Abstract
Human African trypanosomiasis (HAT) is a vector-transmitted tropical disease caused by the protozoan parasite Trypanosoma brucei. High-throughput screening (HTS) of small-molecule libraries in whole-cell assays is one of the most frequently used approaches in drug discovery for infectious diseases. To aid in drug discovery efforts for HAT, the SYBR Green assay was developed for T. brucei in a 384-well format. This semi-automated assay is cost- and time-effective, robust, and reproducible. The SYBR Green assay was compared to the resazurin assay by screening a library of 4000 putative kinase inhibitors, revealing a superior performance in terms of assay time, sensitivity, simplicity, and reproducibility, and resulting in a higher hit confirmation rate. Although the resazurin assay allows for comparatively improved detection of slow-killing compounds, it also has higher false-positive rates that are likely to arise from the assay experimental conditions. The compounds with the most potent antitrypanosomal activity were selected in both screens and grouped into 13 structural clusters, with 11 new scaffolds as antitrypanosomal agents. Several of the identified compounds had IC50 <1 µM coupled with high selectivity toward the parasite. The core structures of the scaffolds are shown, providing promising new starting points for drug discovery for HAT.
Introduction
Human African trypanosomiasis (HAT), also known as sleeping sickness, is a vector-transmitted disease caused by 2 subspecies of the digenetic protozoan parasite Trypanosoma brucei, namely, T. b. gambiense and T. b. rhodesiense. The disease is endemic in 36 Sub-Saharan Africa countries where the vector, the parasite, and the animal reservoir coexist. HAT has a fatal outcome if left untreated. Although the number of reported cases dropped from 37,385 in 1998 to 9689 in 2009, many remain unreported—and therefore untreated—due to the lack of specificity of the clinical diagnostic and the limited access to the infected populations. 1 In addition, T. brucei has an economic importance, because some subspecies are pathogenic to cattle, causing a wasting disease known as nagana and resulting in annual losses of US$1.5 billion in agricultural incomes. 2
There are 2 T. brucei subspecies responsible for the disease in humans: T. b. gambiense, which is found in West Africa and responsible for more than 95% of the cases of the disease, and T. b. rhodesiense in East Africa, which is responsible for the other 5% of the cases. 3 A third subspecies, T. brucei brucei, is unable to infect primates, although it is genotypically very similar to the 2 pathogenic subspecies, making it a good experimental model.
Because there are no vaccines available, drugs remain the main control strategy for HAT. There are 4 approved drugs for chemotherapy. Suramin, pentamidine, and melarsoprol are the most widely used and were developed before 1950. Eflornithine was approved in 1990, and since then advances in HAT treatment have been slow. 4 Furthermore, current treatments possess several limitations, such as limited efficacy and severe side effects due to toxicity, including mortality due to treatment.1,5–7 Nifurtimox, a drug used for the treatment of Chagas disease, was introduced in 2009 in the World Health Organization’s (WHO) List of Essential Medicines to be used as part of the nifurtimox–eflornithine combination therapy (NECT), providing a treatment less favorable toward the development of drug resistance and simpler to administer than the eflornithine monotherapy.2,8
High-throughput screening (HTS) of compound libraries using whole-cell assays is a well-established approach for drug discovery programs in neglected diseases. This kind of assay normally relies on simple pathogen viability readout, precluding the need for a validated target, which can be difficult to achieve; and, in contrast to a biochemical target-based assay, in this assay active compounds are discovered under physiologically relevant conditions. In addition, whole-cell-based assays have been quite successful in resulting in approved drugs for infectious diseases in general.3,9,10 In spite of its compelling advantages, only recently were whole-cell-based assays reported in HTS format for T. brucei. Luciferase and resazurin-based cell viability assays were proposed in a 384-well format.4,11–13 The latter was recently used to screen 87,296 compounds, resulting in 6 hits from 5 new chemical classes with activity confirmed against T. b. rhodesiense. 14
In spite of its versatility and widespread use, the resazurin assay does possess some important limitations. 15 Resazurin accepts electrons from free radicals and from the electron transport chain within the inner mitochondrial membrane, generating ROS and interfering with energy homeostasis.16,17 In addition, the resazurin screening assay for assessment of T. brucei viability in 384-well format assays has additional issues that might lead to misinterpretation of the results. Specifically, the assay requires long exposure of live parasites to resazurin at room temperature. 13 The long exposure to resazurin coupled with room temperature incubation could compromise T. brucei viability and could potentially interfere with compound activity during drug screening.
As an alternative to the resazurin assay, a SYBR Green–based whole-cell assay was developed for T. brucei in a 384-well format. SYBR Green is a cyanine dye that binds to nucleic acids, preferably to double-stranded DNA (dsDNA), thus providing an indirect assessment of cell number in a population. A SYBR Green–based whole-cell assay has been intensively used in antimalarial drug discovery.18–22 Because T. brucei is an exclusively extracellular parasite, whole-cell assays based on quantitative detection of nucleic acids could be a feasible alternative.
Material and Methods
In confirmatory counter-screening, hit compounds had their potency against THP-1 cells determined in dose–response curves as described above for trypanosomes. Briefly, compounds were diluted and plated as described above, followed by plating of 1×104 THP-1 cells/well in RPMI containing 10% FBS, totaling a final volume of 60 µL/well. Plates were incubated at 37 °C and 5% CO2 for 60 h prior to the addition of 5 µL of 130 µM resazurin in PBS, and they were incubated further for 12 h before reading.
Results
SYBR Green Assay Development and Comparison to Resazurin Assay
The resazurin assay has been so far the simplest and most affordable assay for HTS using African trypanosomes. While performing this assay in 384-well plates, we observed that T. brucei bloodstream forms presented a strongly reduced motility at the assay endpoint, even in untreated controls (
During assay development, experimental conditions such as the initial inoculum of trypanosomes in 384-well plates, the incubation time to achieve optimal trypanosome growth, and the volume and the composition of the lysis solution were optimized. Growth curves for different initial inoculums (several densities ranging from 0.5 to 676.6 × 103 trypanosomes/mL) were performed to determine the most suitable inoculum density to achieve optimal trypanosome growth in 384-well plates.
To ensure the optimal conditions to expose dsDNA after cell lysis, 4 different formulations of lysis buffer were tested on several inoculums that simulate the maximal range of trypanosome densities found in the well at assay endpoint (
Overview on the Performance Parameters for Both SYBR Green and Resazurin Methods for Final Assay Conditions.
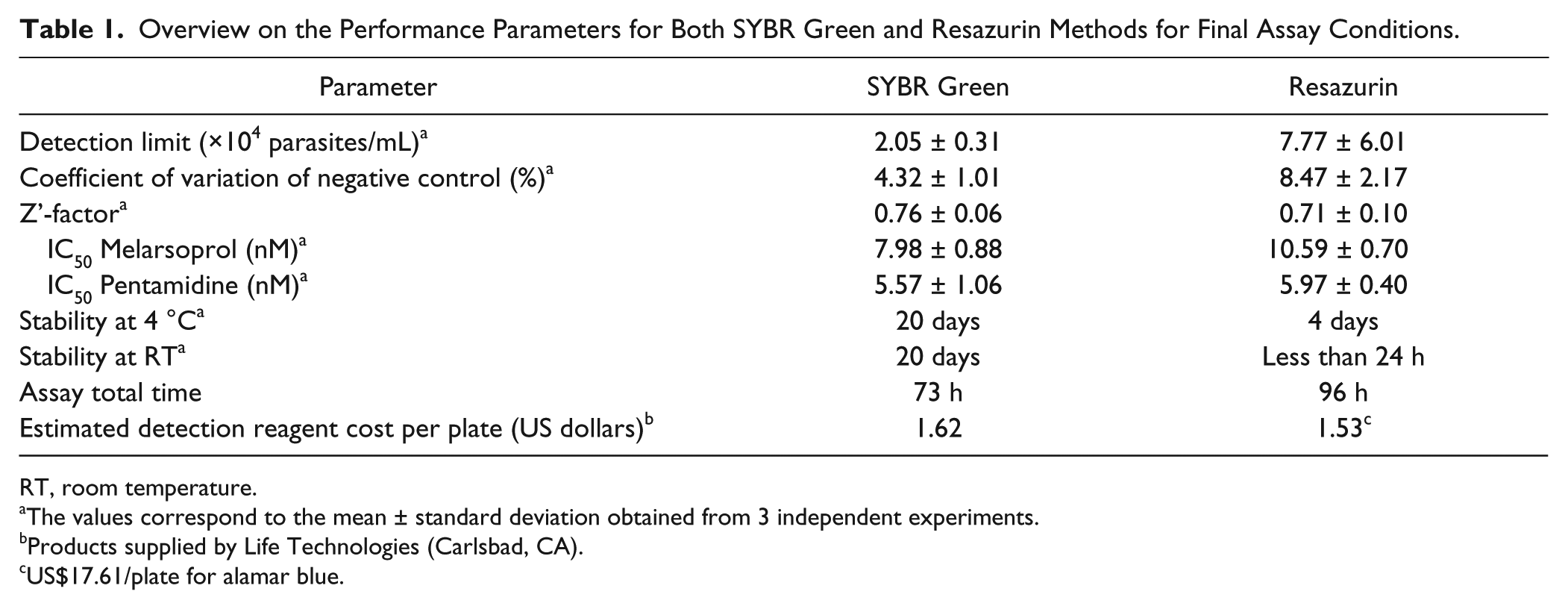
RT, room temperature.
The values correspond to the mean ± standard deviation obtained from 3 independent experiments.
Products supplied by Life Technologies (Carlsbad, CA).
US$17.61/plate for alamar blue.
SYBR Green Assay as an Alternative to Resazurin Assay
After establishing the final experimental parameters, the SYBR Green and resazurin assays were compared regarding the main parameters analyzed in HTS campaigns: the Z’-factor, the coefficient of variation, and the IC50s of reference drugs—in this case, melarsoprol and pentamidine. The results are summarized in Table 1 .
The SYBR Green assay presented lower variability, as indicated by the coefficient of variation of the negative control (4.32% for the SYBR Green assay vs. 8.47% for the resazurin assay) and a considerably more stable signal at both 4 °C and room temperature. In addition, the SYBR Green assay is faster, requiring a total of 73 h for completion—saving around 24 h in comparison with the resazurin assay. Both assays are comparable in terms of assay Z’-factor; IC50 values for melarsoprol and pentamidine, which are in accordance with the literature;13,28 and estimated cost of detection reagent (dye only) per plate. When alamar blue is considered instead of resazurin, however, the reagent cost per plate increases approximately 10 times ( Table 1 ).
The SYBR Green Assay Detects Activity of Both Fast- and Slow-Killing Compounds at the Assay Endpoint
One limiting factor for SYBR Green assay sensitivity in monitoring antitrypanosomal activity would be the required time for DNA degradation after cell death. Time-kill experiments for reference drugs were performed using the SYBR Green assay protocol, and developed with either SYBR Green or Cell Titer Glo as detection reagents to better understand the dynamics of the SYBR Green assay for the detection of fast- and slow-killing compounds and the sensitivity of the method when compared to Cell Titer Glo. The activity of melarsoprol and pentamidine, fast trypanocidal compounds, and eflornithine, a slow-killing compound, was monitored throughout time (
Comparison of the SYBR Green and Resazurin Assays in HTS of a Kinase-Focused Library
The primary screen of a commercial kinase-focused library of 4000 compounds was performed at 10 µM and in duplicate (2 independent runs) to minimize variation and permit robust comparison of both assays. A summary of the results is shown in both
Figure 1
and
Table 2
. The SYBR Green assay screen had a slightly better performance in terms of quality control parameters (
Table 2
), and the IC50s for melarsoprol and pentamidine were within the expected range (
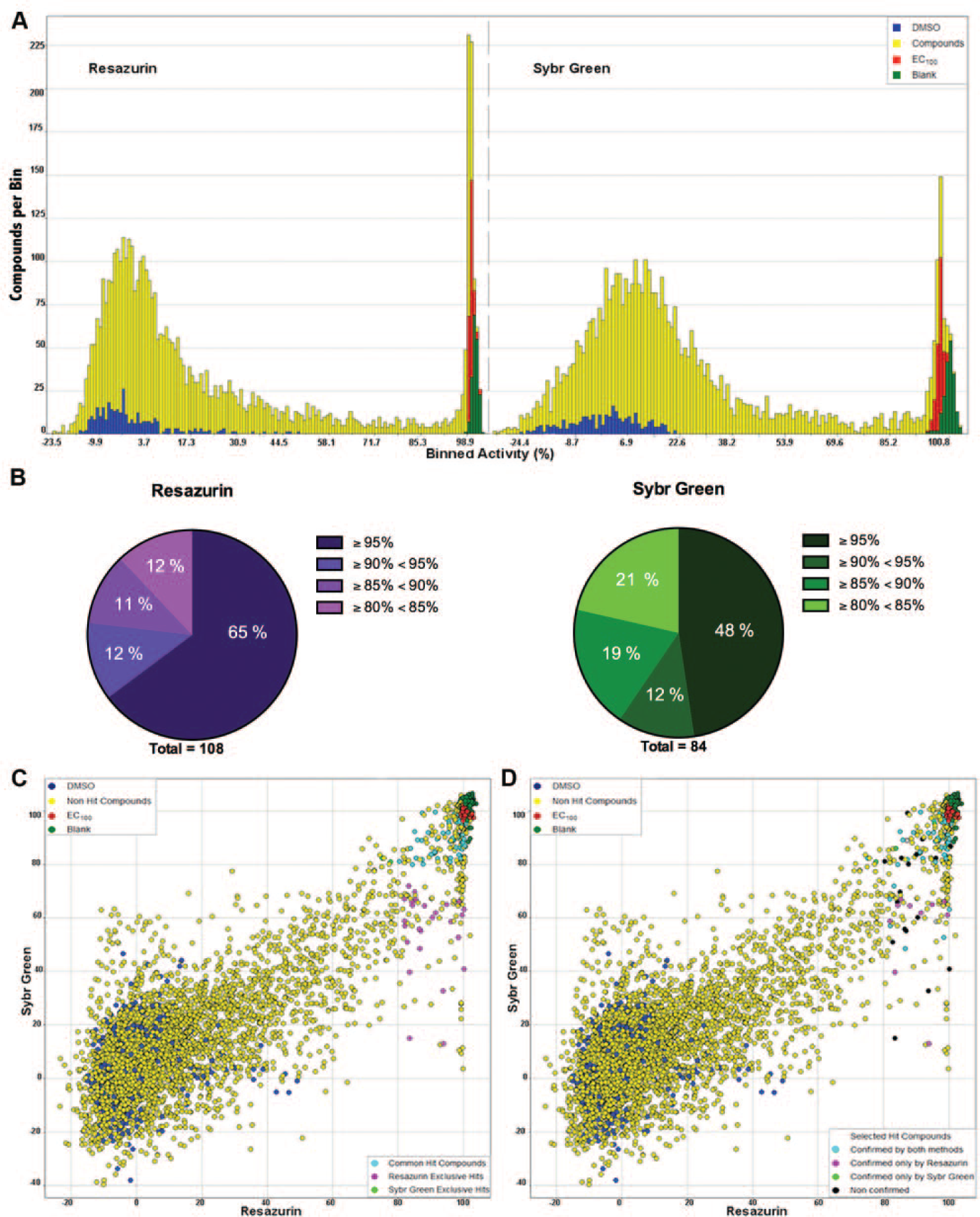
Comparison of antitrypanosomal activity profiles between the resazurin and SYBR Green assays. (
Comparison of SYBR Green and Resazurin Assays’ Quality Control Parameters in the High-Throughput Screening of 4000 Compounds.
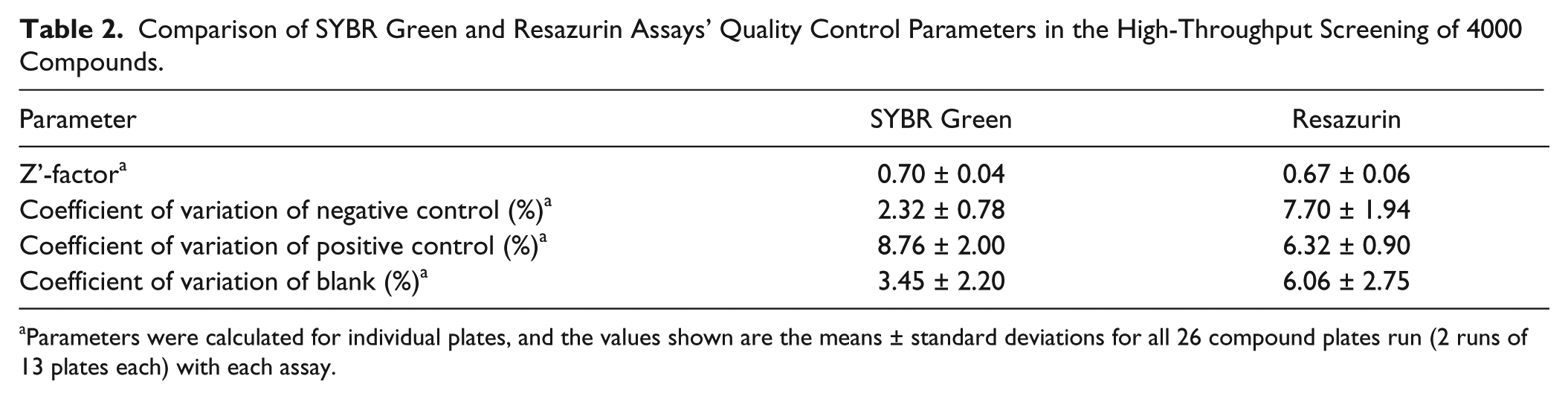
Parameters were calculated for individual plates, and the values shown are the means ± standard deviations for all 26 compound plates run (2 runs of 13 plates each) with each assay.
The binned distribution of compounds and controls per mean value of normalized activity showed a markedly different distribution pattern between the resazurin and SYBR Green assay screens ( Fig. 1A ). The greater variability of the negative control population in the resazurin assay was clear, because it was also evident that, for this method, the activity of compounds was mostly distributed in extremes (i.e., most compounds displayed either very high or very low activity); fewer compounds displayed activity in the intermediate range when compared to SYBR Green assay screen compound distribution.
The selected hits were further filtered for the removal of compounds that showed promiscuous cytotoxicity against human cell lines. Because this library had been previously screened against a panel of human cell lines (data not shown), only compounds that had a human cell line viability index between 1.0 and 1.5 were further progressed for antitrypanosomal activity confirmatory screening.
The difference in activity pattern between the 2 assays reflected the number of hit compounds selected from both screens: 108 compounds with normalized activity ≥80% were selected as hits in the resazurin assay, whereas 84 compounds were selected as hits in the SYBR Green assay following the same selection criteria. This difference was also evidenced with hit compounds distribution per activity bins of 5% increments, showing the increased proportion (65%) of hits with very high activity (≥95%) in the resazurin assay screen, whereas the SYBR Green assay screen had 48% of the hits with activity ≥95% ( Fig. 1B ). Among the hit compounds, 82 were common to both resazurin and SYBR Green assays, 2 compounds were selected as exclusive hits of the SYBR Green assay, and 26 as exclusive hits of the resazurin assay ( Fig. 1C ).
Regarding the whole population of compounds and control wells, both assays were robust and resulted in reproducible data as shown in
Confirmatory and Counter-Screening for the Determination of Compounds With Selectivity Toward T. brucei
Among the 110 selected compounds, 3 compounds were not available for further studies: 2 hits were common to both assays, and 1 hit was exclusive of the resazurin assay. Thus, 79 compounds selected by both methods, 25 compounds selected only in the resazurin assay, and 2 selected only in the SYBR Green assay were tested for activity confirmation against T. b. brucei in dose–response curves to determine the compound potency. In parallel, the hits were also assayed against THP-1 cells to determine compound selectivity toward the parasites. A hit was considered confirmed when the normalized activity of the compound at 10 µM was ≥80% in the confirmatory screening.
Considering only the common hit compounds, the hit confirmation rate was 82.3% (65/79); when considering only the exclusive hits of this assay, the hit confirmation rate dropped to 64% (16/25). For the SYBR Green assay screen, the total hit confirmation rate was 86.4% (70/81) and thus higher than that of the resazurin assay screen. Both exclusive hit compounds for this assay confirmed, and regarding the common hits, the confirmation rate was 84.8% (67/79). One of the 2 SYBR Green exclusive hits was also confirmed by the resazurin assay, thus representing a missed hit or a false negative in the resazurin assay primary screening. From the 16 compounds selected as resazurin assay exclusive hits, 6 were also confirmed by SYBR Green assay. These compounds were also active in the SYBR Green assay primary screen, albeit with lower activity than in the resazurin assay primary screen, and therefore were excluded as hits in the SYBR Green assay screen due to the stringent activity cutoff applied in hit selection ( Fig. 1D ).
Potentiation of Compound Activity in the Resazurin Assay
The resazurin assay screen had a higher hit rate, and the hits had higher activity in comparison with the activity found in the SYBR Green assay screen. Furthermore, the resazurin assay screen had a higher rate of exclusive confirmed hits that could not be detected by the SYBR Green assay. These results could be explained by the fact that the SYBR Green assay may not detect slow-killing compounds within the assay time window; however, this claim is not supported by the time-kill assays performed with eflornithine (
Representative examples of profiles obtained with hits common to both resazurin and SYBR Green assays (
Fig. 2A
), and hits exclusive to the resazurin assay (
Fig. 2B–D
), are shown. The plot for the common hit showed that at 10 µM (the concentration applied in the primary screening), this compound was able to kill 100% of the population with as little as 48 h of exposure, whereas 10 µM of resazurin assay screen exclusive hits plotted in
Figure 2B
could only reduce population viability to 0% after 72 h exposure, suggesting that indeed the SYBR Green assay screen preferentially detected fast-killing compounds. The other common hit compounds were also identified as fast-killing compounds, and in some cases they showed 100% antitrypanosomal efficacy after 24 h exposure, regardless of their potency (data not shown). Among the 7 compounds that had been confirmed only by resazurin assay, 2 other compounds presented the same pattern observed in
Figure 2B
, further suggesting that a potential limitation of SYBR Green assay may be less sensitivity toward slow-killing compounds. It is also important, however, to emphasize that these compounds can also be detected as active in the SYBR Green assay, albeit with reduced potency at the assay endpoint, presumably due to incomplete DNA degradation. The plots in
Figure 2C
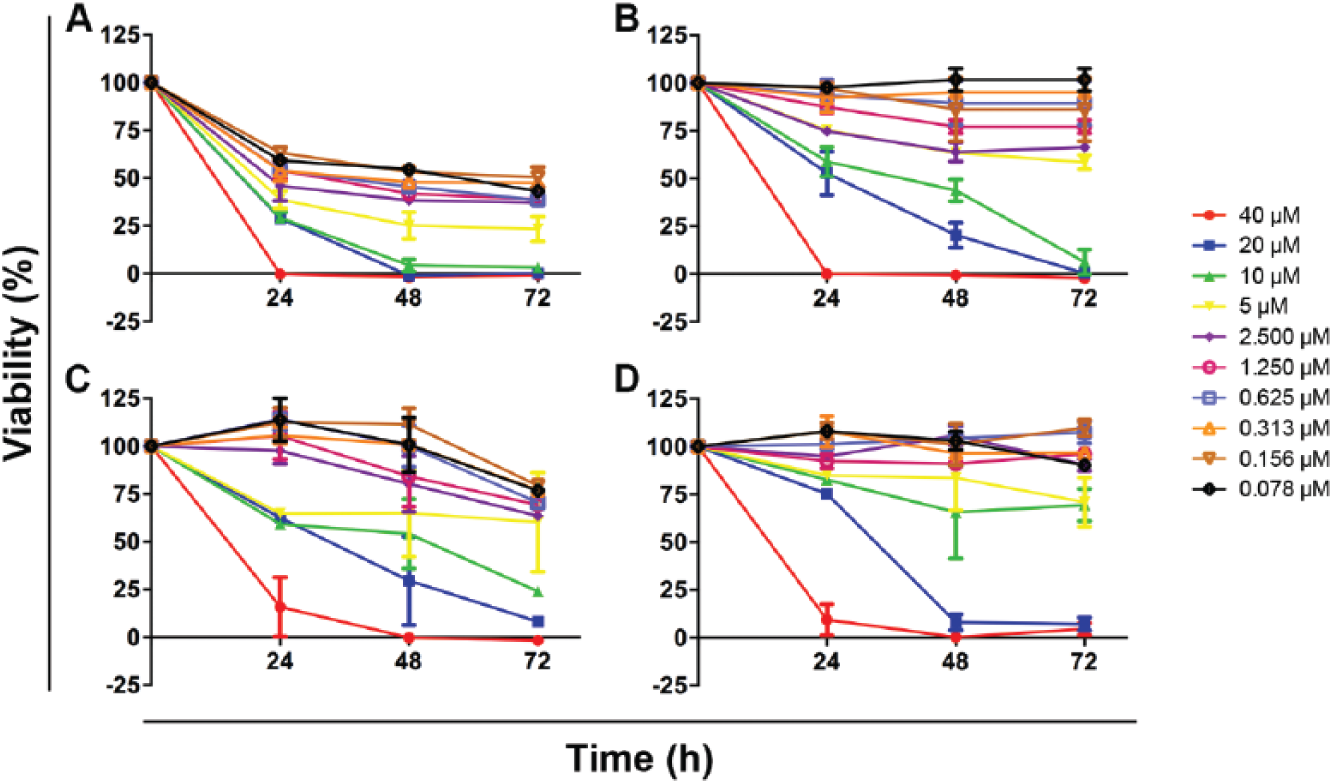
Some of the resazurin assay screen exclusive hits are slow-killing compounds, whereas others kill trypanosomes fast but have only minor activity at screening conditions with the SYBR Green assay. Nonexclusive (
As a counterproof, all of the 16 confirmed hits were tested in a second set of experiments, but this time using the resazurin assay as the screening protocol and the Cell Titer Glo reagent as the reporter of cell viability.
The results summarized in Figure 3 show the activity at 10 µM of these compounds obtained in both sets of experiments described above. Most nonexclusive hits had activity greater than 80% in all tested conditions, except for compound Hit ID 3, in which the activity at 10 µM was approximately 40% in the resazurin assay using resazurin for signal development (RSZ_RSZ). This can be attributed to the higher variation observed with the resazurin assay.
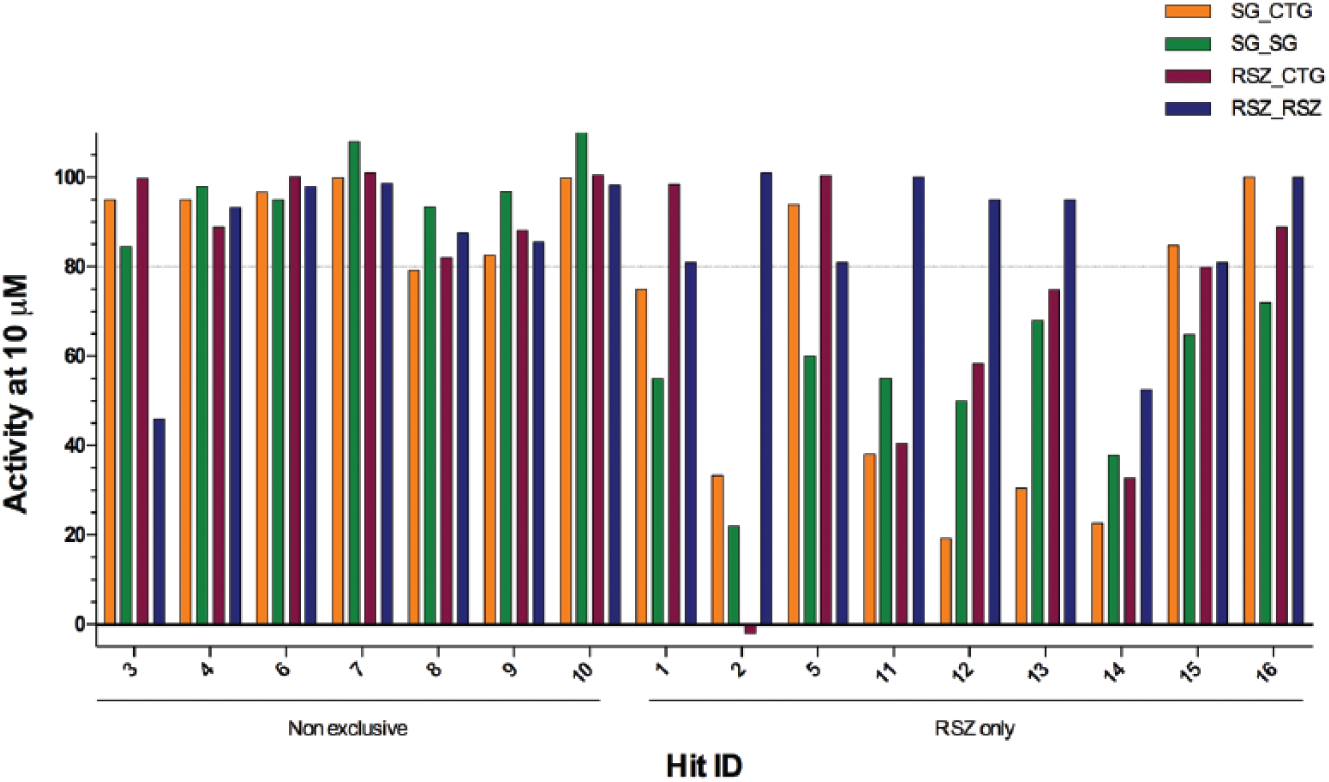
Some resazurin assay screen exclusive hits have pronounced activity in the presence of resazurin. Selected nonexclusive and resazurin assay screen exclusive hits (RSZ only) either were tested in serial dilution following the SYBR Green assay experimental conditions and developed with either Cell Titer Glo (SG_CTG, orange bars) or SYBR Green (SG_SG, green bars), or were tested following the resazurin assay experimental conditions and developed with either Cell Titer Glo (RSZ_CTG, purple bars) or resazurin (RSZ_RSZ, blue bars). A dotted line is shown at 80%, the activity cutoff applied in the selection of hits in the primary screening and used as activity confirmation criteria in the confirmatory dose response screening (shown here). Data refer to 2 independent replicates.
The resazurin assay screen exclusive hits can be classified in 2 major activity patterns: The first group of compounds (Hit ID 1, 5, 15, and 16) was active in all conditions, albeit moderately active (activity ranging from approximately 50% to 70%) only in the SYBR Green assay developed with SYBR Green (SG_SG) assay ( Fig. 3 ), suggesting that they are slow-killing compounds, and in these cases there is a delay in the detection of cell death by the SYBR Green assay. A second group of compounds (Hit ID 2, 11, 12, and 13) were markedly highly active in the presence of resazurin only (RSZ_RSZ) ( Fig. 3 ), regardless of the experimental protocol used in compound testing. Importantly, these compounds were only moderately active in the other assay setups tested, including the resazurin assay developed with Cell Titer Glo (RSZ_CTG), suggesting that, in these cases, compound activity was increased only in the presence of resazurin and differences in detection of antitrypanosomal activity between the resazurin and SYBR Green assays are not due to the speed with which the compound exerts its cidal effect.
Novel Scaffolds for Antitrypanosomal Drug Discovery
Despite the differences in the hit profile between the compound found in both assay protocols, the majority of the most potent hits were confirmed in both assays. The selectivity index toward trypanosomes (SI) varied from 3.9 to more than 512; indeed, 72% of the compounds presented an SI higher than 10 (data not shown).
The 72 confirmed compounds were analyzed for chemical clustering based on scaffold similarity, yielding 13 clusters. Among these, 11 clusters consisted of novel scaffolds with previously unknown antitrypanosomal activity ( Fig. 4 ).
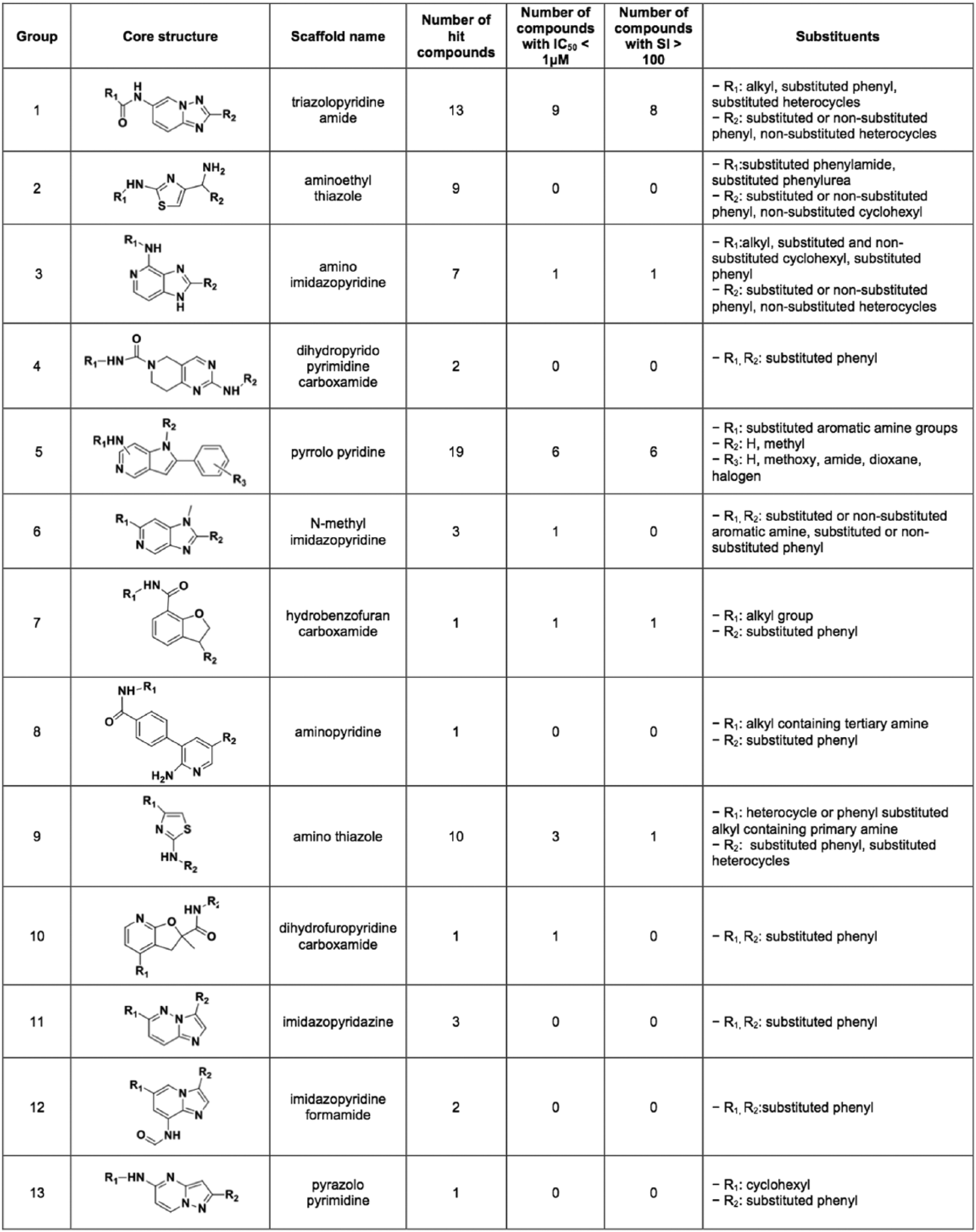
Scaffolds with potent antitrypanosomal activity discovered in the screening of a 4000-compound kinase-focused library against T. brucei. The core structure of hit chemical clusters is shown, together with the number of hit compounds that integrate each cluster, the number of hits with IC50 in the nanomolar range, and the number of compounds with a selectivity index (SI) greater than 100. The substituents for each scaffold are shown in the last column.
Discussion
The aim of this study was to develop a novel and improved drug-screening assay for drug discovery for HAT, and, in doing so, deliver new potential drug candidates. HTS of compound libraries using whole-cell assays is a simple and viable approach for early drug discovery programs in HAT; nevertheless, there are few reports based on whole-cell HTS assays for T. brucei. In the present work, we propose a novel, robust, suitable-to-automation, cost- and time-effective, simple, and reproducible 384-well-format HTS SYBR Green–based assay.
To validate the SYBR Green assay and, most importantly, discover new antitrypanosomal chemical entities, a kinase-focused library composed of 4000 compounds was screened against the T. brucei Lister 427 strain using both the SYBR Green and resazurin assays in HTS mode. The parallel screening of the same library in both assays allowed for comparison of the assays’ performance in real HTS conditions and also whether the hit profiling obtained in both screens would be considerably different.
A thorough comparison demonstrated that the SYBR Green assay is more sensitive, less variable, and faster to develop than the resazurin assay, all features of major importance in HTS. SYBR Green also had a better confirmation rate (approximately 86%) than the resazurin assay (75%), and some of the exclusively identified hits identified in the resazurin assay could not be confirmed or presented lower activity when rescreening with SYBR Green or Cell Titer Glo. This fact, allied with the increased activity of the selected hits in the resazurin assay, raised the hypothesis that this assay could lead to an additive “intrinsic activity” that further debilitates trypanosomes, thus decreasing the population viability and artificially increasing hit rates. Resazurin has been extensively used for monitoring T. brucei viability in drug assays in 96-well plates; however, no data regarding toxicity to trypanosomes are available in the literature. We suppose that in the 384-well-plate format, a longer exposure to resazurin (24 h) in comparison to the 96-well-plate format (4 h) and/or a relatively long incubation period at room temperature (22 h) could account for the reduced parasite motility observed at the resazurin assay endpoint (
The practical effects of using a resazurin assay for HTS for HAT drug discovery can be speculated. Besides offering no advantage in terms of cost and time, the resazurin assay resulted in a higher rate of false positives. The greater assay variability could also result in a higher rate of false negatives in the resazurin assay. The resazurin assay, however, seems to be more sensitive for the detection of compounds that kill trypanosomes slowly. This could be due to cell membrane permeability kinetics, features that can be improved by chemical optimization of scaffolds; alternatively, it could (also) be due to the compound mechanism of action.
The major novelty of this study was the discovery of novel scaffolds with potent antitrypanosomal activity ( Fig. 4 ). Kinase targets have been addressed by the pharmaceutical industry and offer potential for the development of new drugs in several therapeutic areas. 29 The library screened in this study is a focused set of approximately 4000 drug-like small molecules of structural features that favor interactions with kinases and/or phosphatases. Knowledge of the crucial role of protein kinases in cell survival in multicellular organisms has directed attention toward this enzyme class in parasitic protozoa to search for alternative drug targets to treat tropical diseases.30,31
Most of the 72 hit compounds identified here against T. b. brucei presented a high selectivity index toward trypanosomes. Among those, triazolopyridine-based inhibitors ( Fig. 4 , group 1) have been described as atypical kinase inhibitors, applicable to p38 mitogen-activated protein kinase (MAPK). 32 There are 3 major classes of MAPKs: ERK, p38, and JNK. So far, 2 ERK homologues (KFR1 and TbMAPK2) were identified in T. brucei along with 2 kinases that share features of both ERK and cyclin-dependent kinases (TbECK1 and TbMAPK5).30,33 TbMAPK5 is involved in the growth and differentiation of bloodstream forms, and knockdown studies showed a reduction of the peak parasitemia in mice. 33 Extensive studies are needed, however, to determine the targets of the compounds reported in this article. Although not mandatory, knowledge of the target and its structure would facilitate the development of highly antiparasitic selective drugs. And, although some hits already have good potency, further medicinal chemistry optimization is necessary to improve activity and/or pharmacological properties and establish the structure–activity relationship (SAR). In conclusion, the contribution of this study is not only a novel HTS T. brucei viability assay that overcomes the limitations of the standard method but also attractive chemical scaffolds that will be valuable starting points for HAT drug development.
Footnotes
Acknowledgements
We would like to acknowledge Fundação para a Ciência e Tecnologia (FTC) for funding (SFRH/BD/79712/2011), Eunhye Kim and Luís Gaspar for technical support, and Gilles Courtemanche for kindly providing melarsoprol. This work was partially supported by a National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIP; No. 2007-00559), Gyeonggi-do (No. K204EA000001-09E0100-00110), and KISTI.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: We would like to acknowledge Fundação para a Ciência e Tecnologia (FTC) for funding (SFRH/BD/79712/2011), Eunhye Kim and Luís Gaspar for technical support, and Gilles Courtemanche for kindly providing melarsoprol. This work was partially supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIP; No. 2007-00559), Gyeonggi-do (No. K204EA000001-09E0100-00110), and KISTI.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
