Abstract
Hsp90 and its co-chaperones are known to be important for cancer cell survival. The N-terminal inhibitors of Hsp90 that are in ongoing clinical trials as antitumor agents have unfortunately shown disappointing efficacies in the clinic. Thus, novel inhibitors of the Hsp90 machine with a different mechanism of action are urgently needed. We report here the development of a novel high-throughput screening assay platform to identify small-molecule inhibitors of Hsp90 and its co-chaperones. This assay quantitatively measures the ability of Hsp90 and its co-chaperones to refold/protect the progesterone receptor, a physiological client of Hsp90, in a 96-well plate format. We screened the National Institutes of Health clinical collection drug library and identified capsaicin as a hit molecule. Capsaicin is a Food and Drug Administration–approved drug for topical use in pain management. Cell survival assays showed that capsaicin selectively kills cancer cells and destabilizes several Hsp90 client proteins. Thus, our data may explain the seemingly pleotropic effect of capsaicin.
Keywords
Introduction
Hsp90 has been implicated in a multitude of cellular functions and connects multiple signaling networks. It drives the folding, assembly, translocation, and degradation of many regulatory proteins that contribute to the hallmarks of cancer. Many Hsp90 client proteins are primary oncology targets, especially protein kinases (ErbB2, BCR-Abl, Cdk4, PKB/Akt, Raf-1, B-Raf, and FAK) and steroid receptors. 1
Inhibitors of Hsp90, such as 17-AAG and other geldanamycin derivatives, or purine-scaffold derivatives, are effective in different tumor types, including melanoma, multiple myeloma, and breast and prostate cancers. 2 Although there are no Food and Drug Administration (FDA)–approved Hsp90 inhibitors yet, 17 different Hsp90 inhibitors have shown promise in clinical trials for cancer of various origins. Most of these drugs inactivate the ATPase activity of Hsp90, causing proteasomal degradation of its “client” proteins. 3 Unfortunately, these Hsp90 ATPase inhibitors also induce overexpression of Hsp70 and Hsp27, which are apoptosis inhibitors 4 and may contribute to the low efficacy of the drugs. This highlights the need for novel strategies to inactivate the Hsp90 machine. Great effort has been made to develop compounds with reduced stress response induction or to target other domains of Hsp90, such as the C-terminus adenosine triphosphate (ATP) binding pocket. 5 However, alternative strategies to inactivate the Hsp90 machine, including inactivating its co-chaperones, need to be pursued more aggressively.
The proper functioning of steroid receptors, such as progesterone receptor (PR) and glucocorticoid receptor (GR), is strictly dependent on the Hsp90 chaperoning machine. 6 Thus, they have been used as model systems to dissect the Hsp90 chaperoning pathway. Most of these studies used cell-free systems, such as rabbit reticulocyte lysate (RL). This knowledge has been harnessed to develop a PR reconstitution assay using not only RL as a source of molecular chaperones but also purified chaperones. 6 Because of the robustness and reliability of the PR reconstitution assay, we sought to develop it into a high-throughput screening (HTS) format and make it suitable for screening small-molecule libraries to identify new modulators of the core components of the Hsp90 chaperoning machine.
Materials and Methods
PR Reconstitution Assay Using RL
This assay was based on the reconstitution assays previously published. 7 Untreated RL was purchased from Green Hectares (Oregon, WI). Lysates were supplemented with 3 mM MgCl2, 1 mM ATP, and an ATP-regenerating system consisting of phosphocreatine and creatine phosphokinase. Chicken PR-A was overexpressed in SF9 cells. A PR22 antibody was used to adsorb PR complexes to protein A resin. Protein A-PR22-PR resin stripped of endogenous associated chaperones with 400 mM KCl was suspended in 250 µL of RL and incubated for 30 min at 30 °C. Resin was resuspended every 5 min. After incubation at 30 °C for 30 min, the samples were chilled and supplemented with 5 nM [1,2-3H]progesterone (American Radiolabelled Chemicals, St Louis, MO, USA) plus 100 nM unlabeled progesterone. The samples were incubated for 1 h on ice and then washed with 4 × 1 mL of incubation buffer by centrifugation and suspension. At the last wash, 100 µL was removed for measurement of [3H]progesterone using a Microbeta scintillation counter (PerkinElmer, Waltham, MA). The remaining 900 µL was centrifuged. The buffer was removed and the resin resuspended in sample buffer and analyzed by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE).
High-Throughput PR Reconstitution Assay
Chicken PR-A was overexpressed in SF9 cells. A PR22 antibody was used to adsorb PR complexes on the well floor of a high-binding 96-well plate (655094; Greiner Bio-One, Monroe, NC). Briefly, 96-well plates were loaded with 50 µL per well of protein A solution (20 µg/mL) in 1× phosphate-buffered saline (PBS) and incubated overnight at 4 °C. Plates were then washed twice with reaction buffer (200 µL/well; 20 mM Tris-base [pH 7.8], 100 mM KCl, 1 mM dithiothreitol [DTT], 0.03% NP40) and blocked with 200 µL of 5% bovine serum albumin (BSA) in reaction buffer for 1.5 h at 4 °C. Next, 50 µL of 20 µg/mL purified PR22 antibody in 5% BSA was added, and plates were incubated for 1.5 h at 4 °C. Excess PR22 was washed three times with reaction buffer (200 µL/well). SF9-PR lysate (50 µL) in high salt stripping buffer (20 mM Tris-base [pH 7.8], 300 mM KCl, 1 mM DTT, 5 mM MgCl2, 5 mM ATP, 0.03% NP40) was then added, and plates were incubated for 1 h at 4 °C. This was followed by two washes with stripping buffer and three washes with reaction buffer (200 µL each) to remove excess PR and its associated proteins. We then added 100 µL of an RL mixture consisting of 50 µL of pure RL and 50 µL of RL recipe buffer (40 mM Tris-base [pH 7.8], 200 mM KCl, 2 mM DTT, 0.25 mM MgCl2, 0.06% NP40, 0.25 mM ATP, 5% glycerol, supplemented with 178 nM creatine phosphokinase and 1.11 mM phosphocreatine as an ATP regeneration system). DMSO (1%) or a drug at a 10-µM final concentration was added, and plates were incubated for 30 min (shaking every 5 min) at 30 °C in a humid environment. After the incubation, [3H]-progesterone (ART 0063; American Radiolabelled Chemicals, St Louis, MO, USA) was added. Plates were incubated for 3 h at 4 °C. The assembled PR complexes were washed five times with 200 µL per well reaction buffer. Bound [3H]-progesterone was monitored using scintillation liquid and a Microbeta plate reader (PerkinElmer). Each sample was duplicated on another 96-well plate in the same experimental conditions.
MTT Cell Survival Assay
Cells were grown in 96-well tissue culture plates (3599; Corning, Corning, NY) to 60% confluence and treated with indicated concentrations of 17-AAG, capsaicin, or DMSO control (2% total DMSO concentration) for the indicated times. Cells were incubated with 10 µL of the CellTiter 96 AQueous One Solution Cell Proliferation Assay reagent (G3580; Promega, Madison, WI) and 90 µL of culture media/well for 1 h at 37 °C. Absorbance at 495 nm was measured using a SAFIRE-TECAN (Tecan Group Ltd, Australia) plate reader.
Colony Formation Assay
Cells were grown in six-well tissue culture plates to 60% confluence and treated with 200 µM capsaicin or DMSO control for 24 h. Cells were then collected, and 1000 of these cells were replated per 10-cm tissue culture dish (353003; Falcon, Durham, NC, USA) in triplicate experiments. Cells were grown for 15 days in 1× minimum essential media (MEM) supplemented with 10% fetal bovine serum (FBS). Cells were fixed with 6% glutaraldehyde and 0.5% Crystal Violet, and colonies that contained 50 cells were counted.
Results and Discussion
Parent Assay: Reconstitution of the PR Complexes Using RL
PR is a physiological client of the Hsp90 chaperoning machine in cells. Seminal work from the laboratories of David Toft, William Pratt, David F. Smith, and other researchers has led to the reconstitution of PR chaperone complexes in vitro using rabbit RL, which serves as a complete source of molecular chaperones. 7 Properly folded PR binds progesterone with high affinity. The assay is a protein A/Sepharose resin–based immunoprecipitation assay, where recombinant avian PR multichaperone complexes are isolated from SF9 cells using the specific monoclonal antibody PR22. PR is then stripped from SF9 endogenous associated proteins with a high salt buffer. Incubation of the naked PR at 30 °C leads to total loss of PR hormone-binding activity in less than 5 min. However, adding RL in the presence of an ATP regeneration system reconstitutes PR multichaperone complexes and preserves the hormone-binding activity for as long as 30 min at 30 °C (not shown). This assay therefore reflects the ability of Hsp90 and its co-chaperones to protect/refold partially heat-denatured PR to its ligand-binding conformation. As seen in Figure 1 , RL efficiently refolded PR to its hormone binding state. Geldanamycin (10 µM) inhibited the folding process to about 40% ( Fig. 1A ). As expected, analysis of protein complexes using SDS-PAGE followed by Coomassie blue staining ( Fig. 1B ) showed that the active, mature PR complex contained mainly PR, Hsp90, the immunophilin cyclophilin 40, and p23, which is known to stabilize the Hsp90-client protein complexes. Addition of geldanamycin blocked the chaperoning process in an intermediate state, where substantial Hsp70, Hop, and Hsp40 were detected, and the level of Hsp90 was lower than in samples with no drug added. No p23 or immunophilins were detected in geldanamycin-containing samples.
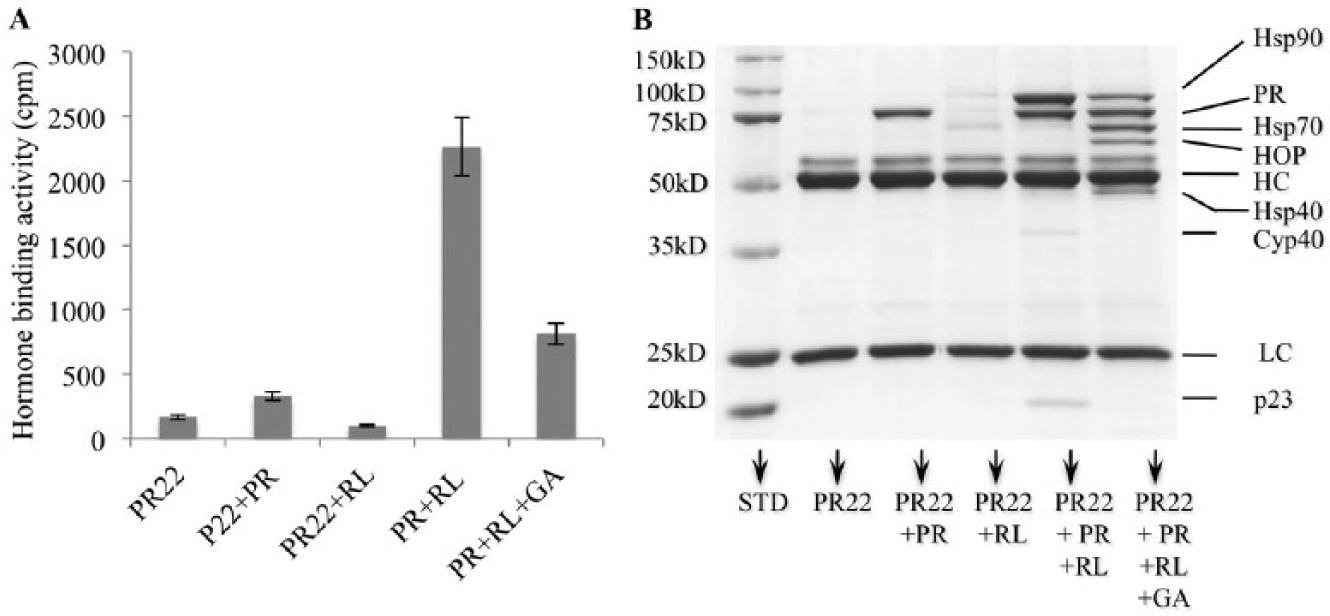
Parent assay: reconstitution of progesterone receptor (PR) complexes using protein A/Sepharose beads, PR antibody, and reticulocyte lysate (RL). PR was incubated in presence of RL alone or with geldanamycin (GA; 10 µM), as described in Materials and Methods. (
Transformation of the PR Reconstitution Assay to 96-Well Platform Screen
Transforming the PR reconstitution assay from an immunoprecipitation-based assay on protein A beads to a microtiter 96-well plate required optimization at every step of the process ( Fig. 2A ). We tested several kinds of plates and optimized the concentrations of protein A and PR antibody, the amount of PR, the stripping conditions, the volume of RL, and the assay buffer additives (glycerol, ATP regeneration system, MgCl2, glycerol, NP40, and DMSO). Our goal was to obtain an optimal hormone-binding activity that was greater than at least six times the background.
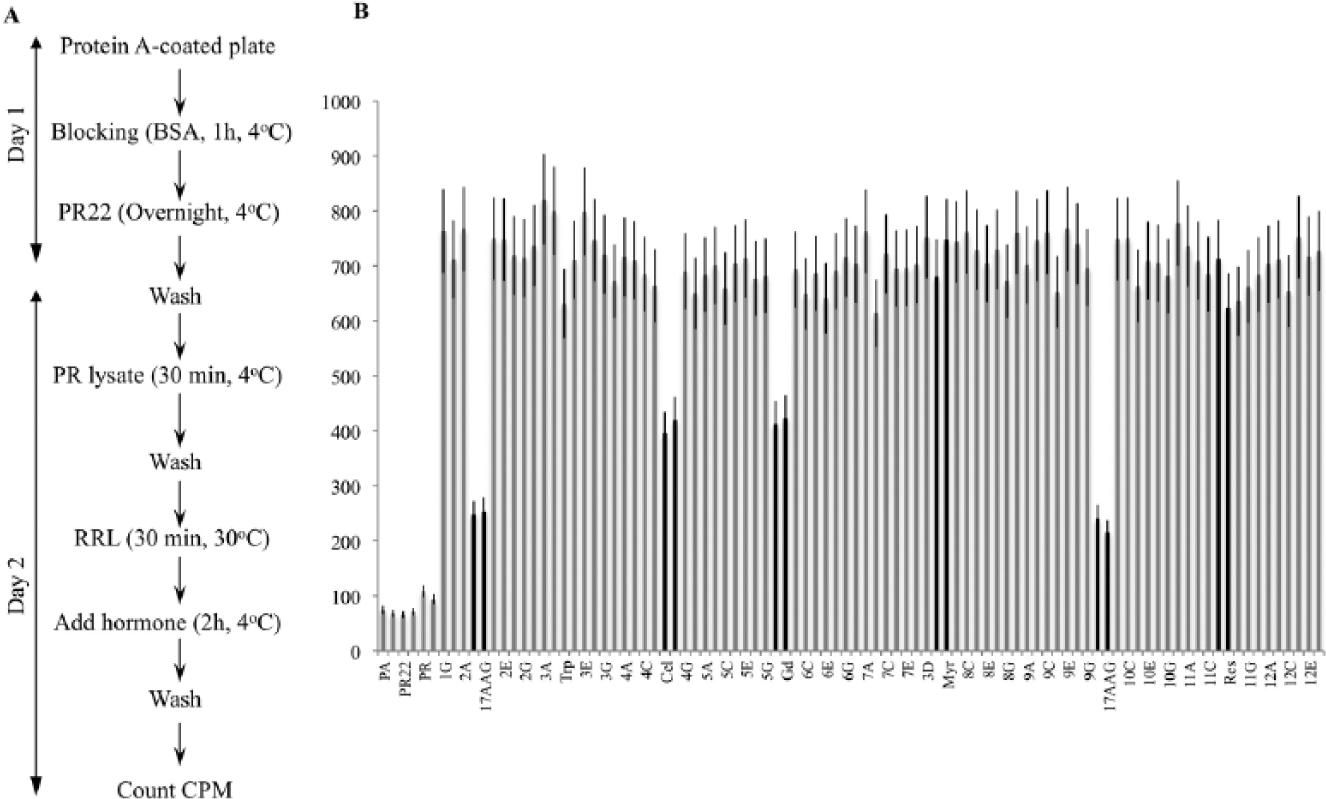
(
To evaluate the suitability of the PR reconstitution assay for HTS, we designed the incubation conditions at 30 °C to minimize the variation between the central and peripheral wells. We investigated the intraplate variability using RL with no inhibitors on a full plate (
The usefulness of the PR reconstitution assay in 96-well plates to detect the Hsp90 machine inhibitors was validated using well-known inhibitors of the ATPase activity of Hsp90, such as geldanamycin derivative 17-AAG ( Fig. 2B ). We also used celastrol and gedunin, which inactivate the Hsp90 machine through interaction with the co-chaperone p23. 8 As negative controls, we used myricetin, resveratrol, and triptolide, which do not show any effect on the Hsp90 chaperoning of steroid receptors.
We further tested the robustness of the assay using multiple plates. We calculated the Z′ factor using the previously published formula. 9 The positive controls were wells with RL without inhibitors or with myricetin, which did not affect the chaperoning of PR; negative controls were wells with RL and 17-AAG or gedunin that inhibited the system to less than 40% or 55% activity, respectively. The Z′ factor was 0.723 for four different plates in 2-day measurements with inhibitors dispersed into end wells, which demonstrated the excellent quality of the assay. The assay is thus robust and can be used as a reliable tool for HTS.
Validation of the PR Reconstitution Assay in a 96-Well Plate Using the NIH Small-Molecule Library
We further tested the usefulness of this assay using a small chemical library consisting of 446 compounds called the National Institutes of Health (NIH) clinical collection (NCC-002). This library consists of FDA-approved drugs or compounds that are in phase I, II, or III of human clinical trials for various diseases and have been evaluated for toxicity, safety, and bioavailability. This library thus offers an attractive set of drugs or drug-like molecules that have shown promising results in human patients. Each compound was tested in duplicate at a 10-µM final concentration. Because inhibitors of the co-chaperone p23, such as celastrol of gedunin, reduced the hormone-binding activity by about 45% ( Fig. 2B ), we consider a “hit” inhibitor a compound that inhibits the hormone-binding activity of PR by approximately 40% or more. Likewise, we consider a “hit” activator a compound that increases the PR hormone binding by at least 40%. The library comes in six 96-well plates. Of 446 compounds tested, we obtained seven primary hits from different plates ( Fig. 3 and data not shown). Inhibitors having chemical structures related to progesterone were not given further consideration, as they compete with 3[H]-progesterone for binding PR. The remaining hits, which are structurally different from progesterone, were rescreened in triplicate at a 20-µM final concentration (not shown). Among these, only capsaicin showed a reproducible effect on blocking the hormone-binding activity of PR by about 40%, which seems to be the maximum inhibition that capsaicin causes in this assay. The hit rate is thus 0.22%, well within 1%, the maximum acceptable hit rate by NIH guidelines for a robust HTS assay.
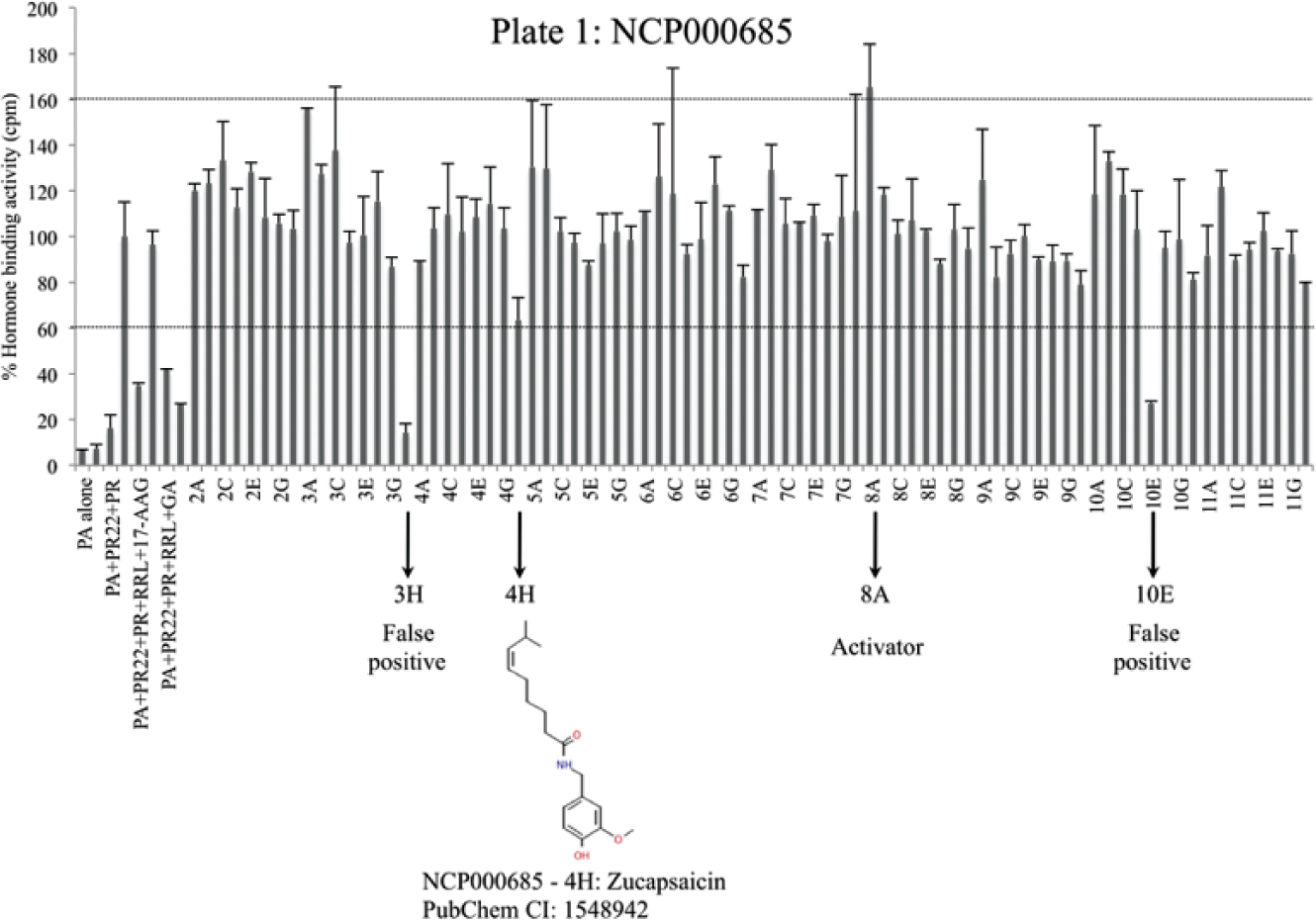
Screening of the National Institutes of Health Clinical Collection drug library. Progesterone receptor (PR) complexes were reconstituted on a 96-well plate using reticulocyte lysate (RL) as the source of molecular chaperones and accessory proteins. Each bar represents percent hormone-binding activity of PR in the presence or absence of chemical compounds. The first samples represent the following internal controls: (1) protein A alone, (2) protein A + PR22, (3) protein A + PR22 + PR, (4) protein A + PR22 + PR + RL, (5) protein A + PR22 + PR + RL + 17-AAG (2 µM), (6) protein A + PR22 + PR + RL + myrecetin (20 µM), (7) protein A + PR22 + PR + RL + geldanamycin (20 µM), and (8) protein A + PR22 + PR + RL + gedunin (20 µM). The remaining samples contain compounds from the NCP000685 plate used at a 10-µM final concentration. Primary hits are represented with arrows on the x-axis. The standard deviation of duplicate samples is shown as error bars.
Capsaicin selectively kills cancer cells of various origins
Since capsaicin consistently and reproducibly blocked PR chaperoning in vitro, we decided to test the effect of capsaicin in killing cancer cells. Using an MTT assay, the cytotoxic effect of capsaicin was assessed with concentrations ranging from 1 to 200 µM and at three time points using MCF7 breast cancer cells and LNCaP prostate cancer cells ( Fig. 4A ). These data suggest that capsaicin starts to cause significant death of LNCaP cells at concentrations higher than 50 µM, confirming the range of doses reported by other investigators. We compared capsaicin’s effect on immortalized normal breast epithelial cells (Hs578Bst) versus breast cancer cells (MCF7) with doses of 50, 100, and 200 µM for 24 or 96 h. As shown in Figure 4A , capsaicin selectively kills MCF7 breast cancer cells in a time- and dose-dependent manner, without affecting Hs578Bst normal breast cells. These results confirm previous reports showing that another nontransformed mammary cell line, MCF10A, is less sensitive to capsaicin treatment. 10 We then assessed the effect of capsaicin on the colony formation efficiency of LNCaP, MCF7, HeLa, and HepG2 cancer cell lines. As shown in Figure 4B , capsaicin efficiently reduces the ability of cancer cells to form colonies.
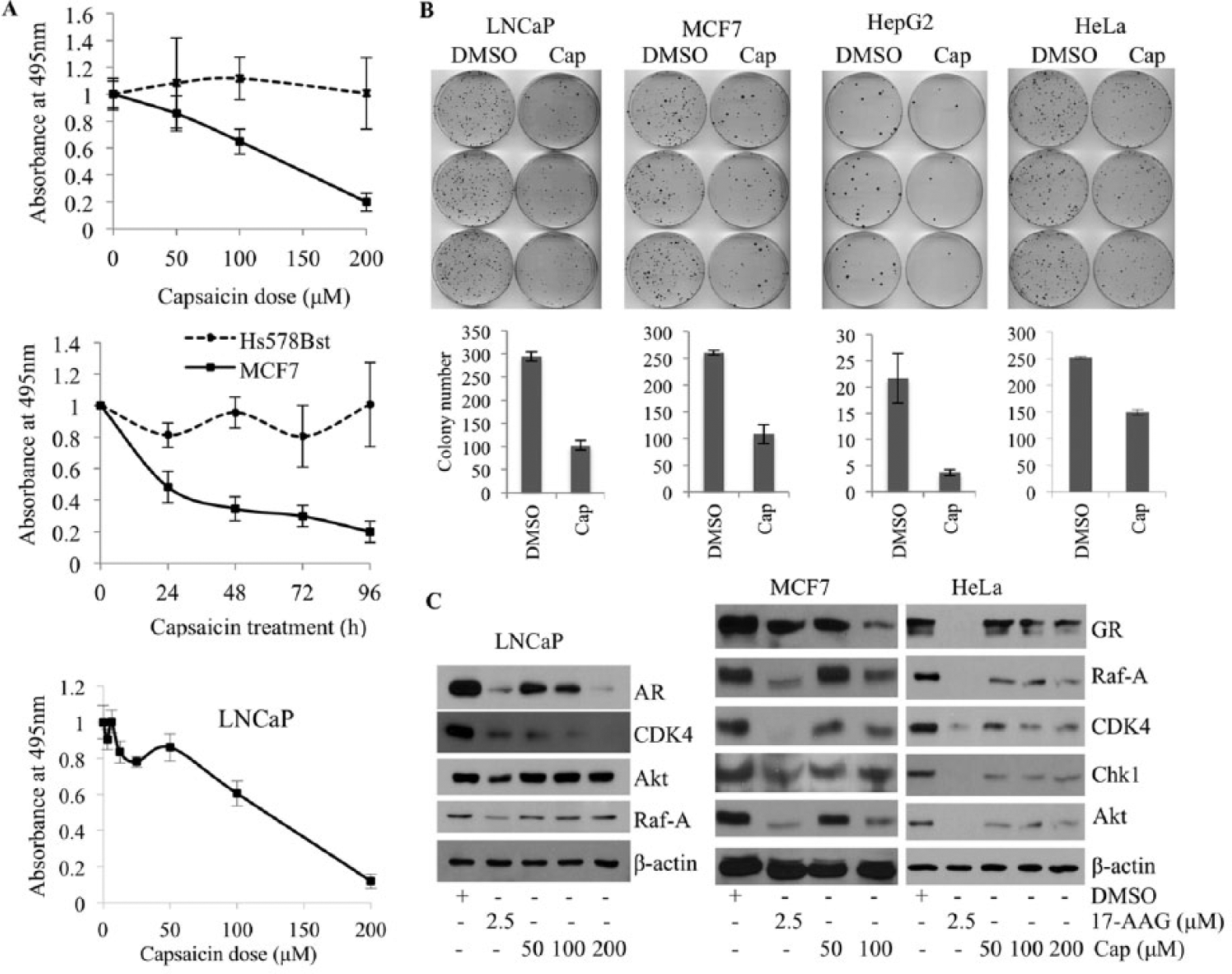
Capsaicin selectively kills cancer cells. (
Capsaicin (8-methyl-N-vanillyl-6-nonenamide), a homovanillic acid derivative, is the main pungent ingredient of red pepper, genus Capsicum. Capsaicin was reported to bind to the transient receptor potential vanilloid (TRPV1) receptor, leading to desensitization of peripheral nerves to noxious stimuli. 11 Numerous recent research reports have convincingly shown that capsaicin induces cell cycle arrest and apoptotic cancer cell death in a variety of cancer cell lines, including colon adenocarcinoma, pancreatic cancer, hepatocellular carcinoma, prostate cancer, breast cancer, and many others. 12 Interestingly, many of the pathways inhibited by capsaicin treatment involve Hsp90 client proteins. These include the EGFR/HER-2 pathway, 10 cyclin-dependent kinase, 13 and Src kinase, 14 among others.
Capsaicin inactivates the Hsp90 chaperoning machine in cancer cells
To verify whether capsaicin targets the Hsp90 machine in cells, we treated LNCaP, MCF7, and HeLa cells with increasing doses of capsaicin and studied the effect of capsaicin on the expression of Hsp90 client proteins. Most Hsp90 client proteins, such as nuclear receptors (GR and AR), and other cell signaling protein kinases, including CDK4, Chk1, Raf-A, and Akt, were downregulated upon capsaicin treatment ( Fig. 4C ). There were, however, some cell-specific effects of capsaicin. Akt and Raf-A, for example, were destabilized in MCF7 and HeLa cells but insensitive to capsaicin treatment in LNCaP cells. Taken together, these data indicate that capsaicin blocks the Hsp90 chaperone system, as reflected by destabilization of protein clients of the Hsp90 machine. Previous reports have shown that capsaicin blocks the activity of many other oncogenic signaling proteins known to be clients of Hsp90, such as NF-κB, ER, EGFR/HER2, Src, VEGF, and PI3K/Akt, among many others. We therefore propose that the seemingly pleiotropic effects of capsaicin 15 may be due to inhibition of the central Hsp90 chaperoning machine.
In conclusion, we have developed a new HTS platform that is sensitive and robust enough to identify small-molecule modulators of the Hsp90 chaperoning machine in vitro. This assay is uniquely qualified to identify compounds that inhibit or activate the Hsp90 chaperoning machine. We have designed a robot capable of executing all the steps of this assay ( Fig. 2A ) automatically. This provides the ability to screen small compound libraries at a rate of 1000 compounds a day.
Footnotes
Acknowledgements
We thank Dr. Lisa Middleton for the critical editing of the manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by NIH R01 grant GM102443-01 for Ahmed Chadli.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
