Abstract
To evaluate yeast as a high-throughput cell-based system for screening chemicals that may lead to drug development, 10,302 full-length human cDNAs (~50% of the total cDNAs) were introduced into yeast. Approximately 5.6% (583 clones) of the cDNAs repressed the growth of yeast. Notably, ~25% of the repressive cDNAs encoded uncharacterized proteins. Small chemicals can be readily surveyed by monitoring their restorative effects on the growth of yeast. The authors focused on protein kinases because protein kinases are involved in various diseases. Among 263 protein kinase cDNAs (~50% of the total) expressed in yeast, 60 cDNAs (~23%), including c-Yes, a member of the Src tyrosine kinase family, inhibited the growth of yeast. Known inhibitors for protein kinases were examined for whether they reversed the c-Yes-induced inhibition of the yeast growth. Among 85 inhibitors tested, 6 compounds (PP2, PP1, SU6656, purvalanol, radicicol, and geldanamycin) reversed the inhibition, indicating a high specificity sufficient for validating this screening system. Human c-Yes was found to interact with Hsc82, one of the yeast chaperones. Radicicol and geldanamycin probably exerted their actions through interactions with Hsc82. These results indicate that when human proteins requiring molecular chaperones for their activities are subjected to the yeast screening system, 2 groups of chemicals may be found. The actions of one group are exerted through direct interactions with the human proteins, whereas those of the other group are mediated through interactions with chaperones.
Introduction
A
The feasibility of such a yeast cell-based screening system has been tested using cDNAs from mainly human and some viral origins. 4 Among 38 cDNAs tested, 12 cDNAs (30%), including those of the mitotic spindle checkpoint kinase BUB1, p38MAPK, aurora-related kinase 1, poly(ADP-ribose) polymerase 1 (PARP1), and PI3 kinase, showed significant interference with the yeast growth. PARP1 is a DNA binding protein that detects DNA strand breaks and also catalyzes adenosine diphosphate (ADP) ribosylation of various proteins. Novel inhibitors of this enzyme have been identified using a yeast strain expressing human PARP1. 5 Its activity has been implicated in diseases such as cancer, stroke, and neurotrauma, and potent inhibitors of this enzyme are now being evaluated clinically. 6-8 Interestingly, PARP activity is not present in yeast, indicating that yeast-based phenotypic screens can even be successful for human proteins whose homologues are not present in yeast.
The completion of the sequencing of the human genome and the progress of the human cDNA projects have made it possible for most, if not all, human cDNAs to be expressed in yeast. Given that a large number of proteins probably remain to be assigned as causes of diseases, more comprehensive screening with a yeast cell-based strategy may cover a far broader range of lead compounds. Furthermore, even if they are not druggable, it is also possible that analyses of the biological alterations induced by novel compounds in human cells might reveal unexpected protein functions. In view of these points, we have designed a research project, designated the “humanized yeast project,” in which comprehensive screening of human cDNAs that inhibit yeast growth is planned. In the present study, we expressed 10,302 human cDNAs (~50% of the total cDNAs) in a budding yeast (Saccharomyces cerevisiae) and found that approximately 5.6% of the cDNAs, including those for protein kinases, interfered with the yeast growth.
Protein kinases are the largest enzyme superfamily involved in cell signal transduction. The human genome contains more than 500 genes encoding protein kinases, 9 and at least 30% of human proteins are phosphorylated by protein kinases, 10 suggesting that protein kinases play crucial roles in human physiology and pathophysiology. Consequently, protein kinases represent therapeutic targets for a range of diseases. Indeed, many successful developments of protein kinase inhibitors and HTS technologies for drug discovery against the human kinome have been reviewed recently. 11 This article reports the feasibility of a yeast cell-based method for surveying inhibitors of human protein kinases. This strategy was validated using human c-Yes, a tyrosine kinase, and known inhibitors of c-Yes. Budding yeast contains more than 100 known serine/threonine kinases, but no typical tyrosine kinases have been found to date. 12
Materials and Methods
Yeast strains
To screen for human cDNAs that repressed yeast growth, an S. cerevisiae (budding yeast) wild-type strain, W303-1A (MATa leu2-3,112 trp1-1 can1-100 ura3-1 ade2-1 his3-11,15), or BY4742 (MATα ura3Δ0 leu2Δ0 his3Δ1 lys2Δ0) was used. For the experiments involving protein kinase inhibitors, a triple-mutant yeast strain deficient in ERG4, PDR1, and PDR3, which are involved in membrane permeability and the efflux of xenobiotics, was generated from the W303-1B strain (MATα leu2-3,112 trp1-1 can1-100 ura3-1 ade2-1 his3-11,15) (Mukai et al., manuscript in preparation). To generate a triple-mutant yeast strain expressing C-terminally HA-tagged Hsp82 (Hsp82-HA) or Hsc82-HA, PCR-based HA tagging was performed according to a previously described method. 13 In these cells, endogenous HSP82 and HSC82 were replaced with HSP82-HA and HSC82-HA, respectively.
Plasmids, transformation, and cultures
Three types of vector plasmids, comprising a multicopy YEp-type plasmid (pYES-DEST52; Invitrogen, Carlsbad, CA), a single-copy YCp-type plasmid, and an integration YIp-type plasmid, were used for expressing human proteins in yeast. The human cDNAs were transferred from the entry clones 14 to pYES-DEST52 using the Gateway LR reaction and a recombinase (Invitrogen) according to the manufacturer’s protocol. To construct the YCp-type plasmid, pYES-DEST52 was digested with Spe I, Sap I, and Stu I, and the resulting 2.7-kb fragment containing the GAL1 promoter and the Gateway cassette was isolated. This fragment was blunt-ended with a Blunting High Kit (Toyobo, Osaka, Japan) and inserted into the Sma1 site of the expression vector pRS316 (provided by the National Bio-Resource Project [NBRP] of the MEXT, Japan). 15 To construct the Yip-type plasmid, pYES-DEST52 was digested with Nhe I and SnaB I to delete the 2µ region, blunt-ended with the Blunting High Kit, and self-ligated.
In all of the expression plasmids, expression of human cDNAs was under the control of the GAL1 promoter. Transformation of yeast with the plasmids was carried out according to a previously described method. 16 Precultures of yeast before galactose-induced expression of c-Yes were carried out in media containing 2% glucose or 2% raffinose. In experiments involving time-dependent induction of c-Yes, the cells were precultured in the raffinose media because the glucose-induced repression of the GAL1 promoter continued for a while after the removal of glucose, whereas raffinose did not have this effect. Therefore, when the precultures were carried out in the presence of raffinose, the GAL1 promoter was immediately activated after the addition of galactose. Induction of c-Yes expression was initiated in media containing 2% galactose after washing out glucose or raffinose in the precultures.
Screening of cDNAs that suppress the growth of yeast
To assess the effects of human proteins on the growth of yeast, wild-type yeast cells harboring pYES-DEST52 containing a human cDNA were grown in the noninducing (glucose) liquid medium at 30 °C for ~12 h until the cultures reached saturation. After 2 washes with sterile water, the cells were resuspended in the inducing (galactose) medium and serially diluted by 5-fold. The resulting cell suspensions were spotted onto plates containing the noninducing or inducing medium and incubated at 30 °C for 2 days. As a control, yeast cells harboring the empty vector were treated in the same way.
Protein kinase inhibitors and their effects on c-Yes-induced repression of yeast growth
The effects of known protein kinase inhibitors on the c-Yes-induced repression of yeast growth were examined using the triple-mutant yeast strain that expressed c-Yes through the YCp-type plasmid. The protein kinase inhibitors were all dissolved in DMSO. The inhibitor solutions were serially diluted by 2-fold with DMSO, and their final concentrations ranged from 100 µM to 1.52 nM (18 different concentrations). The final concentration of DMSO was adjusted to 1%. The restorative effects of the protein kinase inhibitors on the c-Yes-induced growth retardation of yeast were assayed by measuring the cell densities after culture of the cells in the presence of the inhibitors or 1% DMSO alone. Yeast cells harboring expression plasmids for c-Yes were grown in the noninducing (glucose) liquid medium at 30 °C for ~12 h until the cultures reached saturation. The cells were washed twice with sterile water and suspended in the inducing (galactose) liquid medium. The suspensions were diluted with the medium to adjust their optical densities at 600 nm to 0.01. The resulting cell suspensions were dispensed into 96-well plates at 99 µL/well, and 1 µL of an inhibitor was added. The plates were incubated with 30 °C, and the optical densities were determined after specified intervals (24, 48, and 72 h). The ratios of the optical densities between the cultures in the presence of inhibitors and the control cultures (DMSO alone) were calculated.
Cell extracts and immunoblotting
Harvested cells were washed twice with distilled water and lysed using glass beads in a lysis buffer consisting of 50 mM Tris-HCl (pH 8.0), 150 mM NaCl, 5 mM EDTA, 1.0% Triton X-100, 1 mM dithiothreitol [DTT], 1 mM phenylmethylsulfonyl fluoride [PMSF], and a protease inhibitor cocktail (Nacalai Tesque, Kyoto, Japan). After centrifugation of the lysates at 13,200 rpm for 30 min, the supernatants were analyzed by immunoblotting with the following primary antibodies: anti-c-Yes (1:5000; BD Biosciences Pharmingen, San Diego, CA), anti-HA (1:2000; MBL, Nagoya, Japan), anti-α-tubulin (generously provided by Dr. Andrea Baines, Sir William Dunn School of Pathology, University of Oxford), and anti-phosphotyrosine (Millipore, Billerica, MA). As the secondary antibodies, horseradish peroxidase (HRP)–conjugated antimouse IgG antibody and HRP-conjugated antirabbit IgG antibody (1:5000; Invitrogen) were used.
Coimmunoprecipitation of human c-Yes with yeast chaperones
The triple-mutant yeast strains expressing the HA-tagged yeast chaperones (Hsp82-HA or Hsc82-HA) were engineered to produce c-Yes-V5 fusion proteins under the control of the GAL1 promoter. The cells harboring the c-Yes-V5 expression plasmid were precultured in the noninducing medium containing raffinose. The culture medium was then replaced with the inducing medium containing galactose, and the cells were cultured for 2 h. The supernatants of cell lysates were obtained as described above and treated with anti-V5 antibody-conjugated agarose beads (Sigma). The immunoprecipitates were subjected to immunoblotting to analyze whether c-Yes coimmunoprecipitated with Hsp82 and/or Hsc82. c-Yes in the precipitates was detected using an anti-c-Yes antibody, whereas Hsp82 and Hsc82 were detected using an anti-HA antibody.
RT-PCR
The triple-mutant yeast strain harboring the expression plasmid for c-Yes was cultured in the presence of galactose for 2 h. Radicicol (0.78 µM) or geldanamycin (6.3 µM) was added at zero time. After 2 h of culture, total RNA was isolated using an RNeasy Mini Kit (Qiagen, Piscataway, NJ) according to the manufacturer’s protocol. RT-PCR was conducted with ReverTra Ace-α (Toyobo) using 0.8 µg of each total RNA and specific primers that amplified the c-Yes cDNA. As an internal control, the mRNA of α-tubulin was detected. The primers used were 5′-TATGTAGCGCCTGCAGATTCCATTC-3′ and 5′-CCATATCAACCAGCTGTGGAAGCTTCA-3′ for c-Yes and 5′-CAAGAGGCCATTACACCGTTGGTAGA-3′ and 5′-CACCCCTGTATAACAGACAAGTAGCCA-3′ for α-tubulin.
Results
Interference of yeast growth by human proteins
To screen the human proteins that repressed the growth of yeast, the full-length human cDNAs of 10,302 (~50% of the total cDNAs) were introduced into yeast cells using a multicopy-type yeast expression vector in which the expression of the human cDNAs was galactose inducible. Figure 1 shows a representative example of the inhibition of yeast growth. The galactose-induced expression of human c-Yes caused repression of the yeast growth, whereas cultures in the noninducing condition (glucose medium) showed no effect on the growth. c-Yes is a plasma membrane-associated tyrosine kinase that belongs to the Src family as described below.
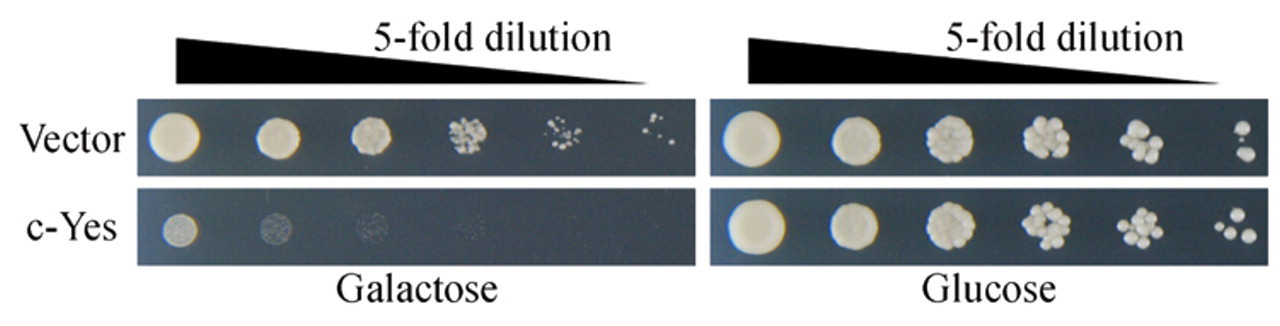

Human c-Yes inhibits the growth of yeast. The yeast strain harboring the YEp-type plasmid engineered to express c-Yes under the control of the GAL1 promoter was cultured in the noninducing glucose medium. The cultures were then serially diluted by 5-fold and spotted onto plates containing the inducing galactose medium or noninducing glucose medium. The data show the growth after incubation for 2 days. Yeast cells containing the empty vector were used as controls.
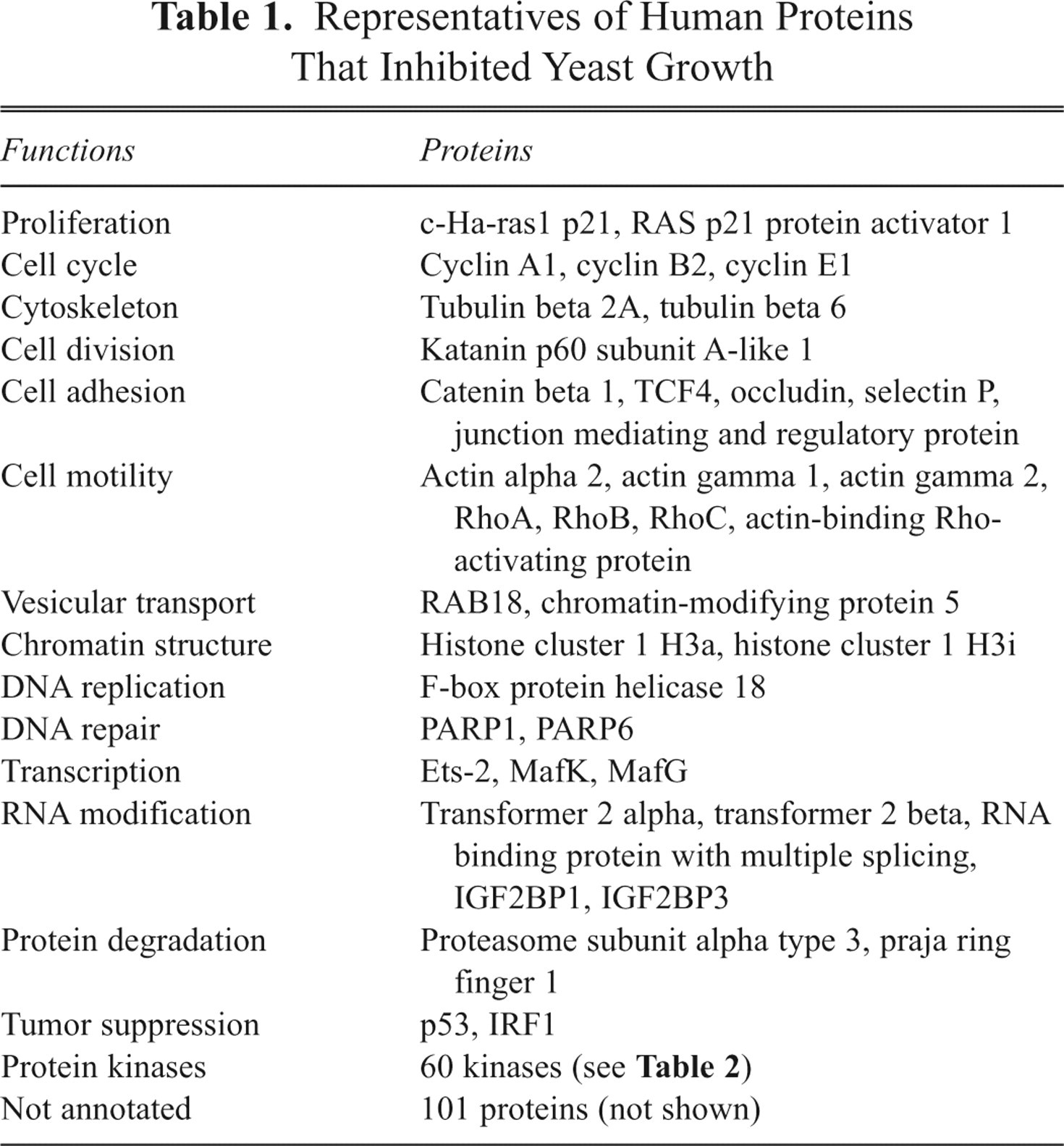
Among the cDNAs, 583 clones (~5.6% of the tested clones) showed inhibitory effects on yeast growth. We did not examine the expressions of mRNAs or proteins encoded by cDNAs that showed no effect on the yeast growth. Table 1 lists representatives of the human proteins that inhibited the yeast growth. These proteins included proteins that play crucial roles in a wide variety of cell behaviors, such as cell proliferation, cytoskeleton, DNA replication, protein kinases, and others, which may be promising targets for screening chemicals. Interestingly, ~25% of the human proteins that showed inhibitory effects on the yeast growth had unknown cellular functions. Studies of the molecular mechanisms by which the yeast growth is repressed by these uncharacterized proteins and the identification of small compounds that can reverse the repression of yeast growth will greatly contribute to clarifying the physiological functions of these proteins.
Representatives of Human Proteins That Inhibited Yeast Growth
Interference of yeast growth by human protein kinases
We focused on protein kinases because they are strongly related to various diseases. Among the 263 (50% of the total) protein kinases tested, 60 protein kinases (~23%) caused repression of the yeast growth ( Table 2 ). Table 3 shows the percentages of the repressive enzymes in different kinase subfamilies. The tyrosine kinase subfamily showed a high score of 35.5%. Hsp90, a molecular chaperone in vertebrates, binds to its client proteins during or immediately after their synthesis and thereby guarantees the proper folding of the client proteins. Because the numerous signaling proteins whose mutations and aberrant expression cause cancers are clients, Hsp90 is an emerging therapeutic target for treatment of cancer. 17 Src family members, including c-Yes, require Hsp90 for maintaining their functions. Hereafter, c-Yes was used for the validation of the yeast-based screening system for small molecules because if yeast homologues of Hsp90 interact with c-Yes, we would be able to identify the molecules that inhibit the function of Hsp90 as well as those that inhibit c-Yes directly.
Human Protein Kinases That Inhibited Yeast Growth
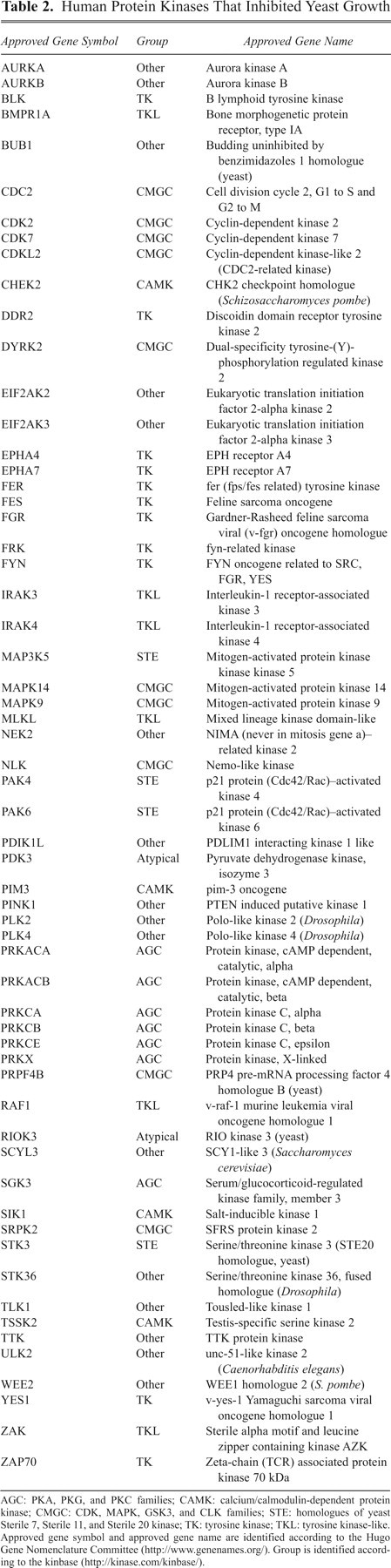
AGC: PKA, PKG, and PKC families; CAMK: calcium/calmodulin-dependent protein kinase; CMGC: CDK, MAPK, GSK3, and CLK families; STE: homologues of yeast Sterile 7, Sterile 11, and Sterile 20 kinase; TK: tyrosine kinase; TKL: tyrosine kinase-like. Approved gene symbol and approved gene name are identified according to the Hugo Gene Nomenclature Committee (http://www.genenames.org/). Group is identified according to the kinbase (http://kinase.com/kinbase/).
Protein Kinase (PK) Subfamilies That Inhibited Yeast Growth
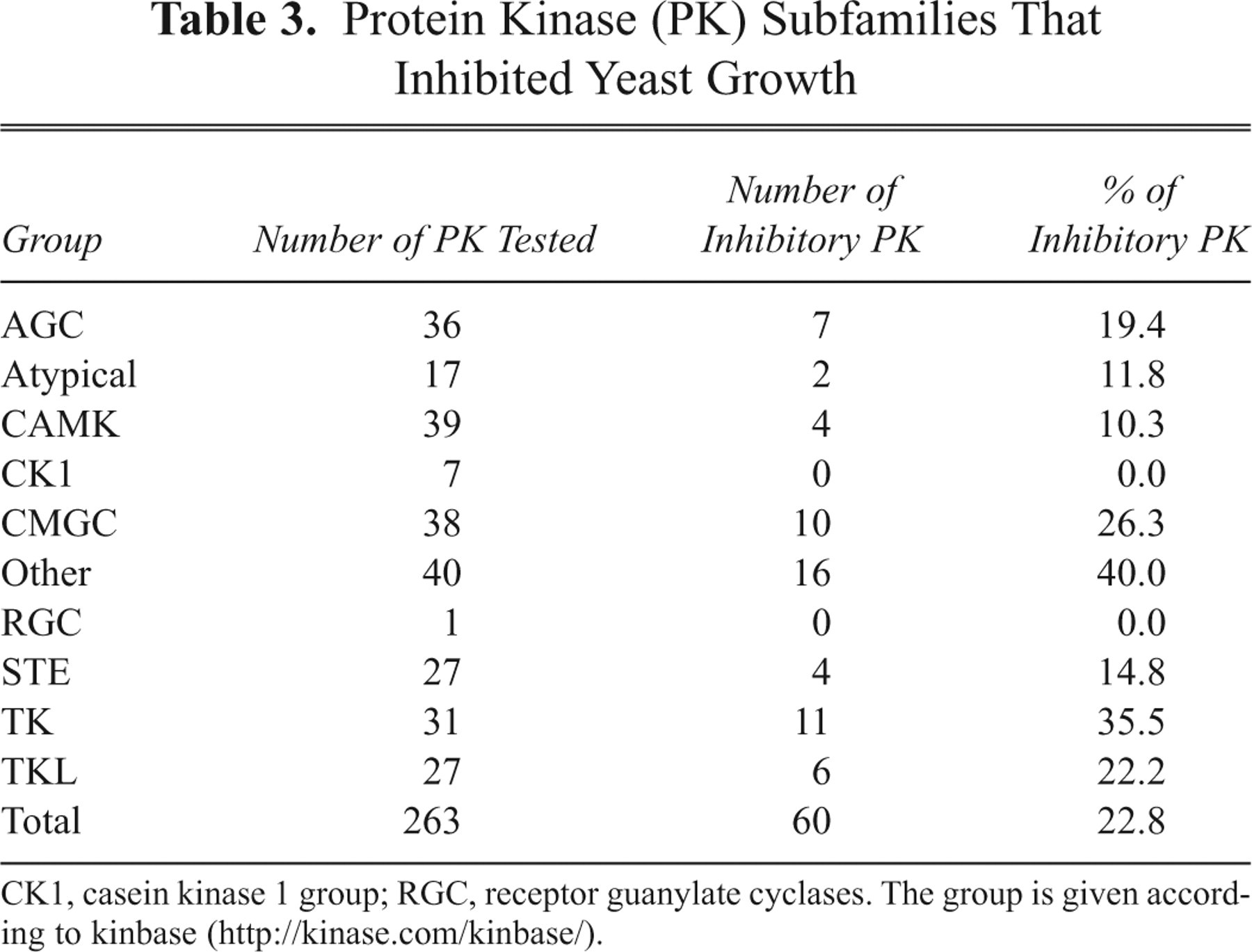
CK1, casein kinase 1 group; RGC, receptor guanylate cyclases. The group is given according to kinbase (http://kinase.com/kinbase/).
Optimization for screening and validation of the screen system
One drawback of using yeast for large-scale drug screening is the presence of the cell membrane, which may function as a permeability barrier and contains efflux pumps for small molecules. Improvements have been attempted by generating yeast strains that lack genes that are probably responsible for these disadvantages. 18,19 To increase the intracellular concentrations of the small molecules to be tested, we prepared a triple-mutant yeast strain that lacked ERG4, PDR1, and PDR3 (Mukai et al., manuscript in preparation). Erg4 is involved in the biosynthesis of ergosterol, which strengthens the permeability barrier of the yeast membrane to small molecules, 18 whereas Pdr1 and Pdr3 are transcription factors that stimulate the expression of pump proteins for the efflux of xenobiotics. 19 As described below, 6 compounds including PP2 were found to recover the c-Yes-induced repression of the yeast growth. PP2 is one of the known inhibitors of c-Yes. 20 Cyclin A1 repressed the growth of yeast ( Table 1 ), and radicicol, an Hsp90-mediated tyrosine kinase inhibitor (see below), reversed this repression by an unknown mechanism. To show that the triple-mutant strain is more convenient for screening chemical compounds than the wild-type strain, we compared these 2 strains with respect to the PP2- and radicicol-dependent recovery of growth. As shown in Figure 2a , the dose-dependent curve of PP2 in the triple-mutant strain was similar to that in the wild-type strain, indicating that the permeability increase by mutations did not improve the sensitivity to PP2 (EC50 app = 0.54 µM in the wild-type strain and 0.29 µM in the mutant strain), but the maximum growth recovery of the mutant yeast was seen at 4-fold lower concentration of PP2 than that of the wild-type yeast. In contrast, the radicicol-dependent recovery from the cyclin A1-induced growth repression was markedly sensitive in the triple-mutant strain as compared with that in the wild-type strain ( Fig. 2b , EC50 app = 2.04 µM in the wild-type strain and 0.06 µM in the mutant strain). Reductions in the growth recovery at high concentrations of PP2 and radicicol are probably brought about by their cytotoxic effects. The relatively narrow concentration ranges of both compounds effective for the growth recovery may result because the concentrations causing their cytotoxicity are close to those required for the growth recovery. Nevertheless, it was an advantage of the strain with an increased permeability that a minimum amount of chemical compounds could be assayed, and therefore we decided to use the triple-mutant yeast for further experiments.
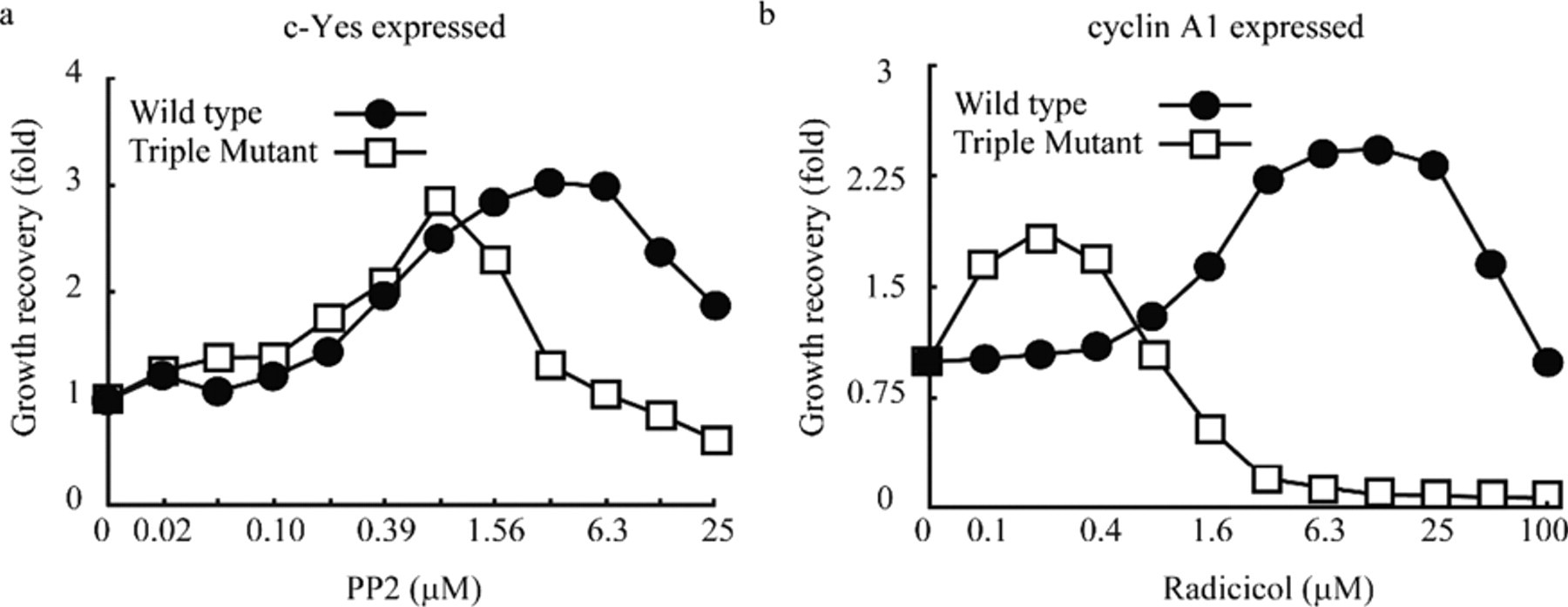
Deletions of genes involved in membrane permeability and efflux pumps provide the mutant yeast cells convenient for surveying small compounds. The triple-mutant yeast strain lacking ERG4, PDR1, and PDR3 and the wild-type yeast strain were transformed with the YEp-type plasmid expressing c-Yes or cyclin A1. The transformed cells expressing (
Next, to validate this screening system, we examined whether the c-Yes-induced retardation of the yeast growth was specifically reversed by known inhibitors for c-Yes. Among 85 protein kinase inhibitors tested, 6 compounds (PP2, PP1 [a PP2 analog], SU6656, radicicol, geldanamycin and purvalanol A) were able to reverse the c-Yes-induced inhibition of the yeast growth. Figure 3 shows the dose-dependent growth recoveries by the 6 positive compounds (PP2 is again included for comparison), among which radicicol was the most potent. At high concentrations, these compounds were cytotoxic, narrowing the effective ranges. All of these compounds could be related to c-Yes. PP2, PP1, and SU6656 are tyrosine kinase inhibitors with broad-spectrum substrate specificities. 20,21 Radicicol and geldanamycin were initially thought to act directly on tyrosine kinase proteins but were later shown to inhibit the enzyme activities via binding with Hsp90. 22,23 As described below, we found that c-Yes is a client protein of yeast Hsc82, a homologue of Hsp90.
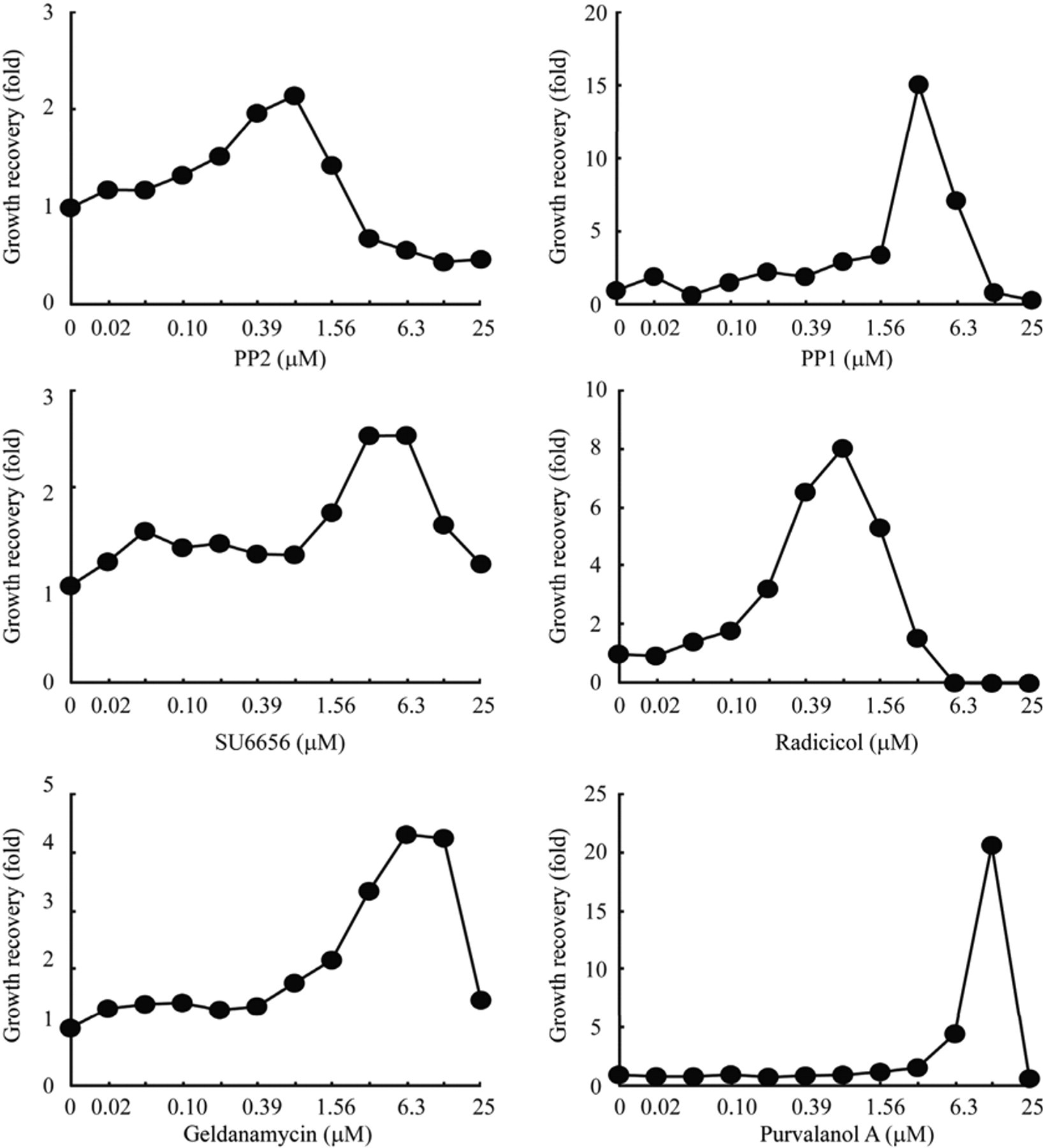
c-Yes inhibitors restore the human c-Yes-induced inhibition of yeast growth in dose-dependent manners. The restoration of the yeast growth was assayed as described in the legend for Figure 2 , except that the YCp-type plasmid was used for the expression of c-Yes. EC50 app values of PP2, PP1, SU6656, radicicol, geldanamycin, and purvalanol were 0.29 µM, 2.26 µM, 1.73 µM, 0.26 µM, 2.20 µM, and 9.24 µM, respectively. EC50 app values were determined as described in Figure 2 .
It was unexpected that purvalanol A reversed the c-Yes-induced growth interference because purvalanol A and B (carboxylated purvalanol A) were thought to be selective inhibitors of cyclin-dependent kinases (CDKs), which are members of the serine/threonine kinase family. 24 More recently, however, purvalanol B was reported to interact with nonreceptor tyrosine kinases, including c-Yes, and receptor tyrosine kinases as well as CDKs. 25 Although we currently have no evidence for an interaction of c-Yes with purvalanol A, it is conceivable that c-Yes expressed in yeast is a target of purvalanol A, thereby leading to the recovery of c-Yes-induced inhibition of the yeast growth. These findings indicate that the actions of the 6 compounds are exerted through c-Yes rather than nonspecific off-targets, thus supporting the validity of the system.
PP2, PP1, SU6656, and purvalanol A inhibit c-Yes activity directly, whereas radicicol and geldanamycin act indirectly
Next, we examined the tyrosine phosphorylation of proteins by c-Yes and the inhibition of phosphorylation by the 6 compounds described above. The tyrosine phosphorylation was detected using an anti-phosphotyrosine antibody. Culture for 2 h after the addition of galactose was sufficient to induce the full expression of c-Yes in yeast ( Fig. 4a ). The expression of c-Yes produced several phosphorylated proteins, and the phosphorylation was reduced by the 6 compounds ( Fig. 4b ). The c-Yes-mediated phosphorylation was severely inhibited by radicicol, geldanamycin, and purvalanol A; moderately inhibited by PP2 and PP1; and slightly inhibited by SU6656. Currently, the phosphorylated proteins responsible for the c-Yes-induced repression of yeast growth are unknown.
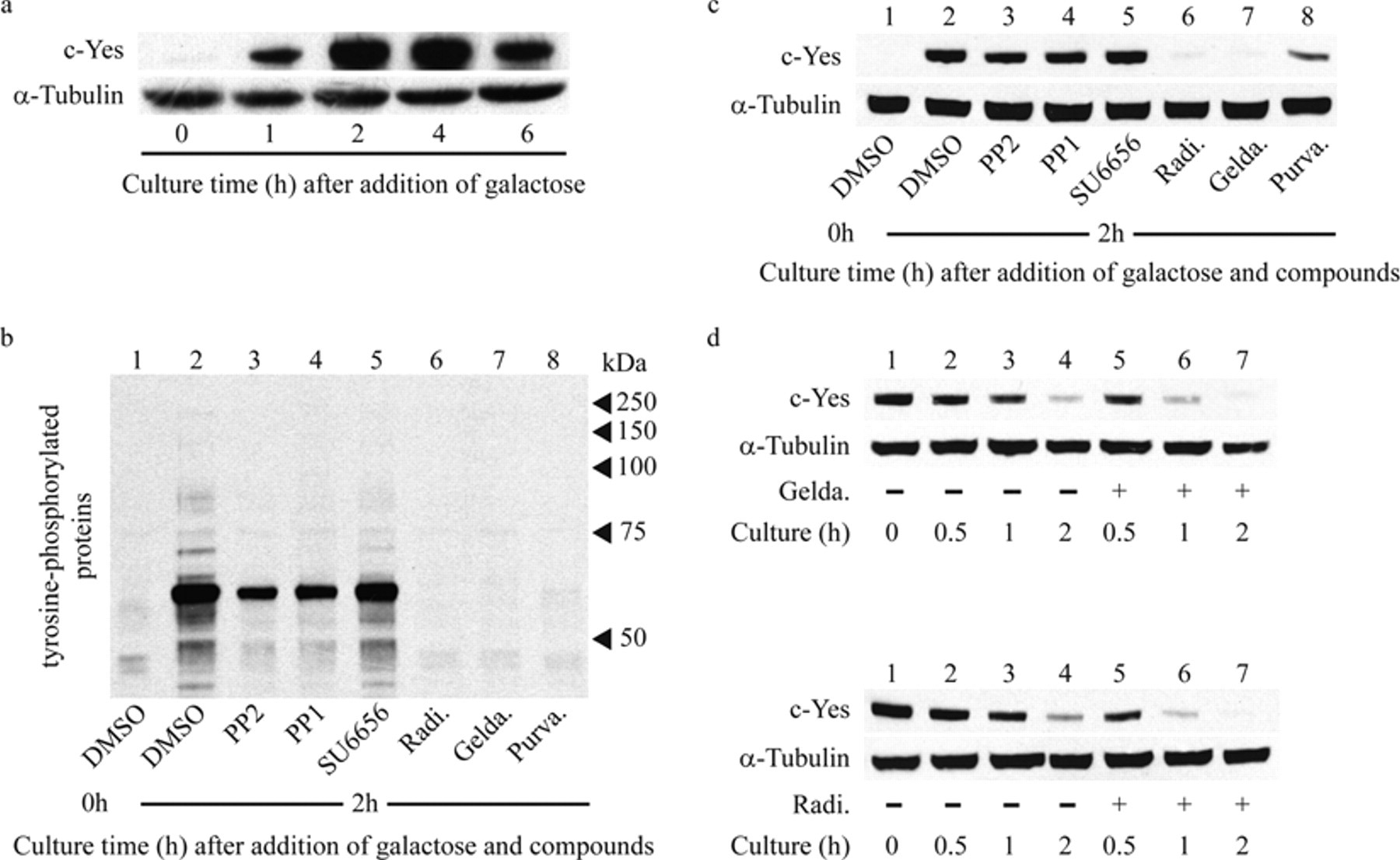
c-Yes inhibitors affect the activity and the stability of c-Yes. (
Next, we examined the effects of the 6 compounds on the expression of c-Yes protein because the growth repression can be caused not only by inhibiting the c-Yes activity but also by decreasing its expression levels. As shown in Figure 4c , PP1, PP2, and SU6656 did not change the c-Yes expression levels, suggesting that the restoration of the yeast growth by these compounds is mediated through direct inhibition of the c-Yes activity. Purvalanol A caused an appreciable reduction in the c-Yes expression level by an unknown mechanism. This reduction may partly contribute to the purvalanol A–derived inhibition of phosphorylation. In contrast, c-Yes was almost undetectable in the presence of radicicol or geldanamycin. The mRNA levels of c-Yes remained unchanged upon treatment with radicicol or geldanamycin (data not shown), excluding the possibilities that these compounds reduced the transcription of the c-Yes gene and/or the stability of its mRNA. Next, we examined the effects of these compounds on the stability of c-Yes protein in the yeast cells. After inhibition of the galactose-induced transcription of the c-Yes gene by the addition of glucose, the decay rates of c-Yes in cells cultured with and without geldanamycin or radicicol were compared. As shown in Figure 4d , both compounds accelerated the disappearance of c-Yes, indicating that these compounds shortened the lifetime of c-Yes in the cells. These results make it likely that the inhibition of phosphorylation by radicicol and geldanamycin was not caused by the inhibition of c-Yes activity but by the rapid degradation of c-Yes.
Src family members, including c-Yes, associate with the intracellular surface of the plasma membrane through the myristoylation of their N-terminal glycine residues. An interaction with Hsp90 is required for the transit of these members to the plasma membrane. 17 Budding yeast has 2 genes, HSC82 and HSP82, that are homologous to vertebrate Hsp90. 26 Hsc82 is expressed abundantly at normal growth temperatures (25-30 °C), whereas Hsp82 is heat inducible. c-Yes may interact with these yeast chaperones, and the accelerated disappearance of c-Yes by radicicol and geldanamycin may be derived from the compound-induced inactivation of the chaperones. To examine the interactions of human c-Yes with the yeast Hsp90 homologues, we prepared 2 types of yeast expressing HA-tagged protein in which the endogenous HSC82 and HSP82 were replaced with HSC82-HA and HSP82-HA, respectively. These yeasts were engineered to express V5-tagged c-Yes in the galactose medium. After culture for 2 h in the inducing media, the cell extracts were treated with an anti-V5 antibody. Hsc82, Hsp82, and c-Yes in the immunoprecipitates were detected by Western blotting. Hsc82 was a major component that coimmunoprecipitated with c-Yes, but a small amount of Hsp82 was also detected in the immunoprecipitates ( Fig. 5a ). Neither the tagged Hsc82 and Hsp82 nor c-Yes was detected in the immunoprecipitates after culture in the glucose medium (not shown). These results indicate that c-Yes binds predominantly with Hsc82. Then we examined whether radicicol and geldanamycin inhibited the interaction between c-Yes and Hsc82. The addition of either compound to the cells after culture for the induction of c-Yes markedly reduced Hsc82 in the immunoprecipitates ( Fig. 5b ). Taken together, we conclude that binding of radicicol and geldanamycin with yeast Hsc82 disrupts the complex of c-Yes and Hsc82 or interferes with formation of this complex, thereby leading to rapid degradation of c-Yes. Currently, it remains unknown whether the degradation is mediated through the proteasome pathway or other pathways.
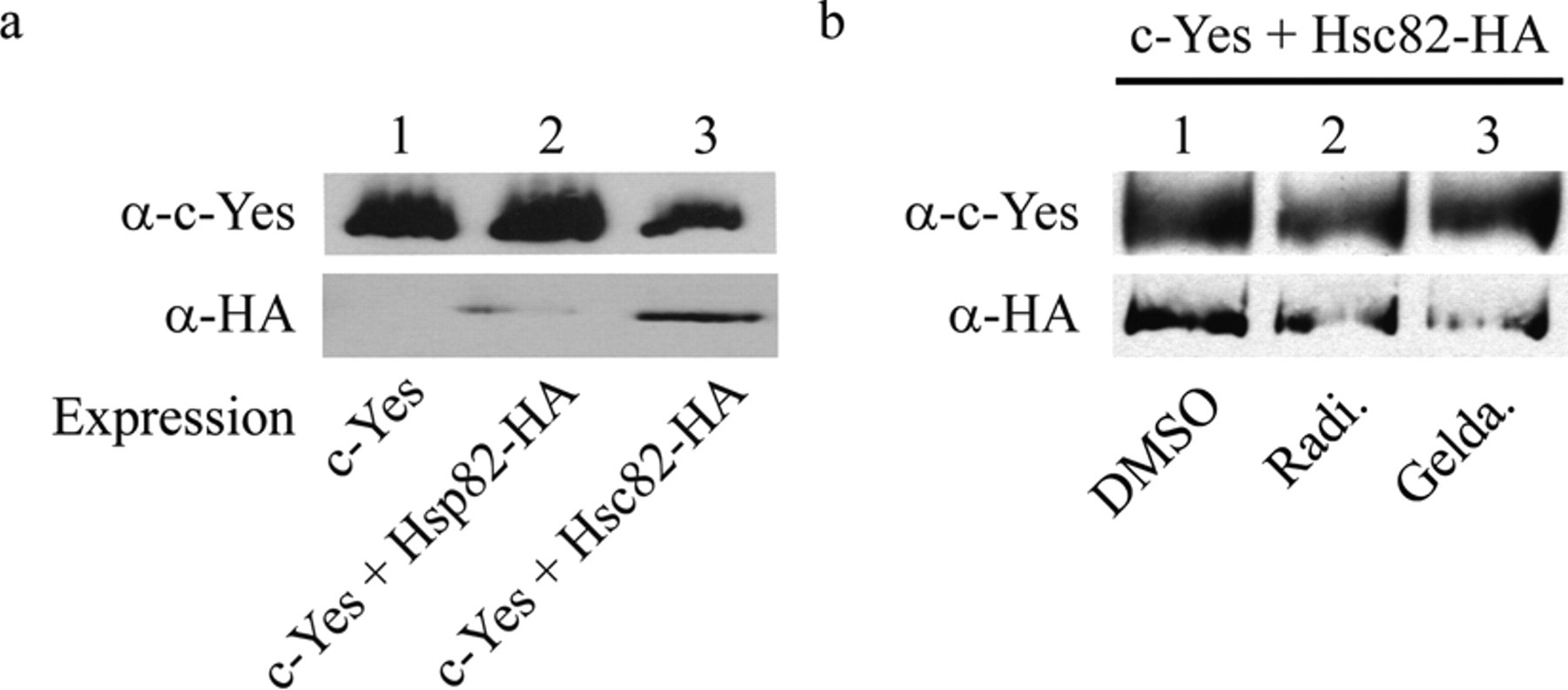
Human c-Yes interacts with the yeast chaperone Hsc82 (
Discussion
In this article, we have reported experimental results that demonstrate the usefulness of a yeast-based screening system for surveying targets that may be druggable. To validate the screening system, c-Yes, a tyrosine kinase, was used as a representative of the human protein kinases that were found to repress the growth of yeast. The examination of 85 known inhibitors for protein kinases identified 6 compounds that were able to restore the c-Yes-induced repression of the yeast growth. The restorative actions of the 6 compounds were ascribable to the inhibition of c-Yes activity or the destruction of c-Yes protein. This high specificity confirms the validity of the screening system, at least for protein kinases.
The first attempt at a yeast-based screening system for drug discovery was made using 38 heterologous cDNAs, and 30% of these cDNAs showed inhibitory effects on yeast growth. 2 This value is much higher than the 5.6% described in this article. This difference may arise because cDNAs encoding proteins involved in signal transduction and cell growth were mainly used in the first attempt, whereas human cDNAs were used randomly in the present experiments. Indeed, when we focused on protein kinases, the percentage of cDNAs with inhibitory effects on the yeast growth increased to an average value of 22.6% with the highest value of 35.5% for tyrosine kinases.
Hsp90 is a molecular chaperone expressed at high levels in vertebrates and supports the proper folding of its client proteins. The Hsp90 client proteins include a wide variety of signal transmission-related proteins such as protein kinases. Hsp90 also binds with mutated and oncogenic client proteins, thereby ensuring their functions and resulting in the promotion of growth of transformed cells. 22 Furthermore, the identification of Hsp90 at the cell surface has led to the possibility that this protein may play a unique role in tumor metastasis. 27 Consequently, inhibitors of Hsp90 may be effective as anticancer drugs, and extensive studies have been devoted to the identification of druggable compounds, including derivatives of radicicol and geldanamycin. 17,23,28,29 In the present study, we have shown that a yeast-based screening system expressing human c-Yes, one of the Hsp90 client proteins, selected 2 types of compounds. One type bound directly to the client protein, whereas the other type bound to yeast Hsc82 to release the client protein, thereby resulting in its rapid degradation. By examining the expression of the client protein (c-Yes in this study) in yeast, one can easily determine whether the inhibition of the client protein activity by the chemical compounds is derived from direct binding to the client protein or inactivation of its chaperone. Therefore, the yeast system is useful for screening compounds that abolish the function of Hsp90 when its client proteins repress the growth of yeast.
Vertebrates contain 2 HSP90 genes, HSP90α and HSP90β. Hsp90β is constitutively expressed at a higher level than Hsp90α, whereas Hsp90α is stress inducible and overexpressed in many cancerous cells. The replacement of both yeast chaperone isoforms with either of the human HSP90s may provide a system with higher efficacy for screening candidates with anticancer actions mediated through Hsp90. Although deletion of both yeast Hsc82 and Hsp82 causes cell death, expression of either of the human HSP90s results in cell survival. Interestingly, expression of HSP90β in the mutant yeast lacking both yeast chaperones rendered the yeast cells highly sensitive to radicicol. 30 In this case, the glucocorticoid receptor, v-src tyrosine kinase, and extracellular signal-regulated kinase-5 were used as client proteins.
Application of the yeast system can be expanded to a mutant yeast that lacks the yeast proteins of interest. In fact, we have found that the human C18orf26 protein with an uncharacterized function inhibited the growth of a mutant yeast lacking Mad2 protein, a component of the mitotic checkpoint complex that monitors the attachment of spindles to kinetochores. The C18orf26 protein had no effects on the growth of the wild-type yeast. The chemical compounds identified by detecting the growth restoration of the mutant yeast induced apoptotic cell death of human cancer cells in a C180rf26 protein-dependent fashion (Kunoh et al., unpublished data, manuscript in preparation). Therefore, it appears that the yeast-based screening strategy has potentials that are almost limitless.
Footnotes
Acknowledgements
We are grateful to M. Yoshida, Y. Yashiroda, Y. Ohya, S. Nogami, M. Takagi, K. Shin-ya, T. Natsume, and M. Hasegawa for helpful discussions and advice. We also thank M. Yoshida, K. Komura, K. Tsuji, S. Hasuike, Y. Fujiishi, Y. Hori, Y. Kunitomo, T. Murakami, Y. Inagaki, and M. Tsuchida for technical support. We are grateful to A. Baines for anti-α-tubulin antibody.
This work was partially supported by a grant from the New Energy and Industrial Technology Development Organization (NEDO) and a grant from Frontier Pharma.
