Abstract
Aminoacyl-tRNA synthetases are enzymes that charge specific tRNAs with their cognate amino acids and play an essential role in the initial steps of protein synthesis. Because these enzymes are attractive targets for drug development in many microorganisms, there is a pressing need for assays suitable for compound screening. We developed (1) a high-throughput assay for measuring aminoacyl-tRNA synthetase activity and (2) an accompanying method for preparing the tRNA substrate. The assay can be performed in 96-well plates and relies on malachite green detection of pyrophosphate (Pi) as an indicator of aminoacyl-tRNA synthetase activity. Analysis of Trypanosoma brucei isoleucyl-tRNA synthetase (IleRS) activity showed that the assay exhibits sensitivity to picomoles of product and yielded a Z′ factor of 0.56. We show that this assay is applicable to other aminoacyl-tRNA synthetases and to enzyme inhibition studies. Using this assay, we found that the compound NSC616354 inhibits recombinant IleRS with an IC50 of 0.6 µM. Enzymology studies were also performed with rIleRS and its Km and kcat determined as 3.97 × 10−5 mol/L and 312 S−1, respectively. This assay will facilitate the screening of compounds to identify inhibitors of aminoacyl-tRNA synthetases.
Introduction
Amino acids are ligated to their cognate tRNAs by aminoacyl-tRNA synthetases, a group of at least 20 enzymes whose members contribute to protein synthesis. 1 Aminoacylation of tRNA is an essential step in translation, and its reaction occurs in two steps: first, the aminoacyl-tRNA synthetase hydrolyzes adenosine triphosphate (ATP) to activate an amino acid with adenosine monophosphate (AMP), thereby forming an aminoacyl-adenylate intermediate and releasing pyrophosphate (E + ATP + AA + tRNA ↔ E:AA-AMP + tRNA + PPi); second, it esterifies the activated amino acid to the terminal ribose of the tRNA and releases AMP (E:AA-AMP + tRNA + PPi ↔ E + AA-tRNA + AMP + PPi). The final product of this reaction is aminoacylated tRNA (AA-tRNA), with AMP and PPi also released as reaction by-products. Aminoacyl-tRNA synthetases are attractive targets for drug development since they have been shown to be essential for growth and survival of many pathogenic microorganisms. In Trypanosoma brucei, the parasite responsible for causing human African trypanosomiasis, aminoacyl-tRNA synthetases have been shown to be essential for parasite growth and survival. 2 Similarly, aminoacyl-tRNA synthetases have been shown to be essential for other pathogens of global importance, including Plasmodium sp., 3 the parasite that causes malaria, and Mycobacterium tuberculosis, 4 which causes tuberculosis.
Despite the importance of aminoacyl-tRNA synthetases as potential drug development targets, few assays have been developed to date. The lack of appropriate assays has been a major factor hindering the identification of aminoacyl-tRNA synthetase inhibitors. Of the few assays that have been developed, most rely on the use of radiolabeled amino acids or tRNAs.5,6 These assays are impractical for high-throughput screening (HTS) and are poor for quantitation. On the other hand, the spectrophotometric assays that have been developed require multiple coupled enzymes and generally exhibit high background and a low signal-to-noise ratio.7,8 Recently, a compound screen using a luciferase chemiluminescence-based assay was reported for methionyl-tRNA synthetase, using Escherichia coli bulk tRNAs, on the PubChem database; however, detailed information on assay quality and validation is not available. 9 Alternatively, a luciferase assay has been reported using bacteria lysates. 10 This method is convenient for HTS, but it is limited to bacterial tRNA synthetases and cannot be used for enzymology studies. To overcome these limitations, we developed a 96-well plate spectrophotometric assay for quantitative measurement of in vitro aminoacyl-tRNA synthetase activity. In this assay, the inorganic pyrophosphatase enzyme was coupled to the aminoacylation reaction to convert the pyrophosphate into inorganic phosphate, which was quantitatively measured via malachite green. We also developed a detailed protocol for preparation of the tRNA substrate, which facilitates adaptation of the assay to any aminoacyl-tRNA synthetase.
Using this method, we showed that recombinant T. brucei isoleucyl-tRNA synthetase (rIleRS) aminoacylates its cognate tRNAIle more efficiently than either total T. brucei RNAs or yeast tRNAs. The assay is sensitive to picomoles of product and has a Z′ factor of 0.56. Using this assay, we identified the compound NSC616354 as an inhibitor of T. brucei IleRS, which shows its application for enzyme inhibition studies. Furthermore, kinetic assays were performed to determine enzymatic parameters (Km and kcat) of rIleRS, demonstrating its use for enzymology analysis. This assay can be easily used in a high-throughput drug development context to identify inhibitors of aminoacyl-tRNA synthetases.
Materials and Methods
Cloning, Expression, and Purification of Recombinant Protein
The DNA sequence of the gene Tb927.10.9190 (which codes for isoleucyl-tRNA synthetase) was amplified by PCR without the mitochondrial targeting sequence (nucleotides 1–198) using specific primers (9820-forward: CCCGAATTCATGACTGGACCACTACAA and 9821- reverse:CCCGCGGCCGCCGATTCACCAGCCGACGG) and cloned into pET28b+ vector (Novagen, Madison, WI) using BamHI and NotI restriction sites with a C-terminal 6xHis-tags. For the gene Tb927.11.7170, which codes for seryl-tRNA synthetase, the PCR product was amplified using specific primers (8307-forward: CCCGGTCTCA-AGCTTCATATGGTGCTTGATATACAGCTGTTTC and 8308-reverse: CCCGGTCTCCTCGAGCTCCCCCTTG-TCGGGT) and cloned into pET29a+ using NdeI and XhoI restriction sites with a C-terminal 6xHis-tag. The constructs were used to transform E. coli Rosetta 2(DE3) pLysS strain (EMD/Merck (Novagen), Madison, WI, USA), and protein expression was induced with 1 mM isopropyl β-D-1-thiogalactopyranoside (IPTG). E. coli were grown in LB media and harvested, and the lysate was prepared with Bugbuster reagent (Novagen). Proteins were purified using nickel-magnetic beads (Millipore, Billerica, MA), dialyzed against 50 mM Tris and 140 mM NaCl, and kept at 4 °C (with 0.05% sodium azide) until use. For the gene Tb927.10.1500 (which codes for methionyl-tRNA synthetase), the DNA sequence were amplified by PCR using specific primers (8478-forward: CCCAAGCTTATGG-CTCTAAAGCTGCTTTCAGA and 8479-reverse: CCCA-GATCTTGTACTCTTTGTATTCTCTGTTGAGCG) and cloned into the pLEW79-MH-TAP vector using HindIII and BamHI restriction sites with a C-terminal tandem affinity purification (TAP)–tag. Plasmids were linearized with NotI digestion and transfected in T. brucei procyclic forms as previously described. 11 Protein expression and purification were performed as previously reported. 11 Briefly, T. brucei procyclic cells were grown in 1 L of SDM-79 medium supplemented with 10% fetal bovine serum (Gibco BRL, Carlsbad, CA) at 27 °C, and protein expression was induced with 100 ng/mL tetracycline (Sigma, St. Louis, MO). For protein purification, 2.0 × 1010 procyclic cells were harvested and the pellet resuspended in lysis buffer (50 mM Tris, 150 mM NaCl, 1% Triton X-100, 0.2% NP40) with EDTA-free protease inhibitor cocktail (Roche, Basel, Switzerland). The cleared lysate was incubated with Protein A Sepharose 4 fast flow (Pharmacia/GE Healthcare Bioscience, Pittsburg, PA, USA) for 2 h rotating at 4 °C. The protein-resin mix was washed with 100 mL wash buffer (50 mM Tris, 600 mM NaCl, 0.2% NP40) followed by wash in 10 mL tobacco etch virus (TEV) protease buffer (50 mM Tris, 150 mM NaCl, 0.2% NP40, 1 mM dithiothreitol [DTT]). The resin was incubated with 100 U AcTEV protease (Invitrogen, Carlsbad, CA) in 1 mL TEV buffer at 4 °C for 2 h. Afterward, the proteins were eluted with 1.5 mL TEV buffer and stored in aliquots at 4 °C until use.
Preparation of tRNA Substrate
The substrates tRNAIle, tRNASer, and tRNAMet were prepared by in vitro transcription using the MEGAScript in vitro transcription kit (Ambion/Life Technologies, Carlsbad, CA, USA). A T7 promoter sequence (TAATACGACTCACTATAGGG) was added at the 5′ of the forward primers used to amplify the tRNAs from genomic DNA. Similarly, the CCA sequence was added to the reverse primers used to amplify the tRNAs (TGG sequence at the 5′ of the reverse primer). The template tRNAIle were amplified by PCR (using Phusion High Fidelity DNA polymerase; Fermentas, Waltham, MA) from genomic DNA using specific primers (9015-forward:
Aminoacylation Assays
Aminoacylation assays were performed in aminoacylation buffer (30 mM HEPES buffer, 140 mM NaCl, 30 mM KCl, 40 mM MgCl2) with 1 mM DTT, 200 µM ATP, 2 U/mL inorganic pyrophosphatase (PPiase; Sigma), 1 mM L-isoleucine (Sigma), 40 µg/mL recombinant IleRS, and 8 µM tRNAIle at 37 °C (unless otherwise stated). Alternatively, 1 mg/mL of yeast tRNAs (Sigma) or 0.4 mg/mL of T. brucei total RNA enriched in microRNAs was used. T. brucei total RNA enriched in microRNAs were prepared using an miRNA kit (Ambion) according to the manufacturer’s instructions. The aminoacylation reactions (50 µL total volume each) were performed in clear, flat-bottom 96-well plates (Costar 96-well standard microplates; Corning, Corning, NY) and incubated for 30 min at 37 °C. Reactions were stopped by addition of 100 µL malachite green (Echelon Biosciences, Salt Lake City, UT) and developed for 30 min at RT. Absorbances were then measured at 620 nm using a Spectramax M2 (Molecular Devices, Sunnyvale, CA).
Assay for calculation of Z′ factor
Reactions were performed with 80 µg/mL recombinant IleRS, 8 µM tRNAIle, 200 µM ATP, 2 U/mL PPiase (Sigma), and 5 mM L-isoleucine (Sigma) at 37 °C for 30 min. The reactions were stopped by addition of 100 µL malachite green. The Z′ factor was calculated as previously reported. 12
Enzymology Assays and Analysis
Reactions were performed in a 50-µL volume with rIleRS enzyme (40 µg/mL), L-isoleucine (0.1, 0.5, 1, 5, 10, and 50 mM), 8 µM tRNAIle, 2 U/mL PPiase, 100 µM ATP, and 1 mM DTT in aminoacylation buffer. Reactions were incubated at 37 °C; at 2.5, 5, and 10 min, 15-µL aliquots were withdrawn and mixed with 10 mM EDTA on ice to stop the reactions. Then, 100 µL malachite green solution was added and the absorbance measured at 620 nm. For the Lloyd et al. 7 method, reactions were performed as described above (for enzymology assays) except that reaction volumes were 40 µL with additional 0.2 mM 2-amino-6-mercapto-7-methylpurine riboside (MESG) and 0.4 U/mL purine nucleoside phosphorylase (EnzChek Phosphate Assay; Life Technologies, Carlsbad, CA). The reactions were incubated at 37 °C and the absorbance measured at 360 nm for every 15 s for a period of 30 min. A phosphate standard (150–1000 pmol) was used for product quantification. Reaction velocities were calculated with data collected at the linear phase of the reaction using GraphPad Prism (version 5; GraphPad Software, La Jolla, CA). Velocities were plotted against the amino acid concentrations, and Michaelis-Menten constants and kcat were calculated using GraphPad Prism (version 5; GraphPad Software).
Identification of IleRS Inhibitor and IC50 Calculation
We searched the National Cancer Institute/Developmental Therapeutics Program (NCI/DTP) database using the PubChem search engine for compounds with structural similarity equal to or higher than 80% of the Ile-AMP intermediate. The Ile-AMP structure (smile: CC[C@@H](C)[C@H](N)C(=O)N[P](=O)(O)NC[C@@H]1O[C@H]([C@H](O)[C@H]1O)[N]2C=NC3=C2N=CN=C3N) was used as a query. The molecule NSC616354 was identified and used for compound inhibition assays. The compound NSC616354 (stock solution prepared at 10 mM DMSO; Sigma) was used at concentrations of 0.001, 0.05, 0.1, 0.5, 1, and 10 µM. Reactions were performed with 80 µg/mL recombinant IleRS, 8 µM tRNAIle, 200 µM ATP, 2 U/mL PPiase (Sigma), and 5 mM L-isoleucine (Sigma) and incubated at 37 °C for 20 min. Reactions were stopped by addition of 100 µL malachite green and absorbance measured at 620 nm. Data points were normalized to a percentage, where 100% is equal to the mean of the data points without inhibitor after background subtraction. Background values were obtained from reactions without enzyme. The IC50 was calculated using a nonlinear regression (dose response vs. inhibitor) from GraphPad Prism (version 5) for Windows (GraphPad Software).
Results and Discussion
A Spectrophotometric Assay for Measuring Aminoacyl-tRNA Synthetase Activity
The aminoacyl-tRNA synthetase reaction produces pyrophosphate (PPi) and AMP as a by-product of ATP hydrolysis during ligation of the amino acid to its cognate tRNA ( Fig. 1a ). Therefore, we couple the inorganic pyrophosphatase (PPiase) enzyme to the aminoacylation reaction for PPi conversion to inorganic phosphate (Pi), which can be easily detected using malachite green ( Fig. 1a ). Addition of malachite green solution to the aminoacylation reaction results in malachite green reaction with Pi. As a result, a change in color from yellow to green can be detected and quantitatively measured using a spectrophotometer at a 620-nm wavelength ( Fig. 1a ). To initially test the assay, we cloned the T. brucei isoleucyl-tRNA synthetase and expressed in E. coli. The enzyme was purified using nickel-affinity magnetic beads ( Fig. 1b ), and enzymatic activity was confirmed in an aminoacylation assay using 32P-labeled tRNAIle as previously described 6 (not shown). To test the assay, we performed an aminoacylation reaction using rIleRS in the presence of PPiase, and the Pi produced was measured using malachite green solution. High levels of Pi (1402 pmol) could be detected in the reaction containing rIleRS ( Fig. 1c ), which resulted from the conversion of the PPi produced during the aminoacylation into Pi. On the other hand, a low amount of Pi (216 pmol) was detected when the assay was performed in the absence of rIleRS ( Fig. 1c ). This amount corresponds to the conversion of the PPi present in the assay components themselves, since almost no Pi could be detected in the absence of both enzymes. The amount of Pi produced in the reactions was quantified using an inorganic phosphate standard ( Fig. 1d ). This is an important part of the assay because accurate reaction quantification is necessary for enzymology studies and compound screening in drug development.
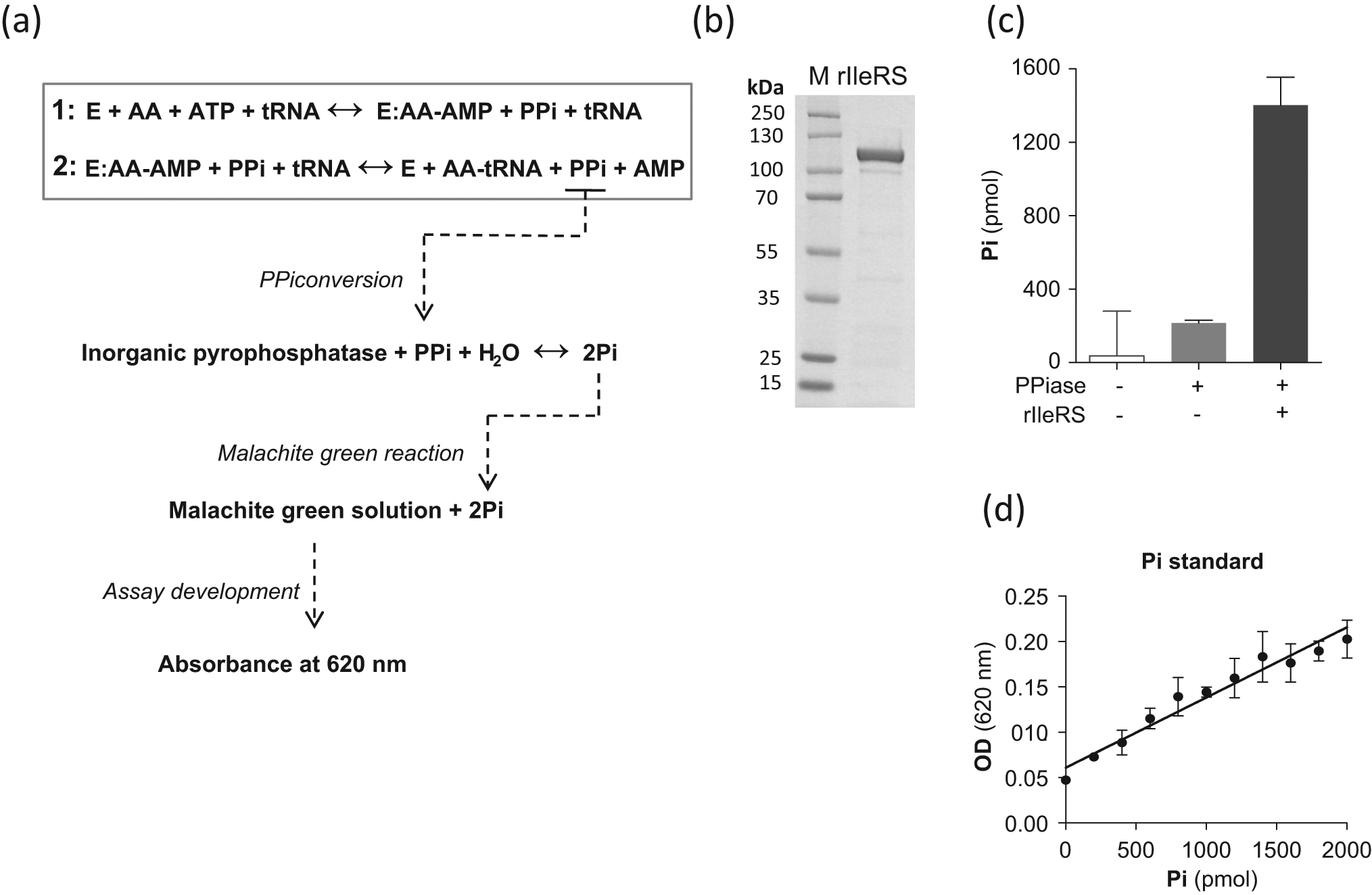
A spectrophotometric assay for measuring aminoacyl-tRNA synthetase activity. (
Preparation of the tRNA Substrate by In Vitro Transcription
An important aspect of the aminoacylation assay is the preparation of its substrate tRNA, which will vary according to the enzyme under study. For better specificity of the assay, we recommend the use of the cognate-specific tRNA rather than a mix of tRNAs (such as total cellular RNAs). The tRNA can be easily prepared by in vitro transcription reactions. The first step in this process is to amplify the tRNA sequence from genomic DNA or other sources (i.e., plasmids, etc.) to be used as a template in the in vitro transcription reaction. We amplified tRNAIle, tRNASer, and tRNAMet from T. brucei genomic DNA using specific primers. The forward primers were designed to have a T7 promoter sequence at the 5′ before the tRNA sequence ( Fig. 2a ). The addition of the T7 promoter is necessary because the PCR product can be directly used for the in vitro transcription reaction, bypassing the need to clone the tRNA into plasmids. During the in vitro transcription reaction, the T7 polymerase produces transcripts containing an extra guanine (from the T7 promoter sequence) at the 5′ of the tRNA; therefore, we have removed the first guanine of the tRNA from the forward primer sequence so that no additional sequence is present on the tRNA after in vitro transcription (see primer sequence in Materials and Methods). When this is not possible (and the extra guanine affects tRNA recognition by the enzyme), an alternative method for tRNA preparation can be used. 13 For the addition of the CCA sequence at the 3′ of the tRNA, we have added the sequence TGG to the 5′ of the reverse primer ( Fig. 2a ). Using the PCR product as a template, we in vitro transcribed the tRNAs. The tRNAs were subsequently purified by phenol/chloroform/isoamyl-alcohol extraction, followed by ethanol precipitation, and resuspended in RNAse-free water. It is important to note that at this stage, the tRNA can be stored at −80 °C. Prior to the assay reaction, the tRNAs were folded by heating at 70 °C for 10 min, followed by the addition of 10 mM MgCl2 and slow cooling at RT. The aminoacylation assay performed with rIleRS and in vitro transcribed tRNAIle resulted in Pi release, which correlates with the amount of tRNAIle in a dose-dependent manner ( Fig. 2b ). Although initial isoleucine-adenylate formation and PPi release by IleRS do not require tRNAIle, 14 its presence in the reaction is required for complete tRNA-aminoacylation. This promotes continuous amino acid activation, resulting in PPi release and formation of Ile-tRNA. Comparison of different sources of tRNAs was performed by measuring the activity of the rIleRS with in vitro transcribed tRNAIle, a mix of yeast tRNAs, and a preparation of T. brucei total RNA enriched in microRNAs. The rIleRS activity was higher when in vitro transcribed tRNAIle was used than when either yeast tRNAs or microRNAs were used ( Fig. 2c ). It is noteworthy that the use of in vitro transcribed tRNAs also avoids any possible misaminoacylation, which can happen when using multiple tRNAs from either total RNAs or yeast tRNA preparations.
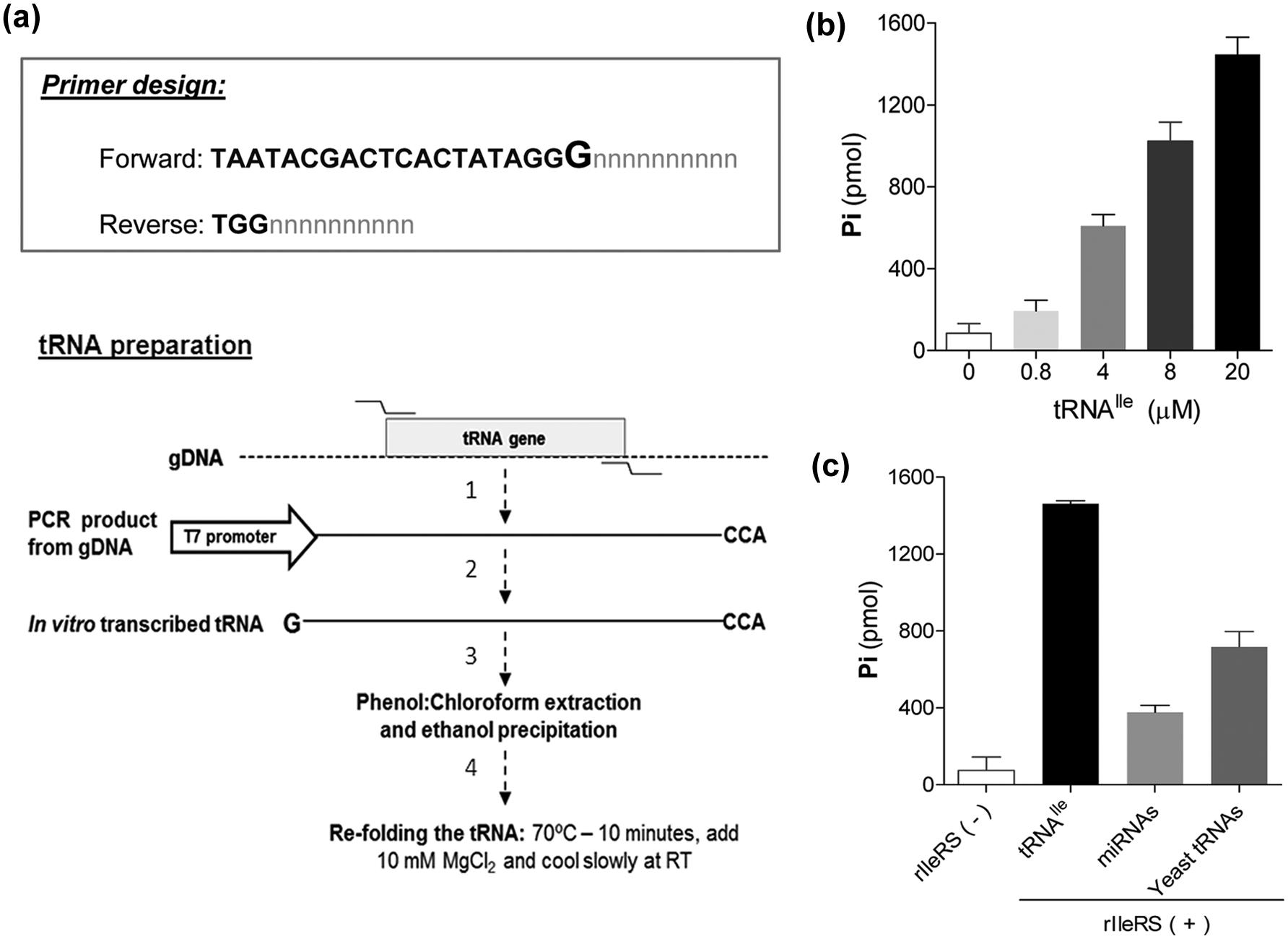
Preparation of the tRNA substrate using in vitro transcription. (
Assay Validation and Enzymology Studies
To further validate the assay, we performed aminoacylation reactions using rIleRS and its cognate in the in vitro transcribed tRNAIle. First, we performed a time course experiment with different concentrations of enzyme to determine the aminoacylation rate over time. Since the assay is performed as an end-point reaction, we used 10 mM EDTA to stop the reaction, which does not affect malachite green detection. We detected an increase in the amount of product formed over time ( Fig. 3a ). Product formation also increased according to the enzyme concentration in a dose-dependent manner ( Fig. 3a ). It is noteworthy that the production of PPi is equimolar to the formation of aminoacyl-tRNAs. 7 Furthermore, Pi conversion by PPiase has also been shown to be linear to the aminoacylation reaction, 7 thereby permitting the use of a combined aminoacylation and PPiase reaction for kinetics studies.
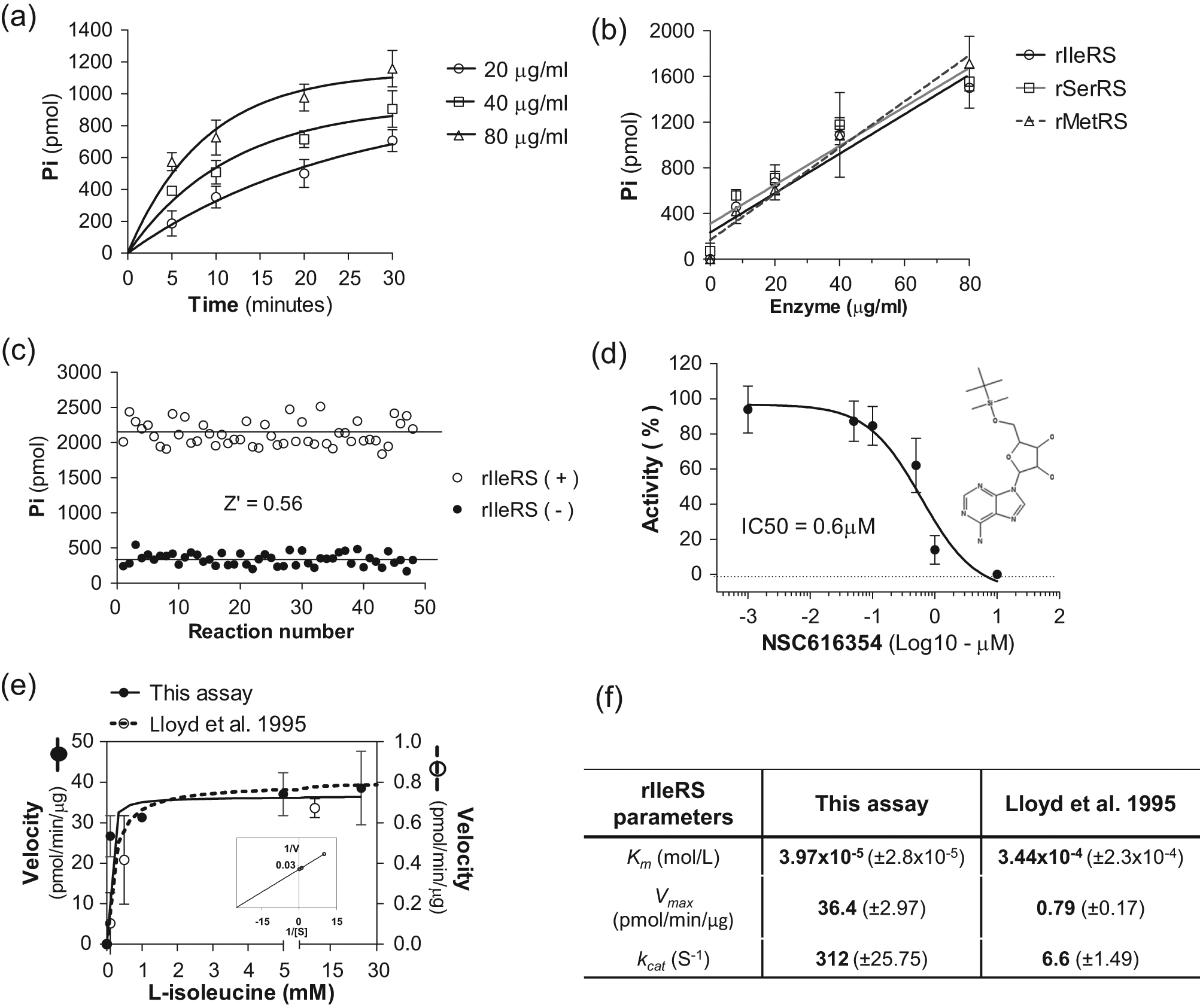
Assay validation and enzymology studies. (
A dose-response assay was performed to determine the minimum concentration of rIleRS required for detection of activity. In this assay, 8 µg/mL rIleRS (corresponding to 400 ng of enzyme per reaction) was sufficient for detecting activity ( Fig. 3b ). To test the assay with other enzymes, we expressed and purified the T. brucei seryl-tRNA synthetase from E. coli. A dose-response assay was performed using the corresponding in vitro transcribed tRNAs, and an increase in product formation was detected in a dose-dependent manner ( Fig. 3b ). Similarly, product formation increased in a dose-dependent manner when using the T. brucei methionyl-tRNA synthetase purified from the T. brucei procyclic stage using a TAP-tag ( Fig. 3b ). It is noteworthy that the enzyme concentrations used for these assays (65.5–655 nM) were less than the concentrations usually used in other methods (from 1–20 µM).5,7 We then evaluated whether this assay is suitable for HTS by calculating the Z′ factor. 12 The Z′ factor is a statistical coefficient that reflects the assay dynamic range and data variation and is used to judge whether an assay is applicable for HTS. 12 We determined the Z′ factor by measuring reactions with and without rIleRS in a 96-well plate. The mean of reactions with rIleRS was 2124 ± 174.4 pmol, whereas the mean without rIleRS was 338 ± 87.7 pmol ( Fig. 3c ). The Z′ factor was 0.56, which, according to Zhang et al., 12 is classified as a suitable (“excellent”) assay for HTS. During the aminoacylation reaction, the IleRS enzyme forms an Ile-AMP intermediate that interacts with the enzyme with high binding affinity (10−9 M).14,15 We searched the NCI/DTP database for molecules similar (≥80%) to the Ile-AMP structure to identify putative IleRS inhibitors. A similar approach has been successfully used for identification of inhibitors against bacterial IleRS. 15 Using this approach, we identified the compound NSC616354 (obtained from NCI) and tested it against the recombinant T. brucei IleRS. The NSC616354 inhibited the rIleRS activity in a dose-dependent manner and with an IC50 of 0.6 µM ( Fig. 3d ). These results indicate that this assay is suitable for compound screening and enzyme inhibition analysis.
To investigate the applicability of this assay to enzymology studies, we performed enzyme kinetics with rIleRS with varying concentrations of L-isoleucine (from 0.1–50 mM), and enzyme velocities were calculated at the linear phase of the reactions. The results showed that the enzyme reaction is dependent on L-isoleucine concentration ( Fig. 3e ). Velocities were slow at limiting concentrations of L-isoleucine (lower than 1 mM); however, as L-isoleucine concentration increased, the reaction velocity also increased until it reached its maximum. At saturating concentrations of L-isoleucine, product formation depended on the enzyme concentration (or enzyme active sites) rather than L-isoleucine availability. This result confirms that Pi production is proportional to L-isoleucine consumption. Using this experiment, we calculated the Michaelis-Menten constant (Km), Vmax, and kcat of rIleRS ( Fig. 3e , f ). The results showed kinetic parameters in a similar range reported for other organisms (i.e., Km of 3.97 × 10−5 mol/L [39.7 µM] for T. brucei rIleRS, whereas for IleRS of other microorganisms, it varied from 10–100 µM 14 ), although differences could also be related to the microorganism-specific biology. To further compare our assay, we performed a similar experiment at the same conditions using the assay published by Lloyd et al. 7 A higher Km (3.44 × 10−4 mol/L, 344 µM) was obtained using their assay ( Fig. 3e , f ). The high Km obtained by the Lloyd et al. 7 assay could be a result of the low signal-to-noise ratio of this assay. This can be observed by analyzing reaction velocities, which indicate apparent low levels of product formed over time compared with our assay. Since both reactions were performed at the same conditions, it is unlikely that the velocities are different. However, this apparent difference could have resulted either because MESG catalysis by the purine nucleoside phosphorylase enzyme is not linear with the aminoacylation reaction or because the assay sensitivity is low, resulting in absorbance values very close to the background noise.
We have developed a convenient and accurate spectrophotometric assay for measuring aminoacyl-tRNA synthetase activity. The assay was used to quantitatively measure enzyme activity, and it is suitable for HTS and enzymology studies. We also demonstrated in detail how to readily and reliably prepare the tRNA substrate, thereby facilitating the application of the assay to other aminoacyl-tRNA synthetases. The assay is very sensitive, detecting picomoles of product, and its performance (Z′ factor of 0.56) is suitable for HTS. The assay has a low cost, and since it is absorbance based, it does not require a sophisticated HTS platform for its use. The assay can also be adapted to 384-well plate formats for higher throughput screening and will facilitate the identification of aminoacyl-tRNA synthetase inhibitors for drug development.
Footnotes
Acknowledgements
We are very grateful to Dr. Lindsay Carpp for revision of the manuscript, Dr. Boniface Mailu for helpful discussions throughout this work and Christina McCormick for administrative support.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by National Institutes of Health [R01 AI078962 (to K.S)]. I.C. was awarded a National Institutes of Health fellowship administered by University of Washington [5T32AI007509-12].
