Abstract
Infection with human rhinovirus (HRV) is thought to result in acute respiratory exacerbations of chronic obstructive pulmonary disorder (COPD). Consequently, prevention of HRV infection may provide therapeutic benefit to these patients. As all major group HRV serotypes infect cells via an interaction between viral coat proteins and intercellular adhesion molecule–1 (ICAM-1), it is likely that inhibitors of this interaction would prevent or reduce infections. Our objective was to use phage display technology in conjunction with naive human antibody libraries to identify anti–ICAM-1 antibodies capable of functional blockade of HRV infection. Key to success was the development of a robust, functionally relevant high-throughput screen (HTS) compatible with the specific challenges of antibody screening. In this article, we describe the development of a novel homogeneous time-resolved fluorescence (HTRF) assay based on the inhibition of soluble ICAM-1 binding to live HRV16. We describe the implementation of the method in an antibody screening campaign and demonstrate the biological relevance of the assay by confirming the activity of resultant antibodies in a cell-based in vitro HRV infection assay.
Keywords
Introduction
Infection with human rhinovirus (HRV) is the cause of the common cold. To most people, this is an inconvenience from which they quickly recover. However, recent studies highlight how respiratory viruses, including HRV, may act as important triggers for respiratory complications in diseases such as chronic obstructive pulmonary disorder (COPD). 1 One study demonstrated that experimental HRV16 infection in patients with mild COPD could induce lower airway symptoms and biomarker changes similar to those observed in an acute disease exacerbation. 2 Such exacerbations are a recurrent feature in COPD and frequently result in hospitalization and increased risk of death. A different study found 64% of COPD exacerbations to be associated with a preceding cold, and 58% of virus-induced exacerbations were found to be associated with HRV. 3
Most viruses infect their host through an interaction of one or more viral capsid proteins with “receptors” on the surface of the host cell. The cell surface receptor for the major group HRV serotypes is the receptor intercellular adhesion molecule–1 (ICAM-1).4–6 To date, 88 of the 109 known HRV serotypes use this molecule for infection. 7 ICAM-1, a member of the Ig superfamily, functions as a cellular adhesion molecule and has a primary role in mediating leukocyte trafficking. 8 It consists of five extracellular immunoglobulin fold domains (domains 1–5), a transmembrane helix, and a C-terminal tail. 9 The rhinovirus binding site is located on the N-terminal, extracellular domain 1 of ICAM-1,9–11 and structural studies with HRV14 and HRV16 indicate that ICAM-1 domain 1 binds within circular canyons on the virus surface formed by the viral capsid protein VP1.12–15 Each canyon contains five ICAM-1 binding sites. 15 Studies investigating the affinity of HRV3 binding to the soluble ICAM-1 monomer have identified a low-affinity biphasic interaction, 16 with estimates for KD1 and KD2 ranging from 0.55 to 0.70 µM and 5.7 to 12.5 µM, respectively.17,18 However, the presence of 60 ICAM-1 binding sites on the viral capsid facilitates a highly avid interaction with membrane-bound ICAM-1. KD values of 43.5 pM and 14.7 pM have been reported for the interaction of HRV14 and HRV15, respectively, with ICAM-1 expressed on HeLa cells. 14 Increased affinity 17 and in vitro functional potency 19 also have been observed in studies with HRV3 and dimeric forms of soluble ICAM-1. The normal role of ICAM-1 in regulating leukocyte trafficking across the endothelium is mediated via an interaction with leukocyte function antigen–1 (LFA-1). 8 It is noteworthy that the binding site for LFA-1 is reported to be proximal to the binding site for HRV and also located within the N-terminal extracellular domain 1 of ICAM-1. Importantly, however, investigations to define the precise binding sites for LFA-1 and HRV have revealed that, although overlapping, the binding sites are nonetheless distinct. 9
Our aim was to isolate a high-affinity fully human monoclonal antibody to ICAM-1 capable of inhibiting the binding of HRV to the adhesion molecule and displaying potent functional blockade of infection by the major serotype group. Such an antibody could be potentially beneficial in the prophylactic treatment of patients with COPD to reduce the number and intensity of exacerbations. To avoid the potential for adverse effects on the normal function of ICAM-1 in leukocyte trafficking, we were also interested in the potential to target ICAM-1 in such a way as to selectively block HRV binding without an effect on LFA-1. Whether this selective profile is possible is currently the subject of ongoing investigation. However, in light of the high potential specificity achievable via an antibody approach and the published data that confirm that the binding sites for HRV and LFA-1 are distinct, we believe such a profile of selectivity is a realistic possibility.
To isolate such antibodies, we used phage display selection techniques with a naive human antibody library to enrich for antibodies binding to ICAM-1. A pivotal factor in our lead isolation strategy was the need for a robust high-throughput screen capable of identifying antibodies with the desired functional profile. However, high-throughput screening (HTS) of phage display–derived antibodies presents a number of key challenges. High-throughput antibody sample material usually takes the form of a crude bacterial periplasmic extract, and such material typically contains relatively high levels of bacterial-derived by-products, including endotoxin. Assay tolerance of this material is therefore important. Furthermore, periplasmic extracts contain relatively low concentrations of test antibody (typically in the region of tens to hundreds of nM), and the affinity of antibodies isolated directly from naive libraries is variable. It is also important to consider the monovalent single-chain fragment variable (scFv) antibody format for HTS, which is a relevant factor when attempting to inhibit avid interactions. Taken together, these factors mean that the sensitivity of HTS assays to detect functionally active scFv is highly critical. The HTS must also be compatible with the anticipated scale of the screening campaign, which typically extends to tens of thousands of scFv.
One possible screening format was a high-throughput cell-based virus infectivity assay, of which several examples have been described previously.20–23 However, virus infectivity assays were considered unlikely to provide the necessary sample tolerance and sensitivity for successful HTS of crude periplasmic scFv antibodies. This influenced our decision to focus on a biochemical screening approach, and we decided to develop the assay using the HRV16 serotype.
Materials and Methods
Source of Soluble ICAM-1 Reagents
Soluble human ICAM-1–Fc chimera was obtained from R&D Systems (Abingdon, UK). Soluble human ICAM-1–FLAG and cynomolgus monkey ICAM-1–Fc chimera reagents were generated in house via expression in transfected HEK-EBNA cells followed by affinity purification and size exclusion chromatography.
Selections
Phage display selections were performed according to the generic methods described by Dobson et al. 24 using naive human antibody libraries. Multiple rounds of phage display selection were performed using either soluble human ICAM-1 only or both soluble human and cynomolgus monkey ICAM-1 reagents at alternate rounds.
Preparation of Bacterial Periplasmic Extracts
The 96-well plates containing bacterial glycerol stocks were replicated into 2-mL 96-well plates containing 500 µL 2TY broth, 100 µg/mL ampicillin (Becton Dickinson, Oxford, UK), and 0.1% glucose (VWR BDH, Lutterworth, UK). Cultures were incubated at 37 °C with shaking at 250 rpm for approximately 5 h and then induced by addition of 100 µL of 0.2 mM IPTG (VWR BDH) in 2TY broth. Incubation was continued overnight at 30 °C with shaking at 250 rpm. Bacteria were pelleted by centrifugation (2500 rpm, 10 min, 4 °C). Supernatant was then removed and bacterial pellets were resuspended in 300 µL of 0.2M HEPES (pH 7.4) (Sigma-Aldrich, Poole, UK) and 0.5M sucrose (VWR BDH). Microplates were placed on ice for 30 min. Resultant periplasmic extract samples were then clarified by centrifugation (4000 rpm, 10 min, 4 °C).
Preparation of Purified scFv and IgG
ScFv (IMAC-only purification) and IgG2 antibodies were expressed and purified as previously described. 25 Molecular reformatting of scFv fragments into human IgG2 format was performed according to the methods described by Persic et al. 26
Europium Cryptate Labeling of Soluble Human ICAM-1–Fc
Recombinant soluble human ICAM-1–Fc chimera was labeled using europium trisbipyridine cryptate N-hydroxysuccinimide (Eu3+-TBP-NHS) (CisBio International, Bagnols, France) to generate Eu–ICAM-1–Fc. A mean incorporation ratio of 2.4 Eu3+ per molecule of ICAM-1–Fc chimera was confirmed by mass spectroscopy.
Preparation of HRV16 Virus as Crude Culture Supernatant
HeLa Ohio cells were seeded into T175 tissue culture flasks the day prior to infection in order to be approximately 80% confluent when used. Then, 0.5 mL HRV16 virus stock was added per T175 flask, and cells were incubated at 33 °C/5% CO2 for 2 to 3 days until 70% to 80% cell death was achieved. Virus was released from the cells by freeze/thawing the flasks. Cell debris was removed by centrifugation, and the supernatant was aliquoted and stored at −80 °C. Each batch of HRV16 culture supernatant was titered in the HRV16 infectivity assay described prior to use.
Preparation of Virus Concentrate
Crude HRV16 culture supernatants were concentrated for use in the homogeneous time-resolved fluorescence (HTRF) HRV16–ICAM-1 binding inhibition assay via centrifugation through Amicon Ultra 30-kDa molecular weight cutoff filter units (Millipore, Watford, UK). Centrifugation (3000 g, room temperature [RT]) was continued until the starting supernatant volume (15 mL per filter unit) had been reduced 10-fold. The extent of virus concentration was confirmed by titer measurement in the HRV16 infectivity assay described and aliquots stored at −80 °C.
HTRF HRV16–ICAM-1 Binding Inhibition Assay
All reagents were diluted in assay buffer comprising 1× Dulbecco’s phosphate-buffered saline (Invitrogen, Paisley, UK), potassium fluoride (0.4M) (BDH, Poole, UK), and bovine serum albumin (0.2%) (Sigma-Aldrich). The assay was performed in black clear-bottom 384-well plates (Corning, Sunderland, UK) by sequential addition of 10 µL anti–FLAG-XL665 (32 nM), 5 µL Eu–ICAM-1–Fc (16 nM), 5 µL ICAM-1–FLAG (48 nM), 10 µL test antibody sample, and, following a preincubation step (30 min, RT), 10 µL prediluted HRV16 concentrate. Final assay HRV16 concentration required optimization on a batch-specific basis and ranged from 5% to 12.5%. Anti–FLAG-XL665 was obtained from Cisbio International. For single-point HTS of crude bacterial periplasmic lysates, final assay sample concentration was 15%. For testing of purified scFv and IgG, duplicate 11-point serial titrations were prepared. The commercially available anti–ICAM-1 antibody, mAb 15.2 (Abcam, Cambridge, UK), was used as a reference inhibitor in IC50 experiments. mAb 15.2 is a neutralizing antibody that targets an epitope within ICAM-1 extracellular domain 1. Assay plates were top sealed and incubated (4 h/RT) before reading on an EnVision plate reader (PerkinElmer, Seer Green, UK) with an excitation wavelength of 320 nm and emission wavelengths of 615 nm and 665 nm. The plate reader was configured in bottom read format, enabling assay plates to remain top sealed during plate reading. Raw 665-nm and 615-nm counts were converted into 665-nm/615-nm ratio values, which were then used to calculate Delta F (%). Delta F (%) is given by the following equation: {[(sample 665-nm/615-nm ratio) − (negative 665-nm/615-nm ratio)]/[negative 665-nm/615-nm ratio]} × 100. Negative 665-nm/615-nm ratio was derived from control wells in which the ICAM-1–FLAG reagent was omitted. Additional data analysis was performed using either Microsoft Excel (single-point HTS; Microsoft Corp., Redmond, WA) or GraphPad Prism software (pIC50 determination; GraphPad, La Jolla, CA), in the latter case using variable-slope four-parameter logistic curve fitting. pIC50 values from multiple experiments are quoted as mean ± standard error of the mean (SEM).
In Vitro Cell-Based HRV16 Infectivity Assay
HeLa Ohio cells (3 × 104/well) were plated into 96-well plates using a culture medium containing minimum essential medium (MEM), 1× nonessential amino acids, penicillin (100 U/mL), streptomycin (100 µg/mL) (each from Invitrogen), and heat-inactivated bovine calf serum (10%) (SAFC Biosciences, Andover, UK). After an overnight incubation (37 °C/5% CO2), medium was removed and cells were preincubated with 30 µL of test sample (or medium for untreated controls) for 30 min in triplicate. The commercially available anti–ICAM-1 antibody, mAb 14C11 (R&D Systems, Abingdon, UK), was used as a reference inhibitor. mAb 14C11 targets an epitope within ICAM-1 extracellular domain 1. Then, 30 µL HRV16 culture supernatant (final multiplicity of infection = 20) was added (or medium for noninfected controls), and virus attachment was allowed to proceed for 2 h at 33 °C/5% CO2. Assay supernatant was then removed and 100 µL of fresh medium added to each well followed by incubation for a further 23 h at 33 °C/5% CO2. Remaining viable cells were fixed in 4% formaldehyde (Sigma-Aldrich), stained (5 min/RT) with 0.5% w/v crystal violet (Sigma-Aldrich), washed three times in water to remove excess stain, and allowed to dry. Plates were read by measuring absorbance at 600 nm, and percentage cytopathic effect (% CPE) was calculated according to the following equation: % CPE = 100 × OD 600 nm [(uninfected control − test sample)/(uninfected control − virus only control)]. Inhibitor pIC50 values were determined using GraphPad Prism software with variable-slope four-parameter logistic curve fitting. pIC50 values from multiple experiments are quoted as mean ± SEM.
Results and Discussion
Assay Design Considerations
The design of the biochemical assay we developed was influenced by several factors in addition to those described in the Introduction. To ensure the biological relevance of the assay, we decided to use live HRV16, and this necessitated workup of robust processes for virus cultivation, harvest, and concentration. A particular challenge lay in the limited options that existed to detect HRV16 given the lack of available anti-HRV antibodies and potential risk of compromising the biological relevance of the assay had we attempted to label the virus directly. Safety considerations around the use of live HRV16 also influenced our preference to avoid processes carrying the risk of aerosol formation. A further technical consideration lay in the very low monovalent binding affinity of the HRV–ICAM-1 interaction.
Assay Principle
To overcome these challenges, we developed a novel HTRF assay in which we exploited the presence of multiple ICAM-1 binding sites on the virus surface. HTRF (CisBio International) is one of a family of commercially available assay technologies based on the principle of time-resolved fluorescence resonance energy transfer (TR-FRET). By differentially labeling two populations of soluble ICAM-1 with TR-FRET donor and acceptor fluorophores and then measuring their simultaneous interaction with HRV16, this enabled the use of live unmodified virus, and the need to detect the virus directly was avoided. Also, exploiting the potential for avid binding of soluble ICAM-1 to the virus in our assay design facilitated the detection of the low µM affinity HRV–ICAM-1 interaction at the low nM reagent concentrations typically required in HTRF assays. To achieve this, we generated two forms of soluble ICAM-1: an ICAM-1–Fc chimera labeled with europium cryptate (Eu–ICAM-1–Fc), believed to exist as a homodimer, and a FLAG-tagged ICAM-1 monomer (ICAM-1–FLAG) that could be detected indirectly with an XL665-labeled anti-FLAG mAb (anti–FLAG-XL665). In detecting the ICAM-1–FLAG monomer indirectly with anti–FLAG-XL665, our aim was to facilitate bivalent binding of both of the soluble ICAM-1 reagents with the virus. When a binding complex is formed between Eu–ICAM-1–Fc, ICAM-1–FLAG (with anti–FLAG-XL665), and HRV16, the europium cryptate donor and XL665 acceptor fluorophores are brought into sufficiently close proximity for TR-FRET to be measured ( Fig. 1 ).
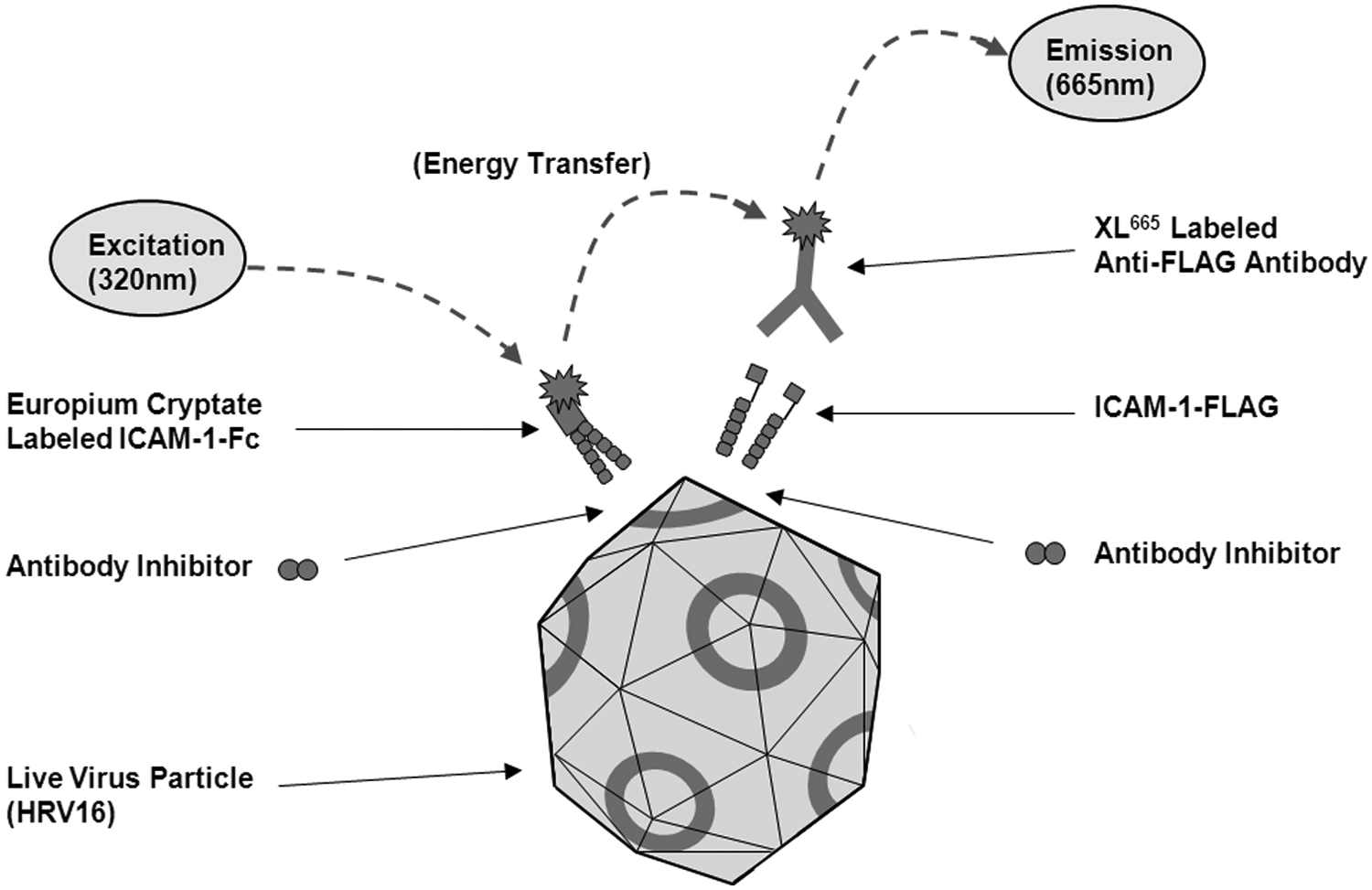
A schematic of the homogeneous time-resolved fluorescence (HTRF)–based assay format designed to identify inhibitors of the interaction of soluble intercellular adhesion molecule–1 (ICAM-1) with live human rhinovirus 16 (HRV16). The virus illustration is a simplified representation of the icosahedral structure previously reported for human rhinovirus 14 (HRV14). 12
Initial Optimization of Soluble ICAM-1 Reagent Concentrations
Initially, a range of different concentrations of Eu–ICAM-1–Fc and ICAM-1–FLAG were incubated (150 min/RT) with a constant concentration of anti–FLAG-XL665 (8 nM) in the presence of HRV16 crude culture supernatant (20%) ( Fig. 2 ). A clear soluble ICAM-1 concentration-dependent TR-FRET assay signal was detected. No significant assay signal was observed in the absence of HRV16 supernatant (data not shown). The optimum concentrations of Eu–ICAM-1–Fc and ICAM-1–FLAG were 2 nM and 6 nM, respectively, and were fixed for future experiments. However, it was of note that the highest Delta F value measured in this experiment was approximately 80.0%, well below what would be considered sufficient for a robust assay.
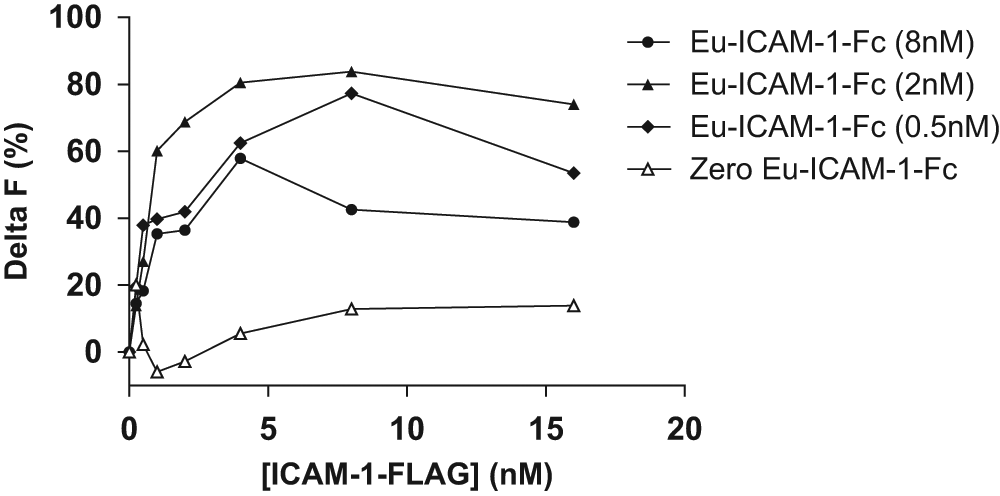
A range of different concentrations of Eu–ICAM-1–Fc and ICAM-1–FLAG were incubated (150 min/room temperature) with a constant concentration of anti–FLAG-XL665 (8 nM) in the presence of crude human rhinovirus 16 (HRV16) culture supernatant (20%). Results are plotted as Delta F (y-axis) versus concentration of ICAM-1–FLAG (x-axis) at each of the indicated concentrations of Eu–ICAM-1–Fc. ICAM-1, intercellular adhesion molecule–1.
Optimization of HRV16 Concentration and Incubation Time
Eu–ICAM-1–Fc and ICAM-1–FLAG were fixed at the concentrations stated above, and a range of crude HRV16 culture supernatant concentrations were investigated over an extended time course (250 min/RT) ( Fig. 3A ). Anti–FLAG-XL665 was used at a fixed concentration of 8 nM based on parallel optimization experiments. Delta F continued to increase over time when the binding incubation was extended, and based on these results, the incubation was therefore standardized to 4 h at RT. In addition, increasing the concentration of HRV16 supernatant resulted in a significant increase in measured Delta F. However, although 50% HRV16 supernatant resulted in a maximum Delta F of just over 200% (which we generally consider to be the safe minimum for HTS), we questioned whether this would be routinely achievable using crude HRV16 supernatants given the likelihood of batch-to-batch variation in titer. To address this, a method for concentration of HRV16 from crude supernatant material was used as described in Materials and Methods. Optimization of this virus concentration procedure enabled titers to be increased approximately 10-fold relative to the source culture supernatant.
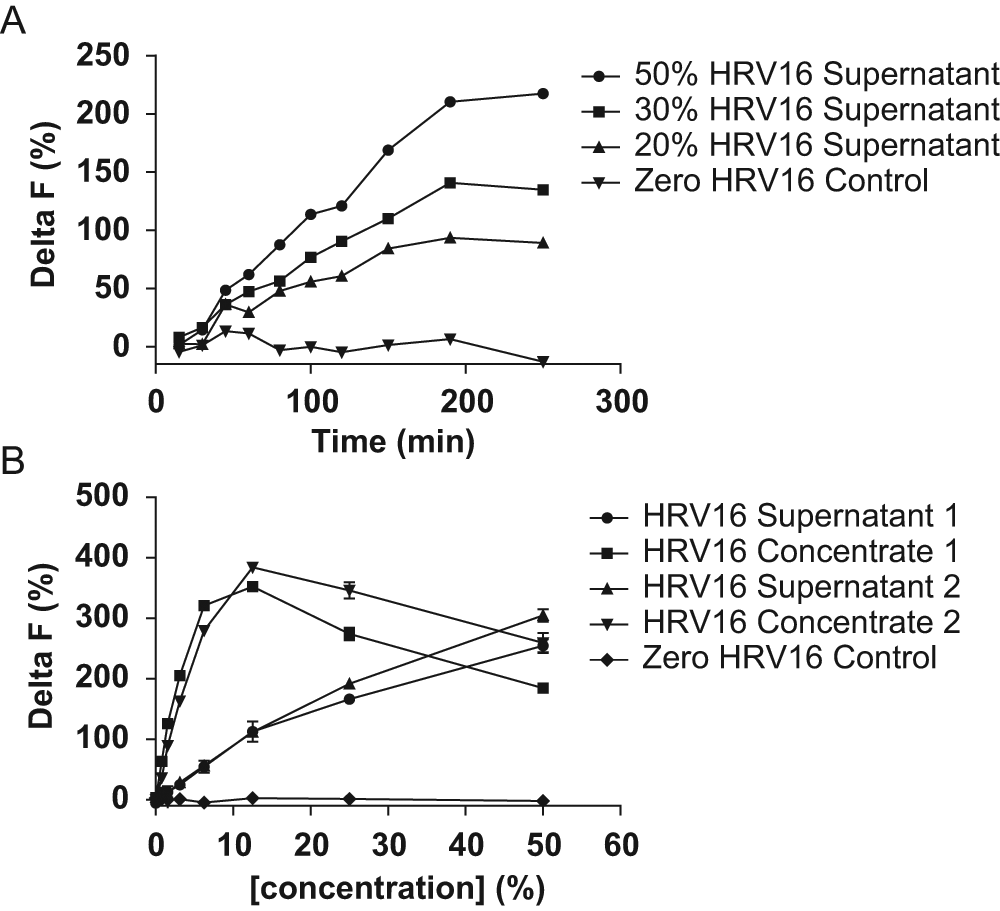
Fixed concentrations of Eu–ICAM-1–Fc, ICAM-1–FLAG, and anti–FLAG-XL665 (2 nM, 6 nM, and 8 nM, respectively) were incubated (room temperature [RT]) with a range of different concentrations of crude human rhinovirus 16 (HRV16) culture supernatant (0%, 20%, 30%, and 50%) over an extended time course (250 min). Delta F (y-axis) is plotted against time (x-axis) for each concentration of crude HRV16 culture supernatant (
Figure 3B shows a titration experiment in which two batches of crude HRV16 culture supernatant were titrated in the assay along side the corresponding HRV16 concentrates. All other assay parameters in this experiment were constant and as defined previously. In agreement with earlier results, both batches of crude HRV16 supernatant showed a concentration-dependent increase in assay signal, reaching a Delta F of approximately 250% at 50% assay concentration. In contrast, for both of the HRV16 concentrates, a Delta F of over 250% was achieved at only 6.25% assay concentration. This result agreed well with the estimated 10-fold virus concentration achieved through molecular weight cutoff centrifugation, which had been confirmed by virus titer measurement.
Routinely, each new batch of HRV16 concentrate was titrated into the assay, and the optimal final assay concentration was defined as that required to give a Delta F of 250% (typically this ranged from 5%–12.5%). The reduced assay signal observed at higher concentrations of HRV16 concentrate ( Fig. 3B ) was not unexpected and can be explained by the HRV16 concentration beginning to exceed the available concentration of the other assay components. In turn, this will progressively reduce the number of HRV16 particles interacting simultaneously with both of the two ICAM-1 binding partners in the system. This phenomenon is frequently observed in homogeneous binding assays and is often referred to as the “hook effect.”27,28
Validation of the Assay
To validate the final assay conditions, a mouse monoclonal anti–ICAM-1 antibody, mAb 15.2, was tested and was shown to give full inhibition (mean pIC50 8.37 ± 0.11nM, n = 3) (
Evaluation of Assay Tolerance to HTS Sample Material
As mentioned previously, HTS of phage display–derived scFv typically necessitates a crude unpurified bacterial lysate sample format. In this case, the HTS sample format was a crude bacterial periplasmic extract, and assay tolerance to this material was therefore important. Adopting the assay conditions defined earlier, the introduction of irrelevant crude periplasmic extract resulted in a moderate dose-dependent reduction in total binding assay signal, but a Delta F value of just over 200% was maintained in the presence of 15% irrelevant sample material (
Implementation of the HRV16–ICAM-1 Binding Inhibition Assay for Single-Point HTS of Crude Periplasmic Extract scFv Samples
Using naive human antibody libraries, phage display antibody selections were performed on soluble cynomolgus monkey and/or human ICAM-1 antigens. Individual antibodies were then prepared as crude periplasmic extract scFv samples in a high-throughput expression format before screening in the 384-well HRV16–ICAM-1 binding inhibition assay according to the protocol as set out in the Materials and Methods section. Figure 4A shows representative single-point HTS data corresponding to 88 sequence-unique confirmed ICAM-1 binding scFv in which the 88 samples were tested in two separate experiments to assess assay reproducibility. A clear positive correlation between the two HTS data sets was observed (R2 = 0.78) (Pearson).
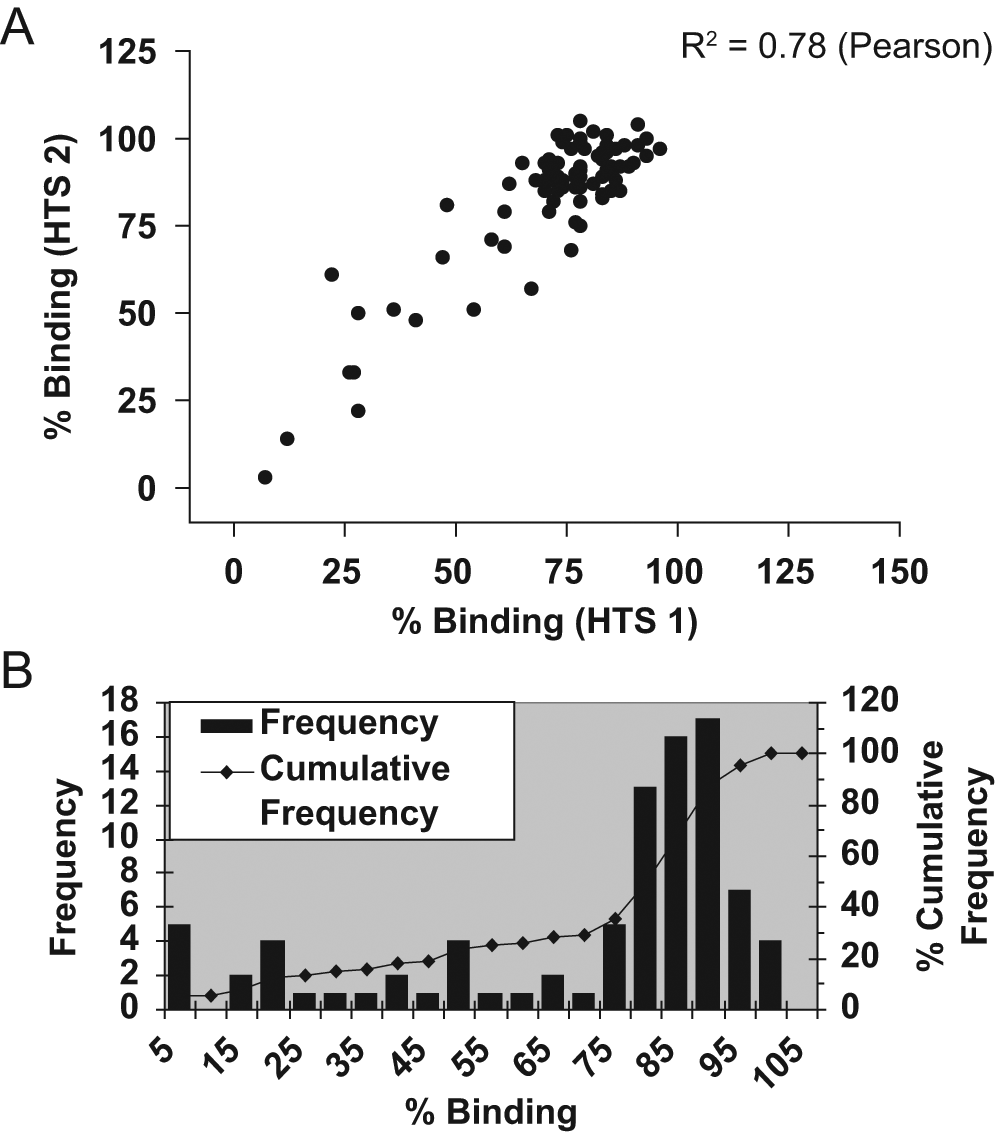
Eighty-eight sequence-unique anti–ICAM-1 scFv derived from phage display selections on soluble cynomolgus and/or human ICAM-1 antigens were tested in the HRV16–ICAM-1 binding inhibition assay (single-point high-throughput screening [HTS] format) as crude periplasmic extracts. Data are presented as a correlation of the resultant % binding values obtained from two separate experiments, HTS 1 (x-axis) and HTS 2 (y-axis) (
Example HTS data from the screening of a phage display selection output directly in the HRV16–ICAM-1 binding inhibition assay are shown in Figure 4B . Here results are shown for 88 crude periplasmic extract scFv samples derived from phage display selections on human ICAM-1. Data are plotted in histogram format, where resultant HTS % binding values are grouped into bins (x-axis) with frequency and cumulative frequency plotted on the y-axis. Although a distribution of noninhibitory scFv is observed, a population of inhibitory scFv hits is also apparent. The Z′ value for the data in Figure 4B was 0.85.
The HRV16–ICAM-1 binding inhibition assay was successfully implemented in single-point HTS format as part of an internal antibody discovery program, and 3344 samples were screened. We identified 52 sequence-unique hits from the primary HTS, which were subsequently generated as purified scFv. The small size of the HTS campaign reflects the efficiency of the phage display selection process, but the throughput capacity of the HRV16–ICAM-1 binding inhibition assay itself could easily have extended to tens of 384-well assay plates per day had this been necessary.
A cutoff of 50% inhibition was used as the criterion for defining HTS hits. This cutoff value was assigned subjectively from HTS results corresponding to irrelevant scFv samples, which would typically be plotted in histogram format (as Fig. 4B ). In setting this cutoff value, our aim was to capture as much diversity as possible within the HTS hits while eliminating the irrelevant noninhibitory scFv population.
Confirmatory Testing of Purified Antibodies in the HRV16–ICAM-1 Binding Inhibition Assay
Inhibitory activity was confirmed in the HRV16–ICAM-1 binding inhibition assay for 43 of the 52 purified unique scFv that had been identified as hits in the primary HTS, reflecting a confirmation rate of 82.7%. Dose-response inhibition plots for two example purified scFv (antibody 1 and antibody 2) are shown in Figure 5A . Clearly, both of these purified scFv show dose-dependent inhibition of HRV16–ICAM-1 binding and produce complete inhibition at high concentration. Based on results from several experiments, mean pIC50 values of 7.91 ± 0.06 (n = 5) and 7.80 ± 0.04 (n = 4) were determined for antibody 1 and antibody 2, respectively, as purified scFv. Antibody 1 and antibody 2 were reformatted to human IgG2, 26 and the inhibitory activity of these leads was reconfirmed in the HRV16–ICAM-1 binding inhibition assay in this format. Example dose-response inhibition plots for antibody 1 and antibody 2 IgG2 are shown in Figure 5B alongside mAb 15.2, which was included as a reference inhibitor. Based on results from several experiments, mean pIC50 values of 8.79 ± 0.03 (n = 7) and 8.82 ± 0.02 (n = 5) were obtained for antibody 1 and antibody 2, respectively, as purified human IgG2. The approximate 10-fold reduced IC50 values (increased potency) for both antibodies in the HRV16–ICAM-1 binding inhibition assay on conversion from scFv to IgG2 format were not unexpected. This likely reflects the potential for avid/bivalent binding of the IgG2 to one or potentially both of the two soluble ICAM-1 components in the system. A mean pIC50 value of 8.37 ± 0.11 (n = 3) was obtained for mAb 15.2. Antibody 1 and antibody 2 were expressed as the human IgG2 isotype to avoid introducing unwanted effector function.
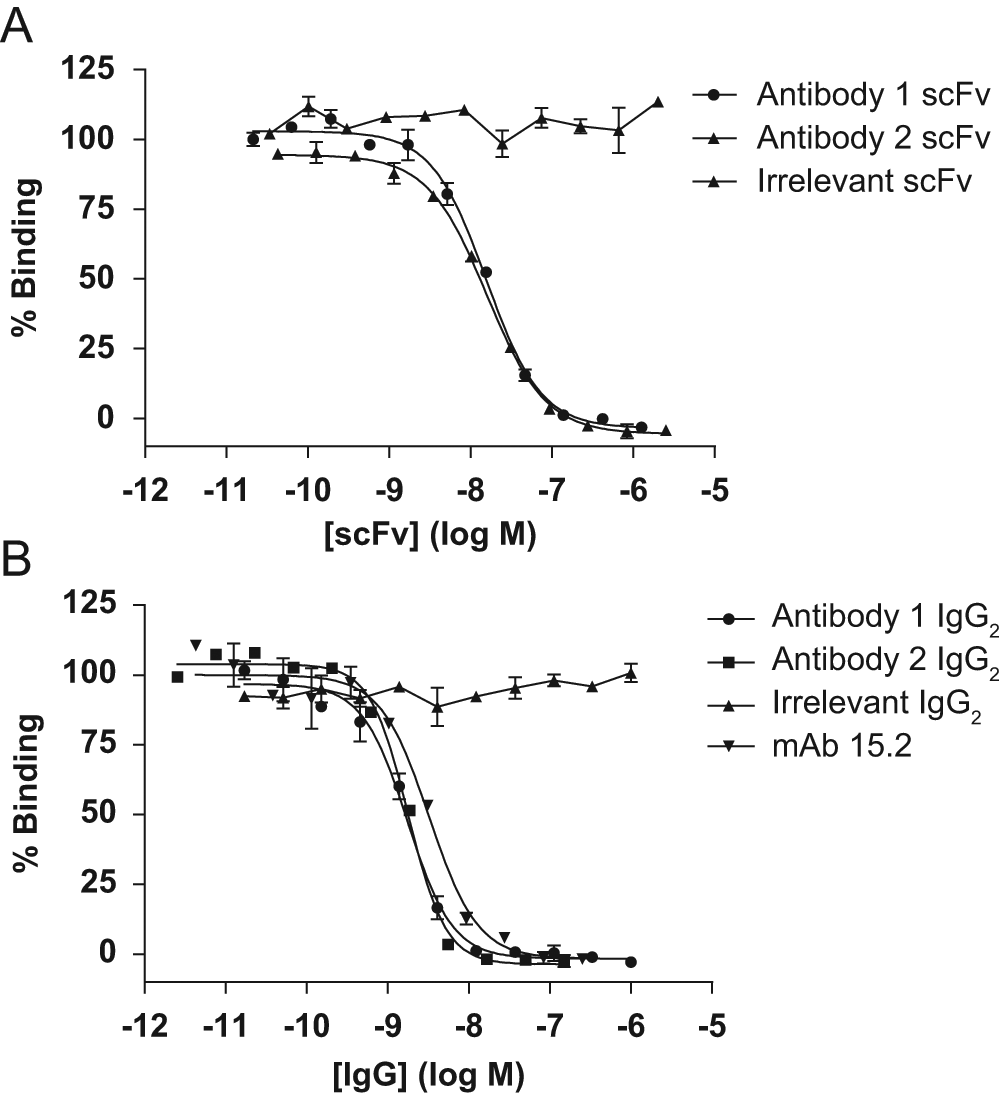
Inhibition dose-response curves are shown for antibody 1 and antibody 2 as both purified scFv (
Evaluation of Functional Potency for Isolated Antibodies in a Cell-Based In Vitro HRV16 Infectivity Assay
To further evaluate the biological relevance of the HRV16–ICAM-1 binding inhibition assay, we were keen to evaluate the functional potency of antibodies identified in the primary screen in a cell-based in vitro HRV16 infectivity assay. Figure 6A shows dose-response inhibition plots for antibody 1 and antibody 2 when tested in an in vitro HRV16 infectivity assay as purified scFv according to the protocol described in the Materials and Methods section. Plotted data represent the mean results from several independent experiments (n numbers indicated). In vitro functional activity was clearly demonstrated for antibody 1 and antibody 2 as purified scFv, although it was notable that the potency of these scFv was insufficient to define a full dose-response inhibition curve in this assay system. No inhibition of infection was observed with an irrelevant negative control–purified scFv. Figure 6B shows corresponding dose-response inhibition curves from the testing of antibody 1 and antibody 2 in the same in vitro HRV16 infectivity assay as purified human IgG2. Again, plotted data represent the mean results from several experiments (n numbers indicated). Both antibody 1 and antibody 2 showed potent inhibition of HRV16 infection as purified human IgG2 with mean pIC50 values of 8.80 ± 0.11 (n = 7) and 8.20 ± 0.40 (n = 5), respectively. No inhibition was observed in the assay with an irrelevant human IgG2 isotype control. The commercially available anti–ICAM-1 antibody, mAb 14C11, was included in these experiments for reference and gave a mean pIC50 of 9.31 ± 0.03 (n = 10).
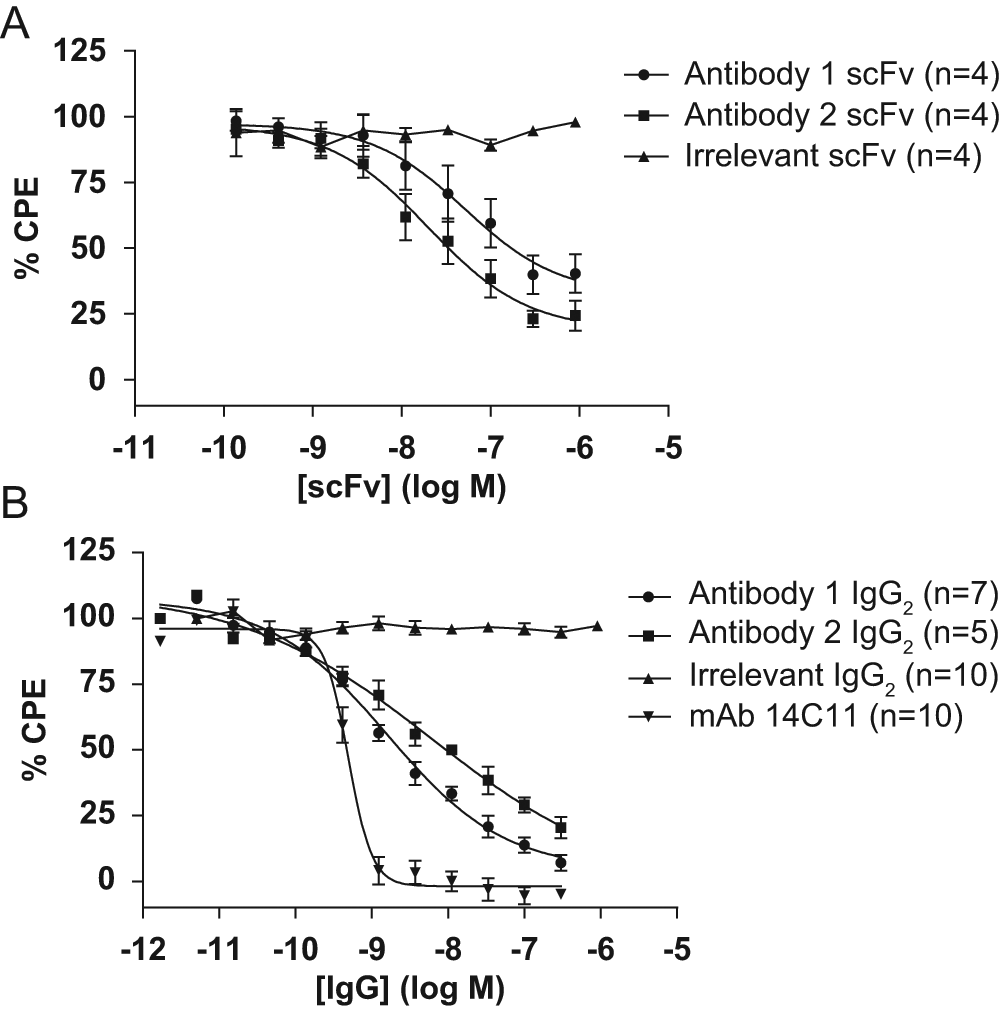
Inhibition dose-response curves are shown for antibody 1 and antibody 2 as both purified scFv (
Akin to the HRV16–ICAM-1 binding inhibition assay results, antibody 1 showed increased potency in the in vitro HRV16 infectivity assay when tested as human IgG2 relative to the corresponding data as purified scFv. This observation was not unexpected and is likely explained by the potential for avid/bivalent binding of IgG2 to membrane-bound ICAM-1. This format-dependent potency gain was also observed with antibody 2, albeit to a lesser degree. It was also noted that antibody 1 and antibody 2 showed greater potency in the biochemical HRV16–ICAM-1 binding inhibition assay relative to the in vitro HRV16 infectivity assay when tested as purified scFv. This likely reflects the greater potential for higher order avid HRV16–ICAM-1 binding in the cell-based assay relative to the biochemical assay (in which only a bivalent interaction of HRV16–ICAM-1 would be predicted) and is in line with previous reports.14,17 These observations supported our initial rationale to focus on a biochemical primary screening approach to ensure sensitive detection of inhibitory scFv under HTS conditions. In addition to this, our experience is that crude bacterial lysate sample materials are generally not well tolerated in cell-based functional assays, particularly over longer incubation times.
From 43 sequence-unique antibodies that gave full inhibition in the HRV16–ICAM-1 binding inhibition assay as purified scFv, 17 of these were subsequently tested in the same assay as IgG2, and 12 showed full inhibition. Five of these 12 IgG2 were subsequently prioritized for testing in the in vitro HRV16 infectivity assay with measurable inhibition confirmed for 3 of the 5. It is likely that a difference in relative sensitivity between the two assay systems was a contributing factor in this observed attrition rate, and this is something we often observe between biochemical and cell-based assays. The HRV16–ICAM-1 binding inhibition assay we have developed has also been applied to the characterization of small-molecule inhibitors of HRV–ICAM-1 binding, 20 and the results were found to correlate well with potency data from in vitro HRV infectivity assays.
In summary, a novel and predictive assay has been developed using HTRF technology that sensitively detects inhibitors of the HRV16–ICAM-1 interaction. The assay design exploits the presence of multiple ICAM-1 binding sites on the virus surface to detect the simultaneous interaction of donor and acceptor fluorophore-labeled soluble ICAM-1 reagents via TR-FRET. The approach avoids the need to detect the virus directly and enables the use of live unmodified virus. Heterogeneous enzyme-linked immunosorbent assay methods measuring the interaction between live HRV and soluble ICAM-1 are not unprecedented,29,30 but we believe this to be the first reported example of a fully homogeneous HTS for inhibitors of HRV–ICAM-1 binding and the first reported demonstration of TR-FRET between donor and acceptor fluorophore-labeled molecules binding simultaneously on the surface of a live virus. The assay format is suitable for both small-molecule and biologics discovery and, in our hands, has proven to be a predictive HTS approach enabling the successful identification of antibodies demonstrating in vitro functional blockade of HRV16 infection. In principle, this assay concept may have a wider application with the potential to enable homogeneous screening of other virus–host cell receptor interactions where multiple host cell receptor binding sites exist on the virus surface.
Footnotes
Acknowledgements
The authors wish to dedicate this publication to the late Dr. Paula Rosamund Harrison, who sadly passed away on September 18, 2012, following a short illness. Paula made a huge contribution to numerous drug screening projects during her 11 years with MedImmune Cambridge. She was a highly respected scientist, was very well liked, and will be greatly missed. Our thoughts are with her family.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
