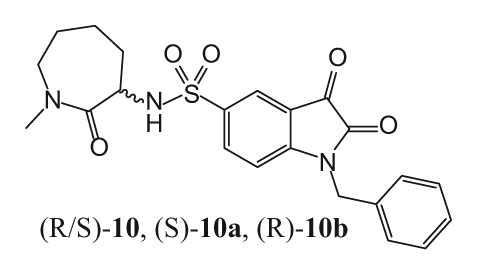
Abstract
The resurgence of interest in caspases (Csp) as therapeutic targets for the treatment of neurodegenerative diseases prompted us to examine the suitability of published nonpeptidic Csp-3 and Csp-6 inhibitors for our medicinal chemistry programs. To support this effort, fluorescence-based Csp-2, Csp-3, and Csp-6 enzymatic assays were optimized for robustness against apparent enzyme inhibition caused by redox-cycling or aggregating compounds. The data obtained under these improved conditions challenge the validity of previously published data on Csp-3 and Csp-6 inhibitors for all but one series, namely, the isatins. Furthermore, in this series, it was observed that the nature of the rhodamine-labeled substrate, typically used to measure caspase activity, interfered with the pharmacological sensitivity of the Csp-2 assay. As a result, a liquid chromatography/tandem mass spectrometry–based assay that eliminates label-dependent assay interference was developed for Csp-2 and Csp-3. In these label-free assays, the activity values of the Csp-2 and Csp-3 reference inhibitors were in agreement with those obtained with the fluorogenic substrates. However, isatin
Introduction
Over the past 40 years, protease research has yielded important drugs, most notably for cardiovascular and viral diseases. 1 In the past decade, several proteases have also been implicated in neurodegenerative disorders, and more recently, caspase (Csp)–6 has been proposed as a target for therapeutic intervention in both Alzheimer 2 and Huntington 3 diseases (HD). The clinical successes in the protease field have been possible because the available peptidomimetic protease inhibitors satisfied the practical pharmacological requirements of peripheral drugs. However, the use of peptidomimetics for CNS indications presents a much greater challenge, particularly for cysteine proteases such as the caspase family members. The first-generation caspase inhibitors were tetrapeptides mimicking caspase substrate cleavage sites and containing a reactive moiety at the C-terminus designed to react with the catalytic cysteine. Although efforts in the de-peptidization of such covalent inhibitors produced highly potent, selective, and cell-active Csp-1 and Csp-3 compounds, these new generations of inhibitors still lack suitable physicochemical properties to achieve acceptable peripheral exposure and, a fortiori, adequate CNS coverage. Consequently, nonpeptidic inhibitors have been sought as unique starting points for the development of CNS-active compounds.
After more than a decade of high-throughput screening (HTS) campaigns, there are still very few reports of nonpeptidic caspase inhibitors, and there is an absence of advanced lead series for the caspases implicated in neurological disorders. Due to this paucity of benchmark nonpeptidic caspase inhibitors and the poor historical HTS tractability of this target class, the current lead identification program for HD focused initially on the in vitro characterization of eight previously published nonpeptidic caspase inhibitor series ( Fig. 1 ) to determine their suitability as lead structures.
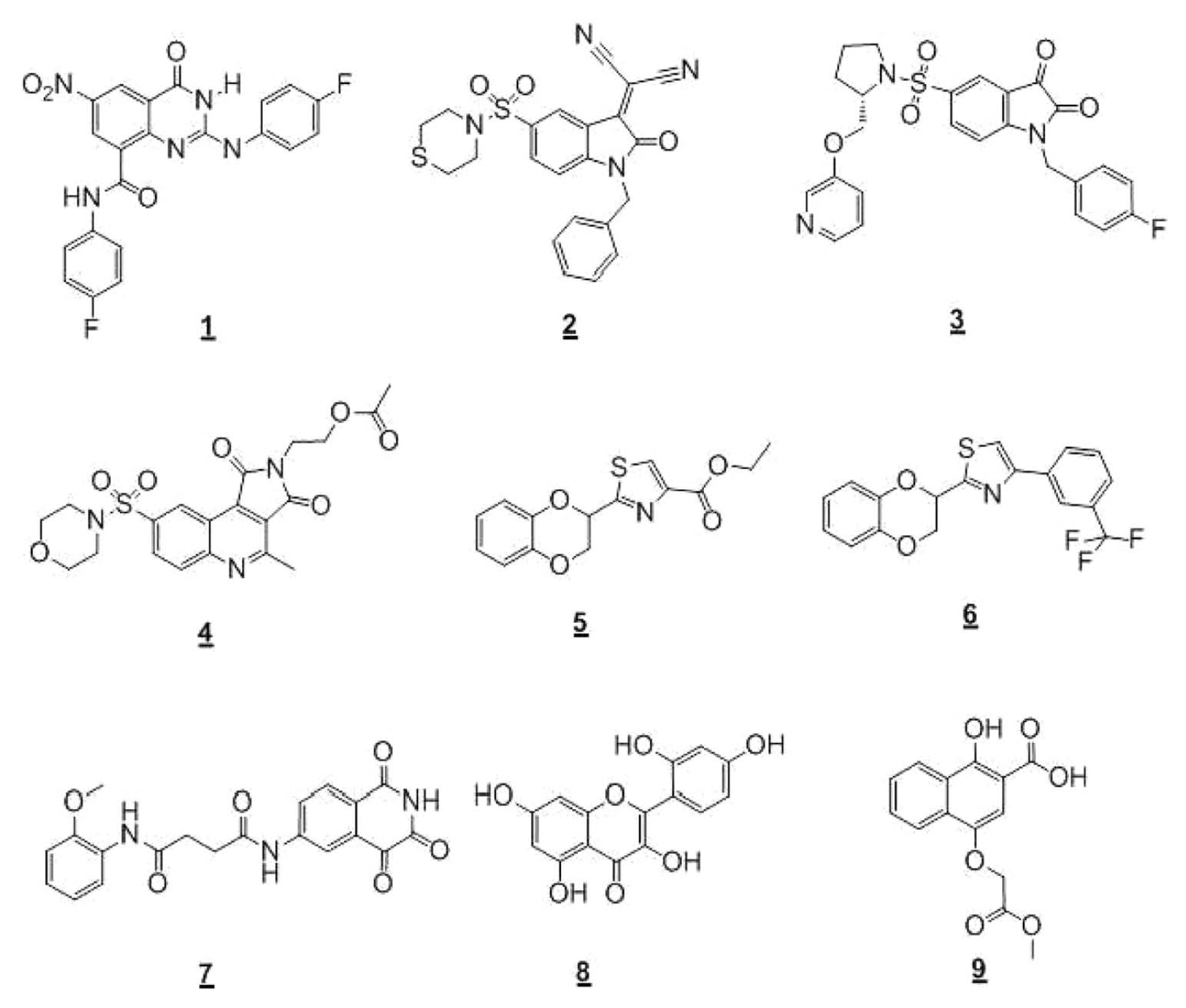
Structures of investigated nonpeptidic caspase inhibitors
This article describes the optimization of Csp-2, Csp-3, and Csp-6 fluorescence-based enzymatic assays and highlights inconsistencies with previous studies. It was found that seven of the eight investigated chemical series did not confirm in the Csp-3 or Csp-6 assays. The development of novel, label-free liquid chromatography/tandem mass spectrometry (LC/MS/MS) Csp-2 and Csp-3 assays as secondary, orthogonal, and label-free readout assays for confirming potency and selectivity of the only validated series is also described, and assay results are discussed in comparison with those obtained by the fluorometric method. Although there is increasing interest in mass spectrometric readouts of catalytic end-point assays, 4 there are only few examples describing such readouts for caspases or cysteine-proteases. 5 The assays described in this publication are designed for compound profiling and could help close that gap.
Materials and Methods
Fluorescence-Based Caspase Assays
These assays are based on the cleavage of a fluorogenic substrate composed of the recognition sequence of the individual caspases and a fluorophore (either rhodamine or coumarin). All assays were performed in a 384-well microplate format (Matrix Screen Mate; Thermo Fisher Scientific, Hudson, NH) using a total assay volume of 20 µL per well. Standard buffer conditions for Csp-2 consisted of 50 mM MES (pH 6.5), 150 mM NaCl, and 1.5% sucrose. Assay buffer consisted of 50 mM HEPES (pH 7.4) and 100 mM NaCl for Csp-3 and 20 mM PIPES (pH 7.2), 100 mM NaCl, 1 mM EDTA, and 10% sucrose for Csp-6. Buffer additives such as the detergents 3[(3-cholamidopropyl)-dimethylammonio]-propanesulfonic acid (CHAPS), Pluronic F-127 (PF127), Triton X-100 (TX100), and the reducing agents dithiothreitol (DTT), glutathione (GSH), and β-mercaptoethanol (β-MCE) were varied as indicated in the Results section ( Table 1 and Fig. 2 ).
In Vitro Inhibitory Potencies of Compounds
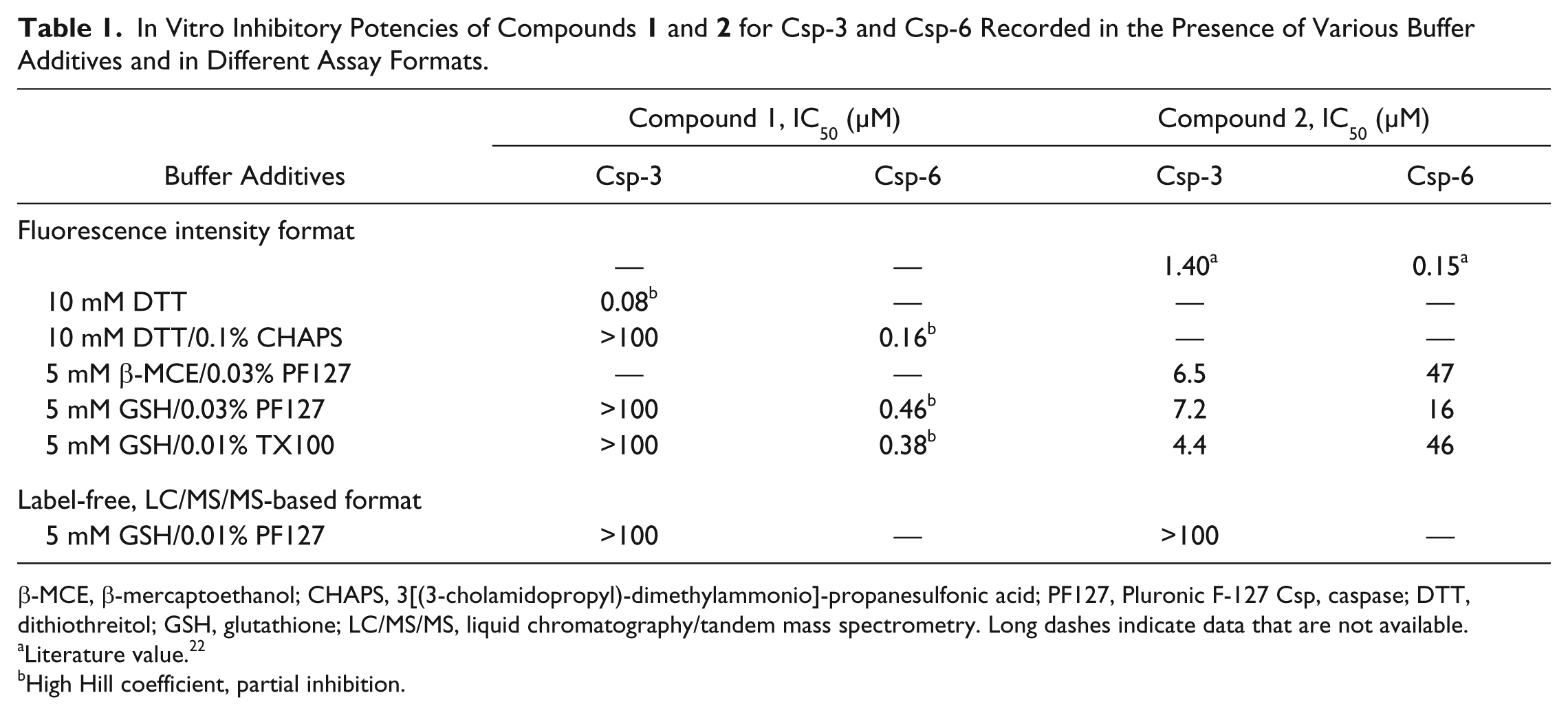
β-MCE, β-mercaptoethanol; CHAPS, 3[(3-cholamidopropyl)-dimethylammonio]-propanesulfonic acid; PF127, Pluronic F-127 Csp, caspase; DTT, dithiothreitol; GSH, glutathione; LC/MS/MS, liquid chromatography/tandem mass spectrometry. Long dashes indicate data that are not available.
Literature value. 22
High Hill coefficient, partial inhibition.

Concentration-response curves of  ), glutathione (GSH)/Triton X-100 (
), glutathione (GSH)/Triton X-100 ( ), GSH/Pluronic F-127 (PF127) (
), GSH/Pluronic F-127 (PF127) ( ), β-mercaptoethanol/PF127 (
), β-mercaptoethanol/PF127 ( ), or DTT without detergent (
), or DTT without detergent ( ). Experimental details are described in the Materials and Methods. Csp, caspase.
). Experimental details are described in the Materials and Methods. Csp, caspase.
The peptide-based inhibitors Ac-VDVAD-CHO (Csp-2), Ac-DEVD-CHO (Csp-3), and Ac-VEID-CHO (Csp-6) were used as reference compounds on each individual assay plate. Reference and test compound stock solutions were prepared in neat DMSO. The 3-fold dilution series with 11 concentrations were prepared in 100% DMSO. Then, 200 nL of the compound dilutions in 100% DMSO were added to each well containing 10 µL of freshly prepared solutions of 20 nM Csp-2, 0.6 nM Csp-3, or 8 nM Csp-6 in the respective assay buffers using a CyBi-Well vario manipulator (CyBio AG, Jena, Germany). Assay plates were incubated at 37 °C for 5 min before the enzymatic reaction was started by adding 10 µL of the substrate solution. Final substrate concentrations were 20 µM for Csp-2 R110- or AMC-based substrates (-VDVAD-) and 10 µM R110- or AMC-based substrates (-DEVD- or -VEID-), for Csp-3 and Csp-6, respectively. These concentrations correspond to the individually determined apparent KM values (
In Csp-2 substrate competition experiments, IC50 values of selected Csp-2 compounds were recorded in standard Csp-2 buffer conditions with 5 mM GSH and 0.03% PF127 (w/v) in the presence of 20 µM, 100 µM, or 200 µM of Ac-VDVAD-AMC to investigate the competing effect between substrate and inhibitor.
Fluorescence was quantified using a Tecan Safire 2 microplate fluorometer (Tecan Austria GmbH, Grödig, Austria) with the following wavelength settings: excitation at 485 nm and emission at 535 nm for R110 and excitation at 380 nm and emission at 440 nm for coumarin 120. Under all conditions, it was ensured that the experiments performed for determining compound potencies were conducted in the linear range of the reaction. The sources of the other reagents used in this study are given in the supplemental information.
LC/MS/MS-Based Caspase Assays
In these assays, the enzymatic activities of Csp-2 and Csp-3 were determined by measuring the N-terminal cleavage products of the unlabeled peptides (Ac-VDVADGTQAS-NH2 and Ac-MDLNDGTQAS-NH2 for Csp-2 and Ac-MDEVDLASCD-NH2 for Csp-3). The assays were run in 384-well plates (Matrix Screen Mate; Thermo Fisher Scientific) in a final volume of 30 µL per well. The buffer consisted of 50 mM MES (pH 6.5), 150 mM NaCl, 5 mM GSH, and 0.03% PF127 (w/v), and the final Csp-2 concentration was 20 nM for the Csp-2 assay. For the Csp-3 assay, the buffer consisted of 50 mM HEPES (pH 6.5), 150 mM NaCl, 0.03% PF127 (w/v), and 5 mM GSH, and the concentration of Csp-3 was 1 nM. Compound handling was conducted in the same way as for the fluorogenic assays. The reaction was started by adding the substrate solution. The final substrate concentrations of 100 µM and 20 µM corresponded to the KM values determined for Csp-2 and Csp-3, respectively (
Substrate and product peptide detection was achieved by multiple-reaction monitoring (MRM) using an Agilent 1100 series chromatography system with a photodiode array detector (Agilent Technologies, Waldbronn, Germany) connected to an API 3000 mass spectrometer (Life Technologies, Darmstadt, Germany). Source parameters were set as follows: polarity ES+, capillary 5.5 kV, desolvation temperature 550 °C, nebulizer gas 8, curtain gas 8, and entrance potential 10; further analyte-specific parameters are shown in
Analytes were separated using an Acquity UPLC BEH C18 column (1.7 µm; 2.1 × 30 mm, Waters, Eschborn, Germany) at 60 °C column temperature. Mobile phases:
Analytes were quantified using MRM methods by comparing the relative intensities (integration of peak areas under the curve, AUC) obtained in incubations with different concentrations. Software used for mass spectrometry data acquisition and analysis was Analyst 1.4.1 (MDS Sciex, Concord, Canada). Transitions monitored are summarized in the supplemental information (
Data Analysis
To confirm the statistical validity of the assays, Z′-factors were calculated using the median values and standard deviations of the negative (1% DMSO) and positive controls (1 µM peptide-based inhibitors Ac-VDVAD-CHO, Ac-DVED-CHO, and Ac-VEID-CHO for Csp-2, Csp-3, and Csp-6 respectively). All screening runs with a Z′-factor >0.5 were accepted for further analysis (
Synthesis of Substrates and Inhibitors
Label-free substrates were synthesized using standard Fmoc solid-phase peptide synthesis protocols (for more details, see supplemental information). Compounds
In Vitro Caspase-3 Inhibitory Potency of Compounds
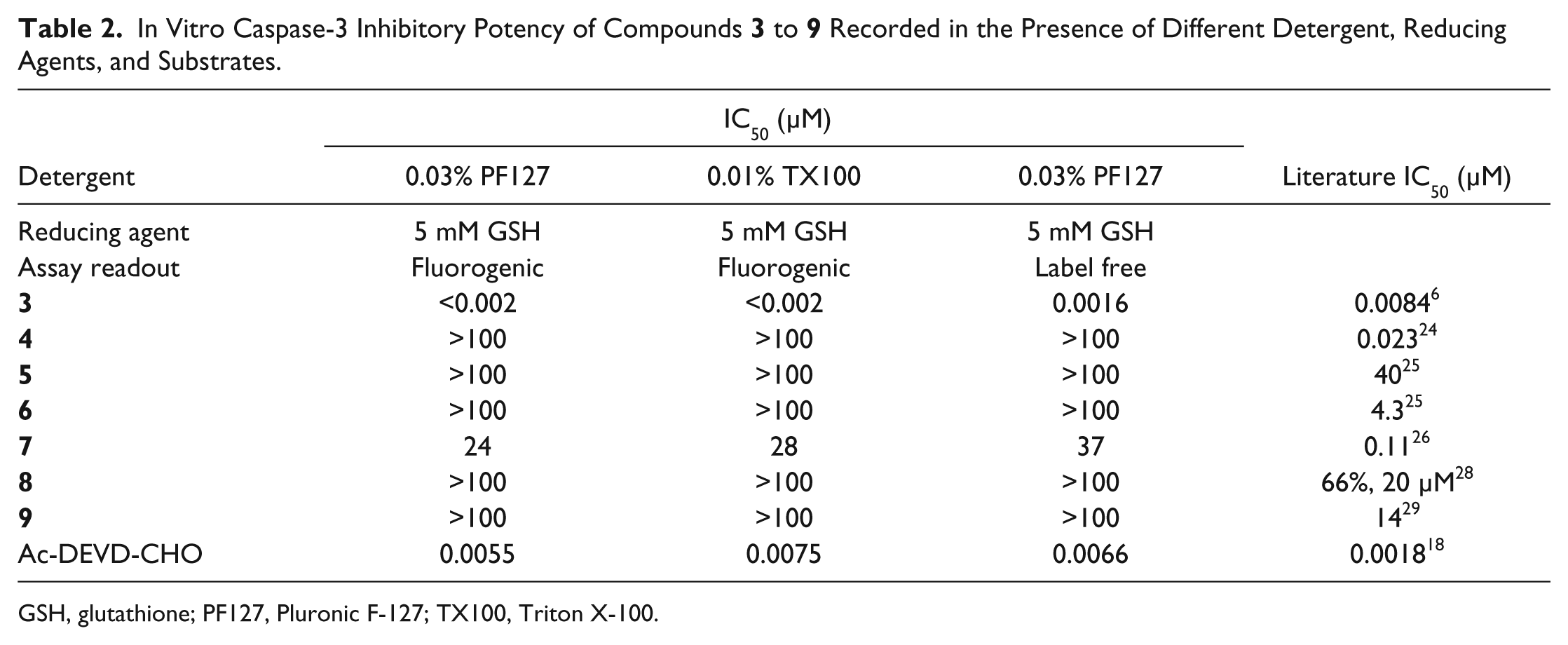
GSH, glutathione; PF127, Pluronic F-127; TX100, Triton X-100.
In Vitro Csp-2 and Csp-3 Inhibitory Potency for the Isatin Series
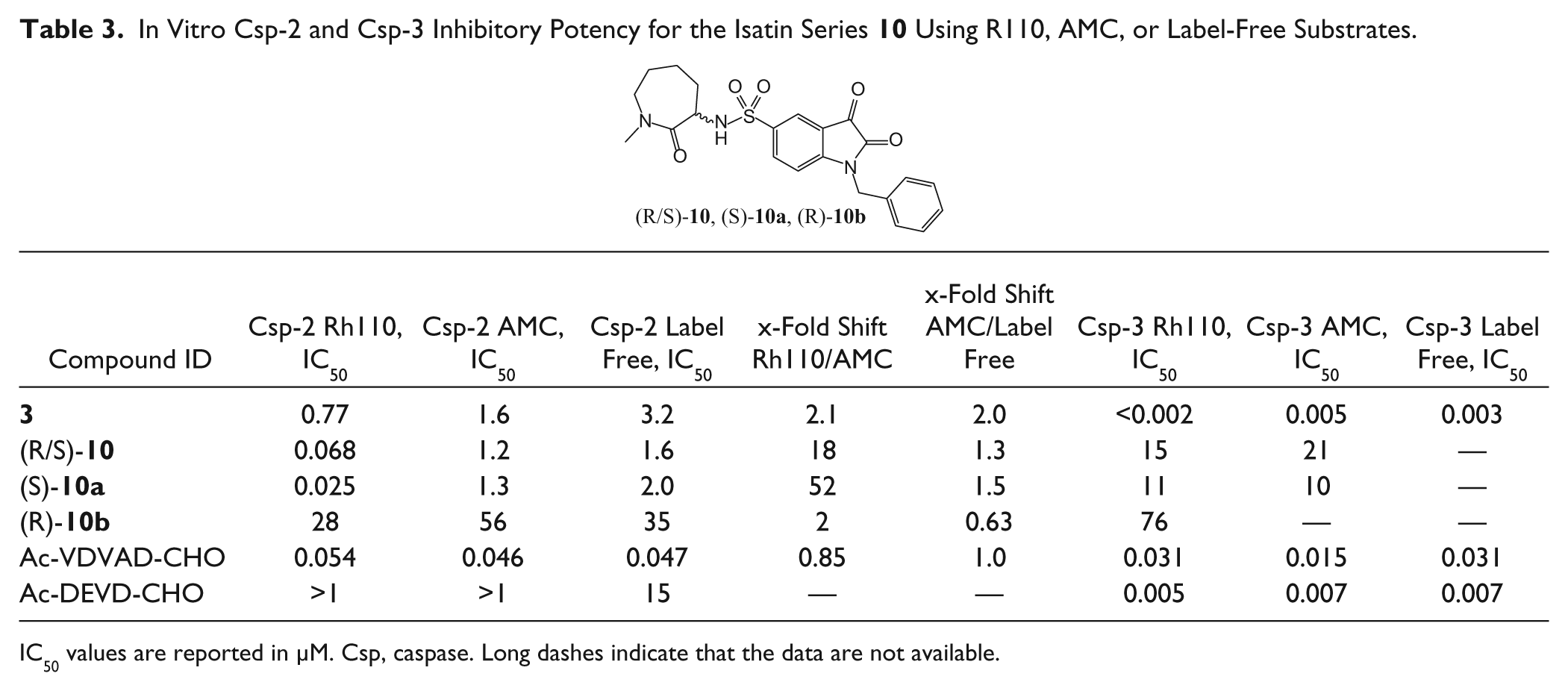
IC50 values are reported in µM. Csp, caspase. Long dashes indicate that the data are not available.
Results and Discussion
Optimization of Fluorescence Assays
Enzymatic assays for Csp-2, Csp-3, and Csp-6 inhibitor screening were implemented using a fluorometric assay principle, which measures the increase in fluorescence after protease cleavage of a specific substrate. Sequence-optimized substrates as previously described were used in these assays.
7
These substrates show micromolar affinities for caspases as confirmed by substrate titration experiments, well suited for the identification of inhibitors during compound screening. Substrate concentrations for each caspase assay were adjusted to match their KM values (
Selection of Reducing Agent
Caspases are sensitive to oxidation and require a reducing environment to preserve enzymatic activity. To maintain the enzyme in a reduced state, DTT is commonly used. However, in an HTS campaign for Csp-8, DTT resulted in an unusually high false-positive hit rate attributed to the formation of reactive oxygen species (ROS).
8
DTT has also been shown to cause indirect inactivation of other enzymes susceptible to oxidation,
9
and two recent publications10,11 have stressed the need, whenever possible, to employ weaker reducing agents that do not lead to ROS generation. As there is no apparent requirement for a strong reducing agent to stabilize or enhance caspase activity, the aim was to assess compounds under more physiological assay conditions. GSH was therefore evaluated in titration experiments to determine a concentration that is effective in stabilizing enzyme activity (
Selection of Detergents
The importance of detergent for inhibitor screening and testing has been extensively studied and described by the Shoichet research group 12 and others.13,14 To summarize, detergents can ameliorate assay performance by stabilizing the enzyme and reducing nonspecific protein adsorption onto the plastic surface. Also, detergent sensitivity has been proposed as a criterion to weed out aggregation-based inhibitors. For caspases, 0.1% (w/v) CHAPS is universally employed for assaying peptidic and nonpeptidic inhibitors because it strongly increases enzyme activity, especially for Csp-6. 15 Nevertheless, zwitterionic detergents such as CHAPS, when used at millimolar concentrations, can negatively affect compound solubility. 16 Furthermore, at such high concentrations, the detergent itself may bind to the target, thereby interfering with the assay readout. Consequently, and since no study has examined the effect of detergents on promiscuous inhibitors for caspases, two nonionic detergents were evaluated during assay development: TX100, which has been reported to be effective in disrupting compound aggregates, 17 and PF127, which is a mild detergent, is compatible with cell cultivation, and has a very high critical micelle concentration. These two nonionic detergents alleviate false-negative inhibition due to compound precipitation because they can be employed at a lower concentration than zwitterionic detergents, which also limits the risk of detergent binding to the protein. As previously reported for CHAPS, 15 a dramatic increase in Csp-2, Csp-3, and Csp-6 catalytic activity was also observed in the presence of TX100 or PF127, reaching a maximum with 0.01% (w/v) and 0.03% (w/v), respectively. The KM values in substrate titration experiments were not affected by the different detergents or different detergent concentrations. Since the GSH/TX100 and GSH/PF127 combinations are less prone to interference by redox-cycling or aggregate-forming compounds, they were preferred over the commonly used DTT/CHAPS combination.
Evaluation of Csp-6 Inhibitors
There is evidence from an HD mouse model that the genetic mutation of the motif in Huntingtin (HTT), thought to be cleaved by Csp-6, alleviates the pathogenic effects of mutant HTT.
3
The replication, in part or fully, of this beneficial effect with a Csp-6 inhibitor would represent a key step toward the validation of this pathogenic mechanism and the development of a therapeutic agent. Therefore, the identification of Csp-6 small-molecule inhibitors was a high priority for the HD program, which started with the verification of the activity of the two published nonpeptidic inhibitors of Csp-6 and their respective selectivity against Csp-3 (
Fig. 1
, compounds
First, the sub-micromolar Csp-6 activity of compound
Compound
Evaluation of Csp-3 Inhibitors as Starting Point for the Design of Csp-2 or Csp-6 Inhibitors
In the absence of validation data for the two Csp-6 inhibitors
Inconsistent SAR in the Csp-2 Inhibitor Series Leads to the Development of an LC/MS/MS Assay
At this juncture, only the isatin
Need for Csp-2 Label-Free Assay: Choice of Substrate
In looking for a more physiologically relevant readout, we attempted to develop a secondary assay based on Csp-2 cleavage of an oligopeptide derived from the residues around the cleavage site of a Csp-2 protein substrate. Quenched fluorescent substrates described for caspases in the seminal work of Stennicke et al. 32 would satisfy the criteria, but the distal fluorophore and quencher could still introduce nonspecific interactions. Instead, we decided to develop a label-free detection method with an LC/MS/MS readout to rule out any reporter interference. In the context of a project whose goal is the identification of tool compounds for HD target validation, the sequence of amino acids around the HTT Csp-2 cleavage site 33 was chosen as the basis for the design of the oligopeptide substrates. In the particular case of Csp-2, it is well documented that a P5 residue is required for efficient cleavage. 34 For this reason, the oligopeptide was extended on the N-terminus side up to the P5 position. In the absence of C-terminus residue requirements for efficient cleavage, 34 the first five amino acids of the HTT sequence beyond the Csp-2 cleavage site were also included, resulting in a decapeptide Ac-MDLND↓GTQAS-NH2. It was to be expected that this decapeptide would be a substrate for Csp-2 as it contained the two key P1 and P4 Asp residues present in the optimal recognition sequence VDVAD. However, the other three residues, particularly the critical P5, were different in HTT than in the optimal sequence. Hence, a hybrid peptide, Ac-VDVAD↓GTQAS-NH2, was also synthesized in which the N-terminus HTT-derived sequence was replaced by the canonical P5–P1 VDVAD motif.
Label-Free Caspase Assay: Development and Validation
Csp-2 activity was determined by quantifying the concentrations of the substrate and the N-terminal cleavage product Ac-VDVAD using LC/MS/MS analysis. Based on the hydrophilic nature of this product, a C18 column was used to separate the N-terminal cleavage product from substrate and the C-terminal side product (
Fig. 3A
). Positive ion mode (ESI+) was selected as a detection method to quantify the amount of the N-terminal cleavage product. Instrument parameters were adjusted so that the LC/MS/MS method allowed monitoring of the substrate (Ac-VDVADGTQAS-NH2) and the cleavage products over a concentration range from 0.1 to 50 µM, with lower limits of detection for substrate, N-terminal (Ac-VDVAD), and C-terminal (GTQAS-NH2) cleavage pro-ducts of 0.1 µM, 0.2 µM, and 1.0 µM, respectively (
Fig. 3B
). Concentration-dependent response of the analytes was linear in the relevant concentration range, thus providing ideal readouts for monitoring the enzymatic cleavage reactions. For product quantification, the N-terminal cleavage product was selected because of the >80-fold higher sensitivity of the MRM method for this cleavage product combined with significant retardation on the chromatography column compared with the early eluting C-terminal cleavage product, which reduces potential matrix effects (

Development of Csp-2 liquid chromatography/tandem mass spectrometry (LC/MS/MS)–based assay ( ), Ac-VDVAD-CO2H (
), Ac-VDVAD-CO2H ( ), and H-GTQAS-CONH2 (
), and H-GTQAS-CONH2 ( ). (
). ( ) and Csp-2 optimized substrate Ac-VDVADGTQAS-CONH2 (
) and Csp-2 optimized substrate Ac-VDVADGTQAS-CONH2 ( ). Insert: Overnight cleavage kinetics of HTT-derived peptide Ac-MDLNDGTQAS-CONH2 in the presence (
). Insert: Overnight cleavage kinetics of HTT-derived peptide Ac-MDLNDGTQAS-CONH2 in the presence ( ) and absence of Csp-2 (
) and absence of Csp-2 ( ). Solid lines represent a fit to first-order exponential equation. (
). Solid lines represent a fit to first-order exponential equation. ( ) and HTT-derived peptide Ac-MDLNDGTQAS-CONH2 (
) and HTT-derived peptide Ac-MDLNDGTQAS-CONH2 ( ). Solid lines represent fits to the Michaelis-Menten equation. Inset: Apparent activity for HTT-derived peptide Ac-MDLNDGTQAS-CONH2.
). Solid lines represent fits to the Michaelis-Menten equation. Inset: Apparent activity for HTT-derived peptide Ac-MDLNDGTQAS-CONH2.
The two selected substrates described above were analyzed for cleavage by Csp-2 (
Fig. 3C
). Whereas the sequence-optimized Csp-2 substrate Ac-VDVADGTQAS-NH2 was cleaved nearly completely (~75%) overnight, only a very small fraction (2.5%) of the HTT-derived substrate was proteolysed during the same period (
Fig. 3C
, inset). The linear range of product formation was confirmed with 10% turnover within an incubation time of 20 min at a 20-nM enzyme concentration. Subsequently, KM determinations were performed by quantifying cleavage product formation within the first 60 min. With the optimized substrate Ac-VDVADGTQAS-NH2, a hyperbolic dependence of concentration against catalytic Csp-2 activity was obtained (
Fig. 3D
), and the apparent KM value was 5-fold higher than for labeled substrates in the fluorogenic assay. (
In this Csp-2 label-free assay, the activity of the reference isatin
To reinvestigate the Csp-3 inhibitors
Various conditions of caspase proteolytic assays in a fluorescence format were optimized and a novel LC/MS/MS-based method to measure the inhibition of Csp-2 and Csp-3 activity was developed. This label-free LC/MS/MS-based assay, used in combination with carefully selected reducing agents and detergents, has provided tools to identify and correct discrepancies in potencies reported for various caspase inhibitors. Moreover, this LC/MS/MS-based method can be used as a secondary screen to confirm hits from an HTS campaign thanks to the 384-well format, which allows for up to 2000 data points per week and thus represents a well-suited alternative to the more artifact-prone fluorescence-based assay. Although currently not of sufficient throughput to serve as a primary screening assay, this protocol could be used as the basis for further developments in the direction of high-throughput mass spectrometry–based screening applicable to large compound collections.
The weak or lack of caspase activity of the compounds presented here illustrates the difficulties seen in conventional caspase screening to identify viable nonpeptidic leads and shows that, due to the challenging nature of caspases as drug targets, assay conditions measuring the inhibitory potency of compounds in vitro need to be carefully selected. Thorough data interrogation is crucial to ensure that accurate structure-activity relationships are developed. Based on this experience, it is recommended that future caspase screening campaigns compare results obtained under different assay conditions and include at least one orthogonal assay. These steps will enable researchers to distinguish genuine enzyme inhibitors from compounds that produce artifactual inhibition due to compound aggregation, redox cycling, or reliance on nonphysiological substrates.
Footnotes
Acknowledgements
We gratefully acknowledge the supply of recombinant human Csp-2, Csp-3, and Csp-6 by Drs. James Wells and Michelle Arkin from the Small Molecule Discovery Center at the University of California, San Francisco. We also thank Drs. Fred Brookfield, Florence Eustache, and Mark Kerry from Evotec for the synthesis of compounds
Declaration of Conflicting Interests
Evotec conducted the research described through a fee-for-service agreement for CHDI Foundation. As employees of CHDI Management and advisors to CHDI Foundation, MCM, CD, HP, and IM-S were all intimately involved in the study design, data collection and analysis, decision to publish, and preparation of the manuscript.
Funding
This work was funded and conducted through fee-for-service contract research on behalf of CHDI Foundation, a privately-funded nonprofit biomedical research organization exclusively dedicated to developing therapeutics that slow the progression of Huntington disease.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.