based assay for ExoS activity, we used the most widely studied toxin fragment (ExoS ART domain residues A233–A453) and its ability to ADP-ribosylate human vH-Ras protein using εNAD+ as co-substrate. First, we assessed the ability of five of the seven human 14-3-3 isoforms to activate the ExoS ART domain. The beta, gamma, epsilon, eta, and theta isoforms of human 14-3-3 displayed similar efficiency in activation of ExoS activity, when present in five-molar excess (
We went on to evaluate whether the N6-etheno group on the adenine moiety of NAD affected the kinetics of the ExoS233–453-catalyzed ADP-ribosyltransferase reaction. First, we determined the catalytic properties of 14-3-3β–stimulated modification of vH-Ras with εNAD+ as co-substrate. Based on five independent experiments, we determined an apparent KM of 22.5 ± 6 µM ( Fig. 1A ; Table 1 ). This is higher than a previous determination using the same method and substrate (9 µM) 23 and lower than the value determined by the use of radiolabeled NAD+ and soy bean trypsin inhibitor as target (185.3 ± 32.5 µM). 24 Together, these data suggested that the etheno ring in εNAD+ might affect the rates of ADP-ribosyl transfer. Therefore, we tested the dependence of the apparent KM on the percentage of εNAD+ in the reaction mixture. As shown in Figure 1B , apparent KM values increased slightly with increasing [εNAD+]/[NAD+]. Thus, our data indicate that εNAD+ is turned over somewhat less efficiently that NAD+. We conclude that differences between kinetic parameters measured here and published previously likely are attributable to assay variations. The simplicity and robustness of the assay warranted consideration of the method for high-throughput screening assays to discover novel inhibitors of ExoS ADP-ribosyltransferase activity.
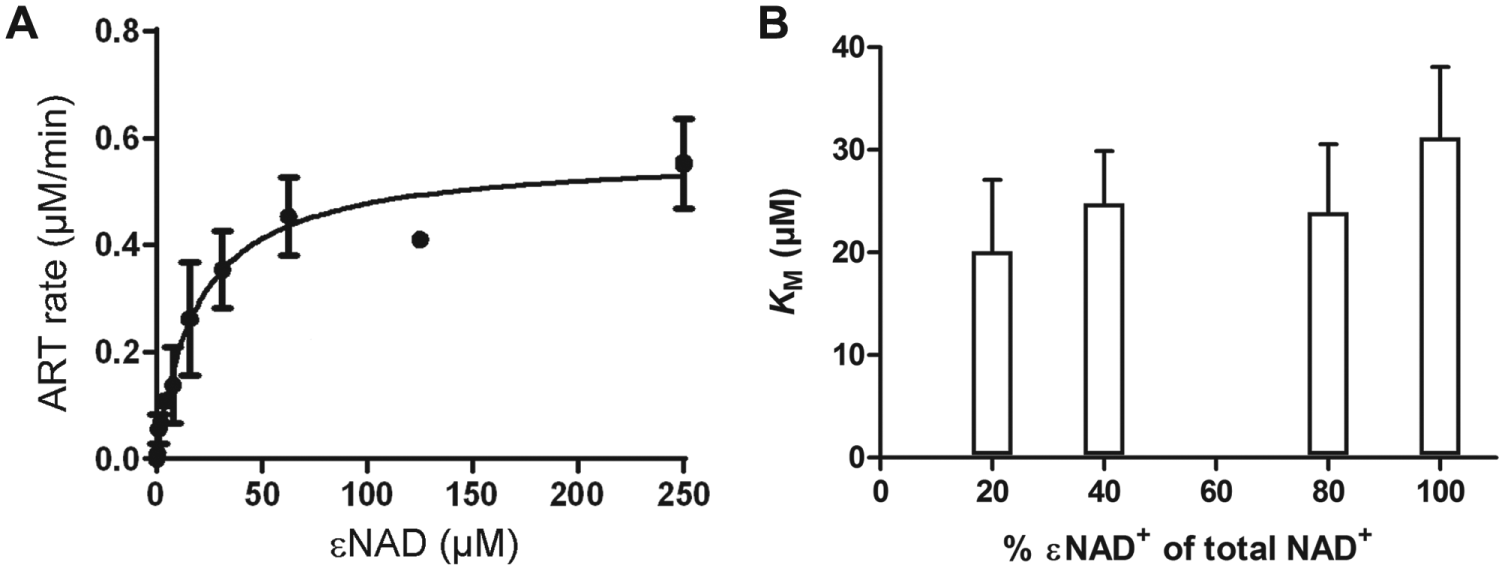
Enzymatic properties of ExoS233–453. (
ExoS assay parameters and statistics.
To test whether this assay of ExoS activity was indeed useful for the identification of ExoS inhibitors, we screened part of a previously described library of PARP inhibitors and chemically related compounds.
21
A screening set of compounds with molecular weights up to 300 Da was chosen from this library (
First, we tested the DMSO tolerance of the assay. Enzymatic activity was unaffected, within experimental error, by DMSO in the range of 0% to 2% (
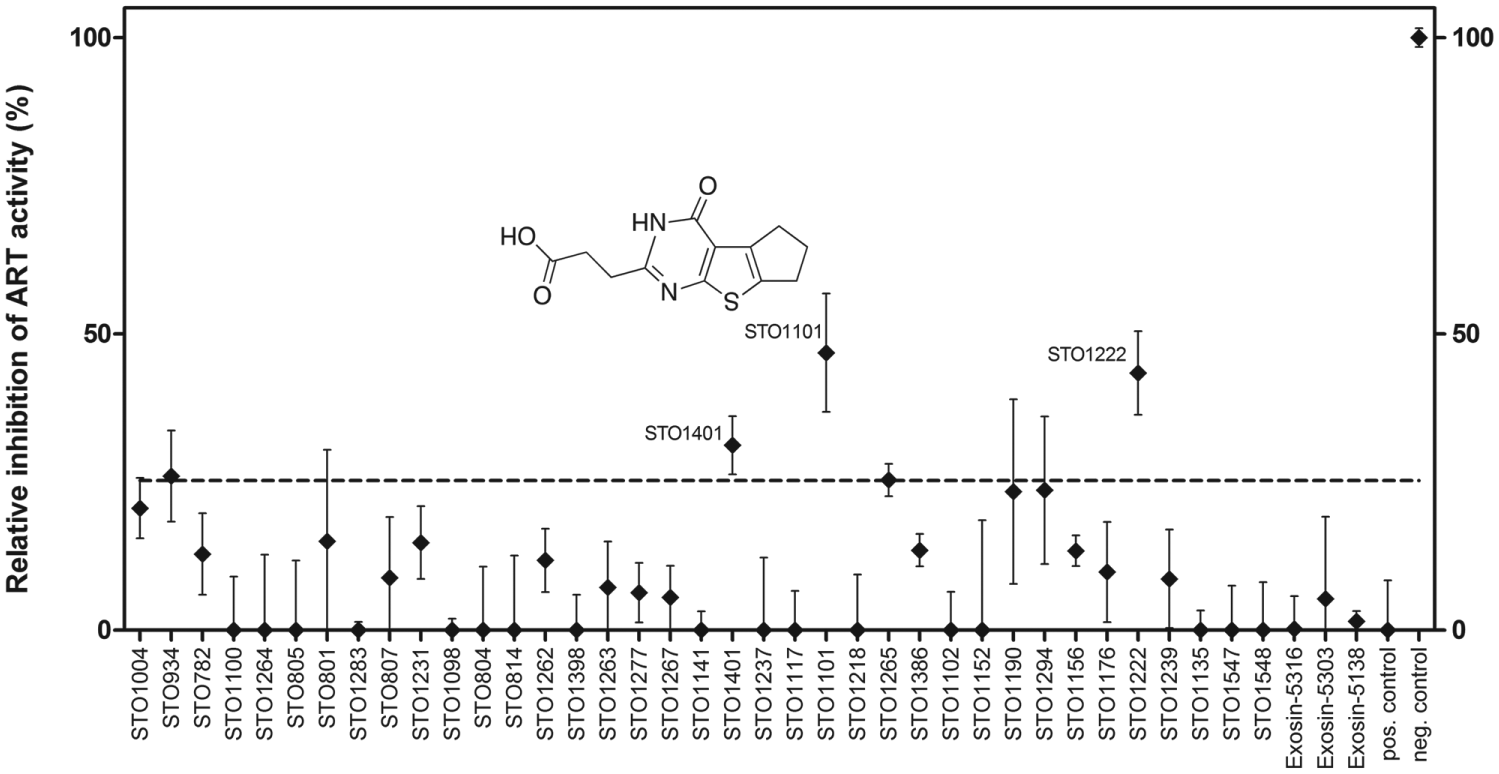
A screen for enzymatic inhibitors of ExoS233–453 using a small collection of PARP inhibitors and chemically related compounds. Correlation plot of data from three independent screens, carried out with independent protein batches. Data are shown as mean percentages ± SD (N = 2, n = 4) of control reactions (containing 1% DMSO), corrected for background signal in the absence of 14-3-3β. The chemical structure of STO1101, the most active compound, is shown. Full data along with compound information are given in
The most potent primary screening hit, compound STO1101, was further characterized in two different inhibition experiments ( Fig. 3 ): (1) KM experiments were conducted in the presence of different concentrations of STO1101. We calculated a Ki of 8.6 ± 3 µM (n = 4) based on the results ( Fig. 3A ). Lineweaver-Burk plots of these data also established STO1101 as a competitive inhibitor of ExoS ( Fig. 3B ). (2) Concentration-response experiments were carried out to determine an IC50 value of 19 µM ( Fig. 3C ). Considering our assay conditions and KM = 22.5 µM ( Table 1 ; Fig. 1 ) and using the Cheng-Prusoff equation, 26 this translates into a Ki value of 9.0 µM, in good agreement with the previous value. For the two less potent inhibitors, we determined IC50 values of 148 µM (STO1222) and 437 µM (STO1401; data not shown).
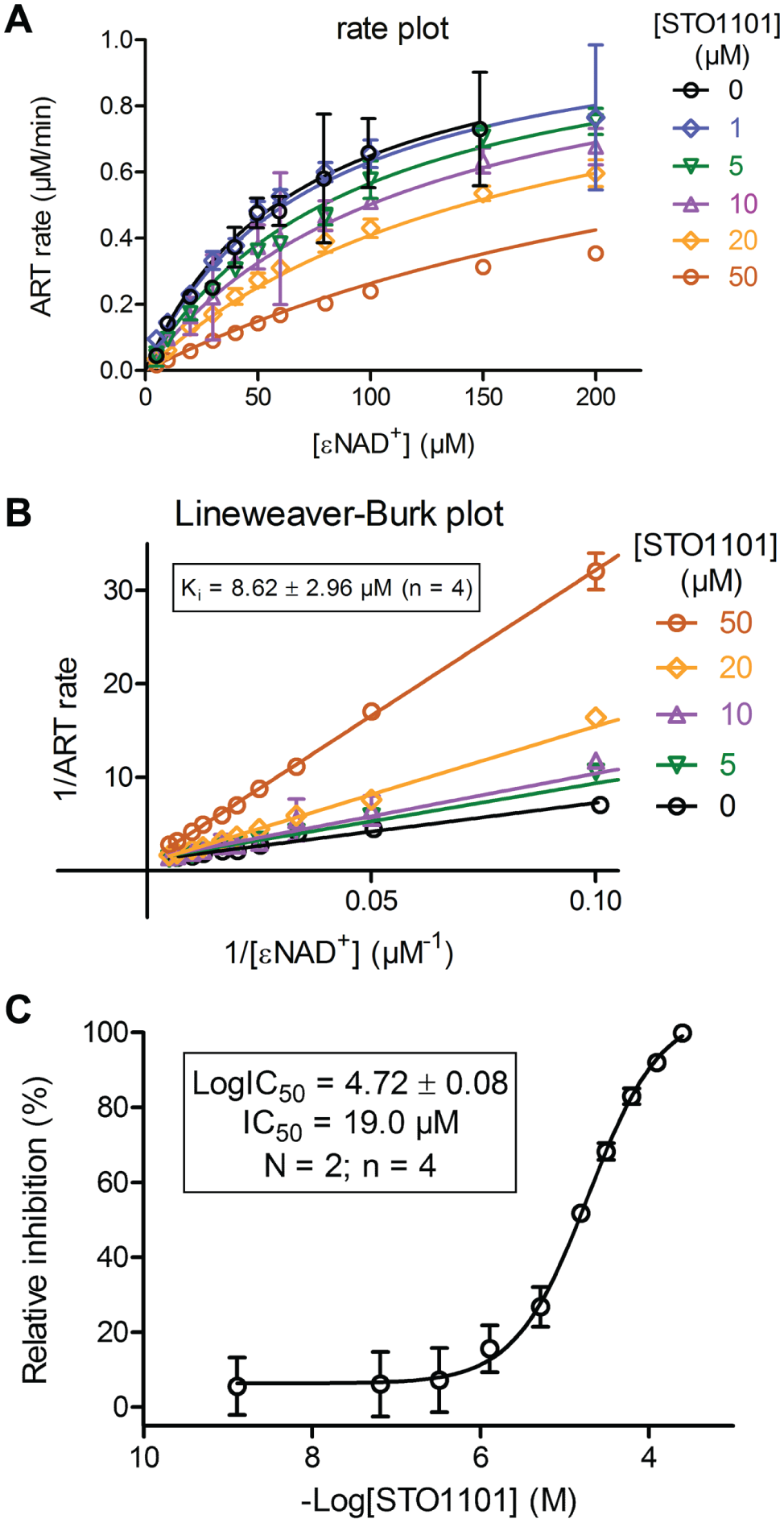
Validation of the primary screening hit. (
Following the finding that STO1101 inhibited ExoS ADP-ribosylation activity with reasonable potency, we resynthesized the compound and confirmed its identity and potency. Furthermore, to establish primary structure-activity relationships (SARs), we synthesized a small set of analogues to investigate the importance of the main functional groups of STO1101 (
Table 2
;
Structure-activity relationship study of the STO1101 scaffold.
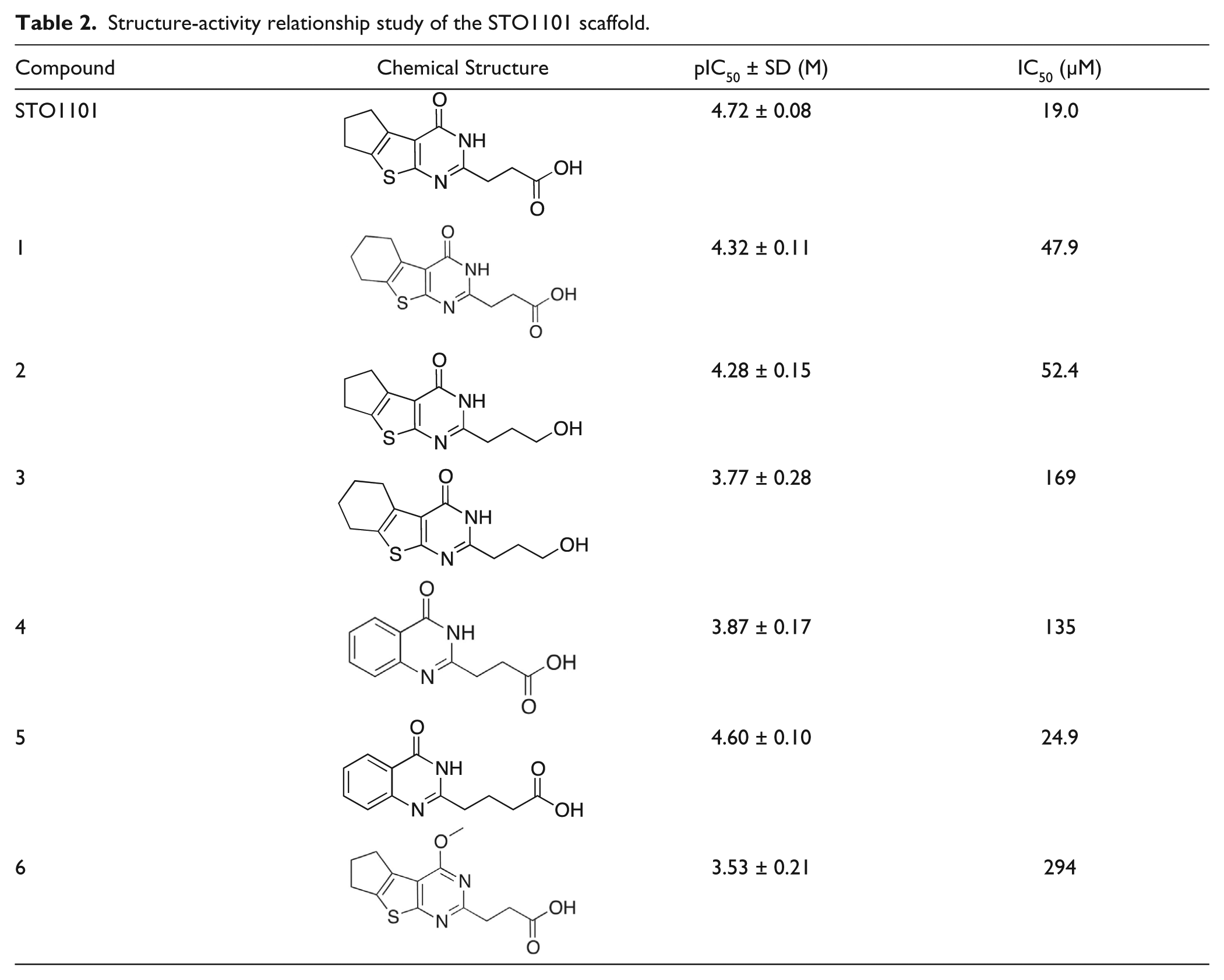
In summary, a real-time fluorescence assay of P. aeruginosa ExoS, using εNAD+ as co-substrate, was optimized for high-throughput screening purposes and used to identify and validate STO1101 as a novel ExoS inhibitor with an affinity (Ki) of roughly 9 µM.
Footnotes
Acknowledgements
We thank the Protein Science Facility at Karolinska Institutet and SciLifeLab for molecular cloning.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Swedish Foundation for Strategic Research (grant SB12-0022; Å.F., M.E., and H.S.), the IngaBritt and Arne Lundbergs Research Foundation and Karolinska Institutet (H.S.), and the Swedish Research Council (grant 521-2012-2802; M.E.).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
