Abstract
CD44, especially the isoforms with variable exons (CD44v), is a promising biomarker for the detection of cancer. To develop a CD44v-specific probe, we screened a 7-mer phage peptide library against the CD44v3-v10 protein using an improved subtractive method. The consensus sequences with the highest frequency (designated CV-1) emerged after four rounds of panning. The binding affinity and specificity of the CV-1 phage and the synthesized peptide for the region of CD44 encoded by the variable exons were confirmed using enzyme-linked immunosorbent assay and competitive inhibition assays. Furthermore, the binding of the CV-1 probe to gastric cancer cells and tissues was validated using immunofluorescence and immunohistochemistry assays. CV-1 sensitively and specifically bound to CD44v on cancer cells and tissues. Thus, CV-1 has the potential to serve as a promising probe for cancer molecular imaging and target therapy.
Introduction
CD44 is an integral membrane glycoprotein and is the principal receptor of hyaluronic acid, which is an important signaling molecule in the extracellular matrix. 1 CD44 also interacts with matrix metalloproteinases, osteopontin, and collagens. Through these diverse interactions, CD44 plays roles in T cell activation, lymphocyte homing, both cell-cell and cell-matrix interactions.1–3 Aberrant CD44 expression is associated with epithelial-mesenchymal transition, structural and functional disorganization; it plays roles in tumor invasion, metastasis, proliferation, and angiogenesis.4,5
The CD44 molecule is composed of two groups of exons. Exons 1 to 5 and 16 to 20 form the standard CD44 isoform (CD44s). Exons 6 to 15 (termed variable exons v1–v10) can be alternatively spliced and inserted between exons 5 and 16. CD44v represents the CD44 proteins that contain variable exons. 3 Both forms of CD44 protein are expressed at high levels in gastric cancer tissues and take part in tumorigenesis and malignancy.6,7 Because of its focal expression in precancerous lesions, CD44s is considered to be a highly sensitive biomarker. 6 However, CD44v is expressed in gastric cancer with higher specificity compared with CD44s, because these molecules are absent in most nontumor tissues. 8 Many types of CD44v can be detected in gastric cancer including CD44v3-v5, CD44v5, CD44v6, CD44v7, CD44v8-10, and CD44v9; some of these types correlate with the pathological type, metastasis, and clinical outcome of stomach carcinoma.9–13 Thus, CD44v can serve as a biomarker for the detection and prognosis of gastric cancer.
The development of specific ligands is a critical goal of CD44-targeted research. Previous studies showed that hyaluronan-based bioconjugates, anti-CD44 antibodies, and anti-CD44v antibodies linked to radioactive isotopes or chemotherapeutic agents displayed antitumor characteristics.14–16 Park et al. 17 developed a peptide with the potential to target breast cancer via binding to CD44; however, to the best of our knowledge, peptide libraries have not been screened for a CD44v-specific probe.
Probes that can target given molecules or cells can be antibodies, antibody fragments, peptides, or other small molecules.18,19 Among these, peptides have several advantages. First, peptides can penetrate and accumulate within the lesion with rapid binding kinetics. 20 Second, peptides exhibit minimal immunogenicity and rapid clearance, which are characteristics that are associated with low toxicity. 21 Finally, because of their simple structure, peptides are easily synthesized and conjugated with drugs or contrast agents. 21
While some naturally occurring peptides have been used for molecular-targeted diagnosis and therapy, 22 phage display offers the advantage of sensitive and specific identification of targets within a library containing a huge number of peptides. 23 Phage display is a high-throughput method used to identify specific ligands for given targets using recombinant DNA technology. 19 With a complexity of 107 to 109 independent clones, a peptide library can be screened against various targets, including proteins, cells, and organs.24–26 Among these, screening against purified proteins can maximize the specificity of the peptide probes.
An improved subtractive phage display method was used to identify a CD44v-specific peptide. The binding affinity of the peptide for cancer cells and tissues was confirmed. This peptide probe may serve as a potential probe for cancer targeting.
Materials and Methods
Cell Lines
The gastric cancer cell line SGC-7901 was purchased from the Cell Bank of Type Culture Collection of the Chinese Academy of Sciences (Shanghai, China) and cultured in RPMI 1640 medium (Gibco BRL, Gaithersburg, MD) with 10% (v/v) fetal bovine serum (FBS; Gibco BRL, Gaithersburg, MD). The human embryonic kidney cell line HEK-293 was obtained from the American Type Culture Collection (Manassas, VA) and cultured in Dulbecco’s modified Eagle’s medium (Gibco BRL), supplemented with 10% (v/v) FBS. Each cell line was maintained in a 5% CO2 incubator at 37 °C.
Selection of Candidate Phages
The Ph.D.-7 Phage Display Peptide Library (New England BioLabs, Beverly, MA), with a complexity of 1.05 × 109 independent peptide sequences, was screened using purified recombinant human CD44 proteins (OriGene, Rockville, MD) and a modified screening method based on the manufacturer’s instructions. CD44v3-v10 or CD44s (5 µg in 100 µL of 0.1 M NaHCO3, pH 8.6) was added to different wells on an enzyme-linked immunosorbent assay (ELISA) plate. After swirled to wet the whole surface of the well, the plate was incubated in a sealed box with damp paper overnight at 4 °C with agitation. Then, the plate was blocked with 0.5% (w/v) bovine serum albumin (BSA) in 0.1 M NaHCO3 (pH 8.6) at 37 °C for 1 h before adding approximately 1 × 1011 plaque-forming units (pfu) of the phages to each well. Unbound phages were transferred into the CD44s well to eliminate phages binding to CD44s. Next, unbound phages were incubated with CD44v3-v10 for 1 h at room temperature. After washing 10 times with Tris-buffered saline (50 mM Tris-HCl, pH 7.5) containing 0.1% (v/v) Tween-20 (TBST), bound phages were eluted with 0.2 M glycine-HCl (pH 2.2) supplemented with 1% (w/v) BSA. This reaction was neutralized by adding 1 M Tris-HCl (pH 9.1). Eluants from the first three cycles were amplified and titered. After the fourth round of screening, the eluted phages were titered without amplification. Twenty clones were randomly selected from the titration plate for further analysis.
DNA Sequence Analysis
For DNA isolation, each individual clone was amplified and dissolved in an iodide buffer containing 10 mM Tris-HCl (pH 8.0), 1 mM ethylene diamine tetra-acetic acid (EDTA), and 4 M NaI. Phage DNA was subsequently precipitated with absolute ethyl alcohol and analyzed using an automated DNA sequencer (Applied Biosystems, Foster City, CA) and the primer 5′-AGT AGC AGA AGC CTGAAG A-3′.
Phage ELISA
To select phage clones with the highest affinity and specificity, CD44v3-v10 (1 µg/well) was immobilized on an ELISA plate. BSA and CD44s (1 µg/well) were used as negative controls. The candidate phages and unrelated phages (URps) were added to the corresponding wells (1 × 1010 pfu/well). After incubation for 1 h at 25 °C and six rounds of washing, bound phages were incubated at 37 °C for 1 h with a goat anti-M13 major coat protein polyclonal antibody (Santa Cruz, Dallas, TX) diluted 1:200 in TBST. Next, the samples were incubated with a horseradish peroxidase (HRP)–labeled rabbit anti-goat antibody (ZSGB-BIO, Beijing, China) diluted 1:5000 in TBST. The 3,3′,5,5′-tetramethylbenzidine (TMB; Amresco, Solon, OH) substrate was applied to a color-developing reaction that was terminated by 2 M H2SO4. Measurements were taken using a microplate reader (Biotek, Winooski, VT) with an optical density (OD) of 450 nm. Selectivity was determined as (OD1v – OD2v)/(OD1s – OD2s). OD1v and OD1s represented the OD values obtained when the candidate phages separately bound to CD44v3-v10 and CD44s. OD2s and OD2v indicated background OD values obtained in the CD44s and CD44v3-v10 wells in which the phages were substituted with TBS.
Western Blotting
Cells were lysed at 4 °C for 30 min in buffer containing 50 mM Tris, 150 mM NaCl, 1 mM EDTA, 1% Triton X-100, 1% sodium dodecyl sulfate (SDS), 5 µg/mL aprotinin, 5 µg/mL leupetin, 1 mM phenylmethanesulfonyl fluoride, and 1 mM Na3VO3. Cell lysates were centrifuged at 13,000 rpm for 10 min at 4 °C. The supernatant containing protein was separated and boiled for 5 min. Samples were run on 10% SDS-polyacrylamide gel electrophoresis gels. Following electrophoresis, proteins were transferred to nitrocellulose membranes (Merck Millipore, Billerica, MA). The membranes were blocked in 3% (w/v) BSA in TBST at room temperature for 2 h and then incubated with the goat anti-CD44v3-v10 polyclonal antibody (AbD Serotec, Oxford, UK), the mouse anti-CD44s monoclonal antibody (AbD Serotec), or the mouse anti-GAPDH monoclonal antibody (Santa Cruz) at 1:400, 1:400, and 1:500 dilutions, respectively. The membranes were incubated with the primary antibodies at 4 °C overnight. Then the membranes were incubated with an HRP-conjugated secondary antibody (ZSGB-BIO) diluted 1:5000 in TBST. The results were obtained by chemiluminescence using the ECL Plus Western Blotting Substrate (Pierce, Rockford, IL).
Immunofluorescence
Cells were grown on coverslips overnight and washed three times with phosphate-buffered solution (PBS) consisting of 137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, and 2 mM KH2PO4 (pH 7.4). After being fixed with 4% (w/v) paraformaldehyde in PBS for 15 min, samples were blocked in 3% BSA in PBS for 30 min at 37 °C. For the detection of bound phages, the cells were treated with candidate phage clones (1 × 1010 pfu/well) and probed using the goat anti-M13 major coat protein polyclonal antibody (1:100 v/v) as described above. After being washed with PBS, the cells were incubated with a fluorescein isothiocyanate (FITC)–labeled rabbit anti-goat antibody (1:100 v/v; ZSGB-BIO). To determine peptide affinity, blocked cells were incubated with 10 µM FITC-labeled peptides for 10 min at 37 °C and washed three times with 0.1% Tween-20 in PBS (PBST). Subsequently, the cells nuclei were stained with 4,6-diamidino-2-phenylindole and visualized using a fluorescence microscope (Leica, Solms, HE, Germany) or a laser scanning confocal microscope (Leica).
Peptide Synthesis
The peptide CV-1, representing the most commonly displayed phage sequence, was synthesized by ChinaPeptides Co., Ltd (Shanghai, China) and purified to greater than 95% purity. A scrambled peptide was also synthesized as a control. All peptides were labeled with FITC and biotin at the N-terminus.
Competitive Binding Inhibition Assay
CD44v3-v10 (1 µg/well) was immobilized on an ELISA plate and blocked with 0.5% (w/v) BSA as described above. The candidate peptide or control peptide at concentrations of 0.01, 0.1, 1, 10, 100, and 1000 µM were incubated in the corresponding wells for 1 h at 37 °C prior to the addition of candidate phages (1010 pfu/well). The bound phages were detected using ELISA as described above. The inhibition ratio was calculated according to the following formula: Inhibition ratio = 100% − 100% × (OD test – OD blank)/(OD inhibition – OD blank). OD test and OD inhibition represented OD values obtained separately with or without peptide competitor; the OD blank control indicated the OD values obtained in the control group lacking the anti-M13 antibody.
Binding of the CV-1 Peptide to the Target Protein
The biotin-labeled CV-1 peptide and control peptide were incubated with immobilized CD44v3-v10 (1 µg/well) or CD44s (1 µg/well) at increasing concentrations ranging from 0.01 µM to 100 µM. After being washed, samples were incubated with HRP-labeled streptavidin (ZSGB-BIO) for 30 min. Then, the TMB substrate was added to the wells for a color-developing reaction terminated by 2 M H2SO4. Measurements were taken using a microplate reader. To analyze the binding affinity of CV-1 to CD44v3-v10 quantitatively, the dissociation constant (Kd) was calculated. 27 The target protein was treated with different concentrations of biotin-labeled peptide probe; the bound CV-1 peptide was detected as described above. OD values obtained from wells containing BSA (control protein) were subtracted from wells containing CD44v3-v10 to measure specific binding. The fitting curves and Scatchard plots were generated using Origin 6.0, and the Kd was determined.
Competitive Inhibition ELISA
CD44v3-v10 (10 µg/mL) was immobilized on an ELISA plate and incubated with mouse anti-human CD44s antibody (AbD Serotec) or goat anti-CD44v3-v10 polyclonal antibody (AbD Serotec) at various concentrations for 1 h at 37 °C. Subsequently, biotin-labeled CV-1 (10 µM) and control peptide (10 µM) were used to probe the immobilized CD44 and were detected by ELISA as described above. The inhibition ratio was calculated according to the following formula: Inhibition ratio = 100% − 100% × (OD test – OD blank)/(OD inhibition – OD blank), where OD test and OD inhibition represented OD values obtained with or without competitor and OD blank control represented the OD value obtained from the control group lacking the peptide probe.
Patients and Immunohistochemistry Analysis
A total of 20 patients with gastric cancer who underwent primary surgical resection at the First Affinity Hospital of Xi’an Jiaotong University were included. None of the patients had undergone prior treatment, and all of the patients provided signed informed consent. The study was approved by the Medical Ethics Committee of the First Affinity Hospital of Xi’an Jiaotong University.
Immunohistochemistry was performed on paraffin sections that were 4 µm thick. After deparaffinization and rehydration, the sections were submerged in TE buffer (pH 9.0) containing 10 mM Tris and 1 mM EDTA. Antigen retrieval was performed using the microwave oven method. To eliminate endogenous peroxidase activity, the sections were treated with 3% (v/v) hydrogen peroxide in methanol. Then the sections were then blocked in 5% (v/v) FBS in PBS prior to incubation with 20 µM biotin-labeled CV-1 and then streptavidin-HRP. The latter component formed a complex with biotin that could detected by the addition of a diaminobenzidine (ZSGB-BIO) substrate. After the color development reaction, all sections were counterstained with hematoxylin.
Staining strength was determined using a previously described guideline that incorporated both the percentage of stained cells (1, 0%–10%; 2, 10%–50%; 3, 50%–80%; and 4, >80%) and the staining intensity (1, weak; 2, moderate; and 3, intense). 28 For each specimen, the mean value of five randomly selected fields of view was calculated blindly.
Statistical Analysis
Statistical analyses were performed using SPSS 11.0 for Windows. Data were expressed as the mean ± SD. Student t test was used to analyze differences between groups. p < 0.05 was considered statistically significant.
Results
Subtractive Selection on CD44v3-v10
After elimination of the phages that could interact with BSA and CD44s, CD44v3-v10 was probed by the remaining phages in the 7-mer peptide library ( Fig. 1B ). Bound phages were isolated and titered during four rounds of panning. The recovery ratio (represented by output phages/input phages) increased in each screening cycle, indicating an enrichment of the phages targeting CD44v3-v10 ( Fig. 1C ).
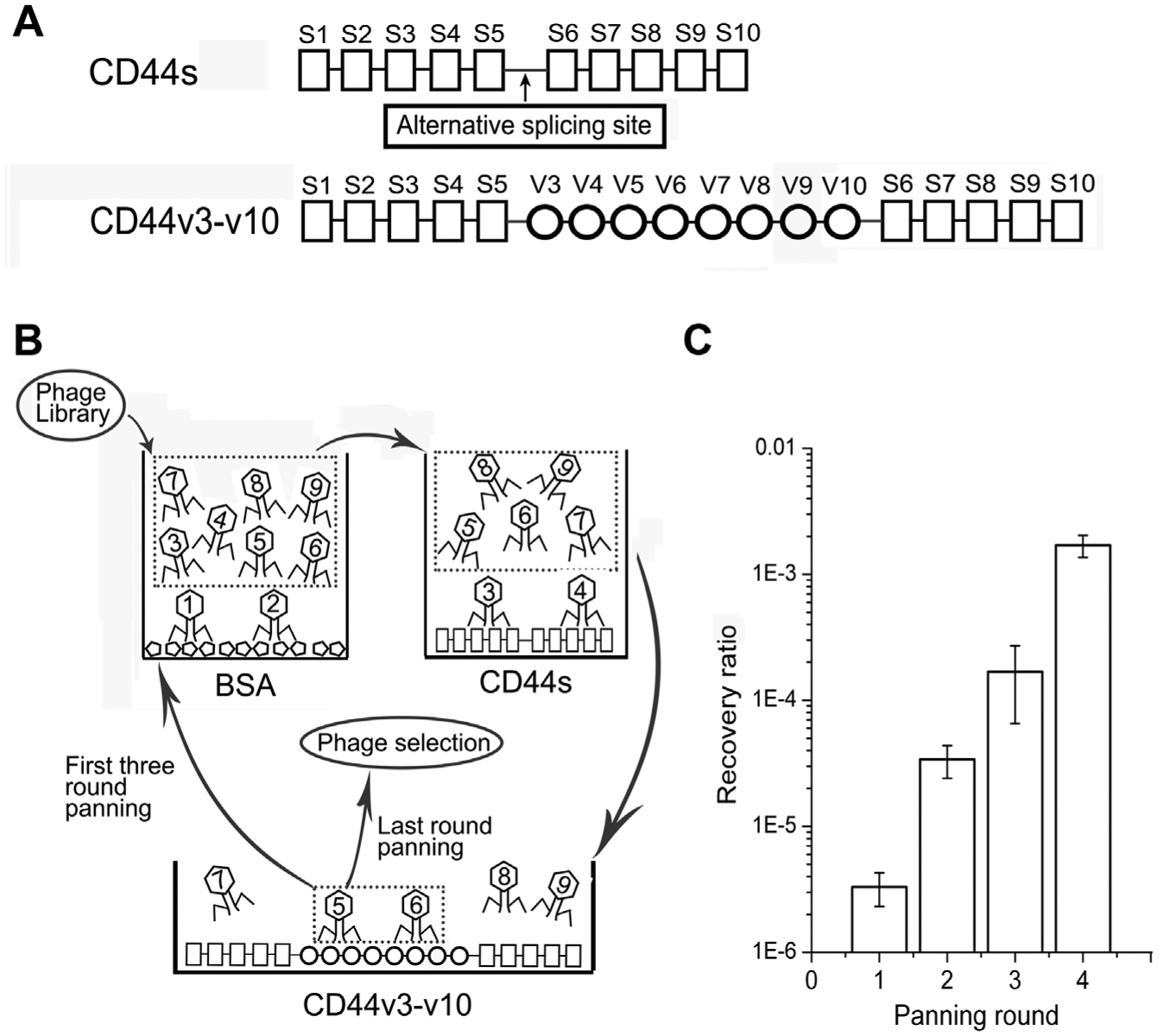
Schematic diagram of the structure of the CD44 gene, the subtractive panning method, and the results. (
Peptide Sequence Analysis
After four rounds of panning, 20 phage clones were randomly selected and analyzed. Three different peptide sequences were obtained (shown in Table 1 ). Sixteen clones contained the sequence THENWPA, designated CV-1. The sequence APQWPAN (CV-2) appeared three times, while another peptide sequence emerged only once. A WPA tripeptide motif was found in CV-1 and CV-2.
Amino acid sequences of screened peptides.
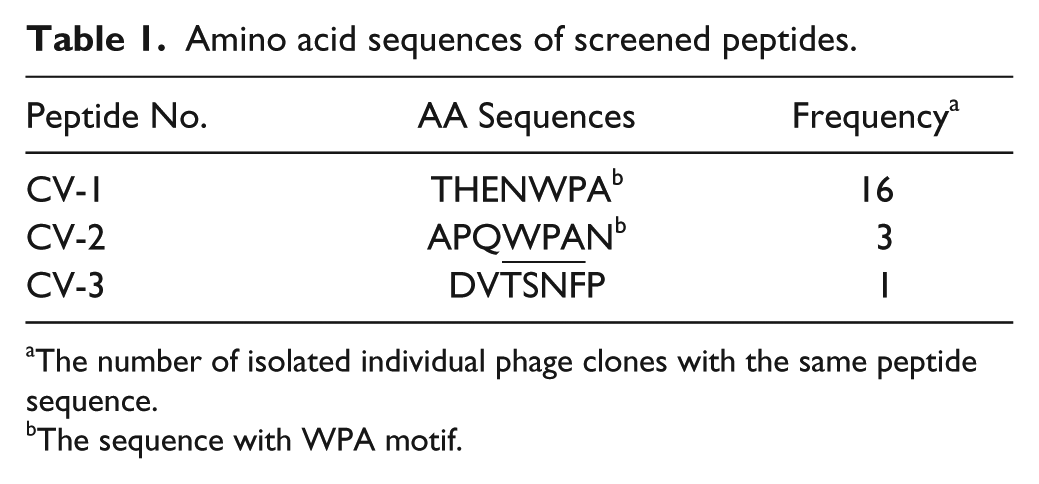

The number of isolated individual phage clones with the same peptide sequence.
The sequence with WPA motif.
Selection of Candidate Phage Clones by ELISA
We performed ELISA on immobilized CD44v3-v10 to select phages carrying peptides with the highest binding affinity and specificity ( Fig. 2A ) using CD44s and BSA as controls. In the CD44v3-v10 group, the OD values of all of the candidate phages were significantly different from the OD values of unrelated phages (p < 0.05). Selectivity was calculated as described above. CV-1 exhibited the highest frequency and selectivity and contained a consensus motif. Thus, CV-1 was selected for further study.
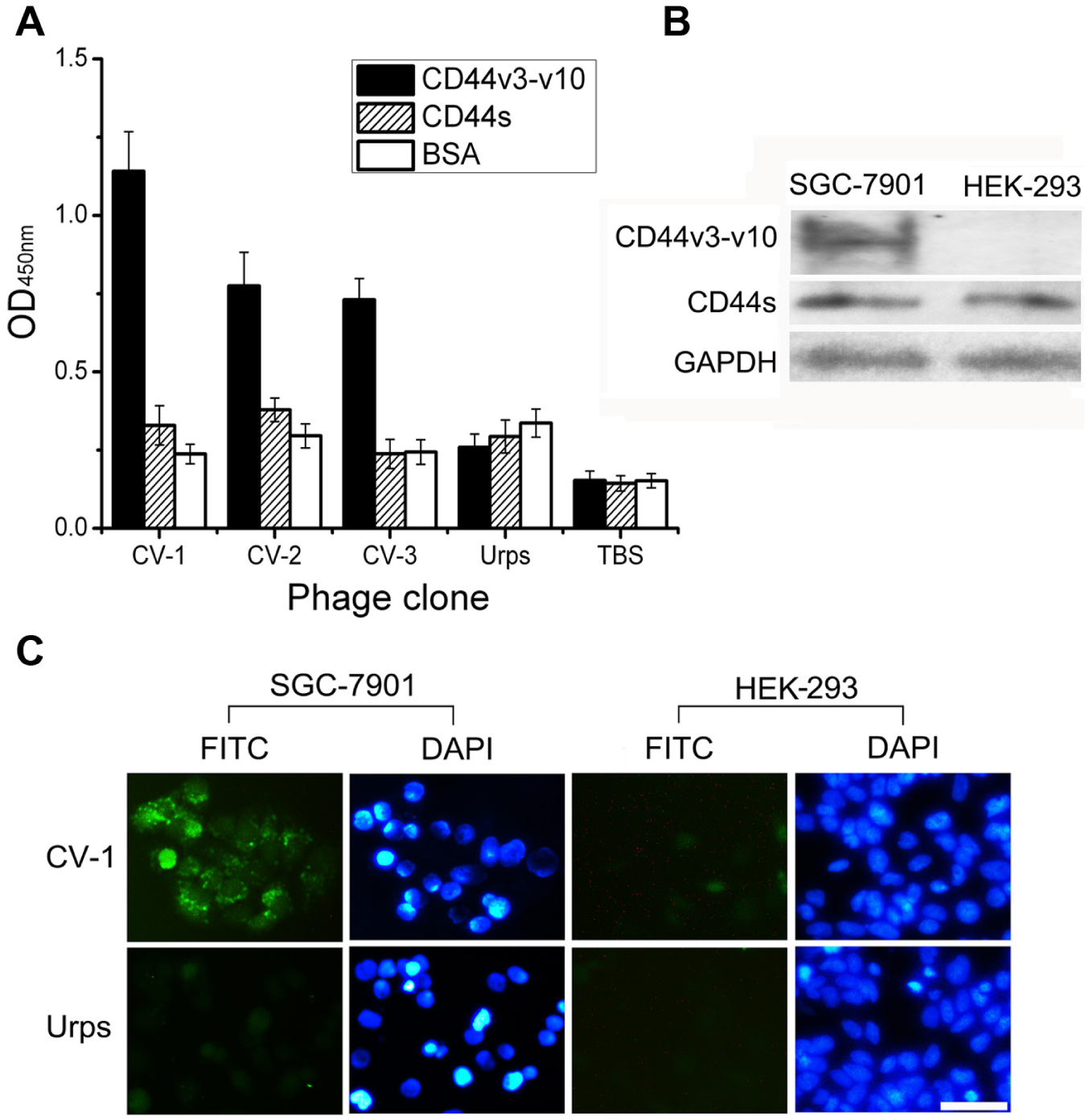
Evaluation of the binding affinity and selectivity of phage clones. (A) Binding of phage clones to CD44v3-v10 (black column), CD44s (striped column), and bovine serum albumin (white column) presented as the optical density value at 450 nm. URps (unrelated phages randomly selected from the original library) and Tris-buffered saline served as the negative and blank controls, respectively. Data are presented as the mean ± SD; four measurements were made for each data point. (
Immunofluorescence Assay of Phage Binding
Although binding of the CV-1 phage to the CD44v3-v10 protein was validated, it was still necessary to confirm the interaction between the CV-1 phage and cancer cells. Expression of CD44s and CD44v3-v10 in SGC-7901 and HEK-293 cells was determined by Western blot analysis. GAPDH was used as the loading control ( Fig. 2B ). As shown in Figure 2C , the candidate phage could bind to the gastric cancer cell line SGC-7901 that expressed CD44v3-v10 but not to HEK-293, a non–tumor-derived cell line expressing only CD44s. However, only background staining occurred when the cells were treated with an unrelated phage.
Binding of the CV-1 Peptide to the CD44v3-v10 Protein
Binding of CV-1 phages to the target protein was investigated using competitive inhibition ELISAs in the presence of the synthesized peptides at different concentrations. The inhibition ratio increased gradually with the concentration of the preincubated CV-1 peptide ( Fig. 3A ). However, no such inhibition was observed in the presence of the control peptide. These results suggested that the CV-1 phage bound to CD44v3-v10 via the displayed peptide but not via the M13 coat protein.
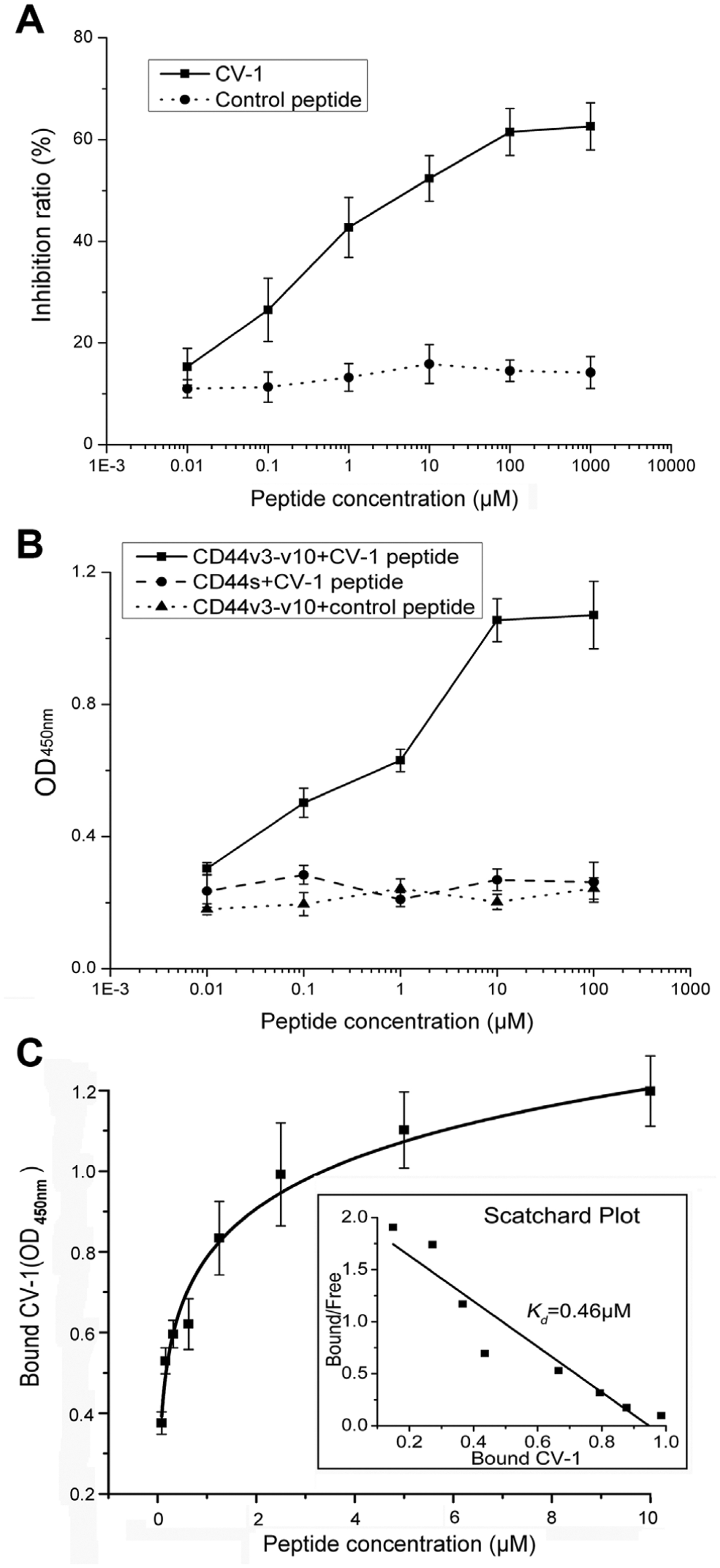
Binding of the CV-1 peptide to the CD44v3-v10 protein. (
Binding of the different concentrations of the CV-1 peptide to CD44v3-v10 was evaluated to determine the specificity and affinity. CV-1 showed a concentration-dependent increase in binding to CD44v3-v10 but not CD44s ( Fig. 3B ). After incubation with CD44v3-v10, the control peptide produced OD values lower than those obtained for CV-1 at each concentration (p < 0.05). These findings indicated that CV-1 bound specifically to CD44v3-v10. The estimated dissociation constant for binding of CV-1 to CD44v3-v10 was equal to 0.46 µM ( Fig. 3C ). Thus, CV-1 also exhibited binding affinity for the target protein.
Investigation of the CV-1 Binding Site on the CD44 Protein
To confirm the presence of the CV-1 binding site on the CD44 protein, we performed a competitive inhibition ELISA with a goat anti-CD44v antibody generated by immunization with recombinant CD44v3-v10 protein. As a control, we used an anti-CD44s antibody that recognized an epitope outside the regions encoded by the variable exons. The binding of CV-1 decreased with increasing concentrations of anti-CD44v3-v10 antibody, while the inhibition ratio rose gradually ( Fig. 4A ). In contrast, no such dose-dependent inhibition occurred in the control peptide group. Neither the CV-1 nor the control peptide exhibited competitive inhibition in the presence of the anti-CD44s antibody ( Fig. 4B ), which indicated that CV-1 was still able to bind to the target protein blocked by the anti-CD44s antibody.
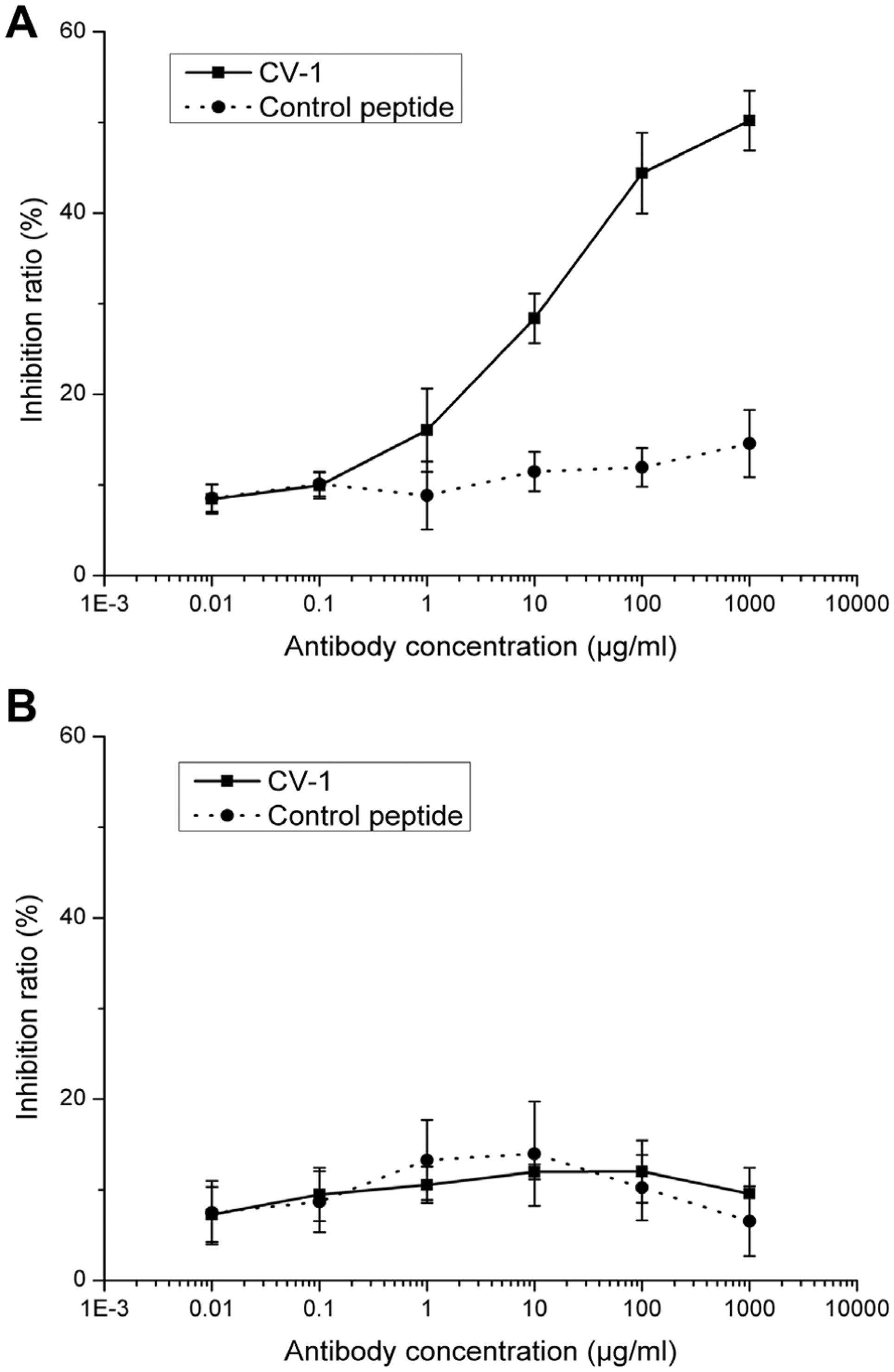
Competitive inhibition of CV-1 binding to CD44v3-v10 by antibodies. (
Binding of CV-1 Peptide to Cells
Binding activity of the FITC-labeled CV-1 peptide to CD44v on cancer cells was investigated using immunofluorescence assays. As shown in Figure 5A , positive staining was observed on SGC-7901 cells but not HEK-293 cells, which did not express CD44v. Specific fluorescent staining was distributed at the cell membrane and in the cytoplasm. Only background staining was observed on SGC-7901 cells incubated with the control peptide.
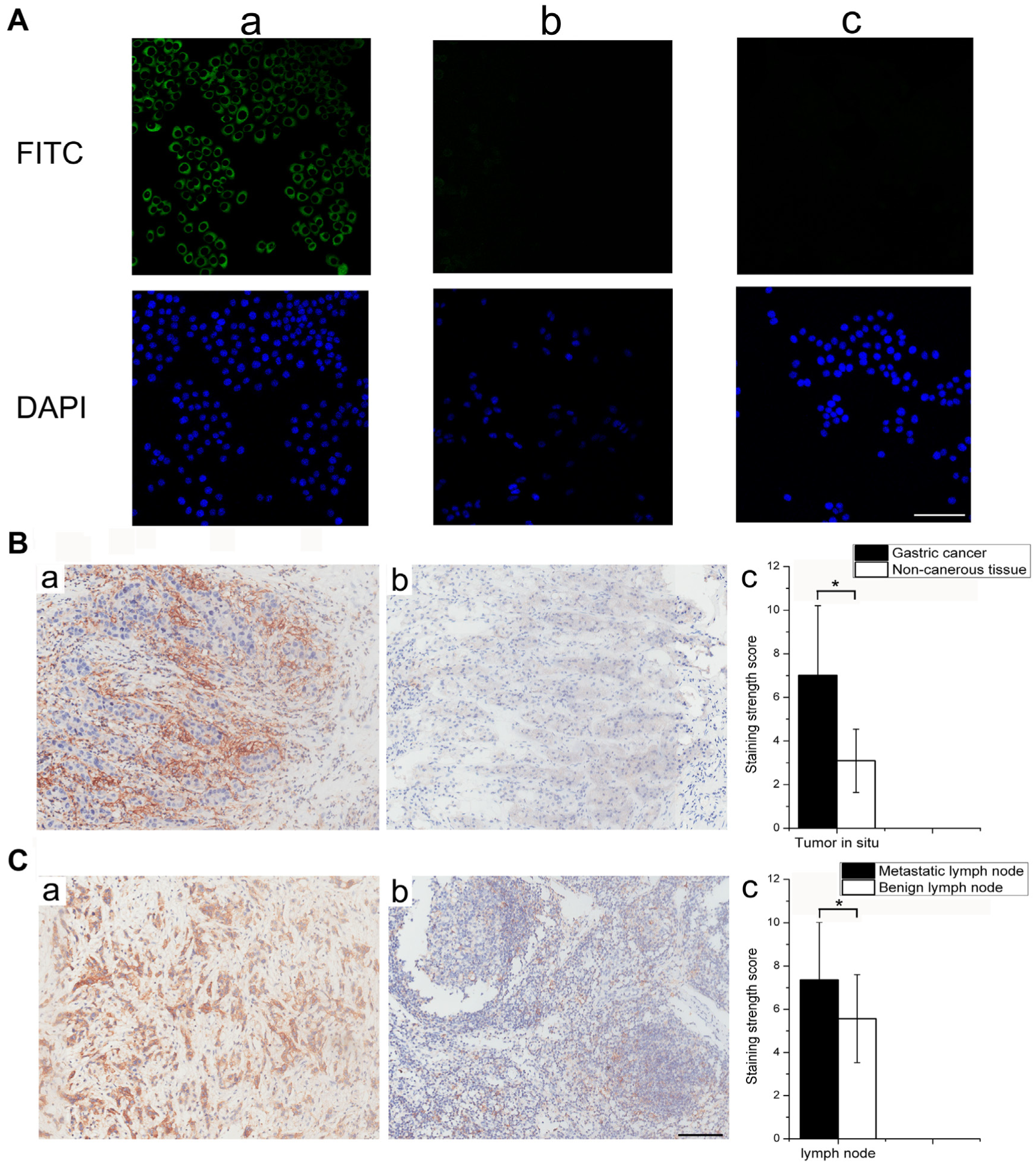
Binding of the CV-1 peptide to cells and tissues. (
Immunohistochemical Staining of CV-1 on Gastric Cancer Tissues
The binding specificity and affinity of CV-1 for the CD44v protein and cancer cells was confirmed. To further evaluate CV-1 as a potential probe for cancer diagnosis and treatment, immunohistochemical staining analysis of biotin-labeled CV-1 was performed on 20 pairs of gastric cancer and adjacent noncancerous gastric tissues, as well as five metastatic lymph nodes. As shown in Figure 5B , positive staining was distributed at the cell membrane and in the cytoplasm. The average score of CV-1 staining in gastric cancer was significantly higher compared with the nontumor tissues (7.01 ± 3.19 v. 3.09 ± 1.45, respectively; p < 0.05). CV-1 staining of the metastatic lymph nodes was significantly stronger compared with staining of the benign lymph nodes (p < 0.05). Thus, CV-1 was implicated as an indicator of not only tumors in situ but also metastatic lymph nodes.
Discussion
Expression of CD44 isoforms that contain variable exons is an important determinant of gastric cancer diagnosis. Many types of CD44v with alternatively spliced v3-v10 exons were expressed at high levels in cancer tissues and correlated with gastric cancer progression. The regions in the whole CD44 molecule maintain a natural structure and do not exist independently. To maximize the opportunities for phages binding to CD44v, a natural large form of the protein (CD44v3-v10) was used. In this CD44 isoform, approximately 50% of all the amino acids were encoded by alternative exons. The recombinant protein was generated in a eukaryotic expression host system (human HEK-293 cells) to allow for more biologically relevant CD44 expression.
To select the phages that bound specifically to CD44v, the phages from the peptide library were incubated with BSA and CD44s prior to being screened against CD44v3-v10. In this way, the phages with the capacity to bind to CD44s were subtracted. Successive rounds of biopanning resulted in significant phage enrichment; 20 clones were randomly selected after the last round of panning. Because some clones displayed the same peptide sequence, DNA sequencing of the phages was performed prior to ELISA to simplify the process of candidate clone selection. Of the three sequences identified, CV-1 occurred with the highest frequency and exhibited the highest selectivity for binding to CD44v3-v10, as well as binding affinity and specificity for cancer cells. Therefore, CV-1 was chosen for peptide synthesis.
CV-1/CD44v3-v10 binding was confirmed using competitive inhibition assays. The inhibition ratio increased in a dose-dependent manner, eliminating the possibility of nonspecific binding of the M13 coat protein to CD44v3-v10. In contrast to CD44s, binding of the CV-1 peptide to CD44v3-v10 increased with the peptide concentration, thereby providing evidence for the specific binding of this probe.
Although screening against recombinant proteins maximizes the specificity of candidate peptides, this approach requires validation in cell-based systems. As expected, CV-1 bound to gastric cancer cells that expressed various exons of CD44 but not an embryo-derived cell line expressing only CD44s. For further confirmation, the CV-1 binding site on CD44 was investigated using an antibody inhibition ELISA. The binding activity of CV-1 was significantly inhibited by the anti–CD44v3-v10 antibody but not by an antibody that recognized only the epitope on CD44s. The results indicated that the binding site was located in the regions encoded by the v3-v10 exons.
Because the binding activity of CV-1 to CD44v was validated, we next investigated CV-1 binding to human cancer and nontumor tissues to evaluate the potential of this peptide to serve as a tool for cancer diagnosis. As expected, CV-1 was more frequently detected in the cancer samples compared with adjacent noncancerous gastric tissues. CV-1 staining was also increased in metastatic lymph nodes, which highly expressed CD44v and were correlated with patient outcomes. 7 Thus, CV-1 not only is a probe to detect local lesions and lymph node metastasis but may also serve as an indicator of tumor progression and recurrence.
In previous studies, CD44-specific ligands were developed for tumor targeting. Verel et al. 29 developed monoclonal antibodies that bound to CD44v with different affinities (Kd ranging from 11 nM to 2.4 nM). Among these, antibodies with lower affinities were advantageous with respect to selective tumor uptake. 29 Peptide probes have several advantages compared with antibodies, including high affinity, diversity, the ability to penetrate deep into tissue, rapid clearance, and minimal immunogenicity. 30 CD44-specific peptides have been previously described for the diagnosis and treatment of some tumors. For example, a peptide binding to CD44 with affinity (Kd 256.5 nM) was used to target breast cancer cells as a potential diagnostic reagent. 17 A CD44-specific peptide was been shown to be effective against melanoma cells by inhibiting tumor cell migration, invasion, and angiogenesis. 31 The binding affinity of CV-1 (Kd 457.5 nM) is lower than the CD44 ligands mentioned above. However, in contrast to the other ligands, CV-1 is a peptide that specifically binds to the region of the CD44 protein encoded by the variable exons. Based on our results, CV-1 has the potential to serve as a determinant of gastric cancer cells and tissues.
In conclusion, we screened a novel peptide specific for CD44v. The in vitro binding assay provided compelling evidence for the affinity and specificity of CV-1 binding to the CD44v3-v10 protein, gastric cancer cells, and tissues, suggesting that this peptide has the potential to be used as a probe to detect and monitor CD44v-positive stomach tumors. In future studies, we will focus on the application of CV-1 for in vivo tumor imaging and treatment of tumor growth and metastasis, thereby making CV-1 more applicable to the diagnosis and treatment of cancer.
Footnotes
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (No. 81172359 and 81472747).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the National Natural Science Foundation of China (No. 81172359 and 81472747).
