Abstract
The hepatitis C virus (HCV) RNA-dependent RNA polymerase (RdRp) plays an essential role in the replication of HCV and is a key target for novel antiviral therapies. Several RdRp inhibitors are in clinical trials and have increased response rates when combined with current interferon-based therapies for genotype 1 (G1) HCV patients. These inhibitors, however, show poor efficacy against non-G1 genotypes, including G3a, which represents ~20% of HCV cases globally. Here, we used a commercially available fluorescent dye to characterize G3a HCV RdRp in vitro. RdRp activity was assessed via synthesis of double-stranded RNA from the single-stranded RNA poly(C) template. The assay was miniaturized to a 384-well microplate format and a pilot high-throughput screen was conducted using 10,208 “lead-like” compounds, randomly selected to identify inhibitors of HCV G3a RdRp. Of 150 compounds demonstrating greatest inhibition, 10 were confirmed using both fluorescent and radioactive assays. The top two inhibitors (HAC001 and HAC002) demonstrated specific activity, with an IC50 of 12.7 µM and 1.0 µM, respectively. In conclusion, we describe simple, fluorescent-based high-throughput screening (HTS) for the identification of inhibitors of de novo RdRp activity, using HCV G3a RdRp as the target. The HTS system could be used against any positive-sense RNA virus that cannot be cultured.
Introduction
Hepatitis C virus (HCV) infection is a major cause of liver cirrhosis, end-stage liver disease, and hepatocellular carcinoma. 1 Currently, there are an estimated 170 million people infected with the virus, which causes a chronic infection in nearly 70% of patients.1,2 There is no vaccine for HCV, and current treatment regimens rely on a combination of pegylated interferon α (PEG-IFN-α) and ribavirin, which are expensive and accompanied by adverse effects. Two protease inhibitors, telaprevir and boceprevir, have recently been approved in Europe and the United States for the treatment of patients infected with HCV genotype 1. When administered in combination with the existing standard of care (PEG-IFN-α and ribavirin), sustained virologic response rates have been shown to increase by around 20% to 30%. 3
HCV nonstructural protein 5B (NS5B) is an RNA-dependent RNA polymerase (RdRp) of 66 kDa that exhibits a high mutation rate of approximately 10−4 substitutions per site. 4 The high mutation rate contributes to the natural genetic diversity observed in HCV, with the virus classified into six distinct genotypes (G1–G6), which differ across the 9.6-kb genome by approximately 35% at the nucleotide level. Genotypes can be further classified into subtypes, which differ by 20% to 25%. Within infected hepatocytes, the viral RdRp replicates the viral positive-sense RNA genome via a negative-sense intermediate. The new strand serves as both a template for protein synthesis and as nascent genomes for packaging into new virus particles.With such a fundamental role in the viral life cycle, the HCV RdRp has become a prime target for the development of direct-acting antivirals (DAAs). DAAs targeting the HCV RdRp are divided into two classes. First, nucleoside inhibitors (NI) are substrate analogues that cause premature termination during synthesis of new RNA molecules. Second, nonnucleoside small-molecule inhibitors (NNIs) bind allosterically to the RdRp and inhibit conformational changes required for initiation or elongation of nascent RNA molecules. 5
High-throughput screening (HTS) campaigns involving recombinant HCV G1 enzyme have been successful in the identification of several NNIs6–8 for HCV G1, nine of which have reached phase II clinical trials. 9 These NNIs bind to one of four allosteric sites within the RdRp; two sites are located in the thumb domain and two in the palm domain of the HCV RdRp. A major advancement in the search for antiviral agents was the development of G1b subgenomic RNA replicons that replicate without producing infectious virus particles. 10 This was followed in 2005 by an infectious cell culture system (HCVcc) from a specific G2a strain (JFH-1). 11 Downstream selection and characterization of NNIs using these model systems has been a vital process to advancing past preclinical studies. However, this strategy has been difficult to replicate for non-G1 HCVs due to the lack of cell culture systems for G3–6, until very recently where replicons for G3a and G4a have been developed. 12 Therefore, the search for non-G1 enzyme inhibitors lags far behind, with the current hope that G1 inhibitors under development may demonstrate cross-genotype inhibition.
Around 20% of the estimated 170 million HCV infections are of G3a, with several countries demonstrating a high prevalence of G3 infections, including India (up to 67% 13 ), Brazil (40% 14 ), the United Kingdom and Finland (40% 15 ), and Australia (34% 16 ). Although G3a patients are better respondents to the current PEG-IFN-α and ribavirin treatment, 17 the newly developed DAAs show lower efficacy for the treatment of this genotype,18–20 including the recently approved protease inhibitor telaprevir. 21
Here we describe a simple, inexpensive, fluorescence-based HTS that was used for the identification of small-molecule inhibitors of the HCV 3a RdRp. The system could easily be adapted for any positive-sense RNA viruses that cannot be easily cultured.
Materials and Methods
HCV G3a RdRp Expression and Purification
Recombinant HCV subtype 3a RdRp was expressed and purified as described previously 22 with modifications. The RdRp-encoding region (GenBank accession number EF189901) of HCV G3a, excluding the C-terminal 21 amino acids, was cloned into the pTrcHis2C expression vector (Invitrogen, Carlsbad, CA). This construct, pVRL69b, was then transformed into Escherichia coli BL21 cells (Bioline, London, UK). An overnight culture of E. coli BL21 (10 mL) was transferred into 200 mL LB broth and incubated for 4 h at 37 °C. Finally, the 210-mL culture was transferred into a 7-L controlled bioreactor (Applikon, Schiedam, the Netherlands) containing 4.7 L of LB with 50 µg/mL ampicillin and grown at 37 °C until the OD600 reached 0.5. Expression of the RdRp was then induced with 1 mM isopropyl-β-D-thiogalactopyranoside and the culture incubated at 22 °C for 16 h. Cells were collected by centrifugation and washed with phosphate-buffered saline, and the bacterial pellet was stored at −80 °C until use.
For RdRp purification, bacterial pellets were resuspended in 1× CelLytic B (Sigma-Aldrich, St. Louis, MO) prepared with binding buffer consisting of 50 mM sodium phosphate (pH 7.5), 0.5 M NaCl, 50 mM imidazole, 10 mM β-mercaptoethanol, 10% glycerol, 0.1% n-octyl glucopyranoside, 0.2 mg/mL lysozyme, 1:200 (vol/vol) protease inhibitor cocktail (Sigma-Aldrich), and 50 U/mL benzonase nuclease (Sigma-Aldrich). The resuspended cells were incubated at room temperature for 20 min to allow for chemical lysis before the lysate was clarified by centrifugation and subsequent filtration. The soluble fraction was then applied to a Ni2+-charged HisTrap HP column (GE Healthcare Amersham, Buckinghamshire, UK) using the ÄKTAprime plus platform (GE Healthcare). Once bound, the column was washed with binding buffer. The RdRp was then eluted by increasing the imidazole concentration to 500 mM. The purity of RdRp was confirmed by SDS-PAGE and Western blotting with an anti–pentahistidine antibody conjugated with horseradish peroxide (QIAGEN, Hilden, Germany). The purified RdRp was dialyzed against storage buffer (50 mM Tris-HCl [pH 7.5], 200 mM NaCl, 0.1 mM dithiothreitol [DTT], and 20% glycerol), and protein concentration was quantified via spectrophotometry and stored at −80 °C.
RdRp Activity Assay Development
De novo RdRp activity assays were performed in 25-µL reactions. Enzyme activity was optimized by a systematic examination of reaction conditions, including enzyme, template, substrate, and salt concentrations, as well as reaction temperatures and incubation times. A standard reaction contained 40 µg/mL poly(C) RNA (average length 300 nt; Sigma-Aldrich), 0.5 mM rGTP (Promega, Madison, WI), 2.5 mM MnCl2, and 5 mM DTT in 20 mM Tris-HCl (pH 7.5) unless stated otherwise. Reactions were initiated by the addition of 300 nM RdRp and incubated at 30 °C for 2 h. The synthesis of double-stranded (ds) RNA was determined using the commercially available fluorescent dye PicoGreen (Invitrogen), which was developed for the quantitation of dsDNA but found to also preferentially bind dsRNA compared with single-stranded (ss) RNA. On completion of the RdRp reaction, contents were transferred into black 96-well microplates (Sigma-Aldrich), and 175 µL PicoGreen (1:350 [vol/vol] diluted in TE buffer, pH 7.5) was added to a final volume of 200 µL. The mixture was covered from light, incubated for 5 min, and the fluorescence intensity measured on a POLARstar Omega microplate reader (BMG Labtech, Ortenberg, Germany) at excitation and emission wavelengths of 485 nm and 520 nm, respectively. A separate control reaction with heat-inactivated RdRp was used to quantify background fluorescence. Standard radiolabeled nucleotide incorporation assays were also run for comparison, as described previously, with the addition of 0.04 µCi/µL [ 3 H]GTP and the GMP incorporation determined on a scintillation counter. 22 The commercially available bacteriophage RdRp, φ6 (NEB, Ipswich, MA), and in-house norovirus RdRp 23 were used as positive controls.
Kinetics of RdRp Activity
The optimum conditions for RdRp activity were examined by titrating enzyme concentration of divalent cations (MnCl2 and MgCl2) and by varying the incubation time and reaction temperature. To determine the kinetic constants of nucleotide incorporation by the HCV 3a RdRp, reactions were run with 40 µg/mL poly(C) for 40 min at increasing rGTP concentrations (0–2.5 mM). In addition, the kinetics of template utilization (poly(C)) was examined with 0.5 mM rGTP substrate and the poly(C) template concentrations ranging from 0 to 80 µg/mL. Molar amounts of double-stranded product were determined from a standard curve generated with a reference 19-bp dsRNA (Sigma-Aldrich). The velocity of the reaction was plotted against substrate concentration and kinetic parameters determined by nonlinear regression in GraphPad Prism, version 5.04 (GraphPad Software, La Jolla, CA).
Assay Miniaturization and Validation
A pilot HTS was conducted to identify novel lead-like small-molecule inhibitors of the HCV G3a RdRp. For this purpose, the assay was miniaturized to a 384-well microplate format in which the RdRp reaction was performed in a total volume of 25 µL, and 65 µL PicoGreen was subsequently added (1:130 dilution) to each well (final volume 90 µL). The miniaturized assay was tested for tolerance to Tween-20 and bovine serum albumin (BSA), which were required for robotic liquid handling and to minimize nonspecific protein adsorption to the plasticware used in the screen, respectively. Tolerance to the compound vehicle, DMSO, was also tested. For that purpose, the effect of increasing concentrations on RdRp activity was measured using a fluorescence output. The nucleotide analogue 3′dGTP (Trilink Biotech, San Diego, CA), which acts as a chain terminator, was used to validate sensitivity for RdRp inhibition. A titration curve was generated for 3′dGTP, and the half maximal inhibitory concentration (IC50) value was determined by nonlinear regression using the four-parameter, variable slope model in GraphPad Prism, version 5.04.
Pilot HTS
A total of 10,208 compounds (>90% purity) were randomly selected from the ~114,000 WECC lead-like compound libraries (Parkville, Victoria, Australia). An extensive battery of functional group filters was developed to remove poorly optimizable and assay interference compounds in creating this compound library. All compounds had a molecular weight between 150 and 450 Daltons, between one and four rings, a cLogP <5, a maximum of 10 rotatable bonds, a maximum of three chiral centers, a maximum of five hydrogen bond donors, and between one and eight hydrogen bond acceptors. Compounds dissolved in DMSO were assayed at final concentration of 10 μM. To that end, 50 nL of each compound was transferred into plates containing the RdRp, while DMSO was aliquoted into control wells. Plates were incubated for 10 min at room temperature, and the reaction was started with the addition of the remaining components (230 µM rGTP, 10 µg/mL poly(C) RNA, 2.5 mM MnCl2, 5 mM DTT, 0.01% BSA, and 0.005% Tween-20 in 20 mM Tris-HCl, pH 7.5). Each test plate contained 16 wells with control (no compound) and heat-inactivated RdRp for positive and negative control measurements, respectively. Final concentration of DMSO in all reactions was 0.2%. Plates were incubated for 1 h at 30 °C, and PicoGreen dye was then added and fluorescence quantified on an EnVision Multilabel plate reader (PerkinElmer, Waltham, MA). Screening data were analyzed in ActivityBase (IDBS, Guilford, UK) and visualized using Spotfire (TIBCO, Palo Alto, CA). Compound activity, equivalent to RdRp inhibition, was calculated as follows: percentage inhibition = 100 - [100 × (sample - meanNSA) ÷ (meanTA - meanNSA)], where NSA is nonspecific activity and TA is total activity. Assay quality for HTS was determined based on signal to background (S/B = meanTA ÷ meanNSA), signal to noise (S/N = [meanTA - meanNSA] ÷ SDNSA), coefficient of variation (CV = 100 × SD ÷ mean), and finally Z′ values, a measure of assay robustness Z′ = 1 - [(3SDTA + 3SDNSA) ÷ (meanTA - meanNSA)], where a Z′ ≥ 0.5 is considered acceptable for HTS.24,25 Each parameter was calculated on a plate-by-plate basis.
Secondary Screening and IC50 Determination
Percent inhibition of RdRp activity for all 10,208 compounds screened was determined to identify leads from the pilot HTS. A frequency distribution was then generated for all screened compounds based on the number of hits at defined percent inhibition bins. Compounds were selected if they showed ≥16.4% inhibition of RdRp activity, which represented inhibition exceeding mean + 2.5 × SD of the distribution. The leads were then assayed in triplicate using the same fluorescence RdRp assay at 10 µM to confirm inhibitory activity. After excluding false positives, hits were reconfirmed using a radioactive GMP incorporation assay. The nonfluorescent nature of the second confirmatory assay removed further false-positive compounds that either interfered with or enhanced the fluorescence signal. 26 Finally, the remaining “top” hits were titrated to a final assay concentration of 60 µM to allow IC50 determination using the radioactive RdRp assay by nonlinear regression using the four-parameter, variable slope model in GraphPad Prism, version 5.04.
Results
Expression and Characterization of HCV G3a RdRp
To search for specific G3a HCV polymerase inhibitors, we needed purified recombinant RdRp and an HTS-amenable assay format to evaluate large numbers of small molecules. To obtain large quantities of purified RdRp for assay development and HTS, we expressed the RdRp from the HCV G3a strain VRL69 in E. coli and purified it by nickel affinity chromatography. The total yield of purified RdRp was 7.5 mg/L culture (>74% purity; data not shown).
A systematic examination of biochemical factors that influenced activity of the HCV 3a RdRp was performed to establish the optimal HTS assay conditions. The effect of enzyme concentration on polymerase activity was examined. Activity increased linearly from 10% of maximal activity at 80 nM RdRp to 40% at 160 nM with maximal activity at 420 nM ( Fig. 1A ). No increase in fluorescence was detected with increasing concentrations of the control heat-inactivated enzyme. A functionally inactive mutant RdRp, with a GAA mutation at the active site, demonstrated similar background fluorescence to the heat-inactivated RdRp (data not shown). We were unable to use our fluorescent assay to quantify the effect of the divalent cation Mn2+ on RdRp activity. At concentrations greater than 2.5 mM, MnCl2 affected PicoGreen fluorescence (data not shown). Alternatively, the optimal concentration of MnCl2 was determined to be 2.5 mM using our radioactive RdRp assay. The enzyme was inactive in the absence of a divalent cation, and no RdRp activity was detected using 0 to 15 mM Mg2+ ( Fig. 1B ). The time course examination of dsRNA formation showed linear accumulation of product up to 1 h, with production formation decreasing after 90 min ( Fig. 1C ). Finally, the optimum temperature for RdRp activity was in the range of 30 to 33 °C, with a 40% decrease in activity at 37 °C (data not shown).
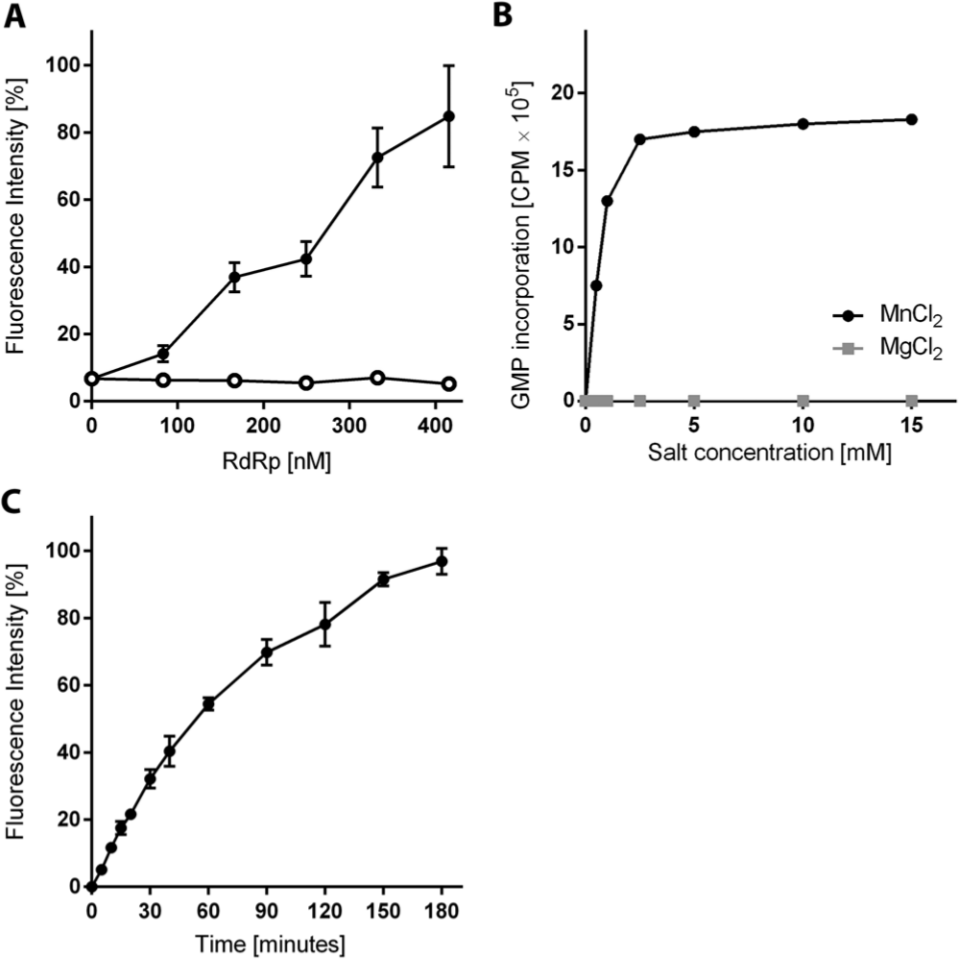
Characterization of biochemical factors influencing hepatitis C virus (HCV) G3a polymerase activity. (
The steady-state kinetics of the HCV G3a RdRp reaction was examined and the Km of the rGTP substrate was calculated to be 237 ± 40 µM, with a maximum reaction velocity Vmax of 52 ± 3 pmol GMP h–1 ( Fig. 2A ). The Km for the RNA template (poly(C)) was 10 ± 2 µg/mL (~33 µM) with a Vmax at 75 ± 5 pmol GMP h−1 ( Fig. 2B ).
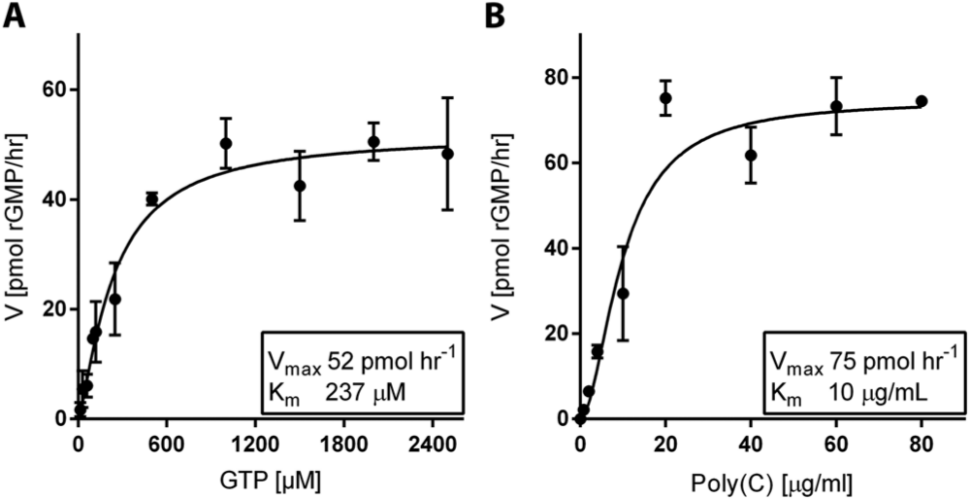
Steady-state kinetics of the hepatitis C virus G3a RdRp using the PicoGreen assay. (
Development of a High-Throughput RdRp Assay
The fluorescent-based RdRp assay was further developed for HTS of small-molecule libraries to identify inhibitors of the HCV G3a RdRp. Initially, the sensitivity of the assay to detect RdRp inhibition was tested by titrating the nucleotide analogue 3′dGTP, a known NI and chain terminator ( Fig. 3A ). Complete enzyme inhibition was reached at 60 µM 3′dGTP, and the half-maximal inhibitory concentration IC50 was calculated at 0.6 µM ( Fig. 3B ).
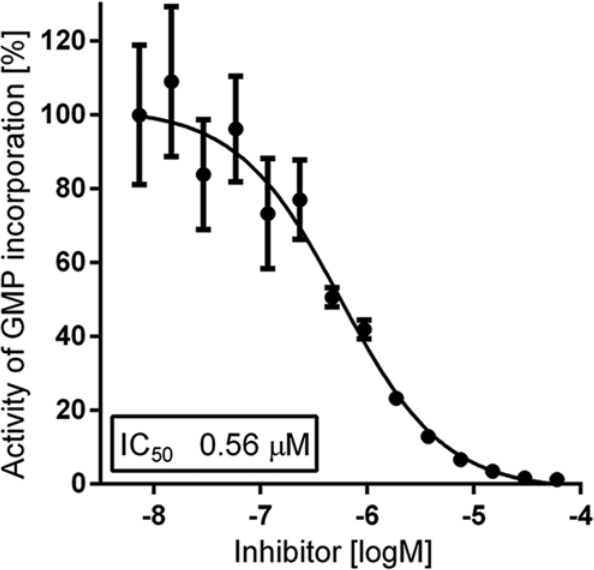
Titration curve of the reference compound 3′dGTP. RdRp polymerase activity was quantified using PicoGreen in the presence of increasing concentrations of 3′dGTP. Inhibition of double-stranded RNA formation was calculated based on control RdRp reactions representing full reaction (DMSO vehicle only) and no reaction (heat-inactivated RdRp).
The assay was then miniaturized for HTS in 384-well microplates and tested for tolerance to Tween-20, BSA, and the compound vehicle, DMSO, and no effect on RdRp activity or fluorescence signal was observed with concentrations up to 0.01%, 0.02%, and 0.6%, respectively (data not shown). Assay robustness and suitability for HTS were examined by calculating signal-to-background and Z′ values, standard statistical measures of assay quality for HTS campaigns. 24 The miniaturized assay demonstrated high sensitivity and robustness, with a mean Z′ of 0.73 and a signal-to-background ratio exceeding 20 in validation screens (data not shown).
Pilot Inhibitor Screening
A pilot screen for NNIs of the HCV G3a RdRp was conducted using a random selection of 10,208 compounds with lead-like characteristics, at a final concentration of 10 µM. The screen was performed in two batches of 384-well plates, and assay control measurements for both batches are summarized in Table 1 . Overall, Z′ values for all test plates ranged between 0.65 and 0.78, with the exception of one plate, which had a Z′ of 0.33 and was therefore excluded ( Fig. 4A ). The frequency distribution of RdRp inhibition, as a percentage of control, for all screened compounds showed a normal distribution ( Fig. 4B ). One hundred fifty compounds that demonstrated ≥16.4% inhibitory activity, or >2.5 × SD above the mean inhibition for HTS, were chosen for follow-up screening. The coefficient of variation (CV) for all controls was 7.8%, which was below the acceptable limit of 15% typically set in HTS assays. 25
Control Measurements of Pilot High-Throughput Screening Assay Performance.
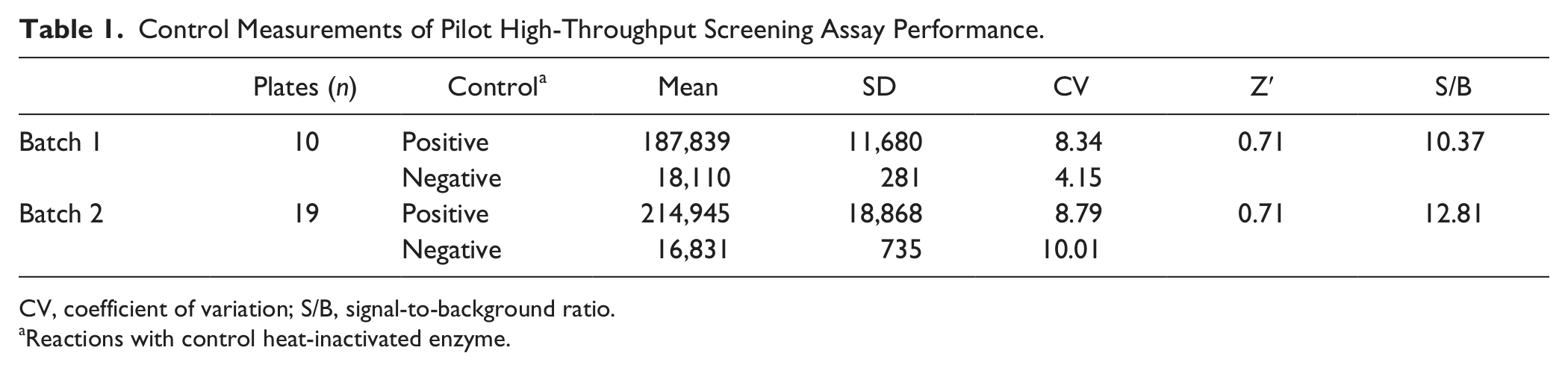
CV, coefficient of variation; S/B, signal-to-background ratio.
Reactions with control heat-inactivated enzyme.
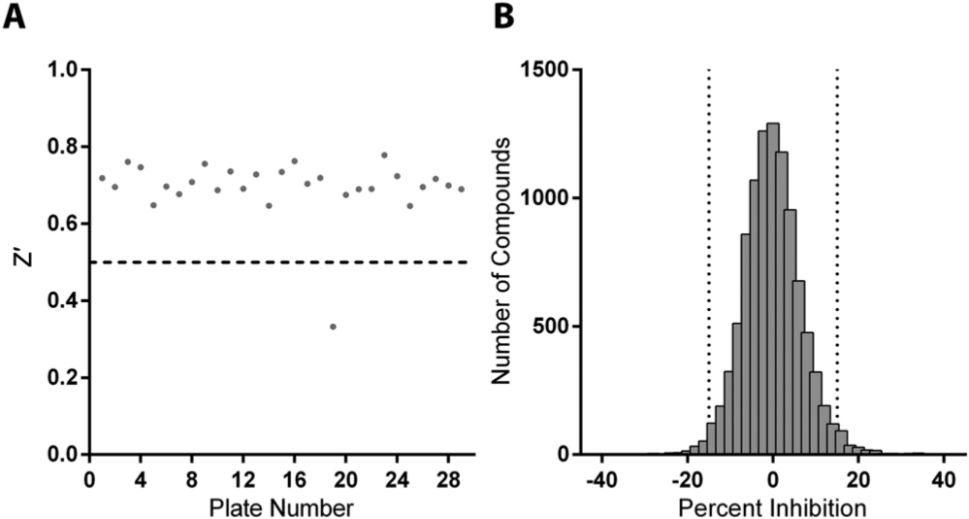
Pilot screening for inhibitors of the hepatitis C virus G3a RdRp. (
Secondary Screening and IC50 Determination
Compounds selected as hits from the pilot HTS (n = 150) were retested using the same fluorescent-based RdRp assay in triplicate. At 10 µM, 85 compounds showed no inhibitory activity against the HCV G3a RdRp, and 55 others showed weak activity (<16.4% inhibition; data not shown). These compounds were excluded from further analysis.
The remaining 10 compounds were subsequently confirmed as inhibitors using both the fluorescent RdRp assay and an additional radioactive-nucleotide incorporation RdRp assay. The latter assay was used to exclude compounds that affect (e.g., quench) fluorescence rather than inhibit enzymatic activity. Six of 10 hits were confirmed in both fluorescent and radioactive assays, and the RdRp inhibition results showed a positive correlation between the two formats, with a coefficient r = 0.8 (p < 0.01; Fig. 5A ). None of the six compounds showed inhibition of a control norovirus RdRp (data not shown). The half-maximal inhibitory concentrations (IC50) for the two most potent hits, HAC001 and HAC002, were 12.7 µM and 1.0 µM, respectively ( Fig. 5B ). The next most inhibitory compounds, HAC003 and HAC004, demonstrated IC50 values of 21.6 µM and 20.7 µM, respectively. These compounds, as well as their chemical derivatives, are currently being examined for their potential as antiviral agents.
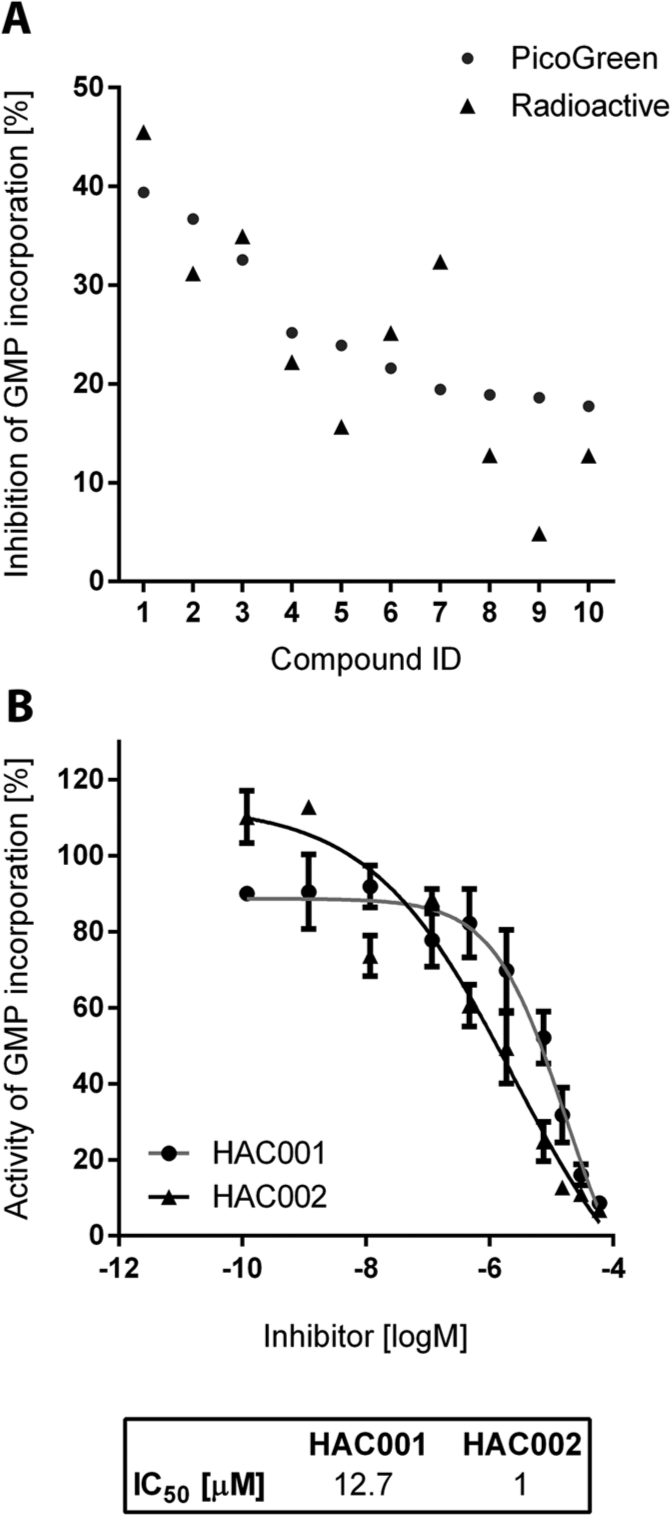
Validation and titration curves for the top hepatitis C virus RdRp hits from the pilot high-throughput screen. (
Discussion
The HCV RdRp is an attractive target for DAA development given the crucial role the enzyme plays in the viral life cycle. Since the majority of global HCV infections are caused by HCV G1, 1 most antiviral campaigns for HCV inhibitors have been developed against this particular genotype, mainly using recombinant enzyme assays6–8 or with the HCV G1b replicon model. 27 Despite the recent development of new replicon systems for G3a and G4a viruses, 12 the long delay (>10 years) in their availability compared with the G1 replicons has hindered campaigns aimed at identifying DAAs that work against non-G1 HCV genotypes. Furthermore, the efficacy of identified G1 inhibitors against non-G1 viruses has been undertaken at later stages of DAA development, with generally poor cross-genotype inhibition demonstrated. For example, DAAs in clinical development have already demonstrated reduced efficacy against HCV G3a, including those that target the viral proteins NS3 (protease), 18 NS5A (phosphoprotein), 20 and NS5B (RdRp).5,19 These finding suggest that a “one drug for all HCV genotypes” approach may not be feasible for HCV and that development of genotype-specific enzyme inhibitors may be essential. No HTS campaigns aiming to identify HCV RdRp inhibitors have thus far been reported where the HCV G3a RdRp is the primary target. Here, we describe a robust in vitro fluorescence-based assay for the detection of primer-independent (de novo) RdRp activity that was subsequently applied in a pilot HTS campaign against the HCV G3a RNA polymerase.
Traditional methods for detecting the activity of viral RdRps have relied on the incorporation of radiolabeled nucleotides into an RNA template.7,22 Although very sensitive, these assays have limitations for HTS because of handling requirements and multiple filter and wash steps. 22 Alternative assays for RdRp activity have been described based on fluorescent labeling of the nucleic acid template 28 or chemically modified nucleotides 29 and/or require the addition of other enzymes into the reaction. 30 These methods are expensive and labor intensive and therefore not ideal for use in HTS campaigns. The fluorescent dye PicoGreen, although originally developed for staining and quantifying dsDNA, has recently been used for detecting dsRNA products catalyzed by viral RdRps31,32 but not for HTS. The assay we describe here uses PicoGreen and a poly(C) RNA template with unmodified ribonucleotides (rGTP) to generate end-point fluorescence-based measurement of the de novo activity of the HCV G3a RdRp. Polymerase activity was then determined by detecting the dsRNA product.
The assay was initially used to examine the steady-state kinetics of the HCV G3a polymerase with quantitation of the amount of dsRNA product formed to define optimal reaction conditions for HTS. Km and Vmax, for rGMP incorporation by the HCV RdRp, were determined ( Fig. 2 ). The affinity for rGTP of the VRL69 G3a RdRp (237 µM) was similar to that reported for the HCV G3a strain, ALB-3a RdRp, at 208 µM. 33 However, the Vmax was lower for the VRL69 RdRp at 74.7 pmol rGMP h−1 compared with 193.8 for the ALB-3a. The requirement of the G3a RdRp for divalent cations was consistent with previous reports, where Mn2+ but not Mg2+ facilitated de novo HCV RdRp activity in vitro ( Fig. 1B ). 33
Upon miniaturization and automation, the assay was shown to be suitable for HTS with minimal handling and small assay volumes. In addition, all measurements for assay performance and sensitivity were within the recommended range for HTS assays (Z′ >0.5, S/B and S/N >2; Table 1 ). 25 In this study, HTS was conducted with a relatively low concentration of compound (10 µM) to avoid unfavorable assay interference possibly caused by high compound concentrations 34 ; for this reason, hits from the pilot HTS were subjected to a secondary screen followed by a counterscreen using a radioactive-based assay. This resulted in a final confirmed hit rate of 0.06%, with four novel compounds that inhibited HCV G3a RdRp activity in the low micromolar range.
In summary, we have developed an HTS format that uses a fluorescent dye, PicoGreen, to identify inhibitors of the HCV G3a RdRp. The assay was robust, cost-effective, and capable of identifying inhibitors of HCV G3a RdRp activity, with IC50s in the low micromolar range. The assay could easily be adapted for any viral RNA polymerase to identify inhibitors by HTS, even for positive-sense RNA viruses that cannot be cultivated in vitro. Hits identified in the present study, as well as their mode of inhibition and their activity on available replicon systems, are being examined.
Footnotes
Acknowledgements
The authors thank Rowena Bull, Scott Minns, and Helene Lebhar for technical support.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors acknowledge the support from the Australian Centre for HIV and Hepatitis Virology Research (ACH2). AAE is supported by and the University of New South Wales International Postgraduate Award.
