Abstract
Histone deacetylase (HDAC) proteins are promising targets for cancer treatment, with several HDAC inhibitors used clinically as anticancer drugs. Most HDAC inhibitors nonspecifically interact with all or many of the 11 HDAC isoforms. Isoform-selective HDAC inhibitors would be useful tools to dissect the individual functions of HDAC proteins in cancer formation, in addition to potentially displaying effective anticancer properties. We report here a robust HDAC activity assay for screening selective HDAC inhibitors, which is inspired by the traditional enzyme-linked immunosorbent assay (ELISA). The key feature of the ELISA-based HDAC activity assay is use of mammalian cell–derived HDAC isoforms instead of recombinant proteins. Importantly, the assay was validated with several known HDAC inhibitors. The ELISA-based HDAC activity assay will facilitate the characterization of isoform-selective HDAC inhibitors against mammalian cell–derived HDAC proteins, which will enhance HDAC-centered cancer research and provide a foundation for anticancer drug development.
Keywords
Introduction
Transcription of genomic DNA is tightly regulated to control the expression of genes for proper cell function. One level of gene expression regulation involves acetylation and deacetylation, which is a dynamic protein posttranslational modification. Histone acetyltransferase proteins catalyze the acetylation of nucleosomal histones, which can enhance gene transcription; histone deacetylase (HDAC) proteins reverse the acetylation of histones, which can lead to reduced gene transcription. 1 Overexpression of HDAC proteins can lead to unregulated transcription, which is observed in some cancer cells. 2 Because of their role in cancer, HDAC proteins have emerged as anticancer drug targets. 3
Eighteen HDAC proteins are known, which are divided into four subfamilies based on size, cellular localization, catalytic mechanism, and homology to yeast HDAC proteins. 4 Class I HDAC proteins include HDAC1, HDAC2, HDAC3, and HDAC8, whereas HDAC4, HDAC5, HDAC6, HDAC7, HDAC9, and HDAC10 are grouped as class II. Based on phylogenetic analysis, class IV includes only HDAC11 because of its similarity to both class I and II. The seven class III sirtuin proteins require NAD+ for histone deacetylation. The 11 class I, II, and IV HDAC proteins are metal dependent and are the targets of the inhibitors discussed here. 4
Among the metal-dependent HDAC isoforms, class I proteins are associated with cell-cycle progression, metastasis, and apoptosis, which are altered in cancers. As illustrative examples, increased expression of HDAC1 was observed in gastric, 5 colon, 6 prostrate, 7 and breast cancer, 8 while HDAC2 was overexpressed in gastric 9 and colorectal cancer. 10 Among the class II proteins, HDAC6 is overexpressed in oral squamous cell carcinoma and breast cancer tumors. 11 Unlike other HDAC isoforms, HDAC6 plays important roles in multiple cancer-relevant biological processes that are not epigenetic in nature.
Given their strong association with cancer-related events, multiple HDAC inhibitors have been developed as therapeutics. 3 SAHA (Vorinostat, Zolinza; Fig. 1 ), romidepsin (FK-228, Istodax; Fig. 1 ), and belinostat (PXD101, Beleodaq) are HDAC inhibitors approved by the Food and Drug Administration (FDA) for the treatment of T-cell lymphoma. Recently, panobinostat (LBH-589, Farydak) was approved for treatment of multiple myeloma. HDAC inhibitors of four structural types are in development, which include hydroxamic acids, benzamides, cyclic peptides, and short chain fatty acids ( Fig. 1 ). SAHA, belinostat, and panobinostat are hydroxamate-containing inhibitors. The benzamide entinostat (MS-275; Fig. 1 ) is currently in clinical trials against both solid and hematological cancers. FDA-approved romidepsin is a cyclic peptide ( Fig. 1 ). Finally, short-chain fatty acid valproic acid ( Fig. 1 ) displays IC50 values in the millimolar range against HDAC proteins but shows promising results in combination treatments with a variety of tumors.
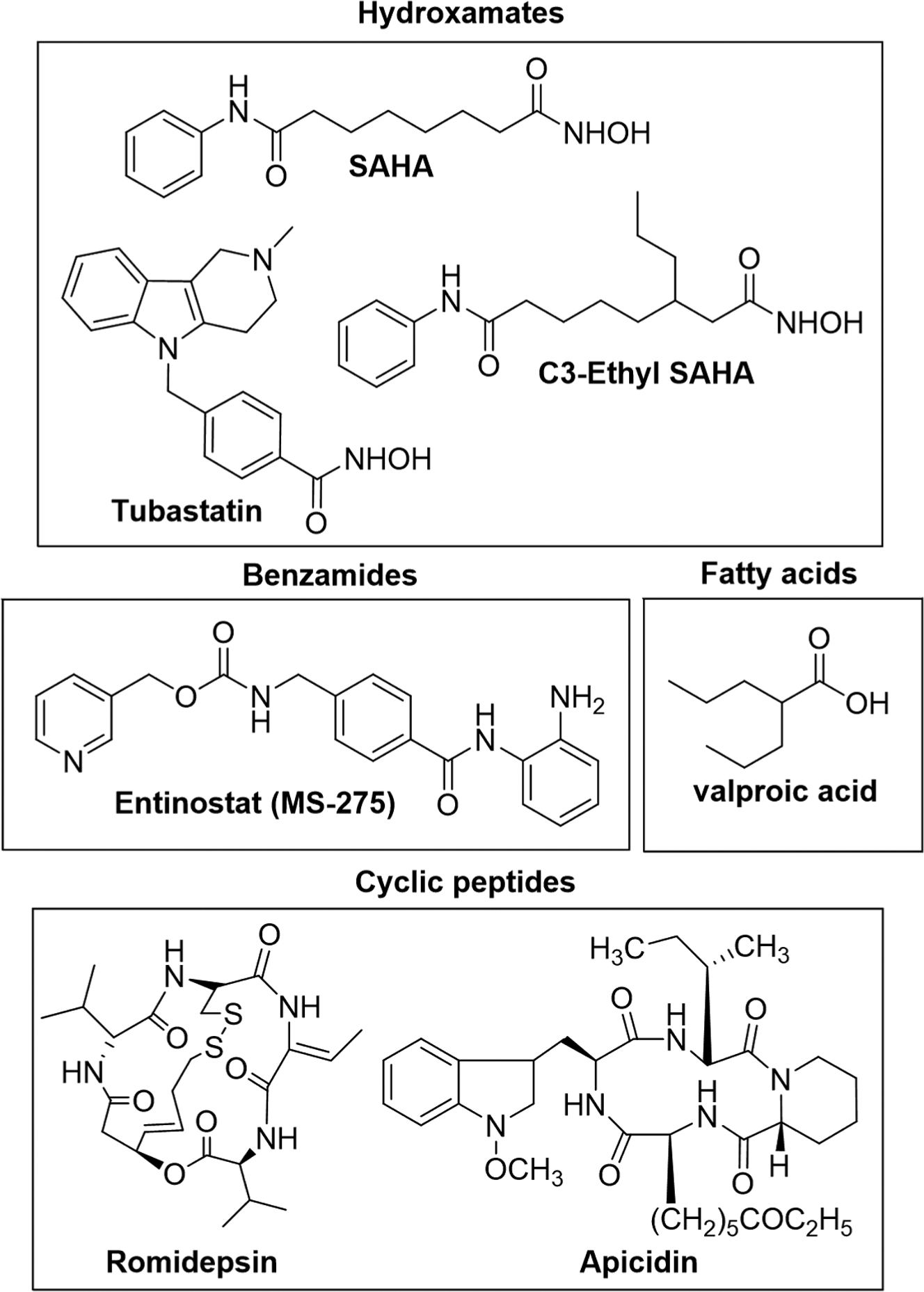
Chemical structures of histone deacetylase inhibitors discussed in the text.
Despite the successes of HDAC inhibitors in cancer treatment, they manifest side effects in patients, including gastrointestinal symptoms, bone marrow suppression, fatigue, cardiac arrhythmia, nausea, dehydration, thrombocytopenia, and anorexia. 12 One hypothesis accounting for the observed side effects is the poor selectivity of the clinically tested HDAC inhibitors; most compounds inhibit all or many of the 11 HDAC isoforms. 13 For example, SAHA, belinostat, and valproic acid are pan inhibitors, affecting all isoforms roughly similarly. 14 Romidepsin displays class I selectivity, inhibiting HDAC1 and 2 more potently than HDAC4 or 6. 15 Panobinostat and entinostat display class I and HDAC9 selectivity, targeting HDAC1, 2, 3, and 9. 14
Isoform-selective HDAC inhibitors, which target only a single HDAC isoform, have the potential to be more effective anticancer drugs. Consistent with this hypothesis, several clinical trials showed reduced toxicity with selective HDAC inhibitors relative to the pan-HDAC inhibitors. For example, hematological toxicities, anemia, and QT prolongation were absent or reduced in clinical trials with class I selective inhibitors entinostat and mocetinostat,16,17 as compared with pan inhibitors such as SAHA. 18 In addition to their possible use as more effective clinical drugs, isoform-selective HDAC inhibitors would be useful chemical tools to study HDAC cell biology.
Although isoform-selective inhibitors would be valuable on the bench top and in the clinic, only a few highly selective compounds have been identified. As illustrative examples, tubastatin ( Fig. 1 ) is a hydroxamate that displays 57- to 2000-fold selectivity for HDAC6 compared with the other isoforms. 19 Apicidin ( Fig. 1 ) is a cyclic peptide inhibitor that maintains 17- to 230-fold selectivity toward class I HDAC2, 3, and 8. 14 MS-275 ( Fig. 1 ) is a benzamide inhibitor that shows 4- to 400-fold selectivity for class I HDAC1, 2, and 3 over the other isoforms.14,20 Whereas some selective compounds have been reported, identifying additional strictly isoform-selective HDAC inhibitors remains a challenge.
One obstacle to identifying isoform selective inhibitors lies with the current compound-screening technologies. The most widely reported in vitro deacetylation assay monitors the fluorescent signal from HDAC-mediated degradation of the Fluor-De-Lys peptide substrate (Enzo Life Sciences). Although the fluorescent assay is robust and straightforward, selectivity screening typically involves use of purified, recombinant HDAC isoforms from baculovirus overexpression systems. 14 To test mammalian-cell derived HDAC proteins, immunoprecipitation of overexpressed isoforms from cell extract has been reported but with typically 30% to 60% error.21,22 Because of the significant error and tediousness of immunoprecipitation, many selective compounds have only been tested against baculovirus-derived isoforms, including tubastatin. 19 However, inconsistencies are seen when comparing inhibition data with the baculovirus- and mammalian cell–derived HDAC isoforms. For example, apicidin showed strong potency against mammalian cell–derived HDAC1 (IC50 = 23 nM) 22 but varied potency against baculovirus-expressed HDAC1 (IC50 >10,000 nM or 0.7 nM in two different reports).14,23 Although the disparities in these studies may be due technical difference, the similar assay formats suggest that the source of the HDAC isoforms can also influence the screening results.
To create a robust assay relying on mammalian cell–derived HDAC isoforms, we report here an enzyme-linked immunosorbent assay (ELISA)–based HDAC activity assay. Screening of several known HDAC inhibitors with different selectivity profiles validated the use of assay for inhibitor screening. These results suggest that the ELISA-based assay will be a useful tool toward characterization of isoform-selective HDAC inhibitors against mammalian cell–derived HDAC isoforms.
Materials and Methods
HeLa Cell Lysates
HeLa-S3 cells (1 × 109; Biovest Inc.) were lysed in lysis buffer (10 mL; 50 mM Tris-HCl, pH 8.0, 10% glycerol, 150 mM NaCl, 0.5% Triton X-100) containing Calbiochem protease inhibitor cocktail set V (100 µL of 100×) with rotation at 4 °C for 30 min. Cell debris was removed by centrifugation at 12,000 rpm at 4 °C for 30 min. Protein content (mg/mL) of the supernatant was determined using Bio-Rad protein assay (BioRad, Bradford reagent).
ELISA-Based HDAC Activity Assay
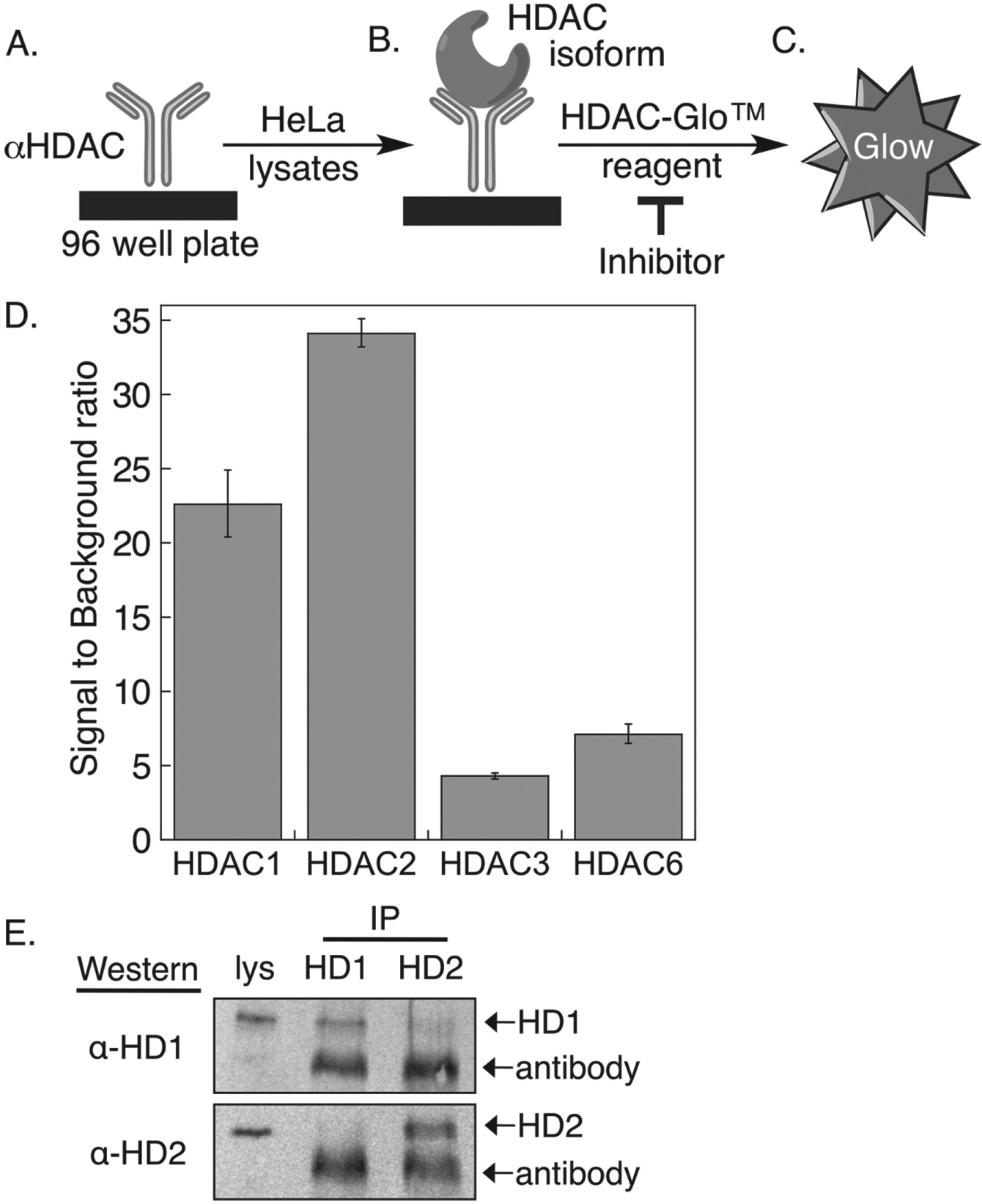
Individual wells of a high-binding polystyrene 96-well white opaque plate (Thermo Scientific) were incubated in binding buffer (100 µL; 0.2M carbonate/0.2 M bicarbonate buffer, pH 9.4) with primary HDAC1 antibody (Sigma Aldrich, H3284, 100 µL of 10 µg/mL), primary HDAC2 antibody (Sigma Aldrich, H3159, 100 µL of 10 µg/mL), or primary HDAC6 antibody (Sigma Aldrich, SAB1404771, 100 µL of 2 µg/mL) with rocking (3 rpm) for 1 h at room temperature. For HDAC3 with MS-275 only, primary HDAC3 antibody (Sigma Aldrich, H3034, 100 µL of 25 µg/mL) was used. For HDAC3 and all other compounds, wells of a secondary antibody-coated 96-well white opaque plate (G-Biosciences) were incubated with primary HDAC3 antibody (Sigma Aldrich, H3034, 100 µL of 1 µg/mL) in TBST buffer (50 mM Tris-HCl, 150 mM NaCl, pH 7.4, 0.05% [v/v] Tween-20) containing 0.1% (w/v) bovine serum albumin (Jackson Immunoresearch) at 4 °C overnight without rocking. For all reactions, unbound antibody was removed by washing quickly three times with TBST buffer (300 µL), followed by a fourth wash under the same conditions but with 5 min incubation and gentle rocking (3 rpm) at room temperature. In the case of high-binding polystyrene plates containing HDAC1, 2, 3, or 6 antibodies, the unbound regions of the well were blocked with 5% nonfat dry milk in TBST buffer (300 µL) for 1 h at room temperature with gentle rocking (3 rpm). Because the secondary antibody-coated plates containing HDAC3 were preblocked by the manufacturer, no additional blocking step was included. To affix HDAC enzyme to the plate, HeLa cell lysates (100 µL of 100 µg/mL for HDAC1 and 2; 100 µL of 1 mg/mL for HDAC6; 100 µL of 1 mg/mL for HDAC3 with MS-275; 100 µL of 100 µg/mL for HDAC3 and all other compounds) in TBST buffer containing 0.1 % (w/v) nonfat dry milk were added to each well and incubated for 1 h at 4°C without rocking, followed by washing with TBST, as described previously. Finally, deacetylase activity was measured using the HDAC-Glo assay kit (Promega) as per the manufacturer’s protocol. Specifically, the HDAC-Glo substrate (1 mL) and developer (1 µL) were first premixed to make the HDAC-Glo reagent mixture, which was stored at −20 °C in aliquots until use. Then, to monitor deacetylase activity, HDAC-Glo buffer (25 µL) and the HDAC-Glo reagent mixture (25 µL) were combined and added to each well (50 µL total volume) and incubated for 30 to 45 min at room temperature without rocking. The deacetylase activity was measured as a luminescent signal using a GeniosPlus Fluorimeter (Tecan) at optimal gain. For each reaction, the luminescent signal was background corrected with the signal from a negative control reaction where the HDAC antibody was absent in the initial antibody binding step. Signal to background ratios are reported in Figure 2D , with the mean and standard error of three independent trials shown.
(
Inhibitor Testing
For inhibitor screening, all steps were identical to the ELISA-based HDAC activity assay protocol described above, up to immunocapture of the HDAC enzyme. Inhibitors in DMSO (1 µL) were added to the HDAC-Glo buffer (24 µL) and then added to the plate after immunocapture of the HDAC isoform and incubated for 15 min at room temperature without rocking. For MS-275 only, a 3-h incubation at room temperature was used with HDAC1, 2, and 3 due to slow association kinetics.
24
A control reaction without inhibitor was also included, which contained DMSO (1 µL) in HDAC-Glo buffer (24 µL). After incubation of the inhibitor with the affixed HDAC isoform, the ELISA-based HDAC activity assay protocol was followed as described above, with addition of HDAC-Glo reagent alone (25 µL). The concentrations of inhibitors reported in the single-dose screen (
Fig. 3
) and dose-dependent studies (
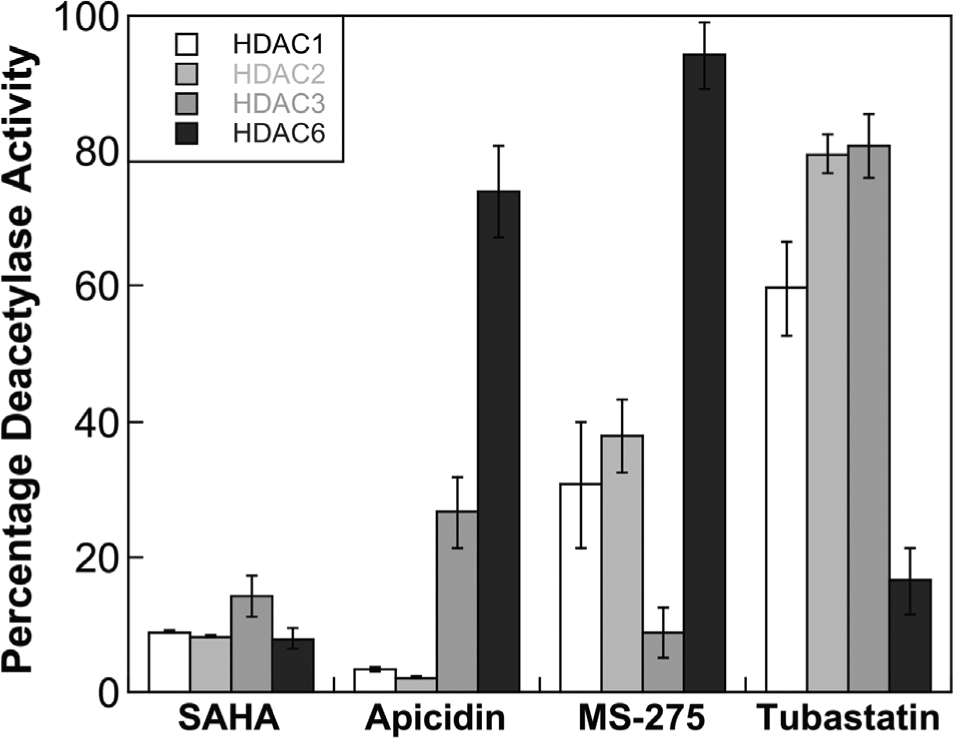
Isoform selectivity screen of SAHA, apicidin, MS-275, and tubastatin against HDAC1, HDAC2, HDAC3, and HDAC6 at a single concentration of 1 µM for SAHA, apicidin, tubastatin, and 4.8 µM for MS-275. The plot displays the mean percentage deacetylase activity remaining compared with an uninhibited reaction from three independent trials with standard errors (
Immunoprecipitation of HDAC1 and HDAC2
The same procedure for the assay was performed for HDAC1 and 2, except that incubation of the well with HDAC1 or HDAC2 primary antibody was done overnight at 4 °C and the Hela cell lysate concentration was 2 mg/mL. After capture of the HDAC isoform, the bound protein and antibodies were eluted from each well with 2× sodium dodecyl sulfate (SDS) loading buffer (20 µL; 200 mM Tris-Cl [pH 6.8], 400 mM DTT, 8% SDS, 0.4% bromophenol blue, 40% glycerol) at 90 °C for 5 min. The combined eluate from eight wells was concentrated under reduced pressure for 30 min at 50 °C. For gel analysis, eluted proteins were separated using 10% SDS polyacrylamide gel electrophoresis (SDS-PAGE) gel, followed by transfer to PVDF membrane (Immobilon P, Millipore) and Western blot analysis with primary HDAC1 antibody (Sigma Aldrich, H3284, 1:10,000) or HDAC2 antibody (Sigma Aldrich, H3159, 1:10000), along with a secondary anti-rabbit IgG, HRP-linked antibody (Cell Signaling, 7074, 7:10,000). The chemiluminescence substrate (SuperSignal West Dura Extended Duration Substrate) was incubated with the membrane for 3 min, followed by detection of the membrane for chemiluminescent signal using a Typhoon 9210 scanner (Amersham Biosciences). HeLa cell lysates (4.5 µg total protein) were loaded on the gel as a control ( Fig. 2E ).
Results
Design of the HDAC Activity Assay
The HDAC activity assay is modeled after the classic ELISA, in which antibodies are used to immobilize proteins from mammalian cell extract onto a plate. In the ELISA-based HDAC activity assay, polystyrene high-binding or secondary antibody–coated 96-well plates were incubated with a primary HDAC antibody ( Fig. 2A ), followed by HeLa cell lysates to immunoprecipitate the respective HDAC isoform ( Fig. 2B ). Once the desired HDAC isoform was affixed onto the plate, deacetylase activity was tested initially using commercially available Fluor-de-Lys substrate and developer. Unfortunately, the fluorescent substrate was not sensitive enough to detect the activity of the immunoprecipitated HDAC enzyme (data not shown). To enhance sensitivity, the commercially available HDAC-Glo I/II assay, 25 which is roughly 85-fold more sensitive than the Fluor-De-Lys assay (data not shown), was used instead. The HDAC-Glo assay uses a proluminogenic substrate, where luciferin is attached C-terminal to acetyl-lysine. Deacetylation and subsequent trypsin digestion of the HDAC-Glo substrate led to release of luciferin, which produced a highly stable and sensitive chemiluminescent signal after reaction with luciferase enzyme ( Fig. 2C ). Screening of HDAC inhibitors involved incubation of compounds with the affixed HDAC protein and monitoring the deacetylase activity remaining in the form of reduced chemiluminescent signal.
To establish the assay, initial experiments focused on HDAC1, HDAC2, HDAC3, and HDAC6 due to their role in various cellular processes and high expression levels in many cancers.5–11 Primary HDAC antibodies recognizing each of the isoforms were immobilized to different wells of a plate, followed by incubation with HeLa cell lysates to affix the respective HDAC isoform ( Fig. 2 ). The HDAC-Glo reagent was then added to monitor the deacetylase activity of the immunoprecipitated HDAC isoform. As a control, a reaction without primary antibody was included to assess background noise due to nonspecific binding of HDAC activity in the well. The signal (reaction with antibody) to background (reaction without antibody) ratios for HDAC1, HDAC2, HDAC3, and HDAC6 were 23 ± 2, 34 ± 1, 4.5 ± 0.2, and 7.2 ± 0.7, respectively ( Fig. 2D ). Notably, HDAC1 and HDAC2 showed the highest signal to background ratio, suggesting that they are highly active in HeLa cell lysates.
Critical parameters of a robust assay are signal dynamic range and variability in the signal. These parameters are assessed using the Z′ factor.
26
The Z′ factor for a robust assays should be within the range of 0.5 to 1.0. With the ELISA-based HDAC activity assay, the Z′ factors for HDAC1, HDAC2, HDAC3, and HDAC6 were 0.69, 0.70, 0.67, and 0.75, respectively. In addition to the Z′ factor, the coefficient of variation (CV) is a dimensionless parameter that defines the extent of variability.
27
A small variability in the data (CV < 10%) suggests a robust assay. The calculated CVs with HDAC1, HDAC2, HDAC3, and HDAC6 were 9.7%, 10%, 9.2%, and 5.2%. It is also noted that the activity of each immunocaptured HDAC isoform fell in the linear range of the assay (
Immunocapture of HDAC1 and HDAC2
The HDAC isoforms can interact in multiprotein complexes in cellulo. The most reported intra-isoform interaction is observed between HDAC1 and HDAC2, which are found in multiple intracellular mammalian complexes. 28 Given the reported interaction between HDAC1 and 2, a concern is that the HDAC1 and HDAC2 activities observed in the assay are due to immunoprecipitation of both isoforms. Although the plate-based assay showed statistically different activities between HDAC1 and HDAC2 ( Fig. 2D ), which suggests that only one isoform is immunocaptured under the assay conditions, an on-plate co-immunoprecipitation and Western blot analysis was performed. Because the quantity of isoform affixed to each well in the assay was too low to be detected by Western blotting (data not shown), 20-fold higher lysate concentrations were used (2 mg/mL instead of 0.1 mg/mL) and immunoprecipitates from eight wells were combined for the analysis. Western blot analysis of the immunprecipitates showed minimal interaction between HDAC1 and HDAC2, even under the high lysate conditions ( Fig. 2E ). HDAC2 is absent in the immunoprecipitates of HDAC1 ( Fig. 2E , lane 2, bottom gel image), whereas only faint levels of HDAC1 were observed in the HDAC2 immunoprecipitates ( Fig. 2E , lane 3, top gel image). Given that the lysate concentration used in the assay is 20-fold lower, these Western blot studies suggest that the activities observed in the assay are due to the immunocapture of a single HDAC isoform.
Screening of Known HDAC Inhibitors
With the optimized HDAC activity assay in hand, we tested several known HDAC inhibitors. We choose three inhibitors with different selectivity profiles. SAHA ( Fig. 1 ) is a pan-inhibitor and shows potency against all HDAC isoforms. 14 To represent selective inhibitors, apicidin ( Fig. 1 ) was chosen for its class I selectivity14,22; MS-275 ( Fig. 1 ) for its HDAC1, 2, and 3 selectivity14,20; and tubastatin ( Fig. 1 ) was selected due to its HDAC6 selectivity. 19
As an initial screen, SAHA, apicidin, MS-275, and tubastatin were incubated at a single concentration (4.8 µM for MS-275 and 1 µM for all others), with each individual HDAC isoform affixed on the plate. The remaining deacetylase activity of the HDAC isoforms was then determined by adding the HDAC-Glo reagents. As a control, a reaction without inhibitor was also included. The percentage deacetylase activity remaining was calculated by comparing the signals from inhibitor-containing reactions to the uninhibited control reaction. In accordance with the reported data, SAHA inhibited the four HDAC isoforms roughly equally at the 1 µM single concentration (
Fig. 3
;
In contrast to SAHA, the four selective HDAC inhibitors showed variable activity depending on the HDAC isoform. Apicidin and MS-275 preferentially inhibited class I HDAC1, 2, and 3 compared with the HDAC6 isoform (
Fig. 3
;
Dose-Dependent Inhibition
The next aim was to determine the IC50 values of the known inhibitors against mammalian cell-derived HDAC isoforms using the ELISA-based assay. In addition to SAHA, apicidin, MS-275, and tubastatin, we also tested a C3-ethyl SAHA analog ( Fig. 1 ) reported by our lab to have HDAC6 selectivity, although with µM potency. 29 Testing these known inhibitors will assess the range of inhibitor potencies and selectivities compatible with the assay.
For dose dependence, various concentrations of each inhibitor were incubated with the individual immunoprecipitated HDAC isoforms before deacetylation was monitored using the HDAC-Glo reagents. Using the percentage remaining deacetylase activity at each small-molecule concentration, the IC50 values were determined. As expected, SAHA inhibited all four HDAC isoforms with similar potency (
Table 1
;
IC50 values of SAHA, apicidin, MS-275, tubastatin, and C3-ethyl SAHA with HDAC1, 2, 3, and 6 using the ELISA-based activity assay. a
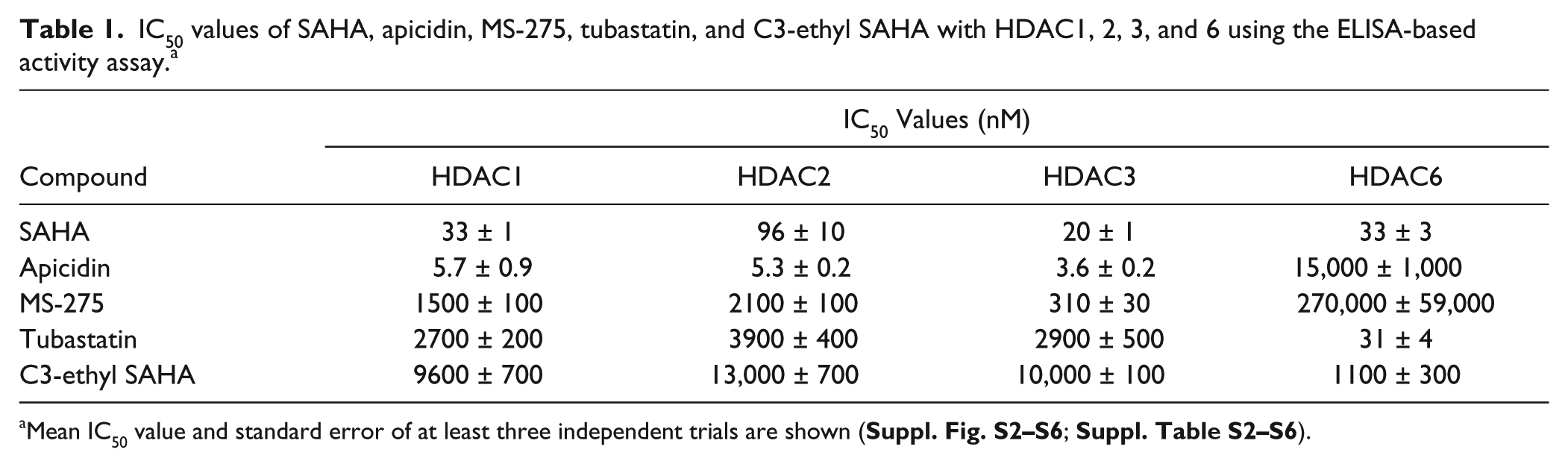
Mean IC50 value and standard error of at least three independent trials are shown (
Apicidin showed selectivity for class I HDACs over the HDAC6 isoform, with roughly 2600-fold selectivity for HDAC1, 2, and 3 compared with HDAC6 (
Table 1
;
MS-275 showed selectivity for HDAC1, 2, and 3 over HDAC6, with 135- to 871-fold selectivity (
Table 1
;
Tubastatin also showed the expected HDAC6 selectivity using the ELISA-based assay (
Table 1
;
Finally, the C3-ethyl SAHA derivative displayed at least ninefold HDAC6 selectivity over the class I HDAC isoforms using the ELISA-based assay (
Table 1
;
Discussion
We describe here an HDAC activity assay using mammalian cell–derived isoforms as an assay platform to test compounds for selectivity. The assay was initially validated using several known selective and pan inhibitors ( Table 1 ). With SAHA, MS-275, and apicidin, results with the ELISA-based assay were consistent with previously reported data with mammalian cell–derived proteins. In the case of apicidin, the ELISA-based assay was used to resolve a discrepancy in the literature where differing HDAC1 sensitivities were reported with baculovirus-derived proteins. The ELISA-based assay confirmed earlier data with mammalian cell proteins showing nM potency with apicidin against HDAC1, in addition to HDAC2 and 3. The potencies and selectivities of tubastatin and C3-ethyl SAHA were assessed for the first time against mammalian cell–derived isoforms. Both compounds showed HDAC6 selectivity, which is consistent with earlier testing with baculovirus-expressed proteins. However, the extent of their HDAC6 selectivities differed between mammalian- and baculovirus-derived proteins. It is noted that technical issues might also explain some of the discrepancy of quantitative selectivity data in the literature. But given that the substrates and assay formats of the prior reports were similar, the source of HDAC activity is also a critical factor in activity assays. In total, the ELISA-based HDAC activity assay will be helpful by allowing screening with mammalian cell–derived proteins to confirm and extend selectivity testing of HDAC inhibitors.
A significant feature of the ELISA-based HDAC activity assay is its excellent reproducibility compared with earlier screening methods using mammalian cell–derived HDAC isoforms. Inhibitor screening using bead-based immunoprecipitates of either endogenous or overexpressed isoforms from cell extract displayed typically 30% to 60% error.21,22 The high error rate using bead-based immunoprecipitates made screening of mammalian cell–derived HDAC isoforms impractical. HDAC inhibitor screening currently relies almost exclusively on baculovirus-expressed isoforms. 14 Given that the source of HDAC isoform activity can significantly affect potency and selectivity, the ability to robustly and reproducibly screen mammalian cell–derived HDAC isoforms makes the ELISA-based HDAC activity assay valuable for selective inhibitor characterization.
In development of the ELISA-based assay, the high-sensitivity signal from the HDAC-Glo substrate from Promega was required to overcome the small quantities of HDAC isoform captured in the assay. As with any enzyme-coupled assay system, reliance of the HDAC-Glo substrate on trypsin and luciferase enzyme activities and light production can lead to false-positive inhibition by a test compound because of trypsin or luciferase inhibition or light absorption. 30 Therefore, when screening for novel HDAC inhibitor compounds using the HDAC-Glo substrate and the ELISA-based assay, counterscreening inhibitors against trypsin and luciferase enzyme alone is recommended, as previously described. 30
In conclusion, we report here an ELISA-based assay to monitor the activity of individual HDAC isoforms from mammalian cell lysates. The robustness of the ELISA-based HDAC activity makes it suitable for future characterization of HDAC inhibitors against mammalian cell–derived isoforms. Importantly, the assay complements the widely used activity assays using baculovirus-expressed HDAC isoforms. Given that HDAC proteins are involved in myriad biological processes and disease states, the ELISA-based HDAC activity assay will embolden the research community to identify highly isoform-selective inhibitors for future drug development and HDAC-centered cell biology studies.
Footnotes
Acknowledgements
We thank the National Institutes of Health (GM067657) and Wayne State University for funding, the Lumigen Instrument Center at Wayne State University for NMR and MS instrumentation, and D. Nalawansha for helpful comments.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: National Institutes of Health (GM067657) and Wayne State University.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
