Abstract
Dengue virus (DENV) is the most significant mosquito-borne viral pathogen in the world and is the cause of dengue fever. The DENV RNA-dependent RNA polymerase (RdRp) is conserved among the four viral serotypes and is an attractive target for antiviral drug development. During initiation of viral RNA synthesis, the polymerase switches from a “closed” to “open” conformation to accommodate the viral RNA template. Inhibitors that lock the “closed” or block the “open” conformation would prevent viral RNA synthesis. Herein, we describe a screening campaign that employed two biochemical assays to identify inhibitors of RdRp initiation and elongation. Using a DENV subgenomic RNA template that promotes RdRp de novo initiation, the first assay measures cytosine nucleotide analogue (Atto-CTP) incorporation. Liberated Atto fluorophore allows for quantification of RdRp activity via fluorescence. The second assay uses the same RNA template but is label free and directly detects RdRp-mediated liberation of pyrophosphates of native ribonucleotides via liquid chromatography–mass spectrometry. The ability of inhibitors to bind and stabilize a “closed” conformation of the DENV RdRp was further assessed in a differential scanning fluorimetry assay. Last, active compounds were evaluated in a renilla luciferase–based DENV replicon cell-based assay to monitor cellular efficacy. All assays described herein are medium to high throughput, are robust and reproducible, and allow identification of inhibitors of the open and closed forms of DENV RNA polymerase.
Introduction
Dengue viruses (DENV) belong to the genus Flavivirus of the family Flaviviridae, and together, its four serotypes (DENV1–4) are responsible for the world’s most prevalent mosquito-borne viral disease. Approximately 3.6 billion people living in tropical and subtropical areas are at risk of dengue infection.1,2 Annually, there are about 390 million cases, of which 96 million infections exhibit disease symptoms. 3 These include a mild, self-limiting clinical infection; a more severe dengue hemorrhagic fever (DHF); or a life-threatening dengue shock syndrome (DSS). Unfortunately, neither effective vaccine nor treatment is available for DENV infection, and the World Health Organization lists dengue as a neglected tropical disease with an unmet medical need. While numerous candidate compounds that target either viral (capsid, envelope, NS2B/3 protease, NS3 helicase, NS4B, NS5 methyl-transferase, and polymerase) or host (translation and pyrimidine synthesis pathway, glucosidase) proteins have been reported, 4 none have as yet progressed into the clinic. Therefore, the development of safe and effective anti-DENV therapy is a pressing concern.
DENV is an enveloped RNA virus with a positive-sense RNA genome that encodes an approximately 3300–amino acid polyprotein chain, arranged in the order of C-prM-E–NS1–NS2A–NS2B–NS3–NS4A–NS4B–NS5. Three structural (C-prM-E) and seven nonstructural (NS1–5) proteins are generated by the actions of both viral (NS2B/3) and host proteases. 5 One of the key steps in the viral life cycle is viral RNA replication, which occurs in two steps: (1) generation of the negative polarity RNA, which serves as a template for (2) synthesis of multiple strands of RNAs of positive polarity. NS5 plays a key role in this RNA synthesis. It is the largest (900 amino acids) and most conserved protein in the Flavivirus genus. More than 65% of NS5 protein sequence is conserved across DENV1–4. It has several demonstrated enzymatic activities: N7 and 2′-O methyl transferase in its N-terminus (MTase, aa 1–2656,7; nucleotide terminal transferase [TNTase]) and RNA-dependent RNA polymerase (RdRp, aa 273–900; EC 2.7.7.48) activities located at the C-terminal domain.8,9 DENV NS5 RdRp performs both minus-strand RNA (replication) and plus-strand RNA (semi-conservative transcription) synthesis via a de novo initiation mechanism. It has been proposed that cyclization of the viral 5′-3′ untranslated regions (UTR) by long-range interactions is necessary for it to initiate de novo RNA synthesis10,11 (reviewed in Iglesias and Gamarnik 12 ). Following the first di-phosphatidyl ester formation, the protein is believed to undergo extensive conformational rearrangements from “closed” to a more “open” form, to accommodate the growing RNA strand.
Previous high-throughput screening (HTS) efforts in our laboratories to identify DENV NS5 inhibitors used either radioactive scintillation proximity 13 or fluorescent-coupled assays 14 that measured the elongation activity of the enzyme. Although several hits were rigorously followed up, we failed to identify specific DENV NS5 inhibitors with good physiochemical properties as starting points from these exercises. One limitation we observed was that the recombinant full-length (FL) DENV polymerase used in the prior screens exhibited poor stability, making the enzyme quite labile and sensitive to nonspecific inhibition. We observed a number of false positives that either aggregated the protein or interfered with the assay readouts. Another limitation was that the assays used were polymerase elongation assays (“open” conformation), which by their nature were unable to capture inhibitors of the “closed” form or initiation step of the polymerase. To overcome these issues and enable successful lead-finding campaigns, we generated new RdRp domain proteins that were more thermostable and catalytically active. 15 In this report, we describe the development of two biochemical HTS assays that are capable of identifying inhibitors of the DENV RdRp initiation and/or elongation phases and compared the inhibition of the FL NS5 and RdRp domain in one of the assays using a small pilot collection of compounds. Both assays employ a DENV subgenomic RNA bearing its complete 5′ and 3′ UTRs, linked via a short nonviral sequence. The first assay is a fluorescent coupled assay using a modified nucleotide analogue (Atto-CTP). The second assay is completely label free, using all four native nucleotides and measuring the pyrophosphate liberated during nucleotide incorporation via liquid chromatography–mass spectrometry (LCMS). We found an excellent correlation between the two assays using dideoxynucleotides and known DENV RdRp tool compound inhibitors. Both assays are robust, reproducible, and compatible for HTS campaigns and allow identification of inhibitors of the “open” and “closed” forms of the dengue RNA polymerase. Furthermore, identification of potential “closed”-form inhibitors was carried out through the use of differential scanning fluorimetry (DSF), which used an apo form of the RdRp (without RNA template), and assessed compound stabilization of the RdRp during a thermal ramp. Last, we used a cellular renilla luciferase–based replicon assay to monitor cell activity of the inhibitors. Several compounds were identified that are under further investigation and optimization as potential DENV RdRp initiation inhibitors.
Materials and Methods
Materials
Microtiter plates used in this study are as follows: 384-well, flat-bottom, black polystyrene, low-volume plates (cat. 3821; Corning Life Sciences, Lowell, MA) and 1536-well, flat-bottom, black polystyrene, high-base plates (cat. 782076; Greiner Bio-One, Monroe, NC) for the de novo fluorescence-based alkaline phosphatase-coupled polymerase assay (FAPA). The 384-well translucent V-bottom plates and small-volume V-bottom plates (cat. 781280 and cat. 784201, respectively; Greiner Bio-One) were used for the LCMS assay. The 384-well plates from Roche (cat. 04-729-749-001; Roche, Indianapolis, IN) were used for the DSF assay. De novo FAPA fluorescence and counterscreen assays were monitored using a BMG LabTech PHERAstar fs microplate reader (Ortenberg, Germany) and EnVision microplate reader (PerkinElmer, Waltham, MA). Differential scanning fluorimetry studies were carried out using a Bio-Rad (Hercules, CA) CFX384 RealTime-PCR Cycler and a fully automated system incorporating a Roche LightCycler 480 II instrument.
Reagents
Reagents for the de novo initiation and elongation biochemical assay were purchased from the following sources: adenosine-, cytosine-, guanosine-, and uridine-triphosphates; nicotinamide mononucleotide; DMSO; magnesium chloride; and purified water were from Sigma-Aldrich (St. Louis, MO). Manganese chloride, potassium chloride, and CHAPS detergent were from Thermo Fisher Scientific (Waltham, MA). Tris-HCl, sodium chloride, and Hepes were from Teknova (Hollister, CA). Calf intestinal phosphatase (CIP) was obtained from New England Biolabs (Ipswich, MA), while the AttoPhos alkaline phosphatase fluorescent substrate system and AttoPhos buffer were obtained from Promega (Madison, WI). Custom enzyme substrates (in vitro transcribed [IVT] DENV4 5′-UTR−3′-UTR RNA; Atto-CTP) and inhibitor dideoxynucleotides (2′3′ dideoxy-ATP, -CTP, -GTP, and -UTP) were all obtained from TriLink Biotechnologies (San Diego, CA). Sybr Gold and To-Pro-3 Iodide for RNA counterscreen studies and SYPRO Orange for the DSF studies were purchased from Invitrogen Life Technologies (Grand Island, NY). The growth media and plating media for the human A549 cell line containing the dengue replicon and renilla luciferase–based reporter were sourced from the following vendors: Hams F12 Medium (cat. 3176500; Life Technologies, Carlsbad, CA), Hams F12 Medium phenol red free (cat. M25450; Atlanta Biologicals, Flowery Branch, GA), fetal bovine serum (FBS; cat. SH30071.03; HyClone, Logan, UT); penicillin/streptomycin (cat. 15150-122; Life Technologies, Carlsbad, CA), and puromycin (cat. A11138-03; Life Technologies, Carlsbad, CA). FL DENV4 NS5 and RdRp were produced at Novartis as described below.
Cloning, Expression, and Purification of DENV4 NS5 Full-Length and RdRp Domain
Complementary DNAs (cDNAs) encoding the full-length NS5 and RdRp (aa 266–900) proteins from DENV4 (MY01-22713 strain) were cloned respectively into a pET28a vector or a modified pET28a vector comprising a PreScission protease cleavage site inserted downstream of the vector N-terminal His tag sequence as described previously.14,15 Protein expression and purification of DENV constructs from BL21 cells were also as described previously.14,15
Synthesis of DENV4 In Vitro Transcript RNA Template
The RNA substrate used for the de novo initiation/elongation assay comprised the DENV4 5′ UTR-core (nt 1–166) linked via a 19-nucleotide nonviral sequence to the 3′ UTR (nt 10264–10653). The entire cDNA sequence was joined by overlapping PCR and cloned downstream of the T7A2 promoter sequence. The RNA was generated via in vitro transcription using a T7 RiboMAX express large-scale RNA production system from Promega.
Compound Collections
The Novartis compounds were provided as 2-mM stocks from the Novartis Sample Management Group, previously stored at 4 °C under 20% relative humidity in a 90% DMSO/10% water mixture in cyclic olefin copolymer plates to avoid freeze-thaw cycles and water uptake. The initial pilot set of structurally diverse compounds for comparison of the DENV4 NS5 FL and RdRp proteins, consisting of 9430 compounds, was provided in 384-well assay-ready plates containing 150 nL prespotted compound (to yield 20 µM final compound concentration in the assay). For the primary HTS campaign, we built a screening set of ~257,000 unique compounds. The main part of the compound set was the “biodiversity set,” a selection of screening plates optimized for chemical and biological activity. 16 Briefly, each plate in the Novartis compound collection was scored and ranked based on the number of annotated biological targets covered by the compounds on the plates. To also increase the chemical diversity of the compounds on plates, redundant scaffolds were eliminated from subsequent scoring of plates lower in the list. The top-ranked plates covering roughly 250,000 compounds were chosen for screening. To complement the biodiversity set, a hypothesis-driven subset of compounds with biological activity annotations <10 µM for past antiviral targets (same organism) or polymerases (same target family) was built in silico. After eliminating compounds already present on the biodiversity set, ~3800 compounds were cherry-picked by the Novartis Sample Management Group and plated for screening. Finally, a Novartis proprietary set of compounds with known, diverse mechanisms of action (~3000 compounds) and the Novartis collection of natural products (~12,000 compounds) were added to the screening set. Compounds for testing were provided as compound source plates for delivery to the 1536-well enzyme assay plates as 50 nL droplets (to yield 25 µM final compound concentration).
Design of Experiment for Assay Optimization
Design of experiment (DoE) technology was used to identify the optimal buffer composition of the enzymatic reaction for the RNA polymerase. Specific details on the DoE methodology can be found in the
DENV Polymerase De Novo Initiation and Elongation Assays for HTS
Counterscreen Assays
Three separate counterscreen assays were developed to identify compounds that would either inhibit the calf intestinal phosphatase (CIP) enzyme or disrupt the RNA substrate. The CIP counterscreen was developed by conversion of AttoPhos substrate BBTP to BBT, which was the end product generated in the DENV FAPA. For RNA disruption, two RNA competition assays were developed to eliminate false-positive inhibitors affecting the RNA itself. The first assay used TO-PRO3 iodide stain, which is a carbocyanine monomer nucleic acid with far-red fluorescence to identify molecules that may interact with RNA by covalent or electrostatic binding, or intercalation, while the second assay used SYBR Gold, an unsymmetrical cyanine dye, to identify double-stranded RNA binders that prevent the release of RNA from the template in the DENV de novo FAPA. All compounds were tested in duplicate at 25 µM concentration, under the assay conditions described in the
Differential Scanning Fluorimetry
Differential scanning fluorimetry was used as a method to identify compounds that bind to the RdRp in the absence of RNA and stabilize the protein from thermal unfolding. Screening of compounds was carried out using a fully automated DSF system under the following assay conditions: 100 µg/mL DENV4 RdRp, 5× SYPRO Orange dye, 20 mM Hepes (pH 7.5), 1 mM TCEP, 150 mM NaCl, 2 mM MgCl2, and 2 % DMSO. The final compound concentration evaluated was 100 µM. To carry out the experiment, 5 µL RdRp protein (stock at 200 µg/mL) and 5 µL of 10× SYPRO Orange dye stock (5000× concentrate in DMSO; Life Technologies) were dispensed into an assay plate (Bio-Rad; HSR-4805) containing 500 nL of compound. The final assay volume was 10.5 µL per well in a 384-well format. Plates were then sealed after reagent addition, mixed for 3 minutes, centrifuged at 1000 rpm for 1 min (VSpin; Velocity11 [Thermo Fisher Scientific, Waltham, MA]), and read on a Roche LightCycler 480 II instrument using an excitation of 465 nm and an emission at 580 nm. The temperature was ramped from 25 °C to 75 °C at 0.11 °C/s. The melting temperature Tm of the raw fluorescence data was identified as the midpoint of the transitions via a semi-parametric fit. The ΔTm was determined by comparing the individual Tm values with the mean Tm apo RdRp protein controls containing DMSO only. Hits from the initial DSF assay were then confirmed using the assay conditions described above with the following changes: the final assay volume was 10 µL per well in 384-well white microtiter plates (cat. 04-729-749-001; Roche), plates were sealed after reagent addition, and readout was on a Bio-Rad CFX384 RealTime-PCR Cycler (filter: fluorescence resonance energy transfer [FRET]; excitation: 450–490 nm; detection: 560–580 nm). The temperature ramp reads were at 0.5 °C increments using a ramp rate of 1 °C per minute from 4 °C to 75 °C. The melting temperature Tm of the raw fluorescence data was identified as the midpoint of the transitions via a Boltzmann fit. The ΔTm was determined by comparing the individual Tm values with the mean Tm of 16 protein controls with DMSO only.
Dengue Replicon Activity and Cytotoxicity Determinations
The activity of compounds against a human A549 cell line containing a Dengue renilla luciferase–based reporter (Rluc) replicon was assessed along with corresponding cytotoxicity measurements as described previously in our laboratory, 17 with the following modifications: DENV replicon A549 cells containing Rluc reporter were cultured in growth media (Hams F12 medium with 10% FBS, penicillin and streptomycin, and 5 µg/mL puromycin) and changed to plating media (Hams F12 medium, phenol-red free with 2% FBS, and penicillin and streptomycin) when ready for plating. Cells were plated at a density of 750 cells per well in 1536-well plates (Corning tissue culture–treated, white polystyrene plates) and incubated for 20 to 24 h with 5% CO2. The compound was then added in 2-fold serial dilutions in the range of 0.234 to 30 µM and allowed to incubate for 24 h. For Rluc-replicon cells, ViviRen (Promega) was added at 10 µM and incubated for 10 min at 22 °C before the luminescence was read on a ViewLux reader (PerkinElmer) with 20-s exposures. Luminescence was plotted against the log transformation of the concentration of the compound, and a curve fit with variable slope was done to obtain the effective concentration (EC50) value using the Novartis proprietary data evaluation software Helios. Using the same range of compound concentration and ViewLux, cell viability was also measured in the DENV replicon cells after compound exposure using a CellTiter-Glo luminescence cell viability assay (Promega) for adenosine triphosphate (ATP) detection. Data were analyzed as above to determine the cytotoxicity concentration (CC50).
Data Analysis
The Novartis proprietary data evaluation software Helios was used to analyze the primary screening data, validation concentration-response curves, and cell-based data. For LCMS analyses, to determine the abundance of pyrophosphate in every LCMS reaction, data were processed through a commercially available data analysis tool (Gubbs Software; Gubbs Inc., Alpharetta, GA). Melting temperatures (Tm) and several other DSF curve characteristics were derived in a fully automated way by Novartis proprietary data evaluation software. Specific details on all of these data analysis tools and their utilization can be found in the
Results
DENV4 Polymerase De Novo Initiation and Elongation Assay Optimization for Screening
To identify inhibitors of the closed and open conformations of the DENV RNA polymerase, both fluorescent-based and label-free LCMS detection assays were developed. The de novo FAPA was designed and adapted from the DENV primer extension assays described previously,13,14 but unlike the earlier elongation assays using poly-U or poly-C RNA templates with exogenous primers, the assay described within this report carried out de novo initiation and elongation. An outline of the entire screening campaign and associated assays is shown in Figure 1 .
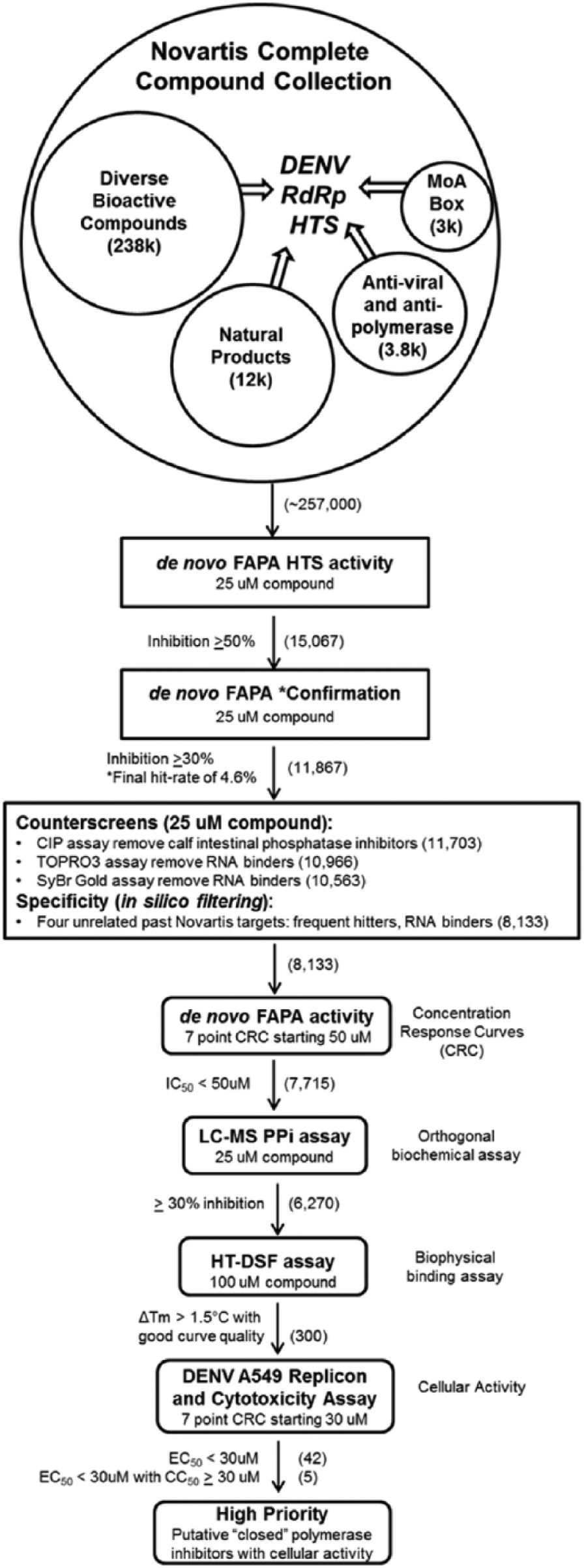
Screening campaign outline of the de novo fluorescence-based alkaline phosphatase-coupled polymerase assay (FAPA) primary high-throughput screening (HTS), liquid chromatography–mass spectrometry (LCMS) orthogonal assay, differential scanning fluorimetry (DSF) biophysical assay, and A549 dengue virus (DENV) replicon assay used to validate hits of the “closed” polymerase. The compound library chosen for the focused high-throughput screen represents a knowledge-based subset of the larger Novartis compound collection and contains a diverse bioactive component, as well as purified natural products, hits from past antiviral and antipolymerase screens, and compounds with known mechanism of action (MoA). The primary HTS assay used a coupled enzyme fluorescent detection format that measured the increase of fluorescence of the Atto-CTP substrate as an indirect measure of RNA-dependent RNA polymerase (RdRp) activity. The orthogonal assay used LCMS detection to measure the ratio of the area for the pyrophosphate (PPi) product ions and the internal standard NMN (nicotinamide mononucleotide) as a direct measure of RdRp activity. The biophysical DSF assay identified compounds that bound to and stabilized the apo-RdRp after a thermal ramp, while the cellular assay identified those compounds that inhibit a renilla luciferase DENV replicon in A549 cells. Compounds remaining at each step of the flowchart are indicated in parentheses to the right of each filter.
The top buffer conditions for the enzymatic assay were identified using design of experiment (
Comparison of DENV Full-Length and RdRp Construct Inhibition in the De Novo FAPA
At the onset of this screening effort, we sought to generate DENV RdRp constructs that were more stable and amenable to prolonged incubation at room temperature, without significant loss in activity. Addition of amino acid residues from the NS5 interdomain linker sequence was found to both increase the stability of and enhance DENV1–4 RdRp activities. 15 These residues facilitated enzyme turnover of the RNA and nucleoside triphosphate substrates during its de novo initiation/elongation process.
To assess if this longer RdRp protein construct would be more suitable for an HTS campaign compared with full-length (FL) NS5 protein, we used a small test set of structurally diverse compounds (9430 compounds; Lib2012D) from the Novartis library. We concurrently ran Lib2012D against both DENV4 FL NS5 protein as well as DENV4 RdRp (aa 266–900) protein in the de novo FAPA. Both runs were performed in a continuous manner; furthermore, compounds in both runs were exposed to the protein alone for at least 20 min followed by reaction initiation with RNA and NTPs. The compounds were screened at a final concentration of 20 µM and the Z′ factor determined as described. The results of the comparison are summarized in
Primary HTS Campaign to Identify Compounds Inhibiting DENV4 RdRp
For the primary HTS campaign, a total of ~257,000 compounds optimized for chemical and biological diversity
16
were obtained from the Novartis Sample Management Group and tested against DENV4 RdRp using the de novo FAPA as the primary screening assay at a final compound concentration of 25 µM. The screen was run with good assay quality, with an average Z′ > 0.7 for all plates tested. In total, 15,067 primary hits were selected at the ≥50% inhibition cutoff and were cherry-picked for confirmation and subsequent counterscreen assays in duplicate. Compounds were considered confirmed if they showed ≥30% inhibition upon retesting at 25 µM in duplicate (3 standard deviations of the control data was 19%), and we found 11,867 compounds confirmed (79%) upon testing (
Counterscreen Analysis of Compounds Inhibiting DENV RdRp
Compounds in the de novo FAPA primary assay could yield false positives due to inhibition of the CIP enzyme used in the FAPA coupled reaction, disruption of the RNA substrate, or sample contamination with inorganic impurities and other reactive chemical species. 19 Thus, three counterscreen assays and in silico filters were developed to identify and remove those compounds from further follow-up.
The primary hits from the DENV de novo FAPA were tested at 25 µM concentration as duplicates in the CIP assay and the two RNA competition assays to flag false-positive hits. All assays performed well, and duplicate runs showed excellent correlation between sets (data not shown), with overall Z′ value for all assays >0.7. Therefore, the average activity from two data sets was used for filtering compounds. Surprisingly, only 1.4% of the hits were active in the CIP enzyme assay. A larger number of inhibitors were active in the two RNA disruption counterscreens of To-Pro-3 and SYBR Gold (6.3% and 3.7%, respectively). Finally, an in silico comparative approach was used to eliminate compounds from the DENV polymerase FAPA hit list that were shown to have high activities in several completely unrelated Novartis HTS assays that had been carried out previously. This approach identified 2430 suspect compounds. Thus, at the end of the confirmation, counterscreens, and in silico analyses, a total of 8133 hits showing activity in the DENV de novo FAPA and passing the various filters were picked for concentration-response curves ( Fig. 1 ) and for subsequent evaluation in the orthogonal LCMS assay of de novo initiation and elongation activity. Concentration-response curves were generated on these hits using seven-point (2-fold) dilution series with a starting concentration of 50 µM in the FAPA.
LCMS Orthogonal Analysis of Compounds Inhibiting DENV RdRp
The LCMS detection of the pyrophosphate assay used the identical enzyme assay conditions as the de novo FAPA with the exception of 20 µM cytosine triphosphate (CTP) as a replacement to the 5 µM Atto-CTP. Enzyme reactions were run for 2 h at room temperature before stopping the reactions and adding internal standard, and the volume of the reaction was designed to enable multiple injections for evaluation on the instrumentation. An overview of the LCMS pyrophosphate detection assay and a typical injection trace are shown in
Figure 2
. To initially evaluate the performance and sensitivity of the LCMS assay compared with the DENV de novo FAPA prior to evaluation of the confirmed and counterscreened primary screen hits, the same tool compounds that were used to evaluate the FAPA were evaluated in the LCMS assay. Compounds were tested in concentration-response curves in the range of 25 µM to 14 nM in a 12-point dose response. IC50 values were calculated and compared with those obtained in the DENV de novo FAPA. With the exception of the 3′-dCTP, IC50 values between the two assays were ≤3-fold of one another (
Fig. 3
). The somewhat higher IC50 found in the LCMS assay for 3′-dCTP is likely due to the higher concentration of that nucleotide found in the LCMS assay (20 µM) compared with the de novo FAPA. Thus, given the very similar IC50 values obtained between the two assays and the good Z′ values obtained on the DMSO control plates, the orthogonal LCMS assay was then used to screen the 8133 compounds that were confirmed from the FAPA by assessing them in duplicate at a single compound concentration of 25 µM. Average Z′ values for all plates after duplicate runs were determined to be 0.68, and the data obtained from the LCMS replicates were very similar to each other (
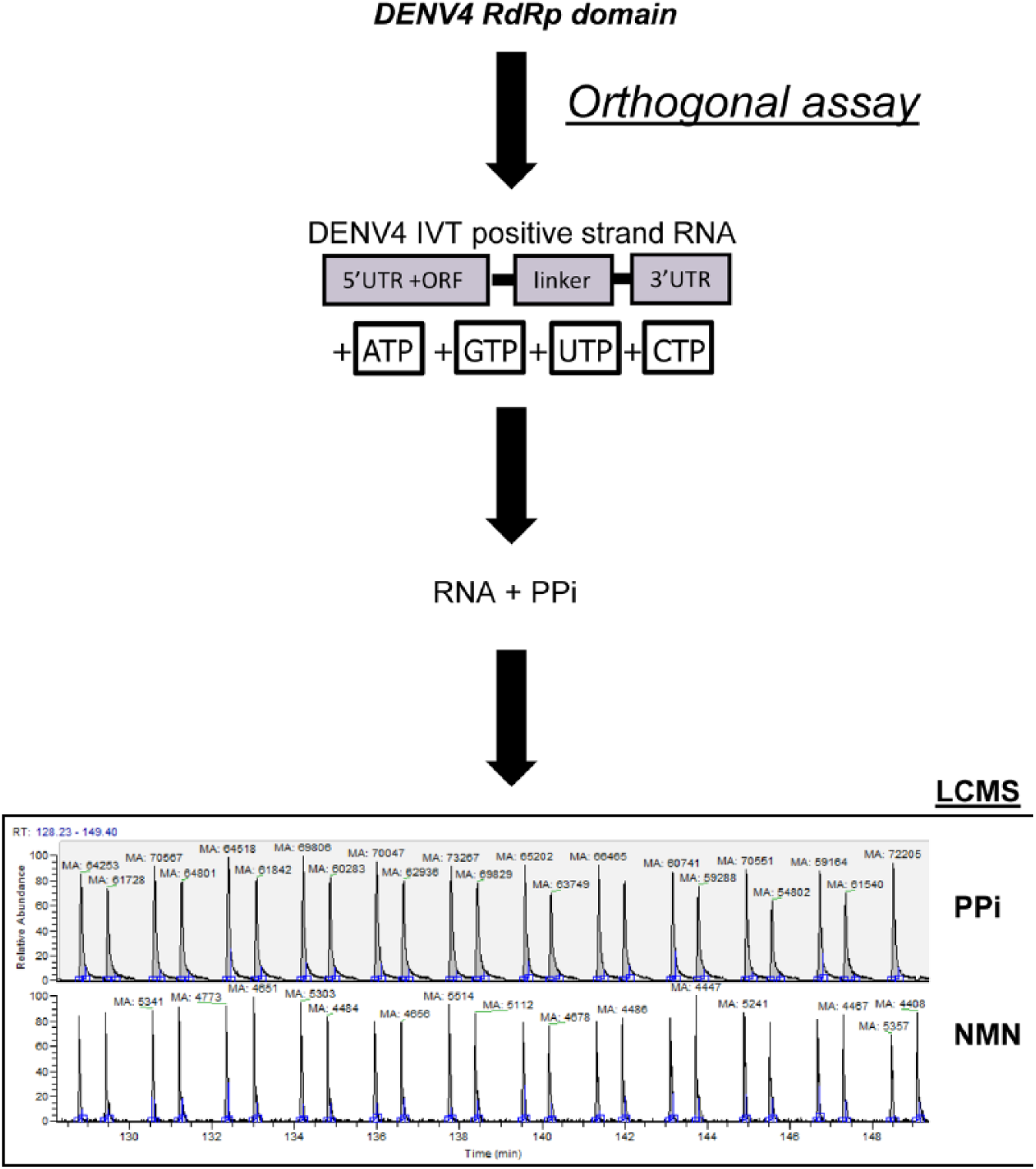
Outline of the orthogonal de novo liquid chromatography–mass spectrometry (LCMS) pyrophosphate detection assay used to measure DENV4 RNA-dependent RNA polymerase enzyme activity. Using the same DENV4 RNA-dependent RNA polymerase (RdRp) and in vitro transcribed subgenomic RNA substrate as the de novo fluorescence-based alkaline phosphatase-coupled polymerase assay (FAPA) primary high-throughput screening (HTS) assay, the orthogonal assay used LCMS detection to measure the ratio of the area for the pyrophosphate (PPi) product ions and the internal standard NMN as a direct measure of RdRp activity. An example of PPi abundance (red trace m/z = 176.9) from DENV4 RdRp control reactions compared with internal standard NMN (green trace m/z = 211.0) is shown along with the reproducibility of the analyte/internal standard injections.
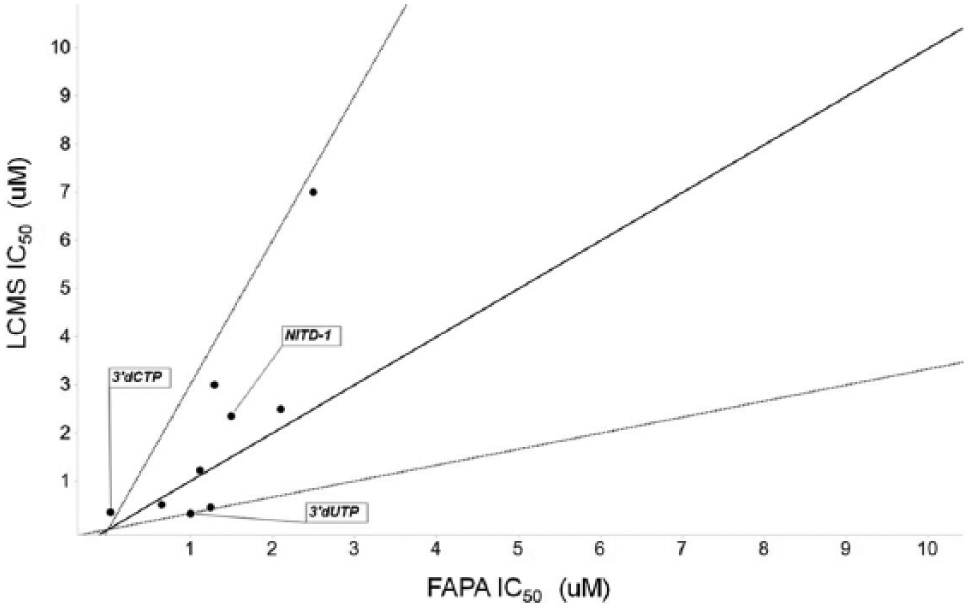
Performance of eight tool compounds and two reference dideoxynucleotide inhibitors in the DENV4 fluorescence-based alkaline phosphatase-coupled polymerase assay (FAPA) and liquid chromatography–mass spectrometry (LCMS) detection assay. The diagonal solid line represents y = x, while the two dotted lines represent ±3 fold change in IC50s. One tool compound was inactive (IC50 > 50 µM) in both assays (not shown). The tool compound labeled NITD-1 is a previously reported inhibitor that blocks the RNA-dependent RNA polymerase (RdRp) activity through binding to the RNA template tunnel of the polymerase. 13
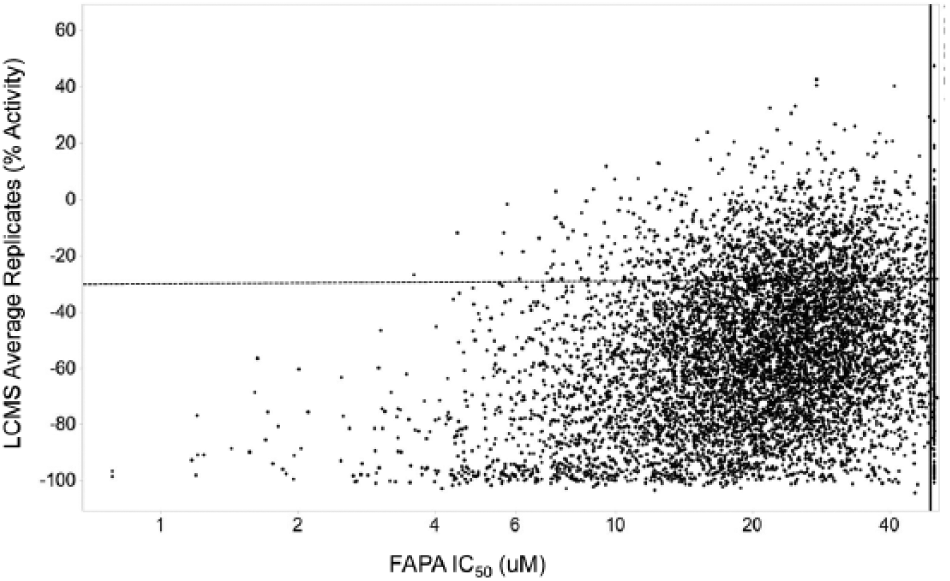
Comparison of average liquid chromatography–mass spectrometry (LCMS) data with fluorescence-based alkaline phosphatase-coupled polymerase assay (FAPA) IC50 determinations. Averages of the LCMS replicate data and FAPA IC50 data were compared with each other to identify overlapping inhibitors. A −30% activity was chosen for the cutoff of hit calling for the LCMS assay (reflected by the dashed horizontal line). Compounds with IC50 <50 µM are to the left of the solid vertical line. Compounds that failed the IC50 determinations in the FAPA (50 µM) are shown at the far right of the x-axis.
DSF of Compounds Binding to DENV RdRp
To identify compounds from the HTS campaign that could bind to the “closed” or initiation form of the RdRp, we used a DSF temperature ramp with the RdRp + inhibitor + SYPRO fluorescent dye and monitored the extrinsic fluorescence as a function of time. As the dye’s emission properties changed during the thermally induced unfolding, the observed Tm parameter was used to measure the RdRp conformational stability in the presence of the inhibitor. Inhibitors that bind the “closed” form of the polymerase have the potential to stabilize this conformation. Profiling the 8133 compounds through the DSF assay identified 300 compounds (approximately 3.7% of the hit list) with a significant thermal shift of ≥1.5 °C ( Fig. 5 ) that were also positive in the LCMS assay (≥30% inhibition) and exhibited de novo FAPA IC50 <50 µM. A typical Tm curve of a stabilizing compound is shown in the inset to Figure 5 .
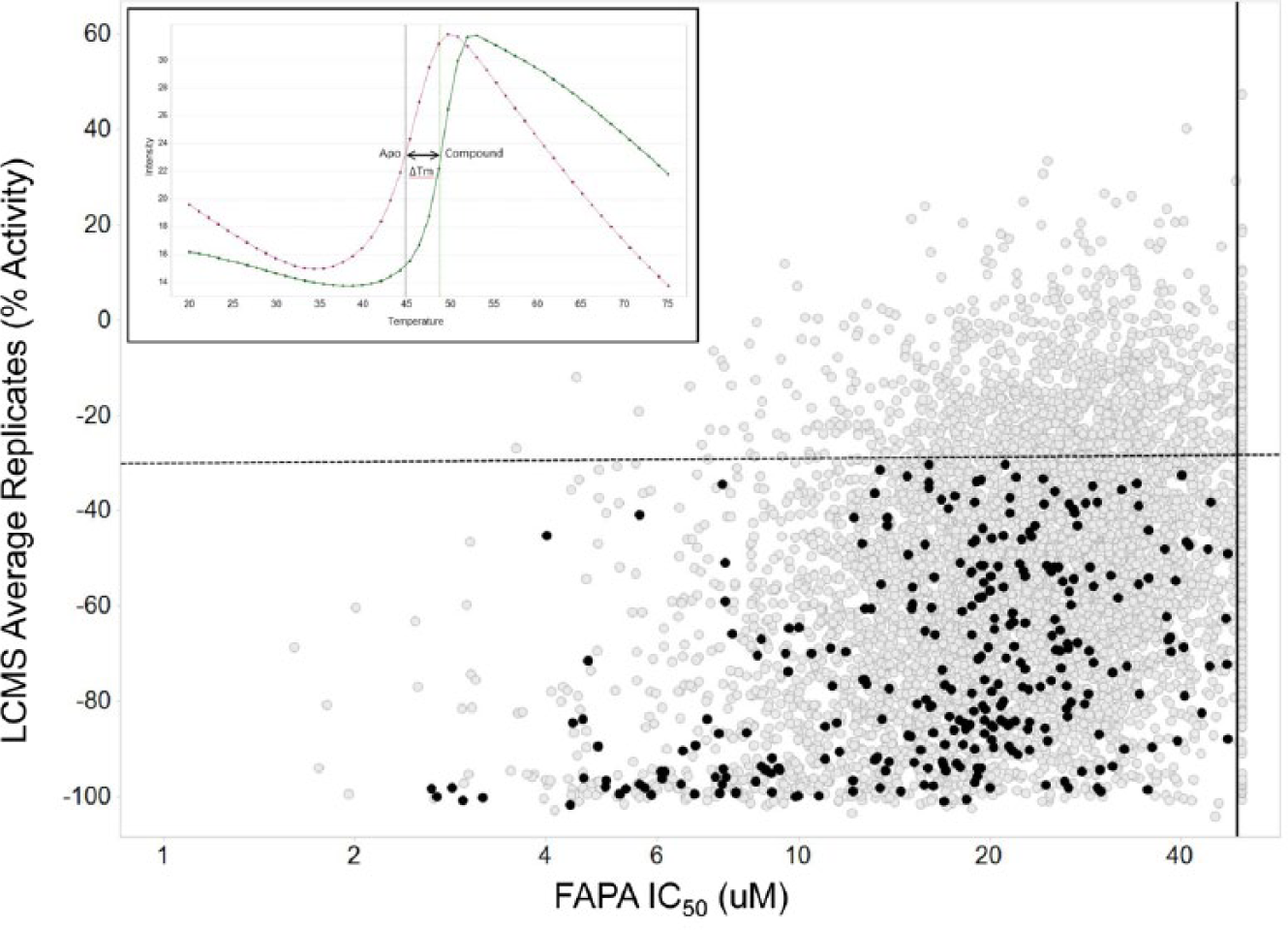
Evaluation of de novo initiation inhibitors by differential scanning fluorimetry. Compounds (gray circles) were assessed for thermal stabilization of the RNA-dependent RNA polymerase (RdRp), and those 300 compounds showing ≥1.5 °C Tm shift, fluorescence-based alkaline phosphatase-coupled polymerase assay (FAPA) activity with IC50 <50 µM (solid vertical line), and liquid chromatography–mass spectrometry (LCMS) activity of ≥30% inhibition (dashed horizontal line) of pyrophosphate generation are highlighted in black. The inset shows a typical Tm shift for RdRp protein with and without a validated compound showing a positive ΔTm of 3.89 °C.
Inhibition of the DENV Replicon Replication and Cytoxicity Determination
Next, DSF-active compounds were tested in the DENV Rluc replicon cell-based assay to determine if they were cellularly active. Compounds were also concurrently checked in this cell line with a commercially available CellTiter-Glo (CTG) assay for an indirect measure of cellular viability (assessed by cellular ATP levels). As shown in Figure 6 , of the 300 compounds assessed for cellular activity, 42 compounds were active in the Rluc replicon assay. In addition, many of these compounds showed some impact on cellular viability as determined in the CTG assay. A total of 42 compounds from the 300 were active in the DENV Rluc replicon assay with EC50s <30 µM, while five of these 42 also showed CC50 ≥30 µM, and one showed EC50s ≤5 µM with >4-fold window of CC50 (highlighted in black, Fig. 6 ).
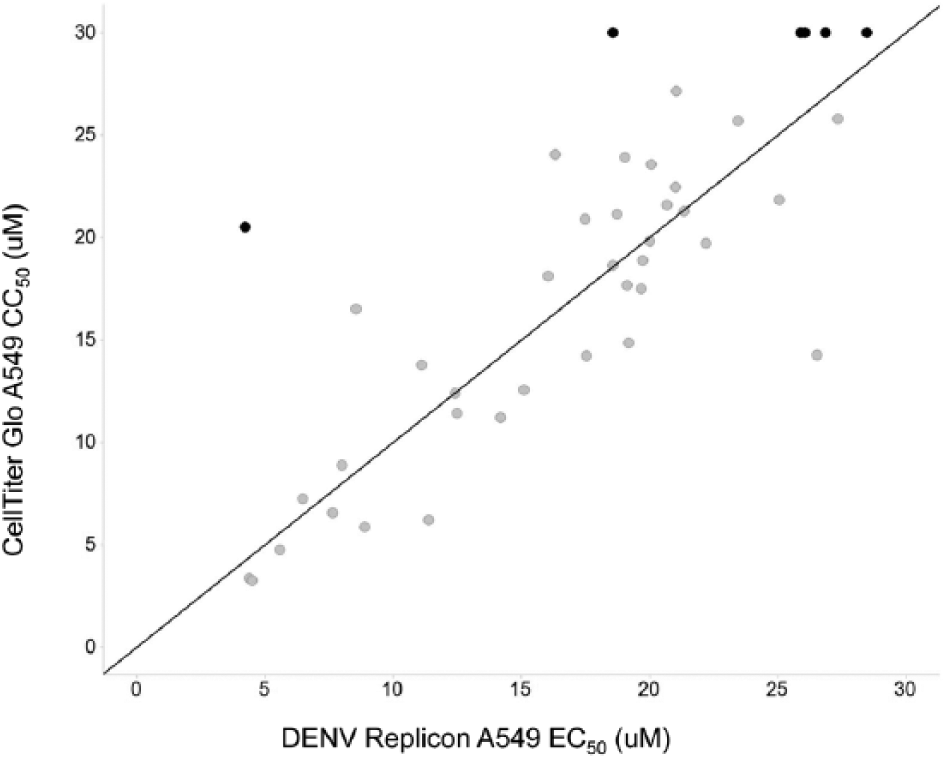
Evaluation of de novo initiation inhibitors using a dengue virus (DENV) replicon assay in A549 cells. Compounds (gray circles) were assessed for DENV replicon inhibition using ViviRen substrate and also cell cytotoxicity using CellTiter-Glo (Promega, Madison, WI). Five compounds that showed DENV replicon EC50 <30 µM with a corresponding CC50 ≥30 µM and DENV replicon EC50 <5 µM with a ≥4-fold window over CC50 are highlighted in black. Concentration-response curves were generated using two-point compound dilutions starting with a 30-µM upper concentration.
Discussion
Being essential to the successful replication of many infectious viruses, viral polymerases are well-validated and proven drug targets, and as such, there are many drugs currently on the market targeting the polymerases of hepatitis C virus, herpes viruses, and human immunodeficiency virus (reviewed in Berdis, 20 Tsai et al., 21 and Oberg 22 ). The absence of RdRp activity in host cells further makes the polymerase an attractive target for anti-DENV intervention. Unfortunately, none of the recently reported inhibitors of DENV NS5 polymerase have been successfully developed into drugs or proven themselves efficacious in patients with DENV infection.4,13,23 Our past efforts at targeting the RdRp used polymerase elongation HTS assays, either in a radioactive scintillation proximity format 13 or a fluorescence-based format. 14 Both approaches used short synthetic poly(C) or poly(U) RNA templates that limited the ability to find inhibitors of alternative conformations (i.e., “closed”) of the polymerase. In addition, in both prior screening campaigns, it was necessary to expose compounds to a precomplex of the protein and the RNA substrate, which helped to retain protein activity. We speculated that this protein-RNA complex altered the protein conformation and rendered sites in the protein inaccessible to compound binding. Thus, our goal was to implement a new approach by using a stable RdRp domain and a de novo initiation and elongation assay that contains critical elements within the 5′ and 3′ terminal regions of the DENV RNA template. This allows both de novo RNA initiation and elongation to occur 24 and captures both “open” and “closed” conformations of the DENV RdRp during RNA synthesis.
During the initiation phase of RNA synthesis, the RdRp domain specifically binds a promoter element, a large stem-loop structure named SLA (stem-loop A) present at the 5′ end of the DENV genome. It then reaches the site of initiation at the 3′ end recruited to the 5′ end via long-range RNA-RNA interactions (reviewed in Iglesias and Gamarnik 12 and Malet et al. 25 ). As a result, the active site of the polymerase switches from the more rigid “closed” form, catalyzing de novo initiation through synthesis of short primers (up to four nucleotides), to a more flexible “open” form for elongation. Such conformational switches allow the single-stranded RNA template to enter the template tunnel and the newly synthesized RNA (together with the template strand) to exit the RNA output channel.13,25 Since the de novo initiation and elongation assay in the present study captures both conformations of the polymerase, it is likely that inhibitors found will act through either blocking the binding of RdRp to the key RNA elements required for initiation, interfering with the conformational changes of the RdRp from the “closed” to “open” form, or blocking elongation of the nucleotide chain.
The de novo FAPA that we employed for hit discovery in this report is an indirect assay in that the detected fluorescent product arose through the action of a coupled alkaline phosphatase enzyme in the reaction. As such, false positives could be expected to be present in the hit list due to the presence of this second essential enzyme as well as due to simple disruption of the RNA template. Hence, we incorporated a phosphatase counterassay and RNA-binding assays to determine the false-positive hits in the HTS campaign and eliminate them from further consideration.
The DENV RdRp de novo initiation and elongation assay was also developed as an orthogonal LCMS assay to the FAPA. LCMS detection of the products carried out in a medium-throughput manner allowed significant numbers of compounds to be processed in a label-free, non–fluorescence-based assay and further validated those compounds that were found active in the fluorescence-based format. Most of those compounds that were hits in the LCMS orthogonal assay (i.e., showing ≥30% inhibition of the RdRp) were active in the FAPA with IC50 <50 µM.
With respect to compound binding to RdRp, we were able to identify 300 compounds from the validated HTS hits that significantly stabilized the RdRp in a differential scanning fluorimetry assay (i.e., thermal shift of ≥1.5 °C). DSF is a relatively inexpensive and lower throughput screening method to identify small molecules that bind to and stabilize purified proteins from thermal denaturation. 26 As these compound bind the apo “closed” form of the RdRp, they may act by blocking the initiation step of the polymerase. It is worth noting that many of the identified inhibitors do not bind the Apo-RdRp enzyme (i.e., “closed” form) in the DSF assay, despite being preincubated with the closed RdRp form prior to reaction initiation with RNA and nucleotide substrates. Whether the closed DENV RdRp presents a conformation less amenable for inhibitors to bind compared with the open form is uncertain, although it is known that Flaviviridae RdRps in a closed conformation possess an active site that is completely enclosed, with two channels, the RNA template binding channel and a second NTP channel, leading to the active site. 27 Thus, Flaviviridae RdRps such as those from DENV that carry out de novo initiation have restricted access to the RNA-binding channel during initiation, allowing only single-stranded RNA to enter the active site. 28 In contrast, RdRps that initiate RNA synthesis via a primer-dependent manner, such as those from the Picornaviridae family, have a wider template-binding channel that can accommodate a larger, template-primer initiation complex, 27 perhaps as a consequence allowing better access for compounds to key inhibitory sites.
Last, we have shown that 42 of the 300 active compounds show cellular activity in a A549 dengue renilla luciferase replicon assay (EC50 <30 µM). Although many of these compounds are cytotoxic, it is not known whether this represents off-target effects in the cell in addition to on-target effects against the NS5 enzyme. Since crystal structures of the DENV3 NS5 RNA-dependent RNA polymerase structures have recently been determined, both in apo and nucleotide-complexed forms, 29 as well as in complex with inhibitors identified from past screening efforts, 30 structural validation studies are ongoing with these unique inhibitors to further define their mechanisms of inhibition. These advances, along with additional structural biology efforts on the DENV1, DENV2, and DENV4 serotypes, should greatly support efforts toward rational drug design and inhibitor development against dengue virus.
Footnotes
Acknowledgements
We thank Chang Bok Lee, Ketan Patel, Chin Chin Lim, Alex Theodore Chao, Hanspeter Gubler, Sandra Kaestner, and Johannes Ottl for technical assistance; Justin Oborski and Chris Hogan for cell culture support; James Koehn and Arunkumar Dubey for DENV protein supply; Paul Smith and Adam Hill for useful discussions and critical review; and Sylvain Cottens and Thierry Diagana for support of this project.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
