Abstract
Sphingomyelin (SM) metabolism deregulation was recently associated with cell metastasis and chemoresistance, and several pharmacological strategies targeting SM metabolism have emerged. The ceramide (Cer) generated in the endoplasmic reticulum (ER) is transferred to the Golgi apparatus to be transformed into SM. CERamide Transfer (CERT) protein is responsible for the nonvesicular trafficking of Cer to Golgi. Blocking the CERT-mediated ER-to-Golgi Cer transfer is an interesting antioncogenic therapeutic approach. Here, we developed a protein-lipid interaction assay for the identification of new CERT-Cer interaction inhibitors. Frequently used for protein-protein interaction by enzymatic and analyte dosage assays, homogeneous time-resolved fluorescence technology was adapted for the first time to a lipid-protein binding assay. This test was developed for high-throughput screening, and a library of 672 molecules was screened. Seven hits were identified, and their inhibitory effect quantified by EC50 measurements showed binding inhibition three orders of magnitude more potent than that of HPA12, the unique known CERT antagonist to date. Each compound was tested on an independent test, confirming its high affinity and pharmacological potential.
Keywords
Introduction
Dysregulated sphingolipid metabolism is widely associated with cell tumorigenicity, proliferation, metastasis, and chemoresistance. 1 Ceramide (Cer) is the backbone molecule of complex sphingolipids such as gangliosides and sphingomyelin (SM), an abundant component of the eukaryotic cells’ outer membrane. Cer was shown to act as a mediator of cell death, proliferation, and survival, whereas SM has been described as a source of fast Cer release, compared with its de novo biosynthesis, through sphingomyelinase activity. Cer is synthesized in the endoplasmic reticulum (ER) upon action of one of the six Cer synthases and the complete synthesis of SM, resulting from the addition of a phosphocholine head group to Cer, takes place in the Golgi apparatus.
Sphingolipids were shown to be associated with cell membrane structural and functional properties, influencing drug influx, sequestration, and efflux from cells. 2 Variations in sphingolipid membrane composition were observed between anticancer drug-resistant cells compared with the drug-sensitive ones, and SM level increases were shown to be associated with doxorubicin resistances in P388, T47D, and MCF-7/ADR cells.3,4 Strategies to interfere with sphingolipid metabolism were largely studied. The inhibition of various enzymatic targets, such as sphingomyelin synthase (SMSs) and Glc-ceramide synthase, have already been proposed as potential anticancer approaches. 5
The CERamide Transfer (CERT) protein is specifically responsible for Cer trafficking from ER-to-Golgi along de novo SM biosynthesis,
6
suggesting that inhibition of CERT-mediated Cer transfer might be a promising strategy for anticancer therapy.1,7 Notably, it was shown that CERT depletion leads to chemotherapy benefit and mediates cytotoxic and cancer cell death through autophagy induction.
8
Because
CERT contains several functional domains, among which is a
We thus sought a more practical and sensitive CERT binding assay that could be amenable to miniaturization and automation. We focused our attention on the fluorescence resonance energy transfer (FRET) detection mode. Because of specific spectral properties, FRET allows homogeneous assays to run without separation steps such as centrifugation, washing, filtration, or magnetic partitioning. 12 Moreover, the fluorescence background produced by chemical samples, buffers, or proteins, generally presenting lifetimes in the nanosecond range, could be eliminated using time-resolved methodology (TR-FRET). Homogeneous time-resolved fluorescence (HTRF 13 ), an homogeneous-phase TR-FRET technology, has been developed for high-throughput screening (HTS) 14 and was already successfully applied to screen diverse compound libraries.15,16
Here, TR-FRET was applied for the first time to monitor a protein-lipid, namely, START-Cer, interaction. Thanks to a specifically designed ω-biotinylated Cer (BioCer) probe, 11 we were able to measure a TR-FRET between an europium kryptate linked to an anti-histidine antibody, as the donor, and d2 carried by streptavidine, as the acceptor. The performance of this miniaturized assay was carefully analyzed and shown to allow large screening experiments. A library of 672 compounds was screened, and seven hits were identified.
Materials and Methods
Products
Streptavidin-d2 conjugate (product No. 610SADLA) and europium kryptate–labeled monoclonal anti-histidine antibody (product No. 61HISKLA) were obtained from Cisbio Bioassays (Codolet, France). Human COL4A3ABP recombinant histidine-tagged protein fragment (START; product No. ab95897) was obtained from Abcam. Biotinylated Cer probe (BioCer, or
Fluorescence Intensity Binding Assay
START was dissolved in Tris-buffered saline solution (TBS) to 1 µM concentration. For negative control, START was not added in this TBS solution. Of the fluorescent Cer probe, 1 µM of was added to the tube, and the mixture was incubated at 37 °C for 30 min. Then TALON metal affinity resin (30 mL of 50% [v/v] preequilibrated with wash buffer) was added to the mixture and incubated for 10 min at room temperature. After centrifugation and washes with TBS (75 mL) containing increasing imidazole concentration, the supernatant was collected as the unbound fraction. To liberate the protein-bound fraction to the TALON resin, the resin was suspended in 250 mM imidazole and incubated for 5 min at room temperature. After centrifugation, the supernatant was collected as the bound fraction. For more details, see Combemale et al. 11
Optimized HTRF Assay
The assay was carried out in a final volume of 20 µL. Six microliters of the START protein solution to a final concentration of 1 µM and 2 µL of 5% of DMSO (or compounds diluted in 5% of DMSO) were incubated in Corning 384-well, low-volume, white, round-bottom polystyrene NBS microplates (product No. 3673). After incubation for 30 min at 37 °C, 2 µL of BioCer solution was added to a final concentration of 1 µM. Five microliters of streptavidin-d2 conjugate and 5 µL of europium kryptate–labeled monoclonal anti-histidine antibody were added in detection buffer (20 mM of HEPES buffer pH 8.5, 800 mM of potassium fluoride, 1 mM of bovine serum albumin in H2O). For the europium kryptate–labeled monoclonal anti-histidine antibody, 2.1 ng per well was added and 50 ng was added for streptavidine-d2. After 18 h of incubation at room temperature, the time-resolved fluorescence was measured using an Envision plate reader (Perkin Elmer; λex = 320 nm, λem = 590 and 665 nm; 100 µs delay time). The donor fluorescence emission was measured at 590 nm instead of 620 nm in accordance with Envision apparatus and Cisbio recommendation. All modifications of this standardized protocol are noted in the legend of the figures.
HTRF Data Handling
HTRF Ratio and FRET Signal
The HTRF ratio readout was calculated as a two-wavelength signal ratio: (665 nm/590 nm) × 104 according to manufacturer recommendations. FRET signal was defined as (HTRF compound ratio – HTRF negative ratio)/HTRF blank ratio. The HTRF negative ratio was (665 nm/590 nm) × 104 of wells containing streptavidin-d2 conjugate and europium kryptate–labeled monoclonal anti-histidine antibody. The HTRF blank ratio was (665 nm/590 nm) × 104 of wells containing europium kryptate–labeled monoclonal anti-histidine antibody.
P/N Signal
The positive control is the maximum HTRF ratio with 1% DMSO. The negative control is the HTRF negative ratio.
Percentage of Inhibition
FRET-positive controls were performed by dispensing DMSO 5%. This maximal FRET signal value obtained was set to 0% inhibition. The FRET negative ratio was set to 100% of inhibition.
The FRET signal calculation was performed as follows:
Percentage of inhibition = 100 − (100* FRET compound) × FRET with 1% of DMSO. All curve fittings, including the one-phase exponential association (defined by the equation: Y = Ymax* (1 − exp (−k*X)), were performed with Prism 3.0 (GraphPad Software, San Diego, CA).
Z′ Factor
This factor was calculated with 32 assays for each control as defined by Zhang et al. 21
Data Reproducibility
Except for the screening experiment itself, all experiments were performed in duplicate or triplicate.
Screening of Chemical Compounds of the Chimiothèque Nationale Essentielle
The Chimiothèque Nationale Essentielle (CNE; an Essential Compound Library) is a subset of 640 compounds, selected according to molecular diversity criteria, representative of the CN (French National Chemical Library; “Chimiothèque Nationale”) and recently used in HTS notably by Ruggiu et al. 22 All structures present in this synthetic products and natural compounds database are accessible on the Website http://chimiotheque-nationale.enscm.fr. CNE was conditioned into 96-well microplates. Compounds were diluted and split in two plates of 384 wells to a final concentration of 50 µM. Thirty-two compounds known or not to inhibit the START-Cer interaction were tested in the same condition as CNE. Two microliters of compound in 5% of DMSO were added to a final concentration of 5 µM in 1% of DMSO.
Results
CERT mediates the nonvesicular ER-to-Golgi trafficking of Cer synthesized de novo at the ER and transferred to the
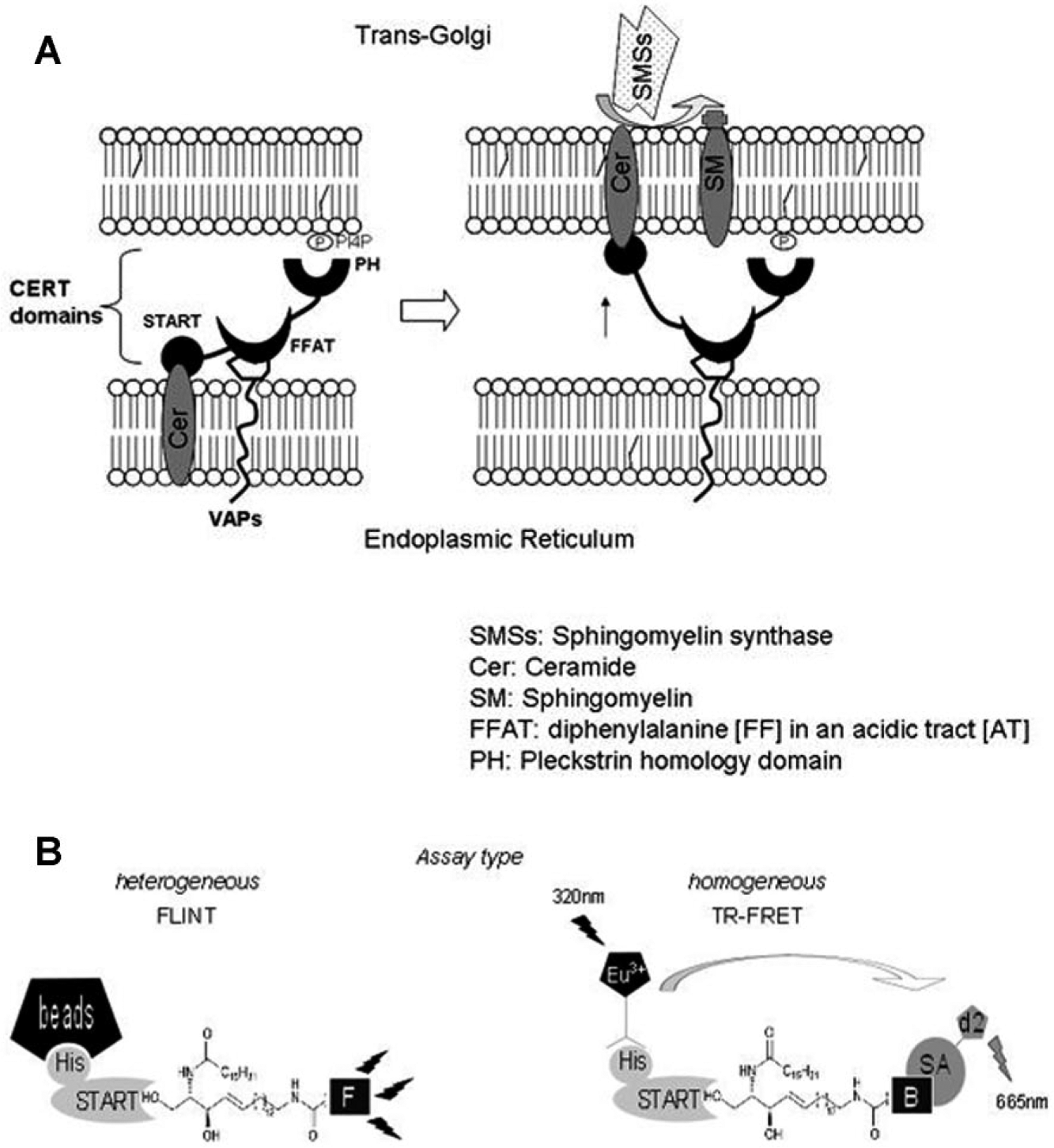
(
Our objective was thus to develop a miniaturized test to assess the START-Cer interaction on the basis of the homogeneous HTRF assay methodology ( Fig. 1B , right).
Development of an HTRF Assay to Measure START-Mediated Cer Binding
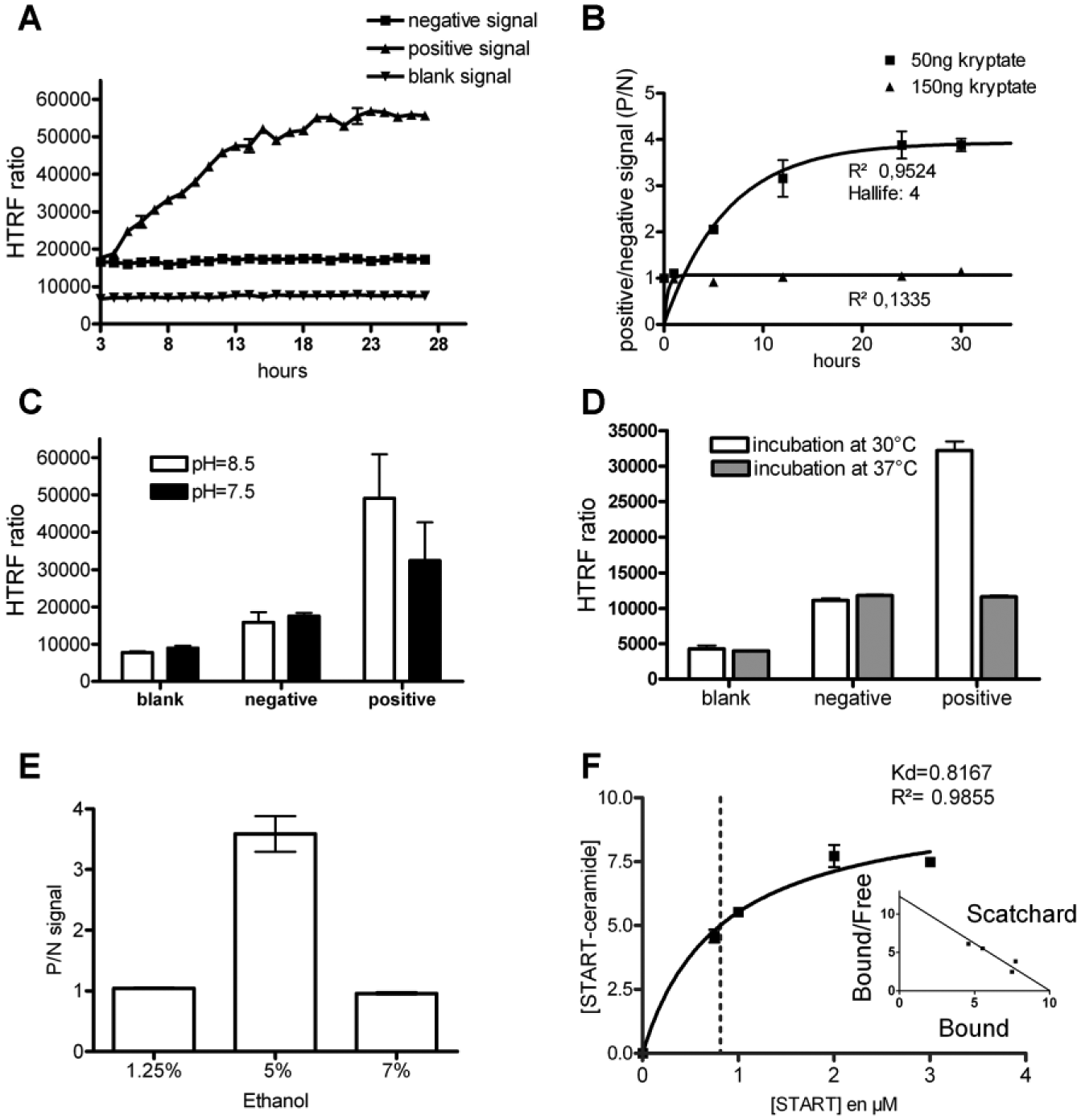
For the FLINT, a 1 µM fluorescent Cer probe solution was incubated with 2 µM of histidine-tagged START protein for 30 min. After resin addition, washing, centrifugations, and elution, fluorescence was measured in the bound fraction ( Fig. 2A ). For the HTRF assay, we started with the same START and biotinylated Cer probe (BioCer) concentrations. Kinetic analysis of the europium kryptate–START/Cer–streptavidin-d2 complex formation showed that a significant HTRF ratio was obtained after approximately 3 h of incubation ( Fig. 2B ). Time-course analysis showed that the HTRF ratio increase was dependent on time but not the increase of fluorescent background signal (as blank or negative signal; Fig. 3A ).
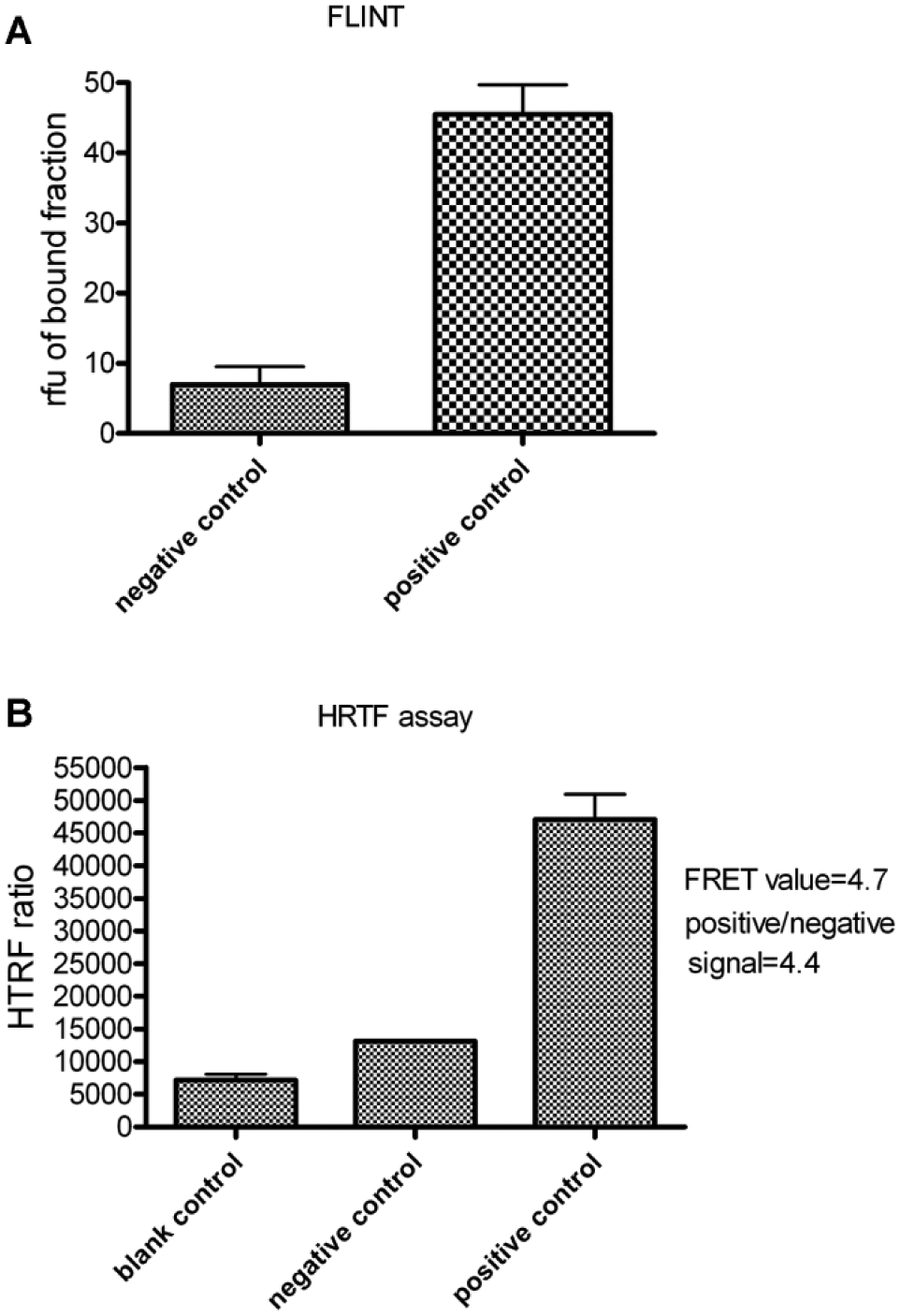
(
(
Various amounts (i.e., 50 ng or 150 ng per well) of europium kryptate–labeled anti-histidine antibody were tested (
Fig. 3B
). At 50 ng per well, kinetic analysis showed a good fit with a one-phase association profile (
Ligand binding depends partially on electrostatic interactions; therefore, pH variations can greatly influence binding parameters by directly altering the ionization state of chemical groups interacting with the ligand. The protein-lipid interaction could be influenced by pH variations as well. We thus investigated the effect of decreasing pH ( Fig. 3C ). The positive HTRF ratio at pH 8.5 is higher compared with pH 7.5. We also controlled the influence of temperature on the assay ( Fig. 3D ) and observed that incubation at 37 °C suppressed the signal compared with 30 °C. This loss of FRET signal at a higher temperature could be associated with faster reagent degradation after this unusually long incubation time. As Cer is dissolved in ethanol, we investigated the effect of ethanol concentration on FRET signal ( Fig. 3E ) and showed that binding measurements tolerates only a narrow window of alcohol concentration.
Finally, before automation, we investigated the binding kinetic parameters. As the curve obtained with 50 ng of kryptate (
Fig. 3B
) described the same reaction scheme as the protein-protein interaction (fitting of the one-phase association curve (
Assay Calibration and Robustness
Experiments performed with HPA12—a competitive inhibitor of a START-Cer interaction 18 ( Fig. 4A )—showed a time- and dose-dependent inhibitory effect of HPA12, confirming the specificity of the assay.
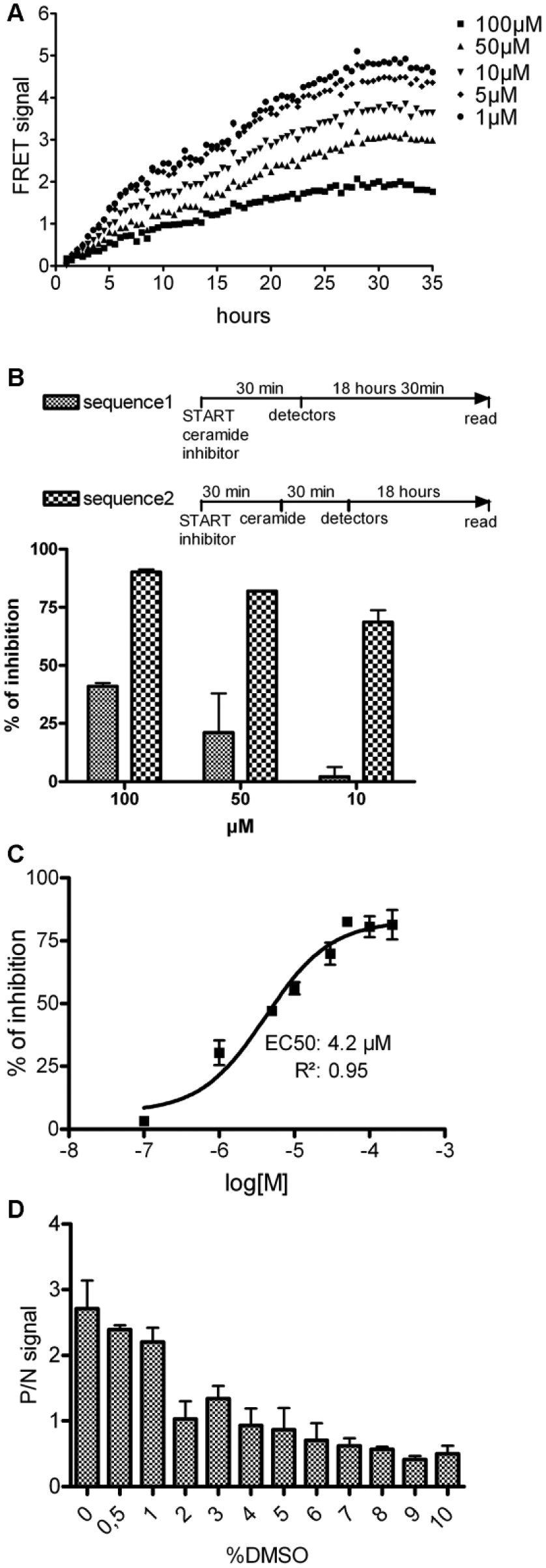
(
As previously observed, the kinetic of the START-Cer interaction is slow. Thus, to favor the identification of inhibitors, we optimized the sensitivity of the assay with a preincubation step (
Fig. 4B
). In sequence 1, START is incubated with the BioCer probe and the competitor. In sequence 2, START is incubated for 30 min at 37 °C with the competitor, and the BioCer probe is added. A higher sensibility to HPA12 inhibition is observed with sequence 2 compared with sequence 1. The EC50 of HPA12, applying sequence 2, was found to be 4.2 µM with the sigmoid dose-response algorithm (
To evaluate the ability of the assay to accurately detect START-Cer binding competitors among a series of molecules, we measured the well-established Z′ factor. 21 As observed in Figure 5A , Z′ factor values were systematically superior to 0.5 after 6 h of treatment, indicating that our assay can robustly detect inhibitors in large compound collections.
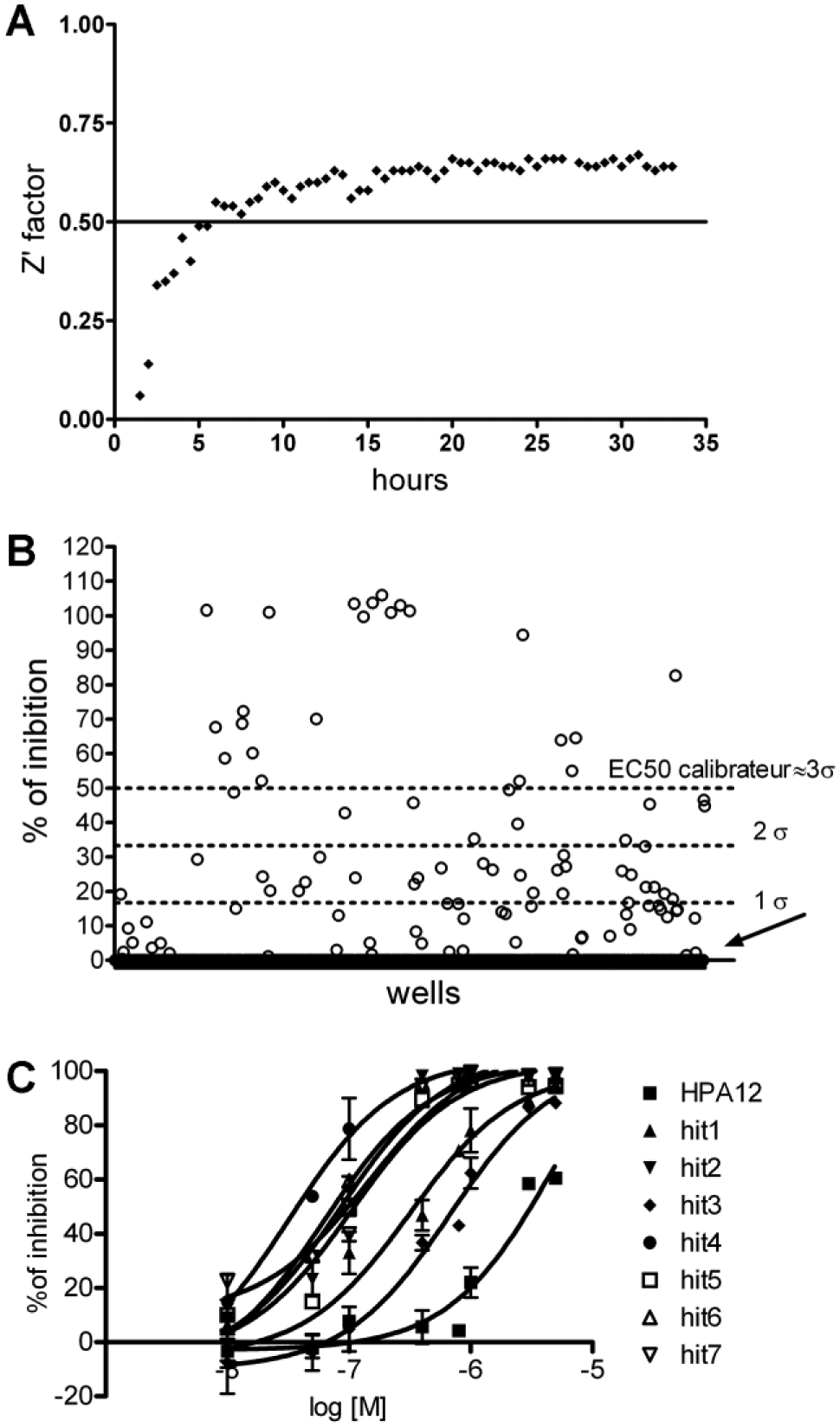
(
Hit Identification from a Set of 672 Molecules
The CNE and 32 original molecules were screened at a 5 µM final concentration ( Fig. 5B ). A total of 22 hits were identified as compounds presenting a percentage inhibition superior to the mean percentage inhibition of the 672 molecules plus three standard deviations (i.e., 49% inhibition of START-Cer binding). Seven hits were selected for dose-response analysis ( Fig. 5C ) and showed an EC50 ranging from 700 to 32 nM, that is, three orders of magnitude more affine that HPA12.
Hit Validation with FLINT
To distinguish valuable hit molecules from false-positives, hits were tested in the FLINT at a 2.5 µM concentration in comparison with HPA12. Results are presented in Figure 6 , showing a maximum level of recovery between both assays and confirming the activity of the seven hits.
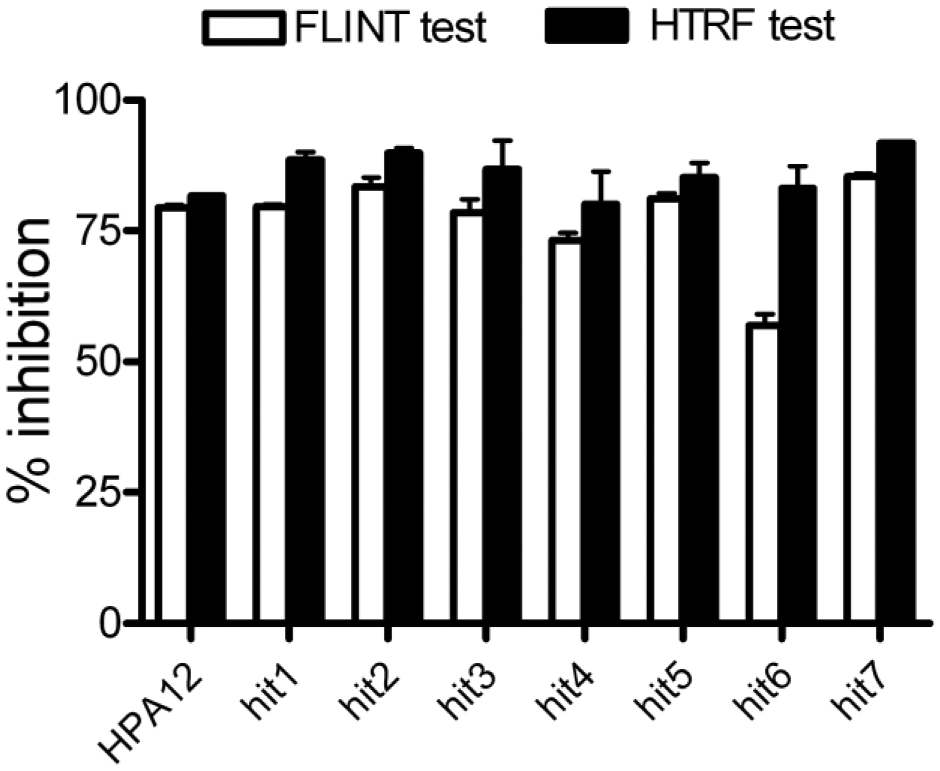
Confirmation of hits. Inhibitory potential of the seven hits discovered with the homogeneous time-resolved fluorescence assays was verified at 5 µM with the FLINT as described in Figure 2A .
Discussion
Blocking SM de novo synthesis via the inhibition of the CERT-Cer interaction is an innovative strategy to restore the intracellular concentration of the proapoptotic Cer metabolite.7,8 To access to CERT-mediated Cer transfer, several technologies were developed. They include the measurement of radiolabeled SM (inter)membrane extraction/transfer by CERT in a cell-free system and chromatographic analysis.18,27 Because they are overly complicated, these technologies could not be applied to a screening strategy. Fluorescent SM detection in cell extracts or in microscopic analysis were developed 28 and a screening of a compound library performed. 24 However, these assays were indirect and did not target CERT trafficking itself, requiring further experimental investigations to identify the active molecule mechanism of action.
Another type of assay was designed to target directly Cer transfer via CERT. It relies on surface plasmon resonance to calculate CERT-mediated extraction of Cer from the reticulum membrane. 29 Even so, automation of this assay is not possible. More recently, phosphorylation of CERT at Ser315 was shown to enhance the interaction of CERT with the ER membrane. 19 Both semi-intact cells and cell-free protocols were used to measure Cer transportation. These two assays rely on radiometric readouts and require extensive ultracentrifugation steps. Here, we propose an easy HTRF assay to measure the START-Cer interaction.
HTRF assays were developed for many targets such as kinases, G-protein–coupled receptors, protein-protein interactions, protein-DNA interactions, a lot of enzymatic assays (see Degorce et al. 13 for review), and lipidic molecule detection. 30 Here, for the first time, we adapted the HTRF technology to a protein-lipid binding assay suitable for large pharmacological studies and HTS. This specific protein-lipid HTRF binding assay required a long equilibration time and displayed a limited DMSO tolerance. The reagent addition sequence also proved to be determinant. Protein and competitor preincubation was shown to have a strong favorable impact on competitor detection sensibility, probably because of the slow kinetics of the formation of the START-Cer probe complex. 15
We identified molecules with a potential inhibitory effect on the START-Cer interaction. To exclude fluorescent false-positive molecules, the time delay of the readout was extended from 60 µs to 100 µs. 31 We further controlled and validated each hit with an independent assay based on the FLINT protocol, showing the good accuracy level of the HTRF assay. Cer competitors discovered in this work are up to three orders of magnitude more potent than the reference antagonist HPA12. An IC50 of 70 nM has been reported for the inhibition of the biosynthetic production of sphigomyelin in a whole living cell system of HPA12. The difference of magnitude between this IC50 and the presently measured EC50 can be explained considering that the observed phenomenon is entirely distinct (the ligand binding versus the inhibition of a biosynthetic pathway including an ER-to-Golgi transport step and an enzymatic transformation), the protein is not entirely the same (the START domain versus the entire CERT protein), and the experimental medium is also fundamentally different (an in vitro assay with a recombinant protein versus a living cell experiment).
Finally, our study paves the way for the development of new series of potential CERT antagonists. Further work is currently under way in our laboratories to elucidate the binding mode and evaluate the effect of the newly identified START domain ligands on the inhibition of the CERT-mediated Cer trafficking. HTRF assay was thus demonstrated to be an effective technology to discover competitors of this lipid-protein interaction.
Footnotes
Acknowledgements
We thank the Research & Development Operational Program funded by the ERDF (ITMS 26220220093) for financial support. The TR-FRET screening experiments were carried out at the “Plateforme Intégrée de Criblage de Toulouse” (PICT, IBISA) facilities. We are grateful to Professor Adam Daïch and Dr. Dušan Berkeš for a generous gift of HPA12 and analogs.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The financial support by CNRS for a “PIR-Therapeutic Innovation” funding, the “Université Paul Sabatier,” and the “Agence National de la Recherche” (ANR) (SphingoDR project) are gratefully acknowledged.
