Abstract
Allophycocyanin and c-phycocyanin have been reported to be potent antioxidants. In this work, the genes encoding the apo-proteins of allophycocyanin α (ApcA), allophycocyanin β (ApcB), c-phycocyanin α (CpcA), and c-phycocyanin β (CpcB) from Spirulina platensis were cloned, and the recombinant proteins were produced in Escherichia coli to study their antioxidant effects. All four recombinant phycocyanins could be produced in the soluble form and purified to more than 97% purity. The results of radical scavenging assays showed that the Trolox equivalent values for peroxyl radical scavenging by the ApcA, ApcB, CpcA, and CpcB proteins were 1.81 ± 0.2 µM, 1.98 ± 0.22 µM, 0.95 ± 0.15 µM, and 1.49 ± 0.15 µM, respectively. The IC50 values for hydroxyl radical scavenging of ApcA, ApcB, CpcA, CpcB, and Trolox were 269 ± 9 µg/mL, 190 ± 5 µg/mL, 129 ± 8 µg/mL, 108 ± 4 µg/mL, and 195 ± 12 µg/mL, respectively. These results indicated that allophycocyanin exhibited higher activity than c-phycocyanin in scavenging peroxyl radicals, whereas c-phycocyanin exhibited higher activity than allophycocyanin in scavenging hydroxyl radicals. All of the apo-phycocyanin subunits possessed strong antioxidant activities and can be further developed and applied to the food and drug industries. However, the selection of the most useful antioxidant should depend on the type of targeted free radical to obtain the highest efficiency.
Introduction
Recently, there has been an interest in the use of antioxidant nutrition to promote health. Antioxidants could help prevent heart disease, cancer, and aging. Spirulina platensis is one supplementary food that is rich in nutrients such as gamma-linolenic acid, linoleic acid, stearidonic acid, docosahexaenoic acid, and vitamins.1,2 In addition, Spirulina platensis contains a macromolecule called phycobilisome that is composed mainly of two phycobiliproteins: allophycocyanin (Apc) and c-phycocyanin (Cpc). Both phycobiliproteins play a role in photosynthesis by absorbing light and transferring the energy to chlorophyll on thylakoid membranes. Phycobiliproteins are composed of a number of α and β subunits assembled in either the trimeric (αβ)3 form or the hexameric (αβ)6 form. Each subunit is attached to at least one cyanobilin chromophore. Cpc is a major phycobiliprotein in cyanobacteria, whereas Apc is found in a lower amount.
It has previously been reported that Apc and Cpc extracted from cyanobacteria exhibit several beneficial activities, including antioxidant activity, 3 antitumor activity, 4 anti-inflammatory effects, 5 and neuroprotective effects. 6 Previously, there was a belief that the bilin chromophore was solely responsible for the antioxidant activities of phycobiliproteins.7,8 Later, much research was conducted to clone and express the genes encoding phycobiliproteins in bacteria. Apo-phycocyanins were produced and tested for antioxidant activities.9–11 Since then, it has been observed that the protein moiety itself also possesses significant antioxidant activity. Ge et al. 12 and Zhang et al. 9 produced the apo-Apc α and β subunits in Escherichia coli as histidine-tagged proteins. They found that the apo-Apc subunit can scavenge hydroxyl and peroxyl radicals. Guan et al. 10 reported that the recombinant apo-Cpc α subunit, produced by a similar cloning strategy, could also scavenge hydroxyl and peroxyl radicals. Our recent work demonstrated potent radical scavenging activity by the recombinant apo-Cpc β subunit. Furthermore, we evaluated the antioxidant effects of recombinant apo-Cpc in a cellular model, human red blood cells, and found that this protein can protect red blood cells from antioxidative damage. 11 Therefore, the apo-phycocyanin subunit seems to be an effective antioxidant worth developing as a pharmaceutical product.
To provide insight into the benefits of phycobiliproteins, we aimed to compare four apo-phycobiliproteins in terms of their production efficiency and antioxidant activity. The genes encoding allophycocyanin α (ApcA), allophycocyanin β (ApcB), c-phycocyanin α (CpcA), and c-phycocyanin β (CpcB) from S. platensis were cloned and introduced into E. coli. Their expression levels, solubility, and antioxidant activities were studied and compared.
Methods
Primer Design and PCR Amplification
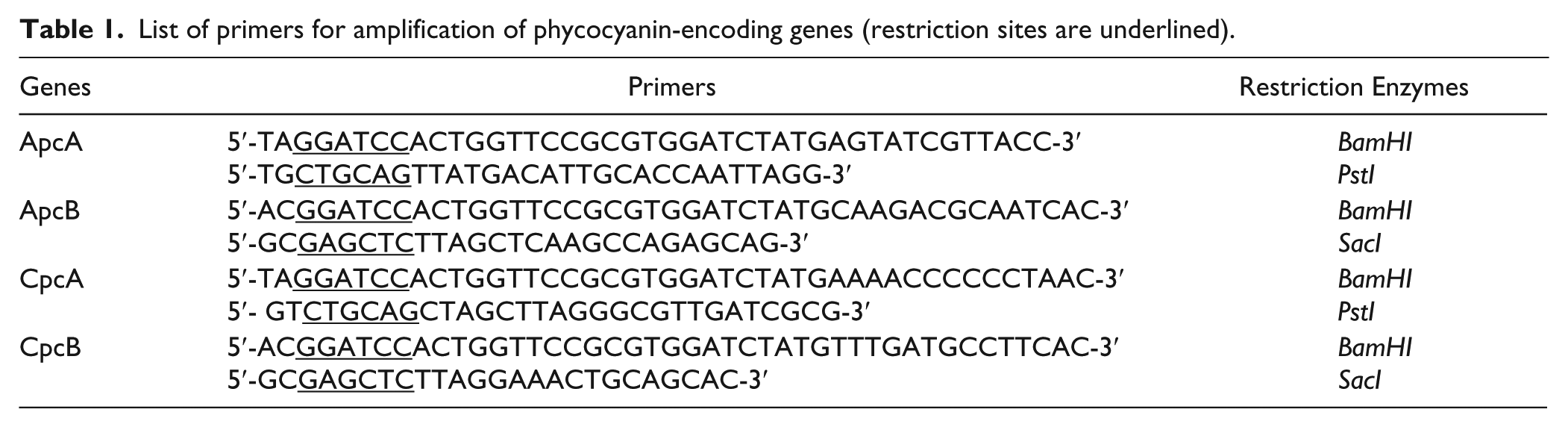
Genomic DNA was extracted from S. platensis using Trizol-reagent (Invitrogen, Carlsbad, CA) and the method recommended by the manufacturer. Primers were designed for the amplification of the genes encoding ApcA, ApcB, CpcA, and CpcB in S. platensis using the GenBank Nucleotide Sequence Database (accession No. AP011615). Restriction sites were added into each primer ( Table 1 ) to facilitate further cloning. Gene amplification was performed by PCR using Phusion Pfu DNA polymerase (Thermo Scientific, Waltham, MA).
List of primers for amplification of phycocyanin-encoding genes (restriction sites are underlined).
Construction of Expression Vectors
To generate the plasmids carrying the genes for the production of the four phycocyanin proteins, the PCR products obtained from the previous steps were inserted into the pGem-T easy vector (Promega, Madison, WI) by the TA cloning procedure. The genes encoding ApcA, ApcB, CpcA, and CpcB were then subcloned into the pETDuet-1 vector (Novagen, Germany) using the restriction endonucleases listed in Table 1 and T4 DNA ligase (Promega). The genes were located downstream of the T7 promoter and the hexahistidine sequence. The resulting plasmids were called pETDApcA, pETDApcB, pETDCpcA, and pETDCpcB ( Fig. 1 ). E. coli DH5α was used for the propagation of the recombinant plasmids. The transformations were performed by the CaCl2 method. 13 The bacteria were cultured at 37 °C overnight in Luria-Bertani (LB) medium containing 100 µg/mL ampicillin (Sigma-Aldrich, St. Louis, MO). The recombinant clones were checked for the presence of the desired phycocyanin gene by restriction endonuclease analysis. The gene sequences were confirmed by DNA sequencing.
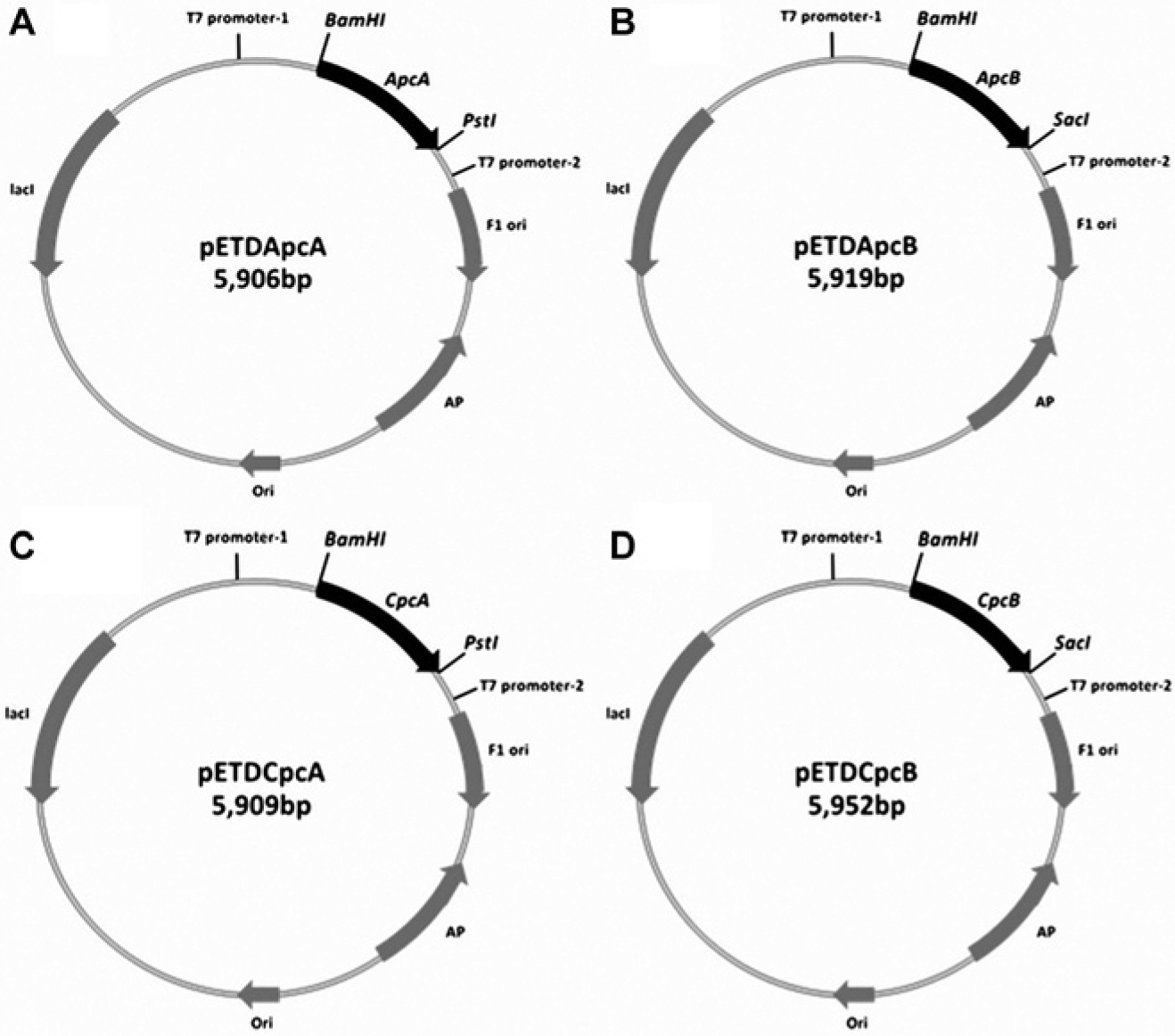

Schematic representations of the expression vectors pETDApcA, pETDApcB, pETDCpcA, and pETDCpcB.
Gene Expression and Protein Purification
Plasmid construction was performed according to Pleonsil et al. 11 For the production of the phycocyanin proteins, E. coli BL21 was transformed with pETDApcA, pETDApcB, pETDCpcA, or pETDCpcB. The cells were grown in LB medium containing 100 µg/mL ampicillin at 37 °C with shaking until the optical density at 600 nm reached 0.40 to 0.50. After that, the expression of the recombinant gene was induced by the addition of 0.5 mM isopropyl β-D-thiogalactoside (IPTG; Sigma-Aldrich). The culture temperature was then reduced to 20 °C, and the incubation continued for 10 h. The cells were harvested by centrifugation at 4500×g for 10 min at 4 °C and washed with IMAC equilibration buffer (50 mM NaH2PO4-Na2HPO4/300 mM NaCl, pH 7.4). The cell pellets were resuspended in phosphate-buffered saline with 0.1 mM PMSF. Cell lysis was performed by ultrasonication (Sonic, Austin, TX) with a program of pulse on for 5 s and pulse off for 5 s for three cycles. The cell lysate was centrifuged at 12,000×g at 4 °C for 20 min. The soluble extracts were collected and analyzed by 12% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE).
Each recombinant protein was purified by immobilized metal ion affinity chromatography (IMAC) with His60 Ni Superflow resin (Clontech, Mountain View, CA) following the procedure provided by the manufacturer. Briefly, the soluble extract was loaded into an IMAC-Ni column. The column was washed with 10 column volumes of equilibration buffer (50 mM NaH2PO4-Na2HPO4/300 mM NaCl, pH 7.4), 2 column volumes of washing buffer 1 (50 mM NaH2PO4-Na2HPO4/300 mM NaCl/20 mM imidazole, pH 7.4), and 2 column volumes of washing buffer 2 (50 mM NaH2PO4-Na2HPO4/300 mM NaCl/40 mM imidazole, pH 7.4). Finally, the recombinant protein was eluted using elution buffer (50 mM NaH2PO4-Na2HPO4/300 mM NaCl/300 mM imidazole, pH 7.4) at a flow rate of 1 mL/min. The 2 mL eluted fractions were collected, and the protein concentration was measured using protein assay reagent (Bio-Rad, Hercules, CA). The fractions with high protein concentrations were pooled and dialyzed against phosphate buffer at pH 7.4 containing 0.1% v/v tween 20/50 mM NaH2PO4-Na2HPO4/150 mM NaCl. The purified proteins were stored at −20 °C until further analysis.
Oxygen Radical Absorbance Capacity Assay
The oxygen radical absorbance capacity (ORAC) assay was selected to assess the antioxidant capacity of the recombinant proteins for peroxyl radical scavenging. This assay was performed according to Dávalos et al. 14 with the following modifications. Briefly, AAPH at a final concentration of 75 mM (Merck, Germany) was dissolved in 75 mM phosphate buffer (NaH2PO4-Na2HPO4, pH 7.4). A 4 mM fluorescein stock solution (Sigma-Aldrich) was made in 75 mM phosphate buffer (NaH2PO4-Na2HPO4, pH 7.4) and stored in the dark at 4 °C. The stock fluorescein solution was diluted 1:500 with 75 mM phosphate buffer (NaH2PO4-Na2HPO4, pH 7.4) immediately before use. A volume of 150 µL of the working fluorescein solution was added to the 96-well microplate, and 25 µL of each sample was then added and mixed well. The plate was incubated for 30 min at 37 °C. After incubation, 25 µL of a 75 mM AAPH solution (freshly prepared) was added into the reaction wells. The fluorescence intensity (excitation at 485 nm and emission at 528 nm) was measured every minute for 40 min using a Synergy Mx microplate reader (BioTek US, Seattle, WA), and 75 mM phosphate buffer (NaH2PO4-Na2HPO4, pH 7.4) was used as the blank. ApcA, ApcB, CpcA, CpcB, and Trolox (Sigma-Aldrich) were diluted to concentrations of 3, 6, 9, 12, and 15 µM and subjected to assays to determine their scavenging activities. The ORAC values were calculated as described by Cao and Prior. 15 The area under the curve (AUC) and the net AUC of the standards and the samples were determined using Gen5 Data Analysis Software (BioTek US).
The net AUC represented the antioxidant capacity of free radical inhibition and was plotted against the concentration of the protein. To determine the Trolox equivalent (TE) of each recombinant protein, the slope of the linear regression of the protein was divided by the slope of the linear regression of Trolox.
Hydroxyl Radical Scavenging Assay (Deoxyribose Assay)
The deoxyribose assay was performed to assess the antioxidant capacity of the recombinant proteins for hydroxyl radical scavenging. This assay was performed according to Ge et al. 12 with the following modifications. Briefly, the reaction mixture contained 20 mM phosphate buffer (KH2P04-KOH, pH 7.4), 2.8 mM deoxyribose, 100 µM FeCl3, 104 µM EDTA (final concentration), and the test sample (25, 50, 100, 200, or 400 µg/mL). The reaction was started by adding ascorbic acid to a final concentration of 100 µM and 1 mM H2O2. The reaction mixture (total of 500 µL) was incubated for 30 min at 37 °C. After incubation, the color was developed by adding 250 µL thiobarbituric acid (1% W/V in 50 mM NaOH) and 250 µL trichloroacetic acid (2.8% W/V in deionized water) and heating the mixture in a boiling water bath for 15 min. The sample was then allowed to cool and diluted twofold with 20 mM phosphate buffer (pH 7.4). The absorbance was measured at 532 nm. The reaction mixture without a sample was used as a blank. The hydroxyl radical scavenging activity was determined by measuring the percentage inhibition.
Statistics and Data Analysis
All measurements were repeated at least 3 times. The data are given as the mean ± SEM. Statistical analysis was performed with SPSS 17.0, and the graph in this article was generated by Microsoft Office Excel 2007 for Windows.
Results
Recombinant Plasmid Construction
Using the designed primers, PCR products of 486 bp (ApcA), 486 bp (ApcB), 489 bp (CpcA), and 519 bp (CpcB) were amplified from S. platensis genomic DNA. Recombinant plasmids carrying the genes encoding ApcA, ApcB, CpcA, or CpcB were successfully constructed. The resulting DNA sequences showed that each gene was fused in frame to the hexahistidine-tagged sequence.
Gene Expression and Protein Purification
SDS-PAGE analysis showed that the genes encoding ApcA, ApcB, CpcA, and CpcB were overexpressed in E. coli, as shown in Figure 2A . Induction with IPTG resulted in visible protein bands of interest for E. coli cells containing the recombinant plasmids, whereas those distinct bands were not observed in noninduced cells. The bands migrated at approximately 19 to 20 kDa, corresponding to the calculated molecular weight of each recombinant protein (ApcA: 19.7 kDa, ApcB: 19.7 kDa, CpcA: 19.8 kDa, and CpcB: 20.4 kDa). Owing to the hexahistidine tag, which was included in the recombinant proteins, the proteins can then be purified to high purity using IMAC ( Fig. 2B ). More than 97% purity was obtained as analyzed by the GeneTool (SynGene, UK) program.
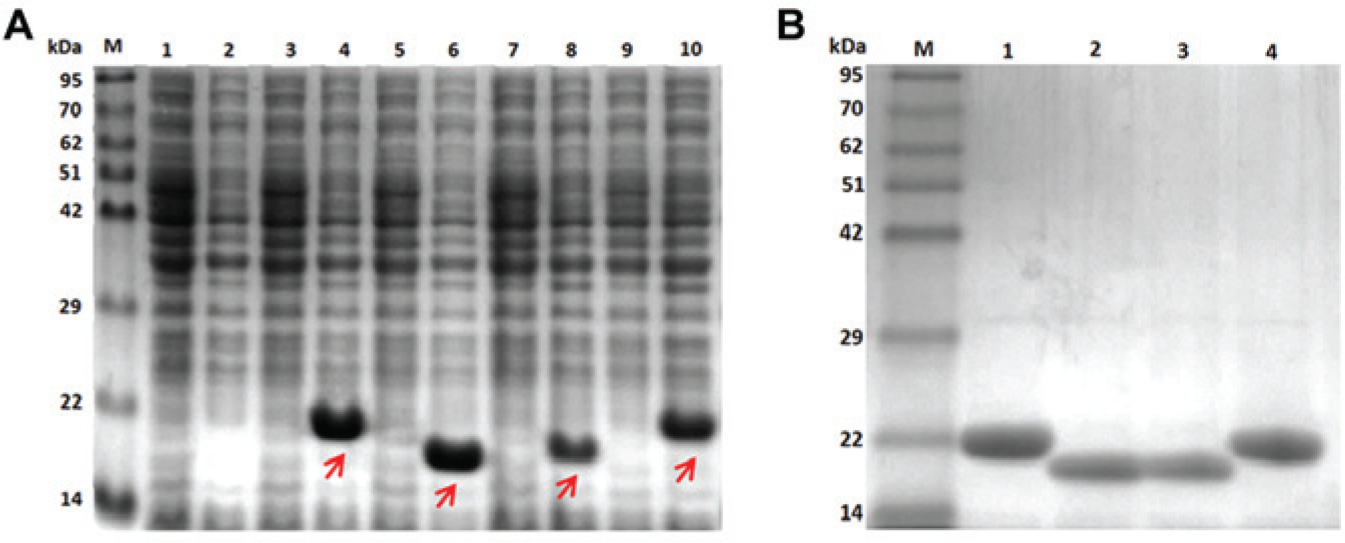
(
Peroxyl Radical Scavenging Activity
An ORAC assay was conducted to study peroxyl radical scavenging activity of ApcA, ApcB, CpcA, CpcB, and Trolox. The radical scavenging capacities of all the recombinant phycocyanins and Trolox are shown in Figure 3 . The TE values were calculated and are listed in Table 2 . The result showed that ApcB possessed the highest activity, followed by ApcA. The TE values differed very slightly between these two subunits. Among the four recombinant phycocyanins, CpcA exhibited the lowest activity but was still comparable with Trolox.
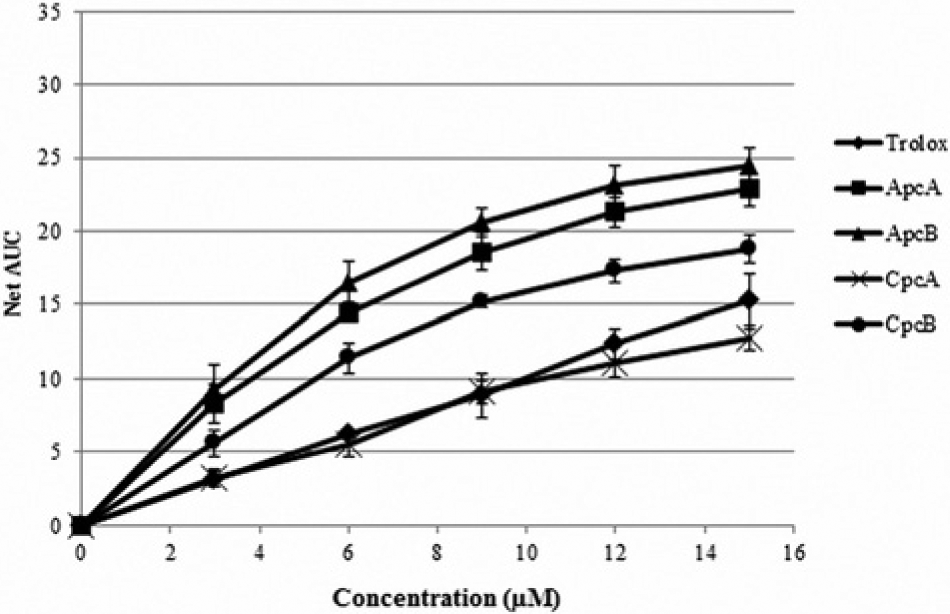
Peroxyl radical scavenging capacity of recombinant ApcA, ApcB, CpcA, CpcB, and Trolox determined by an oxygen radical absorbance capacity assay.
Trolox equivalent (TE) values and IC50 values representing the peroxyl and hydroxyl radical scavenging capacities of various proteins. a
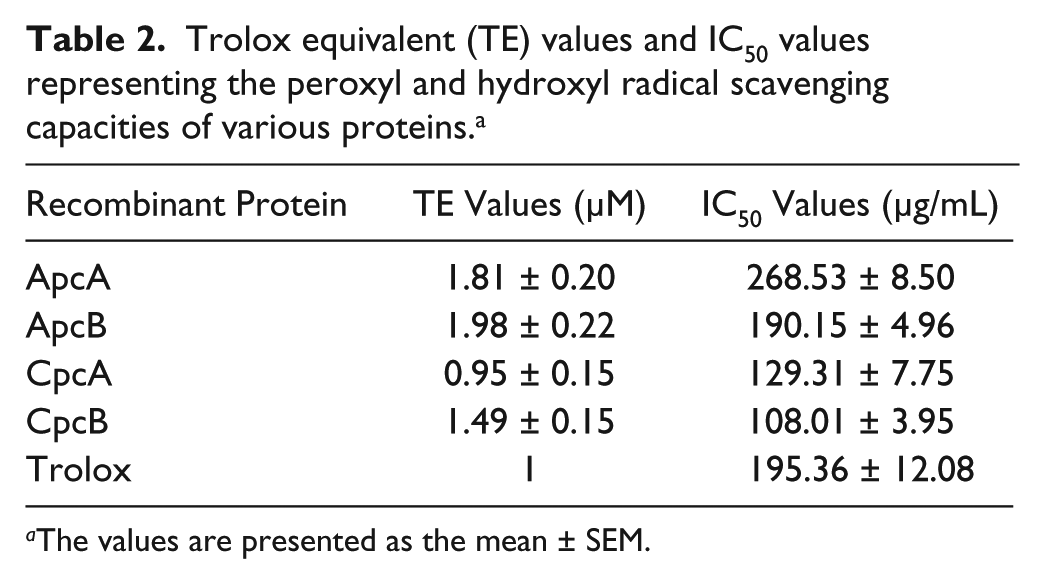
The values are presented as the mean ± SEM.
Hydroxyl Radical Scavenging Activity
The deoxyribose assay was conducted to study the hydroxyl radical scavenging activity of ApcA, ApcB, CpcA, CpcB, and Trolox. The percentages of radical inhibition of all the recombinant phycocyanins and Trolox are shown in Figure 4 . The IC50 values were calculated and are listed in Table 2 . The result showed that the beta subunits of Cpc exhibited higher hydroxyl radical scavenging activity in comparison with Trolox, whereas ApcB exhibited similar activity as Trolox. ApcA exhibited the lowest activity of all four recombinant phycocyanin proteins, and its activity was lower than that of Trolox.
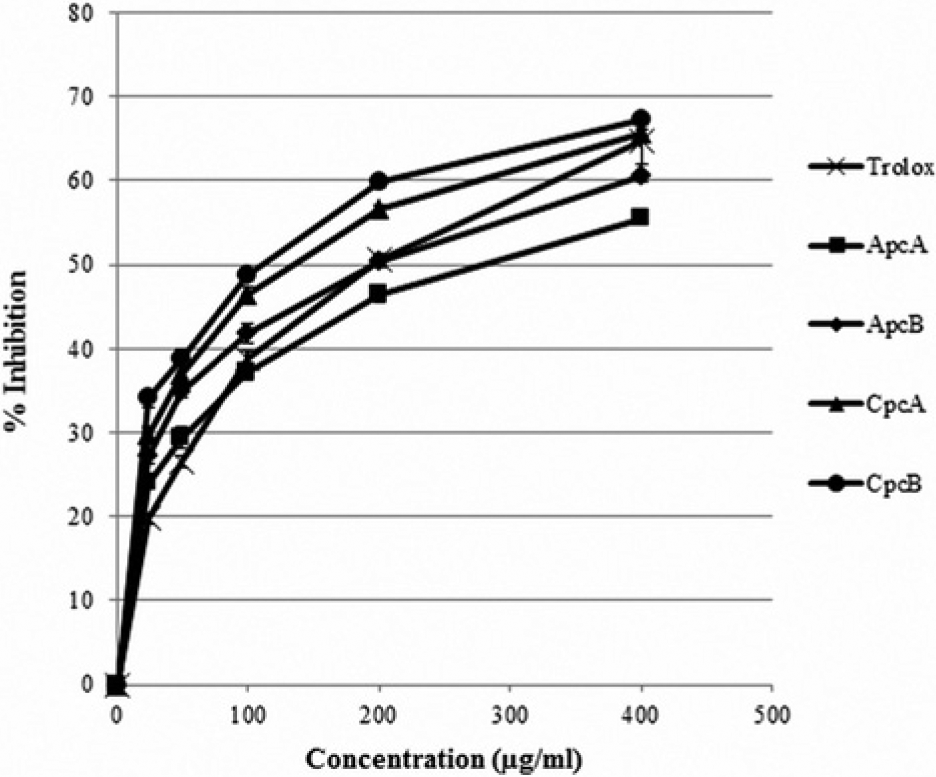
Hydroxyl radical scavenging capacity of recombinant ApcA, ApcB, CpcA, CpcB, and Trolox determined by a deoxyribose assay.
Discussion
Several research groups have reported that phycocyanins possess substantial antioxidant activity against free radicals.3,9,16 Most of the studies paid attention to the action of the bilin chromophore. Although it has been shown that the antioxidant property of phycocyanins could also be attributed to their protein moieties, there were only a few studies focused on the antioxidant activities of apo-phycocyanin.9,12 Molecular cloning has been widely used as a tool for the production of heterogeneous proteins in bacteria because of the rapid growth rate of bacteria and the reliability and the ease of large-scale production.13,17,18 However, the major obstacle that has to be overcome is that the recombinant proteins often aggregate because of their overexpression or unfavorable environments. 19 Previous research has shown that using low IPTG concentrations and low temperatures during gene expression can increase the solubility of the recombinant proteins.18,20 Furthermore, Gräslund et al. 20 reported that using E. coli BL21 (DE3) as an expression host increases the solubility of the recombinant protein. 20 In this work, those strategies were employed for the production of four recombinant phycocyanins. Recombinant ApcA, ApcB, CpcA, and CpcB were expressed in large amounts as soluble proteins. To avoid the complicated process of purification,21–23 hexahistidine tags were included the recombinant phycocyanin sequences. The proteins could therefore be purified using one-step IMAC, which minimized the loss of the protein during the purification process.
When analyzing the proteins with SDS-PAGE, the result showed that the recombinant phycocyanins did not migrate to the exact position expected from the calculated molecular weights. This might be a result of the different hydrophobicities of the proteins as explained in previous reports.24–26 Normally, the binding of SDS to the proteins gives them negative charges and causes them to retain their primary structures. The proteins will then migrate toward the positive pole with the mobility rates depending on their molecular weights. It has been shown that highly hydrophilic proteins and acidic proteins possess low electrophoretic mobility due to their poor binding to SDS. This might be the case for our recombinant proteins. The hydrophilicity of ApcA, ApcB, CpcA, and CpcB were 27%, 23%, 23%, and 21%, respectively (calculated by a peptide property calculator, www.genscript.com). The phenomenon that the β subunit migrated slower than the α subunit in SDS-PAGE, although the molecular weight determined by mass spectrometry was nearly the same, was previously observed for Cpc. 27
Herein, recombinant apo-phycocyanin subunits, which are the major phycobiliproteins in cyanobacteria, were produced, and a comparative study of their antioxidant activities was performed. Although holo-phycocyanins have higher antioxidant capacity than their apo form,9,12 the production of recombinant holo-phycocyanins is tedious. The co-expression of heme oxygenase, bilin reductase, and phycocyanobilin lyase is essential for the synthesis of the bilin chromophore and the incorporation of bilin into phycocyanin in bacteria. 28 In addition, heme supplementation might be required for E. coli to produce enough substrate for bilin synthesis.28,29 By studying apo-phycocyanins, the antioxidant capacity of the subunits can be compared regardless of the number of bilins. The antioxidant activities of the apo-phycocyanins might be attributed to two amino acids that play important roles in free radical scavenging, cysteine, and methionine. Atmaca 30 reported that both amino acids contain sulfur, which can transfer electrons or hydrogen atoms to free radicals. Especially when these amino acids are located on protein surfaces, they become more accessible to free radicals.31,32 Accordingly, the amino acid content and location influence the antioxidant capacity of the protein. In this work, peroxyl and hydroxyl radicals were chosen for the evaluation of the free radical scavenging capacities of recombinant ApcA, ApcB, CpcA, and CpcB. Among tested proteins, ApcB and CpcB have the highest activity in scavenging peroxyl and hydroxyl radicals, respectively. Allophycocyanin exhibited higher scavenging activity of peroxyl radicals than c-phycocyanin, whereas c-phycocyanin exhibited higher scavenging activity of hydroxyl radicals than allophycocyanin. For both allophycocyanin and c-phycocyanin, the β subunit possessed higher scavenging activity than the α subunit, although no significant difference was observed for some pairs. Compared with Trolox, a soluble derivative of vitamin E, the recombinant phycocyanin subunit showed higher or equal scavenging activity. Only ApcA possessed significantly lower scavenging activity of hydroxyl radicals than Trolox. These results suggested that apo-phycocyanins could be excellent candidates for antioxidant food supplements. To provide the most benefit, the selection of a suitable subunit for further applications is important. More information regarding comparisons of production efficiency should be obtained.
In conclusion, this report describes, for the first time, a comparison of the antioxidant activities of four species of apo-phycocyanin. Recombinant α and β subunits of Apc and Cpc were successfully produced in E. coli. ApcB was the most potent antioxidant against peroxyl radicals, whereas CpcB was the most potent antioxidant against peroxyl radicals. Both ApcB and CpcB are small, highly water-soluble molecules, and they can be produced in high yields. Therefore, they should be excellent targets for the development of specific radical scavengers. Future studies should be directed toward improving their antioxidant activities by studying their mechanism of radical scavenging, and genetic modifications could further improve their activities.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The authors are grateful for financial support from the Chulalongkorn Centenary Academic Development Project and the Ratchadaphiseksomphot Endowment Fund on the occasion of the 90th anniversary of Chulalongkorn University (2/2556).
