Abstract
Invasive mycoses have been increasing worldwide, with Candida spp. being the most prevalent fungal pathogen causing high morbidity and mortality in immunocompromised individuals. Only few antimycotics exist, often with severe side effects. Therefore, new antifungal drugs are urgently needed. Because the identification of antifungal compounds depends on fast and reliable assays, we present a new approach based on high-throughput image analysis to define cell morphology. Candida albicans and other fungi of the Candida clade switch between different growth morphologies, from budding yeast to filamentous hyphae. Yeasts are considered proliferative, whereas hyphae are required for invasion and dissemination. Thus, morphotype switching in many Candida spp. is connected to virulence and pathogenesis. It is, consequently, reasonable to presume that morphotype blockers interfere with the virulence, thereby preventing hazardous colonization. Our method efficiently differentiates yeast from hyphal cells using a combination of automated microscopy and image analysis. We selected the parameters length/width ratio and mean object shape to quantitatively discriminate yeasts and hyphae. Notably, Z′ factor calculations for these parameters confirmed the suitability of our method for high-throughput screening. As a second stage, we determined cell viability to discriminate morphotype-switching inhibitors from those that are fungicidal. Thus, our method serves as a basis for the identification of candidates for next-generation antimycotics.
Introduction
Candida albicans is the most common human fungal pathogen, even though it is part of the commensal microflora in the gastrointestinal and urogenital tracts as well as in the oral cavity. Vaginal candidiasis has been described at least once in 75% of all women worldwide.1 Although the disease is not invasive, the quality of life of affected individuals is considerably reduced. Notably, up to 10% of these women additionally experience relapsing incidents of vulvovaginal candidiasis. Bloodstream infections with Candida spp. (candidemia) have been increasing in hospital intensive care units (ICUs) worldwide, with an annual prevalence of 300,000 cases and 30% to 50% mortality.2,3 Among systemic microbial infections in ICU patients, candidiasis ranks second in Europe and North America, causing more than 50% of the cases. 2 Despite the increase in Candida infections, currently available therapeutic agents remain few in number, and of those, several can lead to severe side effects, such as liver damage. 4
The most important antifungal drugs can be classified into six categories with confined fungal targets. 5 Polyenes bind ergosterol and induce pores in fungal plasma membranes. Azoles and allylamines inhibit the synthesis of ergosterol, rendering the membranes unstable, whereas morpholines and antimetabolites prevent nucleic acid production. Many currently applied antifungal drugs have severe side effects. 4 The ergosterol-attacking agents, for instance, can additionally interfere with the human analog cholesterol, resulting in host cell damage. Thus, long-term use of these drugs can cause renal dysfunction, liver toxicity, or bone marrow depression. Finally, echinochadins constitute a new group of antifungal agents. They are inhibitors of glucan synthetase in the cell wall of fungi, which has no counterpart in human hosts, resulting in less severe side effects. However, emergence of echinocandin-resistant strains is a cause of clinical concern.
Systemic and superficial candidiasis is strictly associated with the reversible morphotype switching from budding yeasts to the filamentous hyphae (Y-H). 6 The yeast is a unicellular morphotype and considered the commensal form of C. albicans. 7 Derogation of innate or adaptive immunity can induce adherence to epithelia and conversion from yeast to hyphal growth. 6 Hyphal growth results in a filamentous morphology initiated by germ tube formation at a yeast mother cell. Apical growth at the tip of the filament continues with occasional branching events. Hyphae have been reported to be essential for invasion and dissemination to noncommensal niches as well as for biofilm formation and escape from host immune cells. 6 Notably, biofilms are a major cause of medical device failure and a frequent source of relapsing infections. 8 C. albicans transcription factor knockout mutants (for instance, Δedt1 and Δefg1) that are locked in the yeast morphology have been shown to be nonvirulent in animal models and to be unable to form biofilms, supporting the concept of morphotype transition as a virulence trait.8–10 We reasoned that identification of compounds that inhibit the Y-H transition can be exploited for application as antifungal therapy. Fungal growth should not be altered by these agents but rather redirected into a commensal state that can be controlled by the immune system. According to these assumptions, we hypothesize that during treatment with such agents, selection pressure on fungal pathogens is low and, thus, in turn the possibility for resistance development decreased. Therefore, we aimed to develop a method to screen large chemical compound libraries for Y-H transition inhibitors. The ideal compounds are those that prevent C. albicans morphotype transition without affecting the cellular viability of fungal cells.
The method is based on automated microscopic imaging of labeled fungal cells and thereafter quantitative image analysis, referred to as high-content analysis (HCA). Using HCA, we calculated the mean object shape (MOS) and length/width ratio (LWR) of fungal cells. MOS and LWR were selected to define and quantify the Y-H transition, as calculated by eq 1 (see the Materials and Methods section). To identify compounds that are fungistatic or fungicidal, we quantified fungal viability by measuring adenosine triphosphate (ATP) levels, an assay we previously applied to screen drug libraries for antifungal activity. 5 Viability was calculated using eq 2 (see the Materials and Methods section).
To verify our method, we used farnesol, a natural quorum-sensing molecule secreted by C. albicans. Farnesol blocks hyphal growth without affecting the proliferation as yeast-form cells. 8 In addition, we used knockout-mutant strains that are restricted to yeast-form growth, namely, Δedt1 10 and Δefg1. 9 These transcription factor knockout strains are unable to switch from yeast form to hyphal growth, even when growing in otherwise hypha-inducing conditions. Thimerosal served as a reference for fungistatic or fungicidal agents. The mercury compound kills fungal cells by disruption of mitochondria. Furthermore, the Z′ factors for quality assessment of the method were calculated for both LWR and MOS at 6 h using eq 3 (see the Materials and Methods section), defining our method as valid and suitable for high-throughput screening. 11
Materials and Methods
Media and Cultivation of Fungi
Conditions and cell concentrations were based on the antifungal susceptibility testing (AFST) guidelines of the European Committee on Antimicrobial Susceptibility Testing (EUCAST). 12 C. albicans (SC5314) and C. albicans engineered gene knockout mutant strains locked to yeast-form growth, Δedt1 10 and Δefg1 9 (derived from parental strain C. albicans CAI4), were used in this study.
Yeast cells were grown overnight by shaking at 30 °C in synthetic complete dropout medium with 2% glucose (SC). Subcultures were inoculated from an overnight culture to a final concentration of 107 cells/mL in SC and incubated for 4 h at 30 °C. Cells were washed twice in phosphate-buffered saline 1× (PBS) and adjusted to 2.0 × 105 cells/mL in PBS. To each well of a black, 96-well plate with transparent bottom (Costar, Fisher Scientific, Waltham, MA), 150 µL RPMI 1640 with 0.5% DMSO (0.5%) followed by 50 µL yeast suspension were added using a WellMate Matrix (Thermo Scientific, Waltham, MA) to give a final volume of 200 µL in each well. The plate was then incubated at 37 °C at 5% CO2 for 3, 6, or 24 h. Under these conditions, C. albicans yeast cells start to germinate and to grow as hyphae. To keep the cells at the bottom of the well for better microscopic imaging, the wells were pretreated with 0.1% w/v poly-L-lysine (Sigma-Aldrich, St. Louis, MO) for 30 min and washed three times with PBS. In addition to the above conditions, we added 250 µM farnesol (Sigma-Aldrich) or 0.8% w/v thimerosal (Sigma-Aldrich) to SC5314 as references for morphotype switching inhibition and fungicidal activity, respectively.
Determination and Quantification of Y-H Switch Using HCA
After incubation for 3, 6, and 24 h, C. albicans cells were fixed with 2% paraformaldehyde and stained with 0.1% calcofluor white (CFW; Sigma-Aldrich). We used the chitin-specific fluorescent dye CFW to analyze the fluorescent signal of stained fungal cell walls. Images were captured with an automated microscope (HCA-Cellomics ArrayScan VTI, Thermo Scientific) and the C. albicans cell morphology analyzed. Based on HCA information, the individual fungal cell morphotype was determined by means of LWR and MOS, respectively. These two parameters were sufficient to reliably discriminate between yeast and hyphal cultures (eq 1). LWR determines the average ratio between length and width, which indeed changes considerably during apical growth of a filament versus division of ellipsoid yeast cells by budding. MOS refers to the average measure of detected objects based on the formula MOS = [(c2/4π)*area], which is the ratio of circumference squared to 4π*area (MOS = 1 = circular object).
Determination of Cell Viability Using ATP Quantification
We performed the cellular viability test prior to cell fixation and chitin staining. The percentage of cellular viability was determined using the CellTiter-Glo luminescent cell viability assay (CTG; Promega, Madison, WI) to identify compounds that are fungistatic or fungicidal. A volume of the CTG reagent equal to the cell volume per well was added. After 15 min at room temperature, the luminescence signal was quantified in a luminometer (Infinite F200, Tecan, Männedorf, Switzerland). The luminescence signal corresponds to ATP values and thus to cellular viability. The percentage of cellular viability was calculated for the four tested conditions, C. albicans either with farnesol (250 µM) or thimerosal (0.8%) and the two mutant C. albicans strains in comparison with C. albicans in DMSO (0.5%) as the 100% hyphal growth control (eq 2). The assay was performed at least to three biological replicates in triplicate (n = 3[3]) in 96-well plates with clear bottoms. Liquid handling, plate reading, and automated microscopy were performed at Laboratories for Chemical Biology Umeå (LCBU), Chemical Biology Consortium Sweden (CBCS).
Calculations and Statistical Analysis
Growth inhibition (GrIn) was determined, after 3 and 6 h, from the MOS and LWR values for all conditions. MOS and LWR calculations are derived from the average number of fluorescent pixels from at least 100 cells. The percentage of GrIn (%GrIn) was defined as eq 1: [%GrIn =100 – (
Results and Discussion
A crucial virulence trait of polymorphic fungi is their ability to reversibly switch from yeast-like to filamentous growth. Hence, the aim of the study was to develop a reliable high-throughput screening method for the identification of molecules that break the Y-H transition without disturbing cell viability. Images gathered from an automated fluorescence microscope were analyzed on the basis of fluorescent pixels. From the substantial amount of parameters created by HCA, we chose LWR and MOS, because these values were sufficient to reliably distinguish between yeast and hyphal morphotypes ( Fig. 1 ). This means in particular that C. albicans samples with LWR and MOS values less than 1.5 are defined as yeast cells ( Fig. 1 ). After a 24 h incubation, LWR and MOS values from the hyphal reference samples cannot be taken into account, as confluent growth renders analysis unfeasible. Microscopic images are nevertheless available in substantial amounts for cell morphotype evaluation (data not shown).
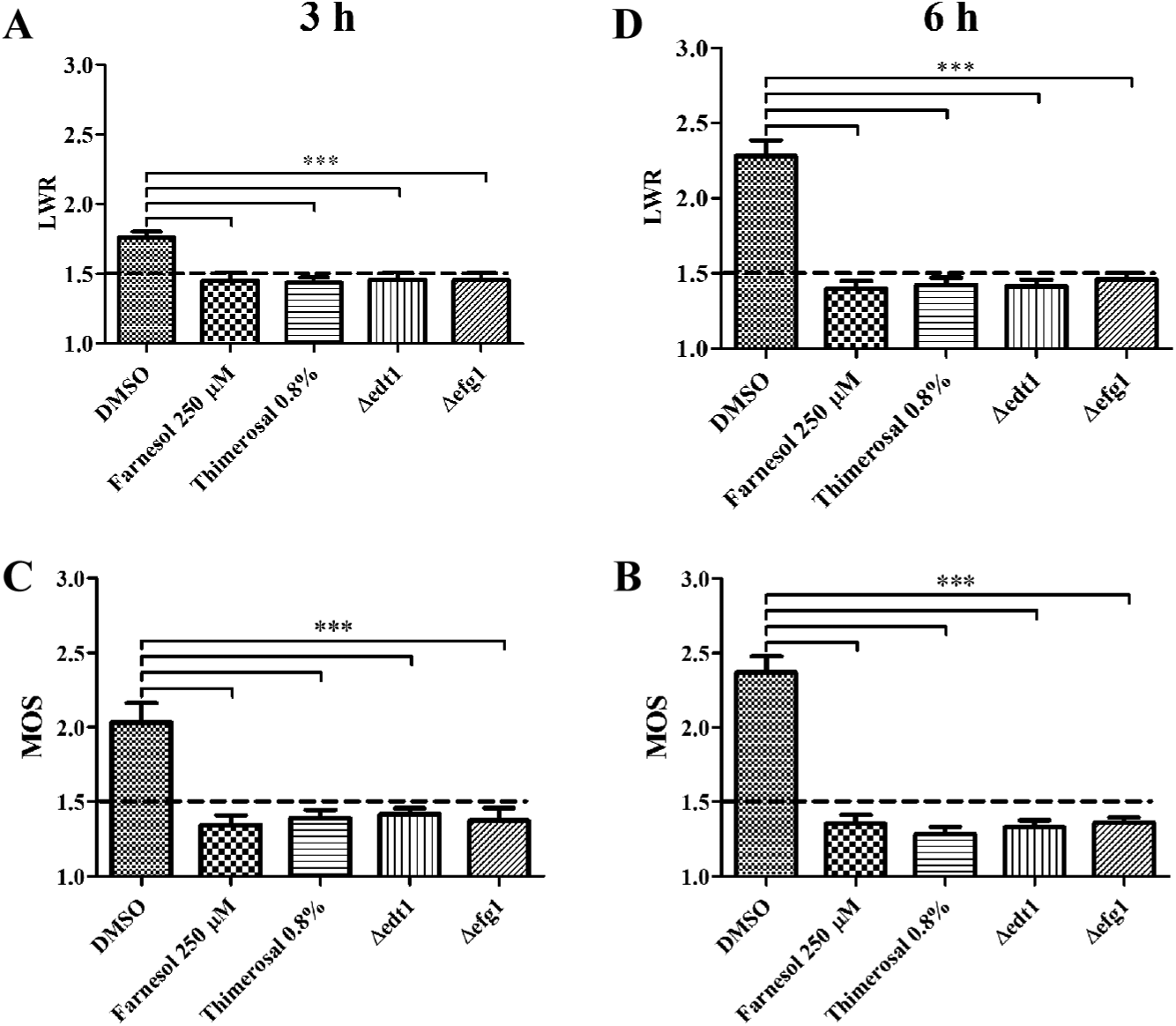
Length/width ratio (LWR) and mean object shape (MOS) define cell morphology. The cells were incubated at 37 °C and 5% CO2 for 3 and 6 h (
To validate whether our method is suitable for identifying switching inhibitors, we used the quorum-sensing molecule farnesol. This natural compound prevents hyphal growth of C. albicans under otherwise hyphae-inducing conditions ( Figs. 1 – 3 ). After 24 h of incubation with farnesol, however, C. albicans yeast growth was additionally reduced to low levels, indicating that over long incubation times, farnesol has growth-inhibitory activity. This is in good agreement with a previous report that showed that farnesol challenge of yeast cells prevented hyphal growth but at the same time significantly reduced cellular viability. 8 Farnesol is nevertheless a suitable reference for morphotype switching inhibitors, because it does not affect C. albicans growth within 6 h incubation periods ( Fig. 3 ). We next used thimerosal to kill off C. albicans cells, which afterward remain as dead and thus nonswitching yeasts. LWR and MOS obtained from HCA data confirmed that thimerosal-treated C. albicans remained as yeasts, because values were less than 1.5 and cellular viability was close to background levels ( Figs. 1 – 3 ). Thus, thimerosal could be used as a reference for fungicidal or fungistatic compounds.
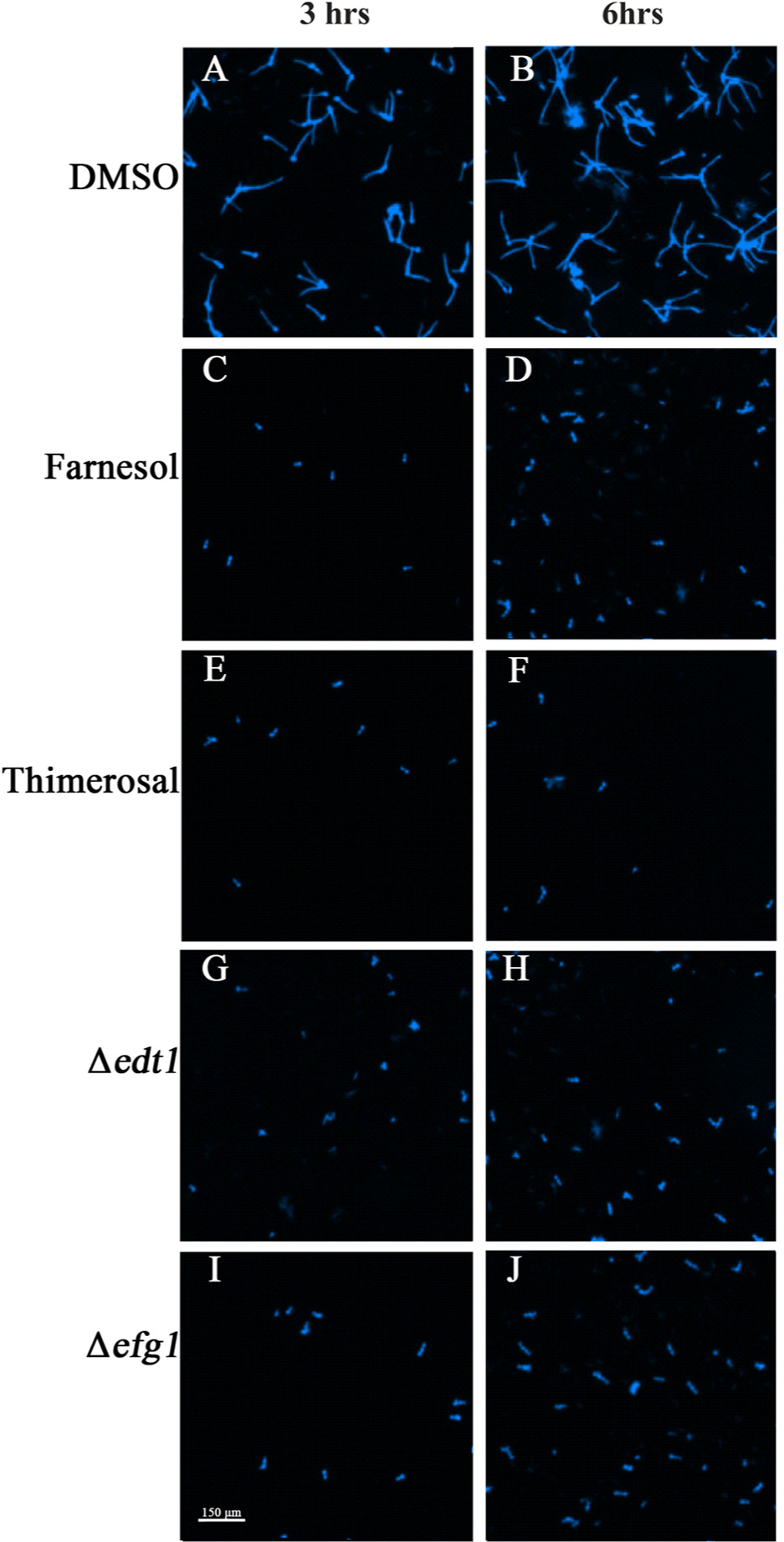
Microscopic images confirm quantification results via length/width ratio and mean object shape. The cells were incubated at 37 °C and 5% CO2 for 3 and 6 h (
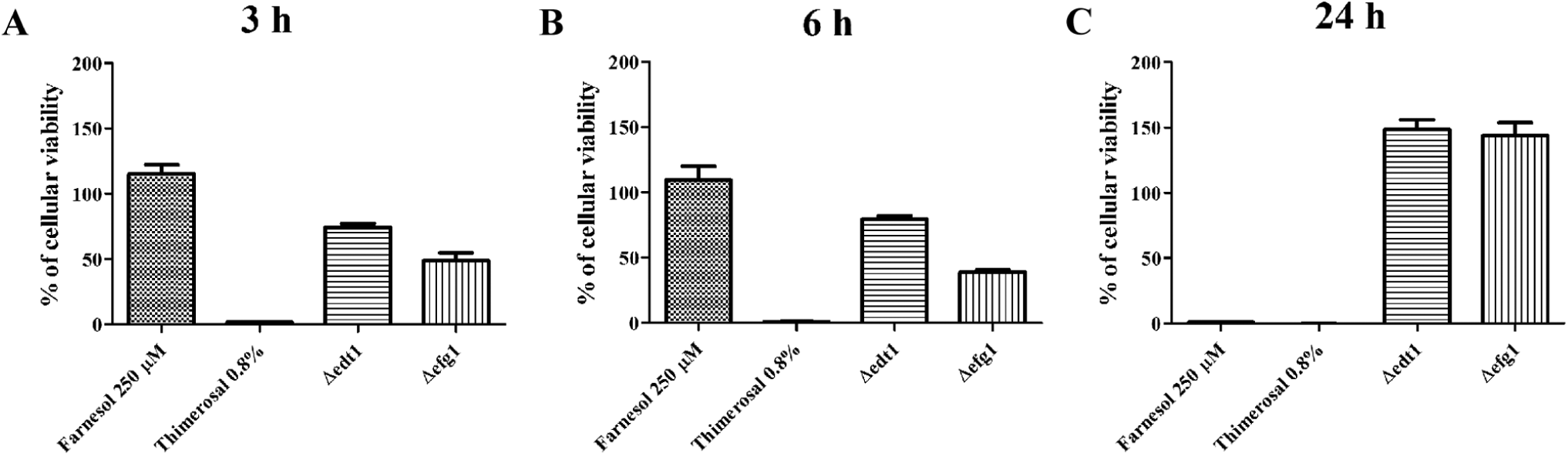
Cell viability determined using adenosine triphosphate (ATP) levels. The cells were incubated at 37 °C and 5% CO2 for 3, 6, and 24 h (
We furthermore assayed two knockout C. albicans mutants Δefg1 and Δedt1,9,10 both yeast-locked strains. They serve as additional key references for determining the accuracy of discrimination between yeast growth versus hyphal growth. This was confirmed by LWR and MOS values, which remained less than 1.5 independently of time points. However, after 24 h, the Δefg1 grew as elongated yeast cells. These elongated cells additionally strongly adhered to each other, resulting in large clumps, which could resemble hyphal growth. This complicated the MOS and LWR analysis. For this reason, we focused on the analysis of 3 and 6 h. Moreover, according to percentage of cellular viability, both mutant strains are metabolically active and grow at these time points, although Δefg1 does so to a slightly lower extent than Δedt1 ( Fig. 3 ). This most likely stems from different growth rates of the two mutant strains. In the stationary phase (after 24 h), ATP amounts of the mutant strains were more equal again and even exceeded those of the wild-type strain, presumably due to an increased number of metabolically active yeast cells as compared with hyphal growth. Hence, the mutant strains can be used as references to screen for Y-H transition inhibitory compounds.
Assays for the identification of morphotype inhibitors have been described previously.13–15 These assays are dependent on fluorescent reporter strains based on the promoter of the hyphal wall protein HWP1, which is hypha specific. The tag was introduced downstream of the promoter either with green fluorescent protein (GFP) or beta-galactosidase enzyme (lacZ).13–15 Methods based on reporters, however, may also identify compounds that interfere directly with GFP or beta-galactosidase rather than influence filamentous growth. Moreover, it might be possible that upon activity of a potential compound, hyphal growth is blocked and the promoters are still active, giving rise to a false-negative signal. 15 In addition, the incubation time with 4 h is shorter than our analysis spanning from 3 h to 6 h for the identification of switching blockers and up to 24 h for the identification of fungicidal compounds. Thus, using the previously presented methods, the possibility remains that effects of the compounds are only temporarily or that the compounds are fungistatic at later time points, as it is the case for farnesol. 8
Our method is based on the type strain C. albicans SC5314 and importantly is applicable to any other wild-type strain from other fungal species. It is optimized for automated liquid handling using small volumes and can be carried out according to AFST guidelines. In this one-step screening approach, the positive hits are detected by means of LWR and MOS. An advantage of this microscopic assay compared with microplate reader assays is that actual image information from each calculation is stored and allows verification of different parameter measurements at later time points. As a second step, we tested positive-hit compounds from the primary screening for fungistatic/fungicidal activity by measuring cell viability via ATP levels. As demonstrated by the use of yeast-locked mutant C. albicans strains, this approach is suitable for distinguishing fungistatic or fungicidal compounds from morphotype-switching inhibitors ( Fig. 4 ). The Z′ factor was introduced as a valuable tool to validate the quality of screening assays. 11 A Z′ factor value between 0.5 and 1 defines the method as an excellent assay, suitable and valid for high-throughput screening. Notably, we found that the mean Z′ factor for LWR and MOS at 6 h was 0.513, confirming that our method is a highly suitable screening assay. All Z′ factor values were calculated from at least three biological replicates. At 3 h, the mean Z′ factor for LWR and MOS reached only a value of 0.1. These Z′ factor values define a smaller separation band at 3 h than at 6 h. Nevertheless, the trend is already confirmed at 3 h.
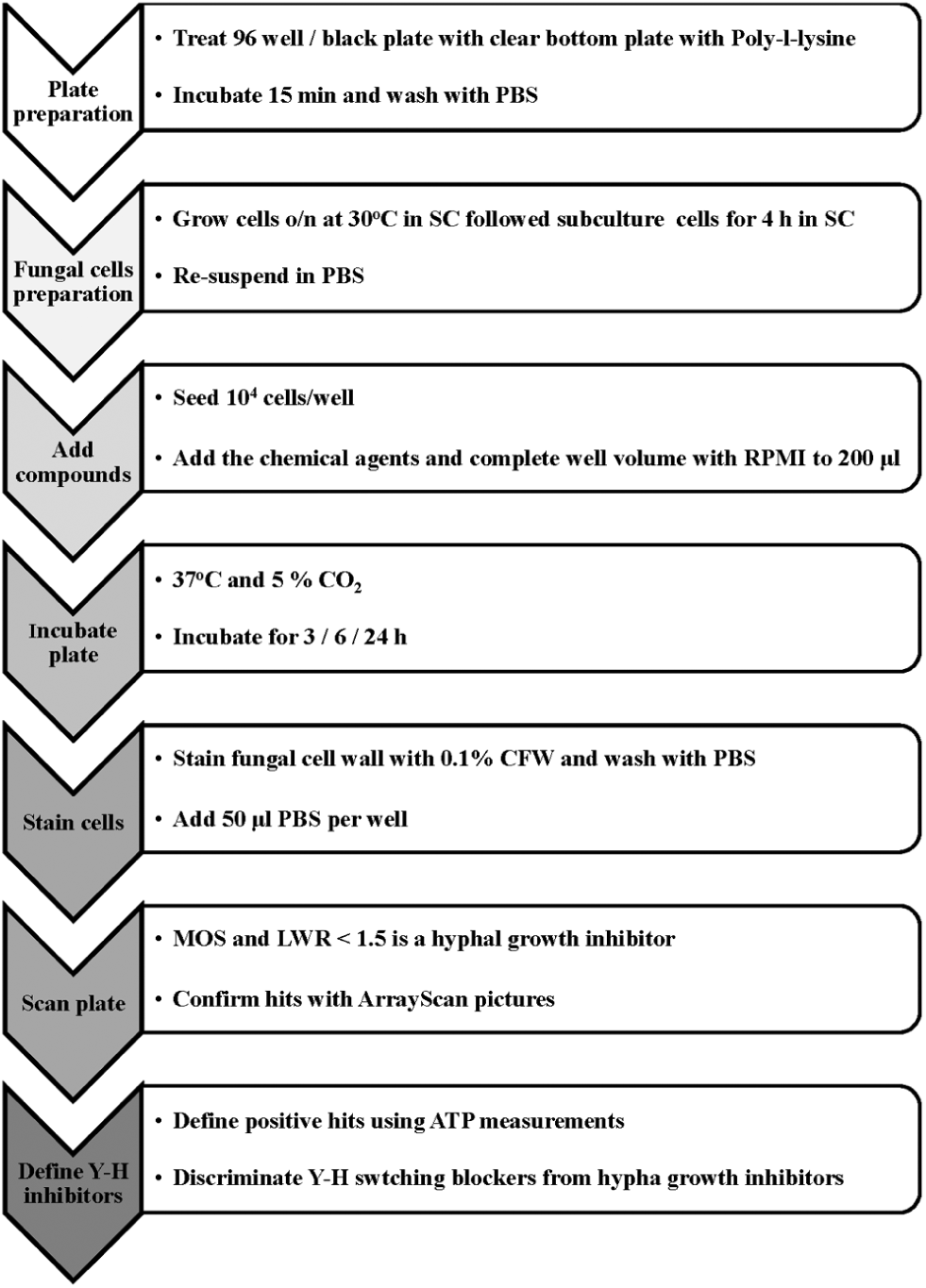
Schematic presentation of the suggested high-throughput screening method for identification of fungal dimorphism blockers.
In summary, we aimed to establish a high-throughput screening method to find compounds that break the Y-H switching. The identified compounds will have the potential to disarm the pathogen without disturbing the cellular viability, probably resulting in low selection pressure. A great advantage of our method is that it may serve as blueprint for screening with other polymorphic fungal pathogens, because wild-type strains without genetic modifications are applicable. Conclusively, our proposed method is a valuable tool for the identification of new and more efficient antimycotics.
Footnotes
Acknowledgements
We would like to thank Steffen Rupp and Robert Wheeler for kindly providing C. albicans strains.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants to C.F.U. from the Swedish Research Council VR-M (2011-2393), the Laboratory for Molecular Medicine Sweden (MIMS), the Åke Wiberg Foundation (3772734), and the Medial Faculty Umeå (316-886-10). M.S. acknowledges financial support from the J.C. Kempes Memorial Fund. Laboratories for Chemical Biology Umeå (LCBU) and Chemical Biology Consortium Sweden (CBCS) thank the Swedish Research Council, the Swedish Governmental Agency for Innovation Systems (VINNOVA), the Knut & Alice Wallenberg foundation, the Kempe foundations, and the Carl Trygger foundation for support. The funders had no role in the design or evaluation of the research.
