Abstract
Hepatic lipid droplets (LDs) are associated with metabolic syndrome, type 2 diabetes, hepatitis C, and both alcoholic and nonalcoholic fatty liver disease. MicroRNAs (miRNAs) are small noncoding RNAs that regulate gene expression at the level of translation. Approximately 1000 different miRNA species are encoded within the human genome, and many are differentially expressed by healthy and diseased liver. However, few studies have investigated the role of miRNAs in regulating LD expression. Accordingly, a high-content assay (HCA) was performed in which human hepatocytes (Huh-7 cells) were transiently transfected with 327 unique human miRNAs; the cells were then fixed, labeled for nuclei and lipid droplets, and imaged with an automated digital microscopy workstation. LD expression was analyzed on a cell-by-cell basis, using automated image analysis. Eleven miRNAs were identified that altered LDs. MiR-181d was the most efficacious inhibitor, decreasing LDs by about 60%. miRNA-181d was also confirmed to reduce cellular triglycerides and cholesterol ester via biochemical assays. Furthermore, a series of proteins was identified via miRNA target analysis, and siRNAs directed against many of these proteins also modified LDs. Thus, HCA-based screening identified novel miRNA and protein regulators of LDs and cholesterol metabolism that may be relevant to hepatic diseases arising from obesity and alcohol abuse.
Introduction
L
LDs occur naturally in a number of different cell types such as adipocytes, skeletal myocytes, and leukocytes. The accumulation of LDs within hepatocytes, or hepatic steatosis, is associated with a number of disease states, including metabolic syndrome, type 2 diabetes, hepatitis C, and both alcoholic and nonalcoholic fatty liver disease (AFLD and NAFLD, respectively). 5-7 The liver is mainly involved in the packaging and distribution of lipids as lipoproteins; hepatocytes generally store energy in the form of glycogen and do not contain the type of large lipid droplets typically associated with energy storage. Pathogenic accumulation of lipids within hepatocytes is most likely due to a combination of increases in free fatty acid uptake and de novo lipogenesis, as well as reduced lipoprotein export and a reduction in fatty acid oxidation. 5
The pathways involved in the development of hepatic steatosis are complex and intricately intertwined. Insulin resistance (IR) is a shared phenotype of both diabetes and the metabolic syndrome and has been demonstrated to be a major contributing factor to the development of hepatic steatosis in several animal models. Although IR in the peripheral fat tissue can result in increased circulating free fatty acids, hepatic IR can lead to alterations in a number of signaling pathways in hepatocytes, resulting in the development of hepatic steatosis. This includes a decrease in peroxisome proliferator-activated receptor α (PPARα) levels, which inhibits both peroxisomal and mitochondrial β-oxidation. 8 Inversely, hepatic steatosis results in an increase in PPARγ and a concomitant increase in lipogenic proteins and de novo lipogenesis. 9 The upregulation of a number of other transcription factors involved in lipogenesis, including sterol response element binding protein, carbohydrate response element binding protein, and liver X receptor, are all associated with obesity-related development of hepatic steatosis. 10-13 In addition, there is extensive crosstalk between the pathways involving most of these nuclear receptors and transcription factors.
MicroRNAs (miRNAs) are small noncoding RNAs that have been shown to play a role in regulating gene expression and may regulate as much as 30% of the human genome. 14 More than 700 human miRNAs have been identified to date, and estimates based on in silica analysis suggest upwards of 1000 miRNAs may exist within the human genome. 15 miRNAs regulate gene expression at the level of translation and are unique in that a single miRNA species has the ability to regulate anywhere from a handful of genes to a thousand or more. 16,17 The potential exists for a single miRNA to control the expression of multiple genes within a pathway or with genes across intertwined pathways. This possibility, combined with observations of altered miRNA levels in various disease states, has led to the hypothesis that miRNA misregulation could be a causative factor in the development and progression of disease. This also suggests that miRNAs could be used as potent therapeutics or therapeutic targets for the treatment of complex diseases. The potential for miRNA-based therapies for the treatment of liver disease is of particular interest. 18-21 Several recent studies have highlighted the therapeutic potential of manipulating miRNA levels as a method to combat hepatocellular carcinoma. 22,23
Recent studies have identified more than 100 miRNAs that are misregulated in either animal models of NAFLD or human NAFLD biopsy samples. 24,25 However, these studies do not directly demonstrate that miRNA expression is a causative mechanism in the development of NAFLD. Furthermore, many of these changes are likely due to perturbations in the mutual regulatory network that exists between miRNAs and transcription factors. This makes it difficult to identify specific miRNA species that may have a direct effect on lipid metabolism.
High-content analysis (HCA) is an emerging discipline that takes advantage of robotic microscopy platforms and image analysis algorithms to generate quantitative data, on a cell-by-cell basis, from microscopic images. HCA experiments involve staining or labeling to visualize specific structures or proteins within the cell, followed by image acquisition using either manual or robotic microscopy platforms. Image analysis algorithms, which are designed to extract quantitative data relevant to the disease/model system being studied, are then applied to the images. Current technology, using automated acquisition of images from multiwell plates combined with automated image analysis software, allows experiments consisting of thousands of images representing hundreds of thousands of individual cells to become routine. Automation of the work flow allows HCA assays to be applied to screening chemical, siRNA, or miRNA libraries for their effects in different model systems. We used this ability to design a functional high-content screen (HCS), to identify those miRNAs that regulate accumulation of lipid droplets, in a human liver cell model (Huh-7 cells) and whose misregulation may be involved in the development of hepatic steatosis.
Materials and Methods
Cell culture and transfection
HuH-7 and Hep3B human hepatoma cell lines were maintained in standard Dulbecco’s modified Eagle’s medium supplemented with 10% fetal bovine serum at 37°C and 5% CO2. For miRNA and siRNA screening, 384-well, black, glass-bottom, TC plates (Greiner Bio-One, Monroe, NC) were spotted with 1 µL of 0.5 µM miRNA or siRNA using a Bravo Pipettor (Agilent Technologies, Santa Clara, CA). Then, 0.1 µL of RNAi MAX transfection reagent (Invitrogen, Carlsbad, CA) in 10 µL of Opti-MEM (GIBCO, Carlsbad, CA) was added to each well. Each plate was incubated for 10 to 15 min at room temperature to allow association of the miRNA and the transfection reagent. Following the incubation, a suspension of 6 × 103 cells in 40 µl of media was added to each well, resulting in a final concentration of the siRNA and miRNAs of 10 nM.
Follow-up experiments on miR-181d in 96-well plates used a modified version of the transfection protocol used in the screening experiments. Briefly, bulk mixes were prepared for either the control miR (cat. AM17110; Ambion, Austin, TX) or miR181d (cat. PM12522; Ambion) that would be equivalent to .5 µL of 5 µM miRNA, .2 µL of RNAi Max, and 19.3 µL of standard media (described above) per well. The miRs used in these experiments are the same as those used in the screening experiments. The bulk mix (20 µL total volume) was pipetted into each well and incubated at room temperature for 10 to 15 min to allow association of the miRNA and the transfection reagent. A suspension of 2 × 104 cells, in 80 µL of standard media, was then added to the wells, resulting in a final concentration of the miRNAs of 25 nM. In all cases, media were replaced after an overnight incubation (12-16 h); the cells were maintained under the culture conditions described above for 4 days.
The miRNA library used in these studies consists of 327 human miRNAs (cat. 17300V2, lot AM0013; Ambion). The siRNAs used were picked from the Ambion Human Druggable Genome siRNA library version 3.1 (cat. AM80991V3.1). Complete information regarding the miRNAs and siRNAs used in this study can be found in the MIARE document (data supplement).
Cell staining, imaging, and analysis
Cells were rinsed once with cold phosphate-buffered saline (PBS) and then fixed for 15 min in 3.7% formaldehyde at room temperature. The cells were then permeabilized with 0.1% Triton X in PBS and stained for nuclei and lipid droplets using a lipid droplet imaging kit (Vala Sciences, San Diego, CA) per the manufacturer’s instructions. Wash steps on the 384-well plates were performed using a BioTek ELx405 Select CW (BioTek, Winooski, VT). Plates (both 384- and 96-well) were imaged on an automated microscopy workstation (Beckman Coulter IC100, Beckman Coulter, Fullerton, CA). Four adjacent fields of view were collected per well using a 20× objective. The 4 fields collected represent approximately 7% of the surface area of a 384 well-plate; all images were captured from the center of the well. An average of approximately 1750 cells per well were analyzed in the primary screen and secondary screens. More than 900 cells per well were analyzed for approximately 98% of the wells in the primary screen.
Image analysis was performed by Vala Science’s CyteSeer® software using the program’s Lipid Droplet Algorithm. CyteSeer® analyzes the images by first using the number and location of the nuclei to determine cell boundaries. Each cell in the field is provided with a cell ID number, allowing for cell-by-cell analysis. CyteSeer®’s Lipid Droplet Algorithm then analyzes the lipid (green) channel for each image, assigning pixels where the lipid signal is detectable to the Lipid Droplet Mask (Lm). 26 Every cell in the field of view is analyzed on an individual basis. Total integrated intensity for the lipid channel represents the sum of the pixel intensities for the pixels within the lipid mask. The mean total integrated intensity was calculated for each well and used as the basis for selecting hits.
Measurement of triglycerides
Triglyceride measurement was performed using an assay kit (Zen-Bio, Research Triangle Park, NC) per the manufacturer’s instructions. Cells were transfected with either control miR or miR-181d using the protocol for 96-well plates described above. Cells were then cultured under normal conditions; on the fourth day following transfection, the cells were lysed using the provided lysis buffer. The plates were centrifuged briefly to clear debris from the lysate, and the samples were transferred to a separate plate for analysis. Protein content was determined using a variation of the Lowry assay (BioRad, Hercules, CA). Total triglyceride content was calculated and normalized to total protein content.
Biochemical measurement of cholesterol and cholesteryl esters
Measurement of cellular cholesterol and cholesteryl esters was performed using an assay kit (BioVision, Golden, CO) per the manufacturer’s instructions. To have enough material to perform the cholesterol assay, cells from 12 wells, each transfected with either the negative control miR or miR-181d, were pooled to create a single replicate. Cells were transfected according to the protocol for 96-well plates described above. Pooling of wells was used to circumvent the need to reestablish the transfection protocol in a format larger than 96-well plates.
Results
Validation of the HCA assay for LDs in Huh-7 cells
Huh-7 were chosen as a model system to explore the role of miRNAs in the development of hepatic steatosis because this cell line develops steatosis spontaneously in culture and is lipogenic. 4,27,28 Huh-7 cells also have triglyceride and cholesterol levels several times higher than human hepatocytes immortalized from normal livers. 29 To analyze the effects of miRNAs on LDs, Vala Sciences Lipid Droplet staining kit components and image analysis software (CyteSeer®) were used that have been validated on a variety of adipocyte and hepatocyte cell models. 26 Images were acquired from the cells using a 20× objective, and the images were analyzed using the Lipid Droplet Algorithm contained within Vala’s CyteSeer® cell image analysis program. The data parameter used for this study was Tii Li Lm (total integrated intensity of the lipid image, integrated over pixels that correspond to the lipid mask), which is an estimate of the total cellular LD content.
Because miRNAs could potentially either increase or decrease lipid droplet formation, the high-content assay was validated by treating the cells with agents expected to either increase or decrease cellular lipid droplet content. Control cells typically exhibited numerous LDs in perinuclear locations (
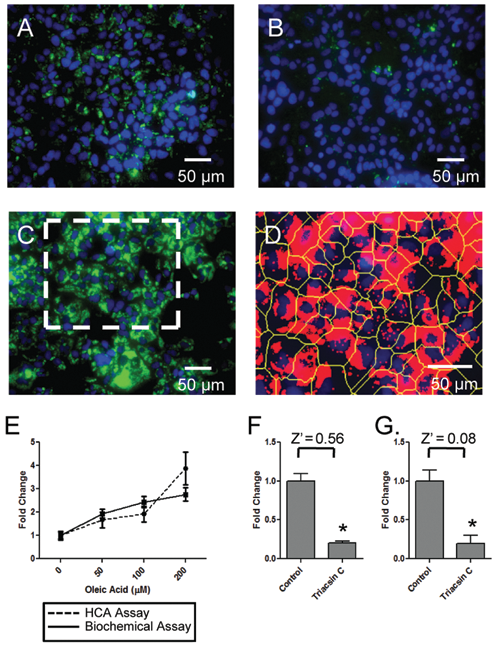
Validation of the high-content assay (HCA) for liquid droplets (LDs) in Huh-7 cells. Huh-7 cells were exposed to either oleic acid or triacsin C for 18 h (overnight), then fixed and visualized for nuclei and lipid droplets; the cells were then imaged with an IC100 workstation and the images analyzed for lipid droplets via CyteSeer®. Alternatively, similarly treated Huh-7 cells were assayed for triglyceride content via a biochemical assay. (
For cells exposed to oleic acid, the HCA matched the sensitivity of the biochemical assay up to oleic acid concentrations of 100 µM (
In previous studies, CyteSeer® quantified oleic acid–induced increases in Tii Li Lm in Huh-7 cells with excellent discrimination between the control and oleic acid–treated cells (Z′ = 0.66 was achieved
26
; Z′ defined by Zhang et al.
31
). To test if changes in Tii Li Lm could also be used to quantify decreases in lipid content elicited by experimental treatment, we compared the HCA and the biochemical triglyceride assay for cells treated with triacsin C. Notably, triacsin C exposure led to very similar reductions (approx. 80%) in Tii Li Lm (
Screening the Ambion miRNA library
The preliminary screen was conducted in 384-well plates using a library consisting of 327 human miRNAs that were introduced into the cells via “reverse transfection” (miRNAs were spotted onto the plates along with transfection reagent prior to seeding with cells). Two identical plates were created and analyzed in parallel. The well plates were incubated in standard media for 4 days prior to being fixed and stained for nuclei and lipids. An average of 1750 cells were analyzed per well. To aid in visualizing the data, results for the Tii Li Lm data parameter for each well were normalized to overall plate average and the results for the 2 plates graphed next to each other (
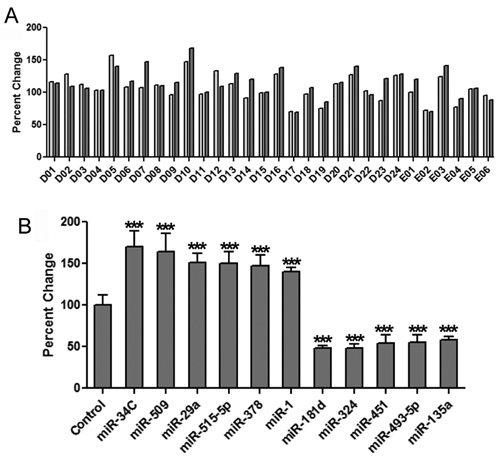
High-content assay (HCA) reveals 11 miRNAs that regulate intracellular lipid content in Huh-7 cells. (
The 75 miRNAs were included in a secondary screen. As with the primary screen, duplicate 384-well dishes were used, but this time each miRNA species was represented by 2 wells per dish (4 wells total, per miRNA). Controls transfected with a scrambled miRNA were also included in the screen (n = 24 control wells), and the lipid content for each well was normalized to the average lipid content of the control wells. A threshold of ±40% from the control mean was used to identify hits in the secondary screen. This resulted in the identification of 11 miRNAs that affected intracellular lipid content, including 6 miRNAs that increased lipid content and 5 miRNAs that reduced the intracellular lipid content (
Follow-up assays for miR-181d
Of the hits that reduced intracellular lipid content, miR-181d was found to have the strongest effect in the Huh-7 cells, reducing lipid content by more than 60%. Follow-up experiments used miRNAs purchased from Ambion separately from the original library and in which the assay was scaled up to the 96-well format. Transfection with miR-181d consistently elicited an approximate 60% reduction in Tii Li Lm (
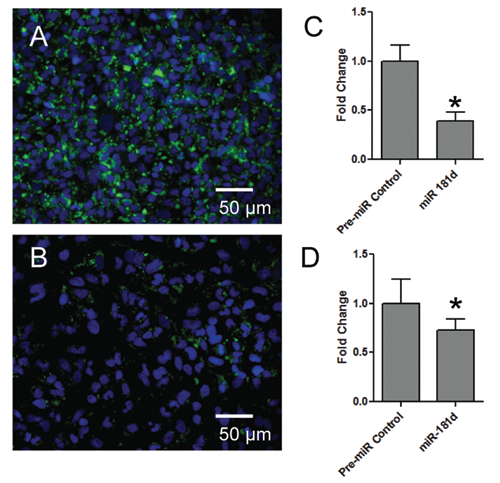
miR-181d is the most effective miRNA at reducing intracellular lipid content. Huh-7 and HEP3B cells were cultured in 96-well dishes, transfected as indicated, fixed, visualized, and analyzed by the high-content assay (HCA). (
Effects of miR-181d on cellular triglyceride levels
To further confirm the effects of miR-181d in modulating intracellular lipid content, Huh-7 cells were transfected with miR-181d and triglyceride levels were measured biochemically. Transfection with miR-181d reduced triglyceride levels by 30% in the biochemical assay (
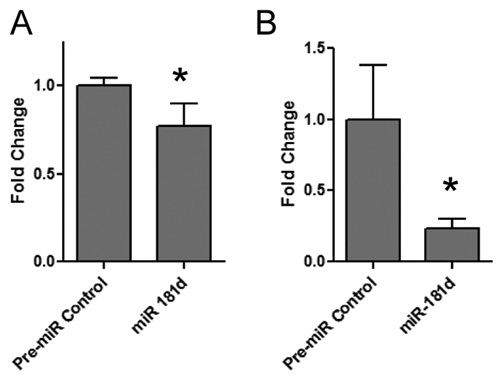
Effects of miR-181d on Huh-7 cellular triglycerides and total cholesterol. The effect of miR-181d on intracellular (
Effects of miR-181d on cellular cholesterol
Lipid droplets within Huh-7 cells contain cholesteryl esters as well as triglycerides.
32
It was therefore of interest to investigate the effects of miR-181d on cellular cholesterol. Accordingly, total cellular lipids were extracted in an organic solvent, dried, resuspended in assay buffer, and incubated with cholesterol esterase (which metabolizes cholesteryl esters to cholesterol). The cholesterol assay revealed that transfection with miR-181d reduced total cholesterol content in the Huh-7 cells by approximately 75% (
siRNA screening to identify novel regulators of lipid droplet metabolism in hepatocytes
miRNAs have the ability to regulate the transcription of many mRNA species simultaneously and influence a number of different signaling pathways. miRNA target prediction algorithms can be used to identify genes that might be targeted by specific miRs. We decided to see if we could use an miR target prediction algorithm and the HCA assay in combination with an siRNA screen to identify novel regulators of lipid droplet metabolism in hepatocytes. To compile a list of target genes, we identified the 9 most effective miRs, 5 that increased lipid droplets (miR-324, -422, -451, -133, 493-5p) and 4 that decreased lipid droplets (miR-29a, -30a, -30b, -520C) in our primary screen. Using the online target prediction algorithm, Targetscan (www.targetscan.org), 111 target genes that were targeted by at least 2 miRs were identified. 16,17 Unfortunately, we only had access to siRNA sequences for 19 of these genes.
Because we were limited to a small number of siRNAs, we expanded the list by including miRNA targets identified from hits in a second miRNA screening experiment (report in preparation), which identified miRs that regulate the expression of the LD-associated protein, adipose differentiation-related protein (ADFP). Here again, the top 9 most effective miRs (increase in ADFP [miR-215, -96, -214, -122, -489] and decrease in ADFP [let7d, let7e, let7g, mir-148a]) were used to generate a list of genes targeted by at least 2 of the miRs. This produced a list of 208 genes, 48 of which were available for screening.
The screen was performed with 4 siRNA species per gene (a total of 268 siRNA sequences were screened), with a single siRNA species used per well; duplicate plates were screened. A scrambled siRNA sequence was used as the control for hit identification, and a threshold of ±20% from the control mean was used for hit identification. For genes where multiple sequences met the 20% threshold, the results of the most effective siRNA are reported. The more moderate threshold was used as the miRNAs typically target multiple genes simultaneously and therefore may produce larger changes in lipid content. siRNAs, on the other hand, target a single gene and therefore may have a more moderate effect on lipid content. In addition, the extent of protein knockdown for each siRNA is unknown, and therefore the lower threshold was used.
Following image analysis, siRNA targeting of 26 of the 67 genes was found to result in alteration of the lipid content by 20% or more. This included 11 genes that decreased lipid content (
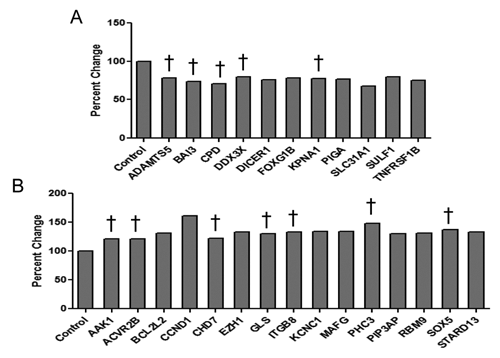
siRNA-mediated effects on lipid droplet content in Huh7 cells. An miRNA target prediction algorithm was used to identify genes targeted by miRs that regulate lipid content and adipose differentiation-related protein (ADFP) expression in hepatocytes. siRNAs directed toward these proteins were then screened for their ability to modify intracellular lipid content using the high-content assay. Of the 67 genes targeted by siRNA, 26 either (
Discussion
The role of miRNAs in hepatic disease has been the topic of many recent studies. The pathogenic accumulation of the lipids within hepatocytes is associated with obesity-related diseases such as metabolic syndrome, type 2 diabetes, NAFLD, and hepatitis C virus (HCV). Altered expression of more than 100 miRNAs has been reported in relation to the development and progression of the NAFLD. 24,25 However, given that many of these changes are likely due to perturbations in the mutual regulatory network that exists between miRNAs and transcription factors, it is difficult to identify specific miRNAs that might have a causative role in the development of hepatic steatosis. Therefore, we designed an HCA that would allow us to screen for miRNAs whose overexpression in human hepatocyte cell lines would influence intracellular lipid content. Our initial experiments demonstrated that Z′ scores of 0.66 and 0.56 were obtainable for the measurement of the increase or decrease, respectively, of intracellular lipid content in Huh-7 cells. With a large dynamic range for both the increase and decrease in lipid content, we could perform screens to identify miRNAs that might have a potent effect on de novo lipogenesis, resulting in either the accumulation or reduction of intracellular lipids.
Using a stringent cutoff of a 40% increase or decrease in lipid content for our secondary screen, we identified 11 miRNAs that altered the intracellular lipid content of the Huh-7 hepatocytes. These 11 miRNAs are a much more manageable list of miRNAs to follow up on compared to the 100 or more miRNAs that were found to be misregulated in animal models and human biopsies of the NAFLD. Comparison of our hits with the miRNAs that were found to be misregulated in these earlier studies showed that 3 of the 11 miRNAs (miR-29a, miR-34C, and miR-135a) were previously shown to be altered in NAFLD livers. In addition, a member of the miR-181 family, miR-181b, was altered in NAFLD. In our screen, miR-181d was the most potent inhibitor of lipid accumulation, reducing intracellular lipids by 60% from control levels. The mature miRNA sequences of mir-181b and mir-181d are identical.
Experiments in the Hep3B cell line demonstrated that the effects of miR-181d are not specific to Huh-7 cells and that miR-181d may play a more general role in regulating lipid content of hepatocytes. The biochemical assays that examined the levels of specific lipids in the hepatocytes were of particular interest. We observed a 30% reduction in triglyceride content in response to miR-181d, confirming the results of the HCA. In addition, because lipid droplets in Huh-7 cells also contain cholesteryl esters, we tested the effect of miR-181d on total cellular cholesterol and found that it reduced cholesterol content by 75%. The combination of triglyceride and cholesterol reduction by miR-181d may have important implications in the treatment of fatty liver disease. Interestingly, misregulation of miR-181 has been implicated in the development of hepatocellular carcinoma 22 ; the development of the hepatic steatosis has long been regarded a significant risk factor for the development of liver cancer.
One aspect of the study of particular interest was the use of miRNA target analysis to identify novel genes and pathways that may be involved in hepatic lipid droplet metabolism. To do this, we compiled a list of genes targeted by miRNAs that had shown an ability to regulate intracellular lipid content in our preliminary screen. To identify individual genes that had the ability to regulate intracellular lipid content, we performed an siRNA screen, in Huh-7 cells, based on these miRNA target genes. siRNA knockdown of transcripts from 26 of the 67 genes tested resulted in changes in intracellular lipid content of >20%. Interestingly, 12 of these 26 proteins are likely targets of miR-181d. Lending credence to this investigative strategy, several of the identified proteins have previously been implicated in lipid metabolism. For example, ADAMTS-5 (A Disintegrin And Metalloproteinase with Thrombospondin motif) is expressed in subcutaneous and gonadal adipose tissue and demonstrates increased expression in obesity. 33 Here transfection with an siRNA against ADAMTS-5 decreased the lipid content of Huh-7 cells. Similarly, an siRNA against DDX3X (DEAD Box Protein 3) reduced lipid content in the Huh-7 cells. DDX3X has been shown to be associated with lipid droplets in hepatocytes that express HCV core protein. 34 In addition, an siRNA directed against the tumor necrosis factor receptor family member TNFRSF1B, which has recently been linked to familial combined hyperlipidemia, was found to reduce intracellular lipid content in Huh-7 cells. 35 The screen revealed a number of hits that will be followed up on in future studies.
Our results, which demonstrate that specific miRNAs can regulate intracellular lipid content, strongly support the hypothesis that miRNAs have a direct effect on lipid metabolism. In comparison to previous reports that identified more than 100 miRs whose expression may be altered in fatty liver disease, we identified a relatively small subset of miRs that can directly regulate intracellular lipid content in hepatocytes. These results suggest that alterations in the expression of a relatively small number of miRNAs may have far-reaching implications on the expression of other miRNAs and their target genes. This emphasizes the need for functional studies to elucidate the roles of miRNAs in the development and progression of disease and also highlights the complex role of miRNAs in regulating gene expression.
Footnotes
Acknowledgements
We also thank Zen-Bio for providing assay kits and expert knowledge regarding their use.
This work was partially funded by the SBIR phase II grant, “HT Image Assay of Lipid Droplet Formation in Human Adipocytes” (2 R42 DK074333-02A1), from the National Institute of Diabetes, Digestive and Kidney Diseases.
