Abstract
The focus of biomarker studies is shifting toward deciphering patterns of biomolecules as they provide a more comprehensive depiction of disease than individual biomarkers. Multiplexing technologies are crucial in deciphering such patterns, but it is essential that they are validated for reproducibility and precision to ensure accurate protein identification. Here the authors examine such properties in Cytokine Bead Array (CBA) and Luminex kits and compare concentration measurements to those obtained using enzyme-linked immunosorbent assay (ELISA). Luminex kits were found to be highly reproducible and reliable; however, CBA kits were not due to aberrant standards. Absolute cytokine concentrations were dependent on the detection kit, but correlations with ELISA were good for all technologies.
Introduction
T
In view of this, focus is beginning to shift toward the identification of patterns of biomolecules for disease identification and characterization. Each disease possesses a unique molecular fingerprint that could be used diagnostically to differentiate it from diseases with closely related phenotypes. Examination of a molecular pattern should provide a more comprehensive indication of pathophysiological conditions and a better understanding of the molecular processes underlying diseases than individual biomarkers. 3 This novel concept, although still in its infancy, is being applied to several complex diseases, including cancer diagnosis, 4 Alzheimer’s disease, 5 polyarticular juvenile idiopathic arthritis, 6 psychiatric disorders, 7 Sjögren’s syndrome, 8 and chronic obstructive pulmonary disease (COPD). 9,10
Multiplexing technologies are crucial to the extremely complex task of deciphering these disease-specific biomolecular patterns as they provide the opportunities for a more holistic approach, not previously possible using conventional technologies such as the enzyme-linked immunosorbent assay (ELISA), to be taken. However, prior to being employed in routine laboratory practice, it is essential that these technologies are investigated for reproducibility, precision, and accuracy. It is also vital that cytokine measurements obtained using multiplexing technologies are comparable to those obtained using single-point measurements (e.g., ELISA). Two commonly used multiplexing technologies are the Cytokine Bead Array (CBA) 11 and Luminex 12 systems, which are bead-based technologies based on the principles behind ELISAs. As with sandwich ELISAs, bead-based multiplexing kits employ immobilized antibodies to capture soluble ligands from within samples. The captured ligand is subsequently identified by a secondary antibody-based detection system. Although the general principles behind ELISAs and bead-based multiparametric technologies are the same, differences occur within the specific methodologies. First, bead-based assays employ fluorescence systems to detect binding of the secondary antibody to the soluble ligand, whereas ELISAs generally use enzymatic amplification of a colorimetric substrate. Second, for bead-based experiments, the capture antibody is immobilized onto spherical beads in suspension rather than immobilized on the flat surface used in ELISAs. It is the fluorescent intensities of these spherical beads that provide the multiplexing capacity of such technologies.
Although ELISAs are a well-established technique, bead-based technologies are relatively new and require thorough validation. Several studies have previously been undertaken to compare CBA kits with ELISAs, 13,14 Luminex kits with ELISAs, 15,16 and Luminex kits from different vendors. 17 However, there has been no experimental comparison between CBA kits and Luminex kits. Here we examine the reproducibility and accuracy of data obtained using CBA and Luminex kits and compare concentration measurements to those obtained using sandwich ELISA. One disease that would benefit from the aforementioned approach to diagnostics is COPD. 9 Although the ideal target for such procedures would eventually be a blood sample, there are problems associated with analysis of the blood proteome 18 that could distort results obtained using commercial cytokine analysis kits. To avoid these potential difficulties, therefore, alternative clinical samples will be used during these validatory investigations. The lung is the representative system for respiratory disorders such as COPD, and clinical samples traditionally examined include induced sputum, bronchoalveolar lavage, lung tissue, and, more recently, exhaled breath condensate. 19 All of these sample types could potentially be analyzed for patterns of biomarkers, but the availability of lung tissue samples has led us to examine these in this study.
Materials and Methods
Sample preparation
The Nottingham Local Research Ethics Committee approved the study protocol, and written informed consent was obtained from the 8 lung cancer patients undergoing surgery. Lung tissue specimens were taken distally from the tumor site and immediately placed into ice-cold RPMI-1640 media. The samples were then finely chopped, vortexed, filtered, and centrifuged. The resultant supernatant was aspirated and stored at –20°C.
Multiplex cytokine detection kits
Three different premixed CBA kits (BD Biosciences Pharmingen, San Diego, CA)—Th1/Th2, Inflammation, and Chemokine—were examined according to manufacturer’s instructions, allowing detection of 14 different cytokines, 3 of which (interleukin-8 [IL-8], interleukin-10 [IL-10], and tumor necrosis factor [TNF]) were duplicated between kits. Flow cytometry was performed using an EPICS Altra Flow Cytometer (Beckman Coulter, Fullerton, CA) or a BD LSR II System (BD Biosciences). Data were analyzed using the BD Cytometric Bead Array software (BD Biosciences). Luminex bead kits (Biosource, Carlsbad, CA) for IL-8, IL-10, and TNF, corresponding to the duplicated cytokines contained within the CBA kits, were examined according to manufacturer’s instructions. Data were acquired on a Bio-Plex 200 system and analyzed with associated software (Bio-Rad, Hercules, CA). All multiplex assays were performed in duplicate. Standard curves for each cytokine were generated using the premixed lyophilized standards provided in the kits, and the cytokine concentrations in samples were determined from the appropriate standard curve. This workflow of the comparisons undertaken is summarized in
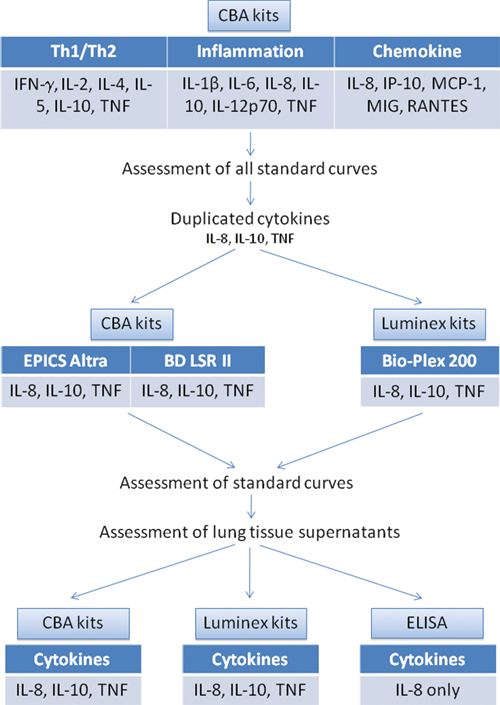
The workflow of the comparisons between multiplex technologies undertaken in this study.
ELISA
IL-8 ELISA was undertaken using commercially available matched antibody pairs according to the manufacturer’s instructions (MAB208 and BAF208; R&D Systems, Minneapolis, MN). Briefly, the wells of ELISA plates were coated overnight with 4 μg/mL capture antibody (MAB208), 100 μL sample was added for 2 h, and then biotinylated detection antibody (BAF208) was added at 400 ng/mL for 2 h. The reaction was developed using streptavidin–horseradish peroxidase (HRP) and tetramethylbenzidine (TMB) substrate, stopped with 1 M HCl, and read at 450 nm. Concentrations of IL-8 in the samples were determined from a standard curve generated with recombinant IL-8 (Abcam, Cambridge, UK).
Results and Discussion
The accuracy and reproducibility of cytokine measurements made using commercial multiplexing kits is highly dependent on the availability of high-quality standard curves. In the course of this study, a variety of problems were encountered during the generation of standard curves contained within the CBA kits (
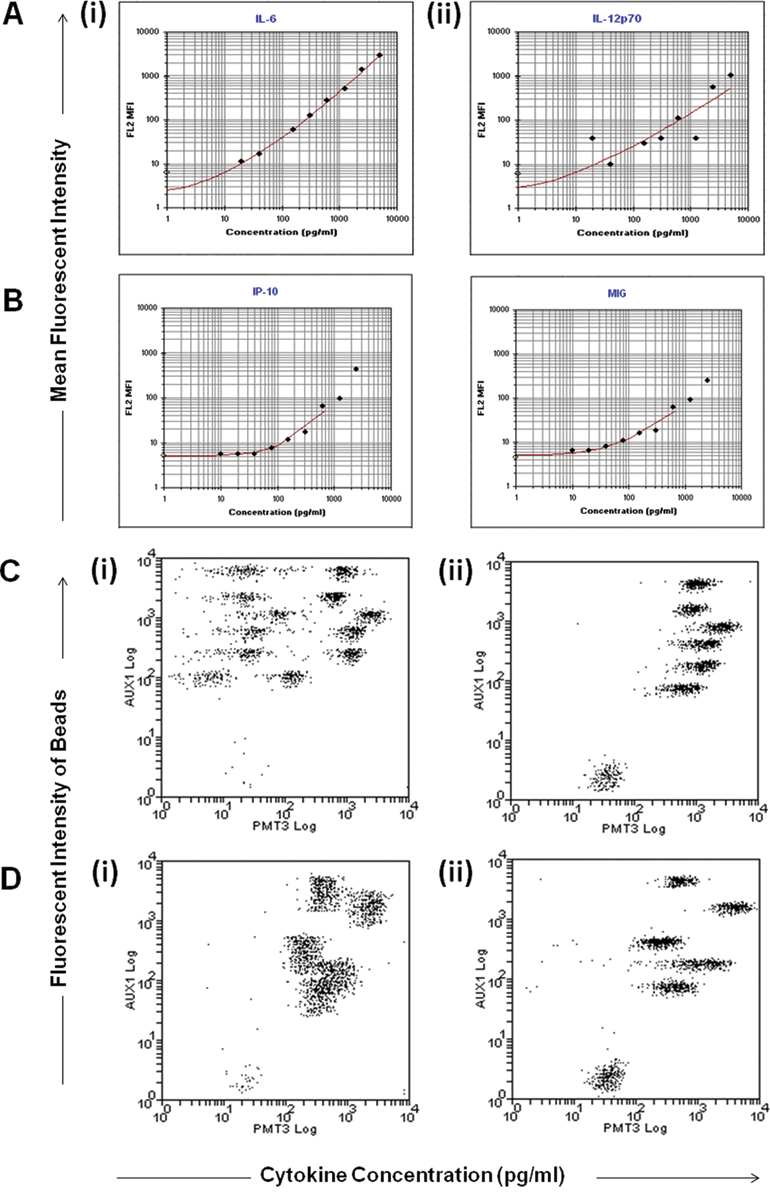
Problems encountered during production of the standard curves contained within cytokine bead arrays: (
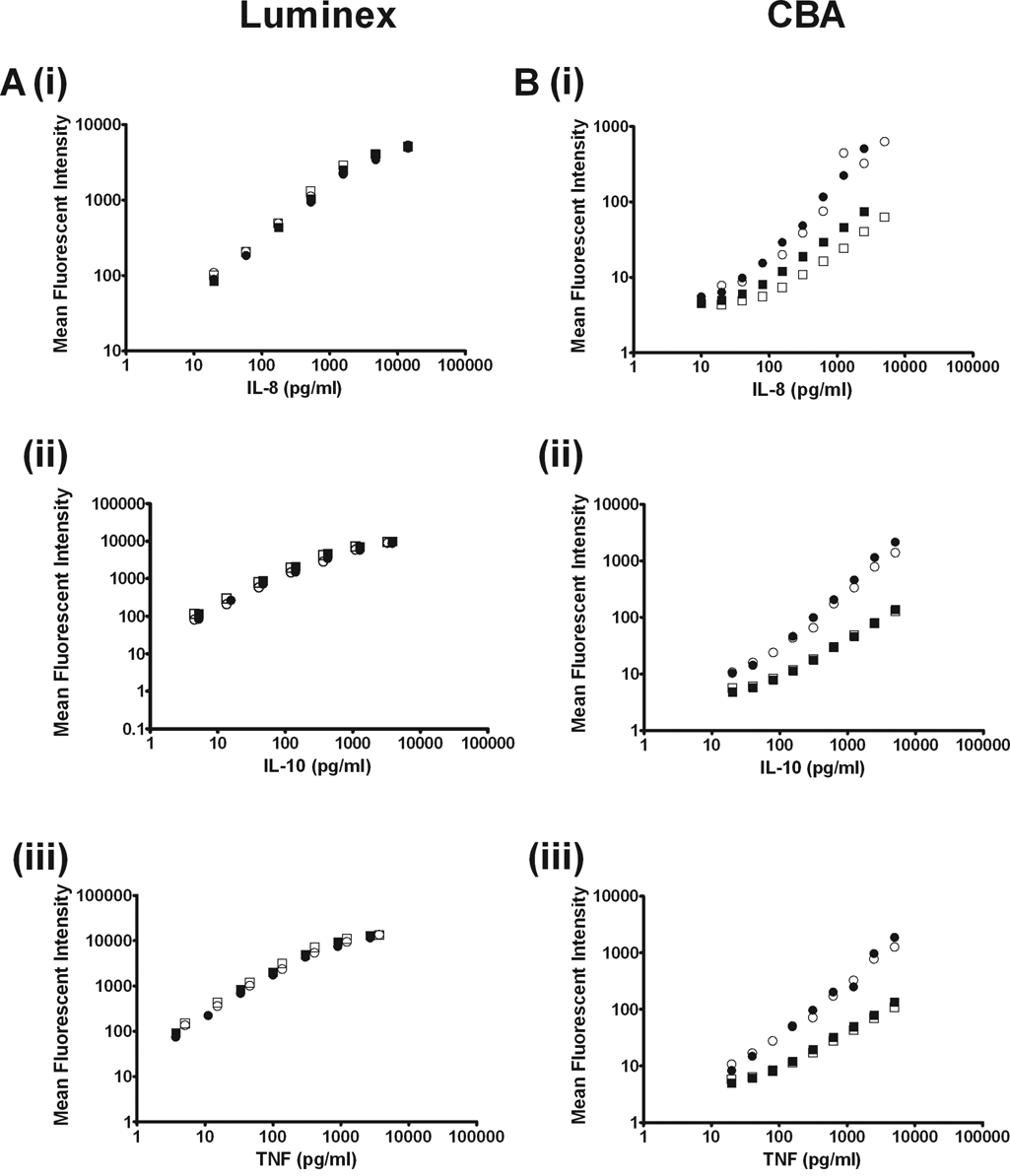
Interkit and intrakit variability of (i) interleukin-8 (IL-8), (ii) interleukin-10 (IL-10), and (iii) tumor necrosis factor (TNF) standard curves generated using (
Such problems will affect the reliability of downstream cytokine measurements made using CBA kits. The majority of studies involving premixed CBA kits use only 1 kit for cytokine measurements, negating the need for validation of interkit variability. However, in cases where maximal characterization of the cytokine profile of a sample is required, it may be necessary to employ multiple kits in conjunction. In such cases, for example, when trying to identify disease-specific biomolecular patterns, it is imperative that cytokine levels determined by individual kits are comparable. In addition, if kits are to be used to characterize patient samples, it is vital that data are consistent on different days. Thus, levels of intrakit and interkit variability in standard curves must be minimal and can be assessed by examination of the cytokines duplicated between kits.
During this study, intrakit and interkit variability was found to be high for the IL-8 standard curves produced using CBA kits (
One advantage of the CBA kits over the Luminex system is the possibility of collecting data on general flow cytometers already installed in many laboratories
22
rather than using a specialized detection system. The manufacturer of the CBA kits recommends the use of a BD flow cytometer for data collection. To ensure that the problems encountered during the generation of the CBA standard curves were not due to the flow cytometer used, we ran samples in parallel on a BD LSR II System flow cytometer and an EPICS Altra flow cytometer. The variations observed in inter- and intrakit variation occurred regardless of the machine used (
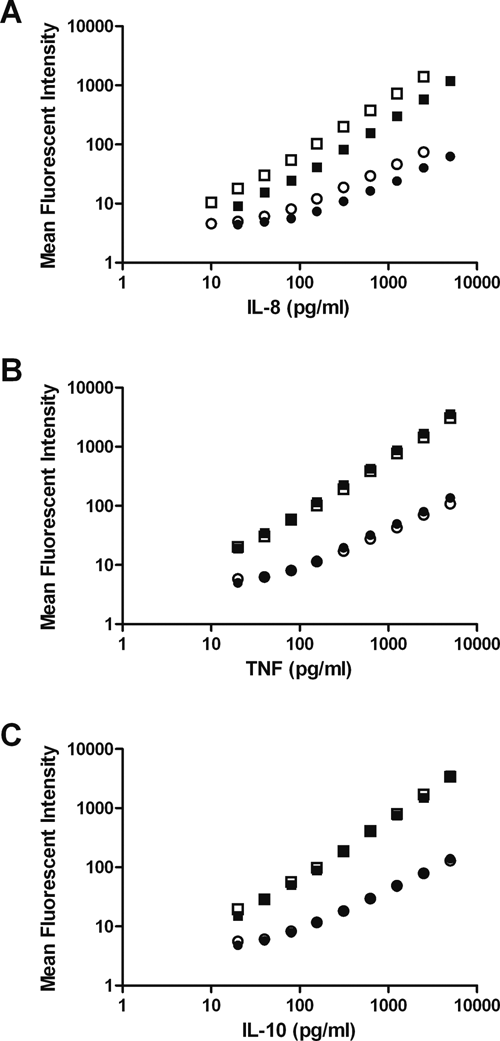
Interkit variation in standard curves generated for (
In addition to investigating intra- and interkit reliability, this study also examined intertechnology reliability. Concentrations of IL-8, IL-10, and TNF in lung biopsy supernatants were measured using the CBA and Luminex kits. Of the 3 cytokines, only IL-8 was detectable in the samples. IL-10 and TNF could not be detected (data not shown) due either to the absence of the cytokines in the samples or to the cytokine concentration falling below the detection limit of the CBA (IL-10: 2.8 pg/mL; TNF: 2.8 pg/mL) and Luminex (IL-10: <5 pg/mL; TNF: <10 pg/mL) technologies. IL-8 was subsequently measured using ELISA for comparison.
The level of IL-8 measured within any 1 sample varied greatly depending on the technique employed in making the measurement, with data obtained for 1 patient’s sample ranging from 288 to 914 pg/mL (
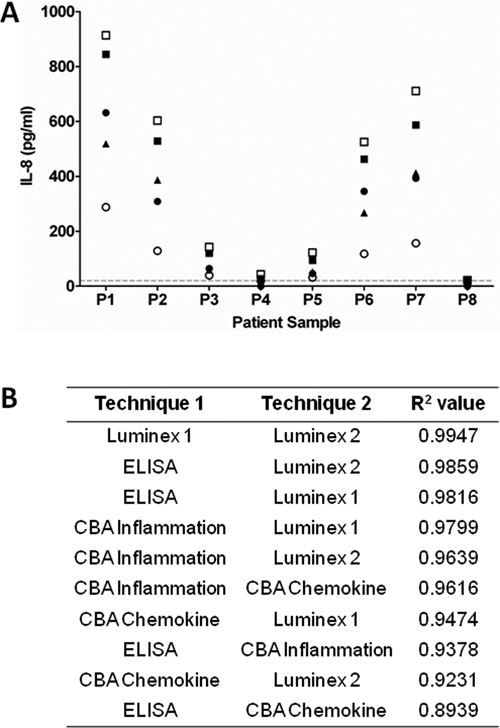
Variations in interleukin-8 (IL-8) measurements in lung tissue supernatants as detected by (▲) enzyme-linked immunosorbent assay (ELISA); (●) Cytokine Bead Array (CBA) Inflammation kit; (○) CBA Chemokine kit; (■) Luminex kit 1; and (□) Luminex kit 2. (
To conclude, cytokine levels determined within a sample are highly dependent on the technology chosen for analysis. When deciding upon the technique to be employed for identification of patterns of biomolecules in biomarker studies, the data obtained during this and previous studies 14 should be considered. Duplication of IL-8, IL-10, and TNF enabled validation of measurements made for these cytokines within the different CBA kits. The problems encountered during CBA standard curve generation were not, however, restricted to these cytokines and affected multiple cytokines, including IL-12p70, IP-10, and MIG. These difficulties, coupled with the variation observed in IL-8 measurements, raise questions about the reliability of concentrations obtained for other cytokines that are not duplicated. In contrast, the Luminex technology was found to be highly reproducible. Despite the aforementioned difficulties, the high correlation between concentrations obtained using the different technologies indicates that biomolecular patterns (i.e., our biomarker panel) should be identifiable using any of the technologies investigated. It is critical, however, that once a technology has been selected, it is used consistently if comparisons are to be made between different data sets. Furthermore, this study highlights the importance of using a control sample of known concentrations to normalize between runs undertaken on different days for quality control purposes.
Footnotes
Acknowledgements
We are grateful to Dr. Adrian Robins (UoN) and Julie Swales (UoN) for help with flow cytometry and to Dr. Simon Hall (GSK), Dr. Ian Spendlove (UoN), and Dr. Richard Bradley (UoN) for help with Luminex. We are grateful to Nick Smith and Niki Wason (BD Biosciences) for helpful discussion.
JLR and RM are funded by the RCUK Basic Technology Programme. RAU is funded by the Jones’s 1986 Charitable Trust.
LF is supported by the Nottingham Respiratory Biomedical Research Unit.
