Abstract
Leukotrienes (LTs) and related species are proinflammatory lipid mediators derived from arachidonic acid (AA) that have pathological roles in autoimmune and inflammatory conditions, cardiovascular diseases, and cancer. 5-Lipoxygenase activating protein (FLAP) plays a critical accessory role in the conversion of AA to LTA4, and its subsequent conversion to LTC4 by LTC4 synthase. Pharmacological inhibition of FLAP results in a loss of LT production by preventing the biosynthesis of both LTB4 and LTC4, making it an attractive target for the treatment of inflammatory diseases in which LTs likely play a role. Small-molecule (SM) drugs often exhibit polypharmacology through various pathways, which may explain the differential therapeutic efficacies of compounds sharing structural similarity. We have profiled a series of SM FLAP modulators for their selectivity across enzymes of AA cascade in human whole blood (HWB), using a recently developed LC/MS (liquid chromatography–mass spectrometry)-based high-throughput lipidomics platform that monitors 122 eicosanoids in multiplex. Highly efficient data acquisition coupled with fast and accurate data analysis allowed facile compound profiling from ex vivo study samples. This platform allowed us to quantitatively map the effects of those SMs on the entire AA cascade, demonstrating its potential to discriminate structurally related compounds.
Introduction
Eicosanoids, including leukotrienes (LTs), eoxins, lipoxins, prostaglandins (PGs), and epoxyeicosatrienoic acids (EETs), are all bioactive lipid mediators derived from the oxidation of arachidonic acid (AA). 1 The synthesis of eicosanoids is initiated by inflammatory stimulation which promotes the translocation of cytoplasmic phospholipase A2 (PLA2) to cell membrane. 2 After its release from membrane by PLA2, AA is then metabolized to eicosanoids by three major enzymatic pathways: lipoxygenase (LOX), cyclooxygenase (COX), and cytochrome P450 (CYP) oxygenase. Because of the critical roles of eicosanoids in diverse pathological and physiological processes, these pathways have become the target for the treatment of inflammation, asthma, pain, cardiovascular diseases, and cancer. 3
In the 5-lipoxygenase (5-LO) pathway, the synthesis of LTs, including LTB4, the cysteinyl LTs (CysLTs), and 5-oxo-eicosatertraenoate (5-oxo-ETE), requires 5-lipoxygenase-activating protein (FLAP) as an essential partner of 5-LO for converting AA into the epoxide-containing first intermediate, LTA4. Identified by Merck scientists as the protein binding target of LT synthesis inhibitor, MK-886, 4 FLAP facilitates this conversion step via an unknown mechanism. Furthermore, FLAP assists in the condensation of LTA4 with glutathione to form the CysLT, LTC4. 5 Silencing of FLAP via pharmacological inhibition or genetic manipulation resulted in complete loss of production of all LTs. 5 LTs exert their biological effects through the activation of specific G protein-coupled receptors (GPCRs). LTB4, generated by LTA4 hydrolase, signals through BLT1 and BLT2, triggering chemo-attraction and activation of neutrophils, monocytes, and eosinophils, as well as the production of proinflammatory cytokines. 6 CysLTs also act through several GPCRs, including Cys-LT1, Cys-LT2, and P2Y9, to trigger bronchoconstriction and increase in vascular permeability. 7 CysLTs are potent inducers of diverse immune responses. 8 Mainly targeting eosinophils, 5-oxo-ETE has been hypothesized to play a critical role in asthma and may contribute to prostate tumor progression due to its proliferative effects. 9
There are several approved drugs on the market targeting the LT pathway, such as montelukast (Singulair), zafirlukast (Accolate), and pranlukast (Onon), all of which are Cys-LT1 receptor antagonists, not directly inhibiting LT production. They have limited efficacy, primarily in chronic and allergic asthmatics. 10 Zileuton (Zyflo), the only commercially available 5-LO inhibitor, provides an additional therapeutic option by its inhibition of all LT synthesis. 10 However, zileuton is sparsely used in the clinic, due to its poor pharmacokinetics 11 and idiosyncratic hepatotoxicity. 12 By blocking the formation of all LTs, FLAP inhibitors such as MK-886, MK-0591, and BAY-X1005 showed promising clinical profiles in small clinical studies for several indications. 13 None of those compounds were brought to the market for undisclosed reasons. FLAP bears no additional functions beyond its role in LT synthesis, and FLAP-deficient mice have no observable developmental or adult detrimental phenotypes. 14 In humans, the gene ALOX5AP encoding FLAP is associated with greater risks of myocardial infarction and stroke by increasing LT production. 15 Recent insights into the important roles of LTs in carcinogenesis have added more fuel to the fire. 16 Through different mechanisms, in colon, prostate, and pancreatic cancers, these bioactive lipids can modulate tumor cell proliferation and apoptosis, and promote cancer growth and metastasis. This provides evidence that certain proinflammatory eicosanoids, such as LTB4 and LTE4, are potential drug targets for prevention of certain types of cancers. Furthermore, the elucidation of the three-dimensional (3D) structure of human FLAP added impetus to the search for its small-molecule (SM) modulators. 17
SM drugs often exhibit polypharmacology (on- and off-target effects) through various pathways that remain hard to predict via computational chemistry, yet it may explain the differential therapeutic index observed with compounds sharing close structural similarity. Evidence for the critical role that off-target pharmacology plays in SM therapeutic efficacy indeed keeps mounting, as more sophisticated ways to detect and quantify it emerge. 18 Here, we demonstrate the power of lipidomic profiling to identify key SMs from a series of FLAP modulators. For this, we performed a comprehensive analysis of their potency at target, and selectivity across enzymes of the AA cascade, using a recently developed liquid chromatography–mass spectrometry (LC/MS)-based high-throughput lipidomics platform that simultaneously monitors 122 eicosanoids derived from AA. 19 Eight novel in-house synthesized FLAP inhibitors and two benchmark compounds (MK0591 and MK886) were evaluated in HWB using calcium ionophore as the stimulus. Rapid data acquisition and fully automated data analysis allowed for facile compound profiling from ex vivo study samples. By systematically quantifying effects on the synthesis of LTs, prostaglandins, thromboxanes, eoxins, and lipoxins, we were able to generate unprecedented maps of the diverse activities of those SMs. Thus, this platform displayed the potential to discriminate structurally related compounds and provide useful information about modulation of the eicosanoid pathway.
Materials and Methods
Reagents and Materials
Fresh heparinized HWB samples (healthy donors taking no current anti-inflammatory medications) were obtained from the Normal Blood Donor Service at the Scripps Research Institute (La Jolla, CA) on the day of experiments, treated with heparin as anticoagulant, and used immediately for experiments. All eicosanoids and their deuterated internal standards (IS) were purchased from Cayman Chemical (Ann Arbor, MI). The calcium ionophore, A23187, and MK-886 were from Sigma-Aldrich (St. Louis, MO). MK-0591 was obtained from ApexBio (Boston, MA). RPMI-1640 medium was from Thermo Scientific (Rockford, IL). The LTB4 enzyme-linked immunosorbent assay (ELISA) kit was from Enzo Life Sciences (Farmingdale, NY). All high-performance liquid chromatography (HPLC)-grade solvents were purchased from EMD Millipore (Billerica, MA). All other FLAP inhibitors (JNJ prefix) were synthesized at the Immunology Discovery Medicinal Chemistry laboratories of Janssen R&D (La Jolla, CA).
Compound Treatment
HWB samples were diluted 1:1 (v/v) with RPMI-1640 medium and aliquoted into 96-well plates at 200 μL/well. Each well was pretreated with or without FLAP inhibitors ( Table 1 ) at different concentrations (0.01, 0.1, 1, 10 μM) for 15 min at 37 °C, followed by a 30-min incubation with calcium ionophore A23187 (final concentration 30 μM). Compounds were tested for samples from each donor in triplicate.
FLAP Inhibitors IC50 against LTB4 via ELISA and Percent Inhibition via Lipidomics.
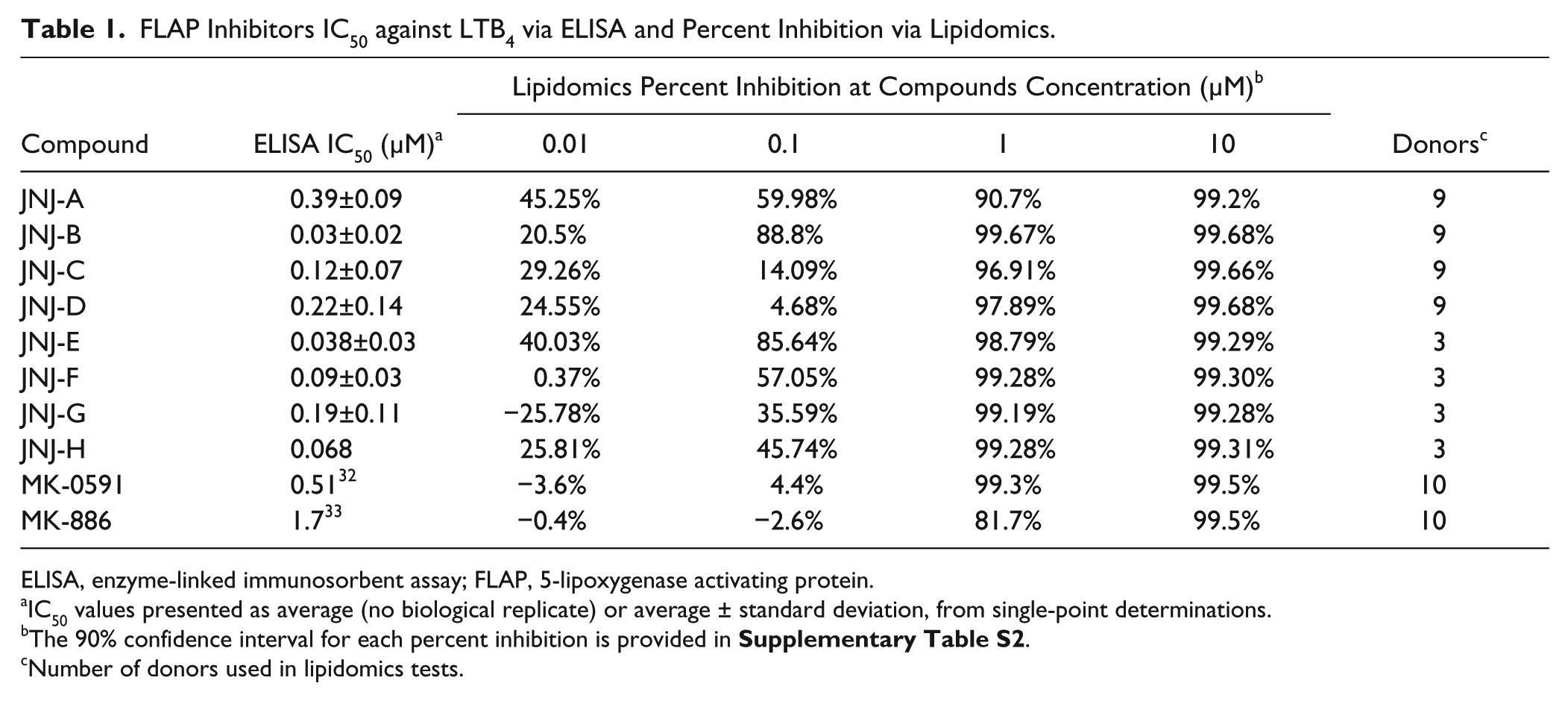
ELISA, enzyme-linked immunosorbent assay; FLAP, 5-lipoxygenase activating protein.
IC50 values presented as average (no biological replicate) or average ± standard deviation, from single-point determinations.
The 90% confidence interval for each percent inhibition is provided in
Number of donors used in lipidomics tests.
ELISA Detection of LTB4
The measurement of LTB4 by ELISA was performed according to the protocol provided by Enzo Life Sciences. Briefly, treated HWB samples were diluted 1:240 (v/v) with assay buffer, followed by the addition of LTB4 conjugate and antibody. The mixture was incubated at room temperature for 2 h before adding p-nitrophenyl phosphate (pNPP) for development. Samples were then incubated in the dark at 37 °C for 2.5 h. Plates were read at 405 nm, and IC50 values were calculated using a macro in Excel.
Sample Preparation and LC/MS/MS Analysis
On treatment, HWB samples and standards (diluted in half-log steps from 300 to 0.1 ng/mL) were processed and submitted for LC/MS/MS analysis as previously described.
19
Briefly, 100 µl HWB was diluted with 100 µl RPMI media, followed by centrifugation at 1300 rpm for 10 min. Then, 20 μL of the IS mix (50 pg/μL, 1 ng of each deuterated standard) was spiked into the supernatants immediately. After diluting with 10% MeOH in water to 1 mL, the lipids were extracted via solid phase extraction (SPE) using Strata-X polymeric reverse phase 96-well plates (Phenomenex, Torrance, CA). The SPE plates were then washed with 10% MeOH, and lipids were eluted by MeOH, and then dried using a Speedvac. After resuspension with 100 µl mobile phase A (water/acetonitrile/acetic acid=70/30/0.1, v/v/v), the samples were injected into an Acquity ultra-performance liquid chromatography (UPLC) system (Waters, Milford, MA) and separated on a Kinetex C18 column (Phenomenex, Torrance, CA). The linear LC gradient, given as percentages of mobile phase B (isopropanol/acetonitrile/acetic acid = 50/50/0.02, v/v/v), was: 0 to 0.24 min, 0.5%; 0.6 min, 30%; 2.75 min, 45%; 3.25 min, 60%; 4.5 min, 75%; 4.62 min, 95%; 5.2 min, 95%; 5.3 min, 0.5%; 6.5 min, 0.5%. The total run time was 6.5 min. MS detection was performed with an AB SCIEX QTRAP 6500 system (AB SCIEX, Foster City, CA) under Analyst 1.6 software. Data acquisition was realized using scheduled multiple reaction monitoring (sMRM) in negative ion mode with the following parameters: −5500 V ion spray voltage, 30 psi curtain gas, 60 psi ion source gas (GS1), 60 psi ion source gas (GS2), 8 psi collision gas (CAD), −10 V entrance potential (EP), −10 V collision cell exit potential (CXP), 600 °C, and 20 s MRM detection window. The optimized MRM fragmentation transitions and MS parameters for each lipid and internal standards are provided in
Data Analysis and Visualization
Absolute quantification was performed using a stable isotope dilution method, in which each lipid was quantified against an IS on a 10-point calibration curve. Data processing was completed automatically using our in-house developed R-based software, including retention time shift (RTS) correction, global normalization of the raw intensities of all eicosanoids across all samples, and monotonic standard curve generation and analyte quantification. 19 Data validation and quality control (QC), including principal component analysis (PCA) to detect outlier samples, differential production analysis of eicosanoid lipids, and hierarchical clustering, were conducted using ArrayStudio V7.0 (www.omicsoft.com). For easy visualization, the absolute concentrations of lipids on different treatments were log2 transformed, and they were presented as heat maps to reflect their production profiles under different treatment conditions.
For each subject, the effect of calcium ionophore induction on individual lipids and the inhibition effect of test compounds at various calcium ionophore induction concentrations were calculated as log2 ratios based on samples assayed on the same plate. In total, 1024 HWB samples from 22 healthy subjects were treated ex vivo with calcium ionophore, in the absence or presence of FLAP compounds at four concentrations (0.01, 0.1, 1, and 10 μM). The lipid profiles of these 1024 samples were assayed on thirty-six 96-well plates. QC among the 1024 samples was performed based on PCA and correlation coefficients among technical replicates; among all samples, 30 were detected as outliers and removed from further analysis. In this way, 291 sets of log2 ratios were generated representing the effects under 41 conditions, including induction of calcium ionophore, and treatment effects of 10 FLAP inhibitors with four concentrations for each compound among 22 subjects. Although the effect of calcium ionophore induction was measured in all subjects, the effect of individual FLAP inhibitors was assayed in subsets of them. The averaged effects under 41 conditions among all tested subjects on individual lipids were further calculated as the averaged log2 ratios and transformed into percentages of induction and inhibition. The 90% confidence intervals of percent induction by calcium ionophore, and of percent inhibition by compound treatments, were calculated based on the 90% confidence intervals of the corresponding log2 ratios, respectively. Lipids that were significantly induced by calcium ionophore stimulation were selected based on the result of one group t-test among all 22 subjects, with a fold change greater than two and a false discovery rate (Benjamini–Hochberg procedure) less than 0.05.
Results
An Automated Lipidomics Platform for the Global Profiling of Eicosanoids Modulation by FLAP Inhibitors
To explore the therapeutic potential of SM AA-pathway modulators, we synthesized various FLAP inhibitors and assessed their activity against the LT pathway by monitoring LTB4 production ex vivo in HWB after calcium ionophore stimulation via ELISA ( Table 1 ). LTB4 ELISA and the detection of urinary LTE4 are validated clinical readouts to assess FLAP target engagement, 20 and it has been a standard approach to measure LTB4 levels to derive the IC50 values of compounds tested. But, by solely relying on the LTB4 to monitor SM inhibitory activities, it was not possible to assess the effect of those SMs on the production of cysLTs, 5-hydroeicosatetraenoic acid (5-HETE), and 5-oxo-ETE, which are all important in the induction of inflammation and bronchoconstriction. Finally, it is impossible to evaluate shunting effects common in the AA cascade with a single readout assay. Our platform 19 enables a “fingerprint” composed of multiplexed quantification of 122 eicosanoids, covering the LOX, COX, and CYP450 pathways. Many other groups have also described LC/MS-based methods to quantify eicosanoids in various biological matrices,21–24 yet the throughput or coverage does not meet the needs for larger-scale analysis. Addition of automation routines not only improves the efficiency of the entire work flow but also enhances the accuracy of quantification due to minimal human intervention. Our software is capable of tracking RT for each IS, generating a functional relationship between the expected retention time and the observed RTS among all ISs in each individual sample, and adjusting the RT for individual peaks of molecules to be measured, based on the functional relationship generated ( Fig. 1A, B ). The RTS threshold to define an outlier is 5 s. Furthermore, summaries of dynamic ranges of IS spikes across all samples on one 96-well plate, as well as dynamic ranges of all primary lipids in a series of diluted samples to generate standard curves, are also provided for easy detection of outliers ( Fig. 1C, D ). Overall, the fast data collection at bench coupled with highly efficient data analysis in silico allow facile compound profiling covering the entire AA cascade.
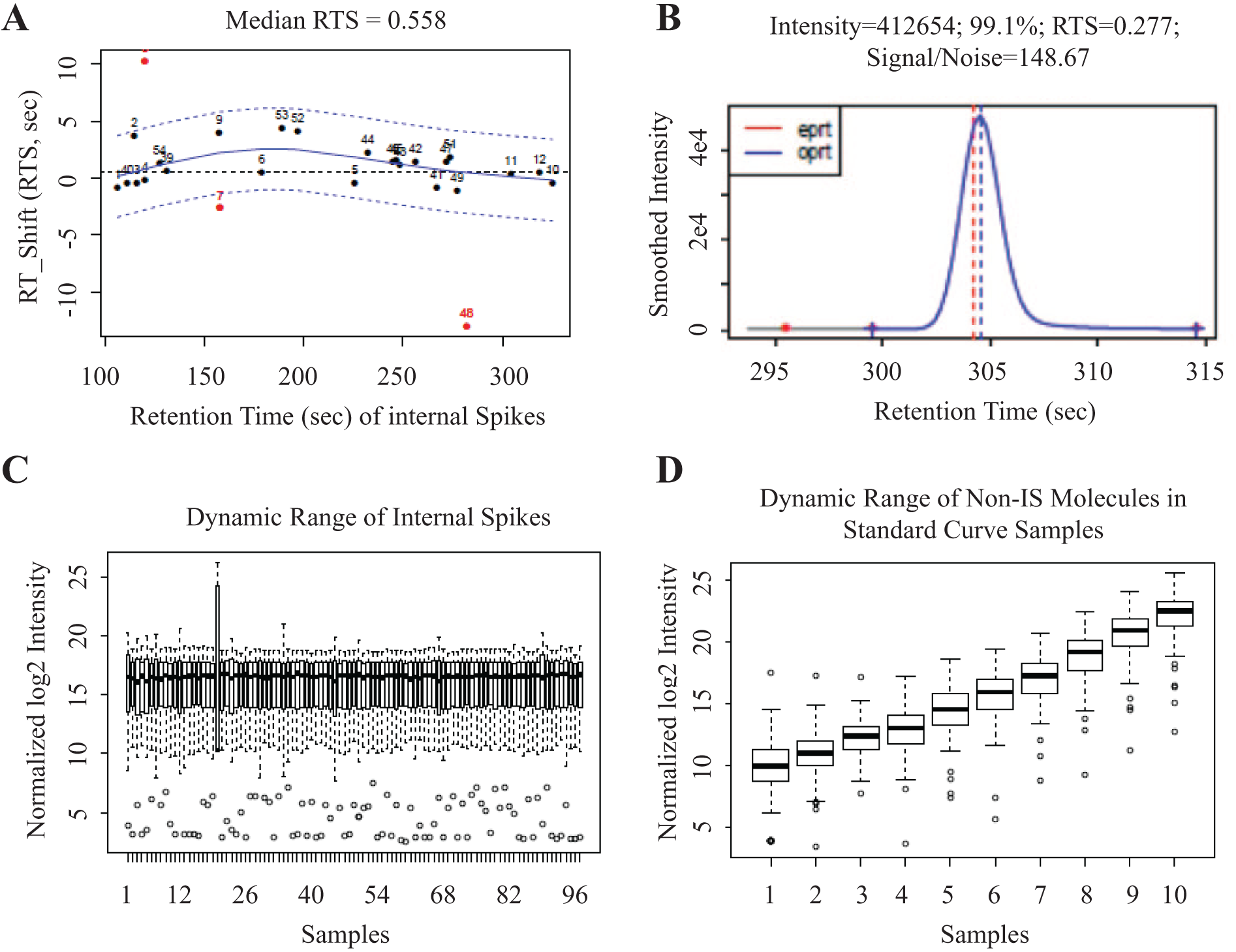
Automated data processing using in-house developed software. (
To better understand the polypharmacology of FLAP inhibitors, we used the lipidomics platform to globally profile their differential modulation of 122 eicosanoids. Eight internally synthesized SMs together with two reference compounds (
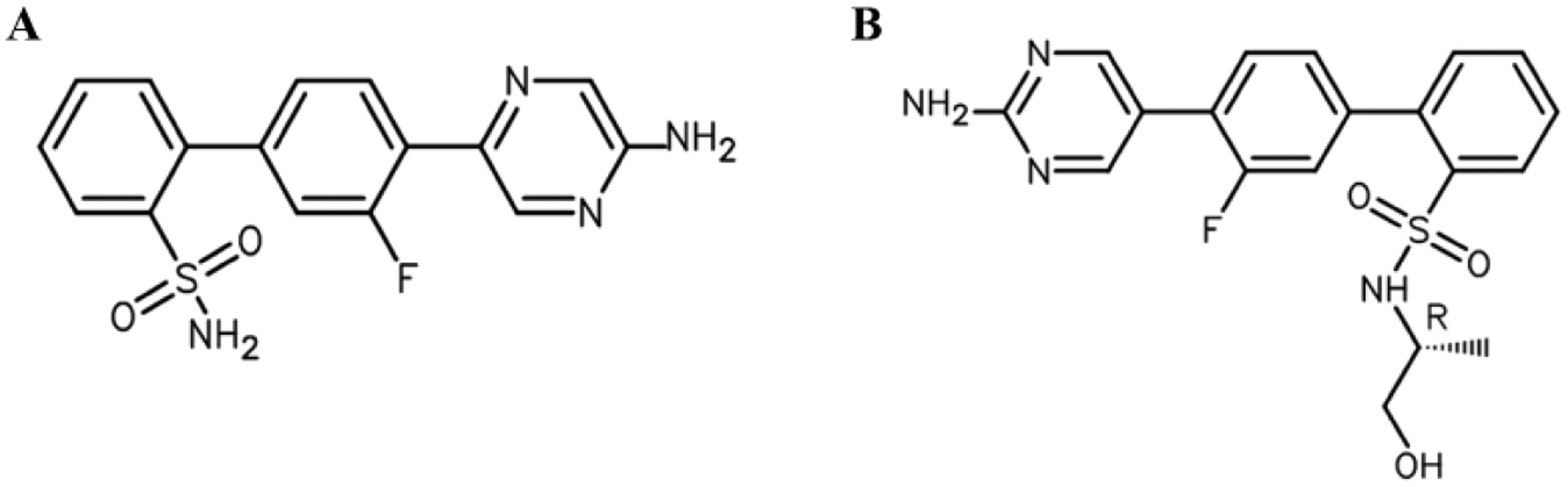
Structures of (A) JNJ-A and (B) JNJ-E.
The percent inhibition values on LTB4 production measured via lipidomics is presented for all compounds, at each tested concentration (0.01, 0.1, 1, and 10 µM), in
Table 1
(the percent inhibition and its 90% confidence interval on all lipids are provided in
The power of this multiprobe platform is further highlighted when data are visualized in the format of a heat map (
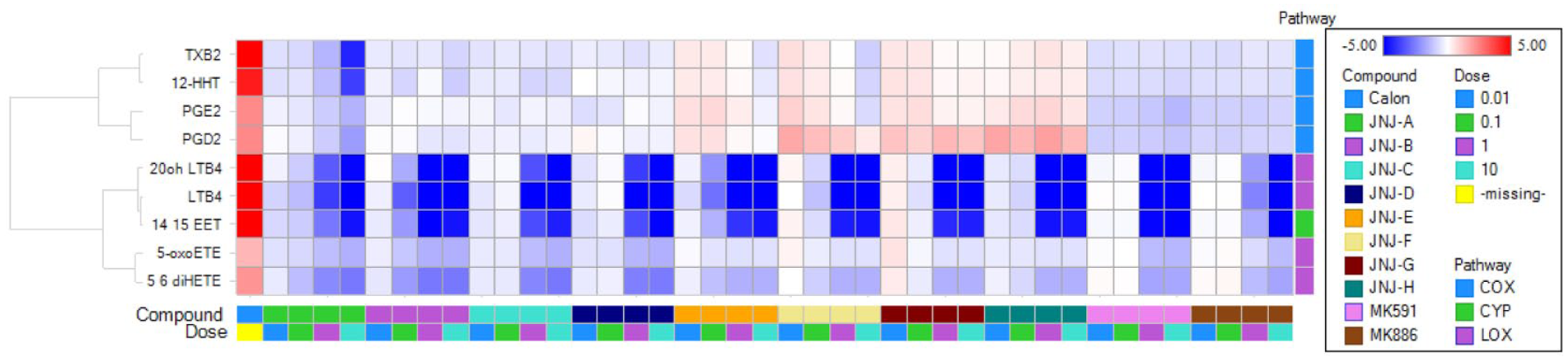
Lipidomic analysis reveals the differential behavior of FLAP modulators on the eicosanoid biosynthetic pathways. Heat map of averaged log2 ratios (among multiple subjects) of nine selected lipids under 41 conditions, including induction of calcium ionophore (labeled as Calon), and treatment effects of 10 compounds, with four concentrations for each compound.
Percentage Inhibition (%) of Eicosanoids by FLAP Inhibitors at 0.1 and 1 μM.
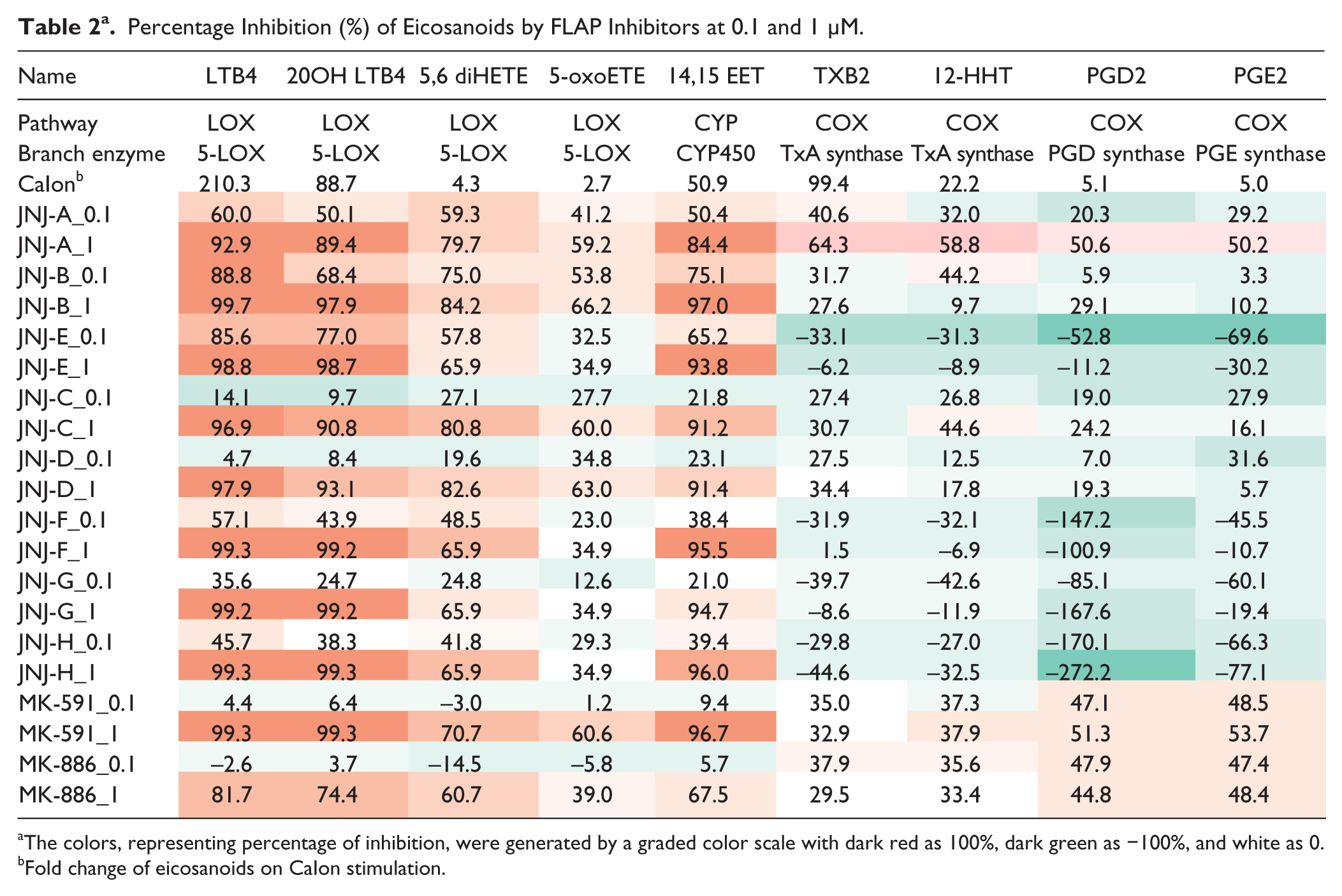
The colors, representing percentage of inhibition, were generated by a graded color scale with dark red as 100%, dark green as −100%, and white as 0.
Fold change of eicosanoids on CaIon stimulation.
Intersubject Variability on Same Drug Treatment Revealed by Eicosanome Profiling
Donor variability is intrinsic to any experiment with human subjects, because of the impossibility of controlling genetic and environmental factors. This is particularly the case with the study of eicosanoid products, in which genetics, gender, age, diet, activity, emotional distress, lifestyles (use of alcohol, tobacco, and recreational drugs), medication history, and many other factors can play major roles in modulating their production. 27 Although we performed very stringent statistical analysis by merging all sample plates and setting high and low detection limits (LLODs) to minimize the effect of intersubject variations, our reporter probes still indicate variability in donor responses to the same drug treatment. Figure 4 depicts changes in nine selected lipids upon pre-incubation of JNJ-B followed by calcium ionophore stimulation in HWB from five healthy subjects. Although similar results were generated from most donors, donor 21 stands out for its overall weak response to the compound at 0.1 µM; likewise, donor 33 (donor #3, batch 3; for coding description, please refer to Fig. 4 legend) drew our attention for its differential responses: instead of blocking the production of 5-LO metabolites, such as 5-oxo-ETE and 20-OH-LTB4, JNJ-B significantly lowered the concentration of COX metabolites, including TXB2, 12-HHT, and PGD2. Furthermore, this compound shows higher potency in subject 32 relative to other donors.
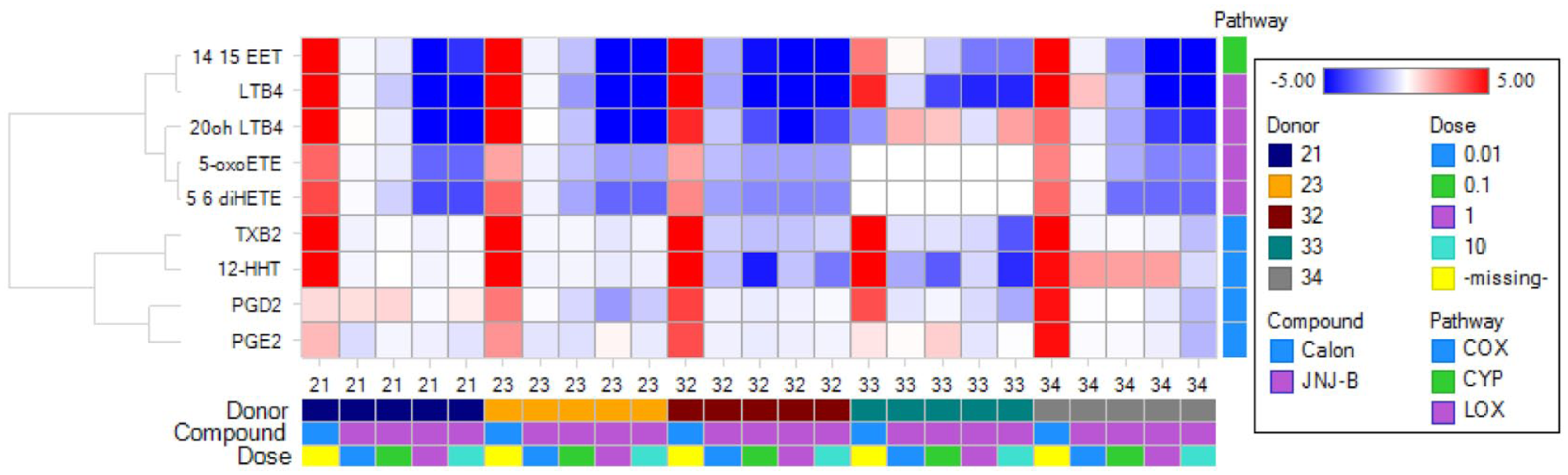
Heat map of log2 of nine selected lipids under five conditions, including induction by calcium ionophore (labeled as Calon), and treatment effects of four increasing concentrations of JNJ-B, among five subjects. The donors were categorized by batch; for example, donor 32 represents the second donor from batch 3.
Discussion
Taken in perspective, our multiplexed lipidomic platform offers advantages over conventional single-readout assays in many aspects. We were able to differentiate FLAP inhibitors sharing structural similarities by comparing their global fingerprints on the eicosanoid pathway, facilitating the identification of the most potent compounds, including JNJ-B and JNJ-E. The highly efficient workflow that we have mapped out also allows for more accurate assessment of activity, using multiple data points. Simultaneous, comprehensive monitoring of multiple biochemical intermediary probes opens the door to an accelerated way to discover unknown pharmacology and targets, while providing confirmation of known targets and mechanisms as well.
The challenge in SM drug discovery lies in understanding their exceedingly complex, often unexpected pharmacology, adding difficulty to attempts to drive structure–activity relationships toward selectivity, or to exploit non-selectivity in the design of drugs that are capable of interacting with more than one molecular target, to achieve better efficacy. 28 To make things worse, traditional target-focused screening methods make it effectively impossible to learn about additional targets underlying either beneficial actions adding to efficacy or unwanted adverse effects. High-content, information-rich analysis of compounds’ effects on biochemical networks offers a potential solution to complement target-directed screening with limited readouts. Here, with the help of a high-throughput lipidomics platform, including rapid LC/MS-based data acquisition and automated data analysis, we have established differentiation at an unprecedented level of resolution for FLAP SM inhibitors by performing this highly granular molecular phenotypic screening, monitoring 122 eicosanoids in a short 6-min run. The highlighted multiprobe feature of this workflow enables the interrogation of polypharmacology to efficiently delineate the compound mode of action. Unlike ELISA, which is restricted by the availability of antibodies, our lipidomics panel covers a larger number of lipids, and it can be easily expanded to other lipids as well. For example, critical bioactive modulators such as 5-oxo-ETE, 20OH-LTB4, and 12epi-LTB4 can only be practically detected by MS. Among all lipid mediators, 5-oxo-ETE most strongly stimulates the migration of human eosinophils, 29 driving inflammatory responses in multiple tissues, including the lung. Even though 5-oxo-ETE is not as sensitive as LTB4 to inhibition within the 5-LO pathway, the relationship between its concentration levels and biological activity at the receptor target may be different from that of LTB4. Another benefit from broad reporter analyte coverage is the improvement of confidence in the mapping of drug pharmacology, because technical noise can be minimized by measuring several derivatives on the same pathway or multiple isomers of the same lipid.
We have recently reported on the application of this approach to examine additional mechanisms other than 5-LO inhibition contributing to the efficacy of zileuton, likely explaining why other compounds with higher 5-LO selectivity and potency failed in proof-of-concept clinical studies in asthma. 30 And, given that dosing of zileuton in clinical practice exerts only partial inhibition of leukotriene production and no drug in the market completely blocks the action of all 5-LO mediators. 31 it is necessary to assess changes of 5-oxo-ETE on compound treatment. We also show that JNJ-A displays much higher potency compared to JNJ-E in blocking the production of COX pathway lipids, although they behave similarly as silencers of LOX pathway metabolites.
Not surprisingly, subjects tested in our study exhibited donor-to-donor variations in terms of responses to compound treatment. Furthermore, we identified four additional donors in whom the production of eicosanoid lipids was induced at the lowest concentration (0.01 µM) for all compounds tested, relative to calcium ionophore stimulation (data not shown). We do not know whether this effect stems from either technical pitfalls or biological variability inherent to human study subjects. Rather than a liability, these differences suggest the utility of this platform for patient screening in clinical settings.
In conclusion, we have successfully used a lipidomics platform to globally profile the differential eicosanoid modulation of internally synthesized FLAP inhibitors and, as a result, identified potent compounds with high selectivity. This multiprobe approach not only enables the differentiation of compounds from within chemical series but also reveals distinct pharmacology inherent to different donors on same treatment. This high-throughput and cost-effective methodology opens a door to more comprehensive screening in early drug discovery, which may eventually replace the conventional single-target and single-readout assays.
Footnotes
Acknowledgements
We thank Drs. Dashyant Dhanak, Tadimeti Rao, and Anne Fourie from Janssen R&D for many insightful comments. We also thank Dr. Pieter Dorrestein and Michael Meehan from University of California, San Diego for discussions, and members of the Janssen Immunology Discovery Team in La Jolla for support. MEM wishes to dedicate this article to Dr. Anthony P. D. W. Ford on his birthday.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
