Abstract
The authors recently reported the development and application of multiple LanthaScreen® cellular assays to interrogate specific steps within the PI3K/Akt pathway. The importance of this signaling cascade in regulating fundamental aspects of cell growth and survival, as well as in the progression of cancer, underscores the need for portable cell-based assays for compound profiling in multiple disease-relevant cell backgrounds. To meet this need, the authors have now expanded their LanthaScreen® assay platform across a variety of cell types using a gene delivery technology known as BacMam. Here, they have demonstrated the successful detection of Akt-dependent phosphorylation of PRAS40 at Thr246 in 10 different cell lines harboring mutations known to activate the PI3K/Akt pathway. In addition, they generated inhibitory profiles of 17 known pathway inhibitors in these same cells to validate the approach of using the BacMam-enabled LanthaScreen® cellular assay format to rapidly profile compounds in disease-relevant cell types. Importantly, their results provide a broad illustration of how the genetic alterations that affect PI3K/Akt signaling can also influence the inhibitory profile of a given compound.
Introduction
N
We recently reported the development and validation of a set of cell-based, time-resolved Förster resonance energy transfer (TR-FRET) LanthaScreen® assays for the interrogation of the PI3K/Akt pathway. 3 This assay platform has proven to be broadly applicable for the cellular analysis of other signal transduction cascades as well, such as NF-κB, JAK/STAT, and MAPK, 4-6 and features cell lines that express green fluorescent protein (GFP) fusions of a kinase substrate and terbium (Tb)–labeled phospho-specific antibodies for the substrate. The cellular phosphorylation event is initiated by stimulating cells with agonist and is then detected upon lysis of the cells and addition of Tb-labeled antibody (in the same step). When the antibody binds to the phosphorylated GFP substrate, FRET between Tb and GFP can be measured. If signaling is triggered in the presence of an inhibitor, phosphorylation is reduced as indicated by a decrease in the TR-FRET signal.
In the past, we have used clonal cell lines that stably express a GFP fusion protein of interest. Despite the reliable and robust nature of a stable clone, their development can consume both time and resources. Moreover, the typical workhorse cell lines from which these clones are traditionally derived (e.g., HEK, CHO, U-2 OS) often lack the disease relevance and may not be the ideal host cell background for primary screening or secondary analysis.
To make the LanthaScreen® cellular assay more readily “portable” into a wide range of different cell backgrounds, we have implemented a transient gene delivery method that employs baculovirus vector DNA modified with a cytomegalovirus (CMV) promoter to drive the expression of protein substrates in mammalian cells. This technology, known as BacMam, provides several advantages over other commonly used methods for heterologous gene expression. 7 Some of these benefits are high transduction rates, variable protein expression levels that can be controlled by viral dose, low cytotoxicity to host cells, safety in production and handling (Biosafety Level 1), compatibility with primary and stem cells, and the ease and convenience of use.
Previous studies with BacMam have used this method mainly as a means to overexpress a recombinant protein (e.g., an ion channel or G-protein-coupled receptor [GPCR]) and subsequently evaluate that target. In contrast, our approach differs in that we are using BacMam to deliver a GFP substrate to support the assay readout. Here, the substrate we have selected as a model to interrogate PI3K/Akt pathway signaling is the proline-rich Akt substrate of 40 kDa (abbreviated as PRAS40). The phosphorylation of PRAS40 at residue Thr246 by Akt is a strong indicator of both PI3K-and Akt-dependent kinase activity in the cell. This site is regulated primarily by serum and growth factors (e.g., insulin and insulin-like growth factor, IGF-1), 8 but mutations in these upstream kinases can also result in constitutive PRAS40 phosphorylation.
To demonstrate the application of the BacMam-enabled LanthaScreen® cellular assay format, we first assessed the transduction efficiency of BacMam virus that drives the expression of GFP-PRAS40 by fluorescence-activated cell sorting (FACS) analysis in a panel of 10 different tumor cell lines, including those with normal/wild-type signaling and others exhibiting commonly observed pathway mutations. We then profiled a set of known PI3K and Akt inhibitors using the LanthaScreen® PRAS40 [T246] cellular assay readout in these same cell backgrounds following BacMam transduction. Interestingly, we observed cell type–dependent differences in the activity of many of these compounds, most significantly evident in PTEN mutant cell lines. Taken together, our results validate the approach of combining BacMam and LanthaScreen® technologies to rapidly generate compound profiles in disease-relevant cellular backgrounds in a high-throughput screening (HTS)–compatible manner.
Materials and Methods
General
Unless stated otherwise, all reagents for molecular biology, cell culture, and LanthaScreen® assays were from Invitrogen, part of Life Technologies (Carlsbad, CA). All small-molecule kinase inhibitors were from Calbiochem® (San Diego, CA), Cayman Chemical (Ann Arbor, MI), LC Laboratories® (Woburn, MA), or Axon Medchem BV (Groningen, The Netherlands). Stock solutions of compounds were dissolved in DMSO (≥10 mM) and stored at −20 °C until the day of the assay.
Plasmid construction and generation of BacMam virus
The expression vector for GFP-PRAS40 was generated via Gateway® cloning technology according to standard protocols. The entry clone IOH27857 for full-length AKT1S1 (PRAS40) was recombined via LR reaction with a pDEST8 vector that was modified to express CMV-driven N-terminal GFP fusion proteins in mammalian cells. This construct was transformed into DH10Bac™ competent cells for the production of recombinant bacmid DNA. PCR-qualified bacmid DNA was then transfected into SF9 insect cells to prepare BacMam virus stocks according to the Invitrogen/Life Technologies Bac-to-Bac® expression system manual.
Cell lines and cell culture
Cancer cell lines (BT474, HepG2, LNCaP, MCF7, MDA-MB-231, MDA-MB-468, PC-3, SKBr3, and T47D) evaluated in this study were purchased from ATCC (Manassas, VA). HEK 293E cells were from Invitrogen (Carlsbad, CA). All cell lines were grown as an adherent monolayer and maintained in 37 °C/5% CO2 in a humidified incubator with complete growth media (Dulbecco’s modified Eagle’s medium [DMEM] or McCoy’s 5A medium) supplemented with GlutaMAX™, 10% dialyzed fetal bovine serum (FBS), 100 U/mL penicillin, 100 µg/mL streptomycin, 25 mM HEPES, and 0.1 mM nonessential amino acids (NEAA).
Transduction of mammalian cell lines with BacMam reagent
Near-confluent cell cultures were harvested on the day prior to the transduction. Following trypsinization, cells were resuspended to a density of 2-3 × 105 cells/mL in complete growth media containing 10% BacMam virus (v/v). In most instances, trichostatin A (0.5 µM) from Sigma Aldrich (St. Louis, MO) was added to enhance the expression of GFP-PRAS40. For a representative experiment, 10 mL of this cell suspension was added to a 10-cm culture dish, and the transduction was carried out overnight in a humidified incubator set to 37 °C and 5% CO2. The following day, the transduction efficiency was visualized under a fluorescence microscope or quantified by flow cytometry using a FACSDiVa instrument from BD Biosciences (San Jose, CA). Alternatively, cells were cryopreserved 24 h posttransduction using Recovery™ cell culture freezing medium for assay at a later date. Analysis of transduced cells by Western blot and fluorescence microscopy is provided as supplementary data (
LanthaScreen® cellular assay protocol
The generic protocol for performing the LanthaScreen® cellular assay has been described previously. 3,5,6 Briefly, cells were harvested following overnight transduction and plated at a density of 10,000 to 20,000 cells per well into white 384-well flat-bottom, tissue culture–treated microplates (Corning #3570). Cells were then incubated overnight in Opti-MEM® reduced serum medium supplemented with charcoal/dextran-stripped FBS (0.5%), NEAA (0.1 mM), and sodium pyruvate (0.1 mM). The following day (48 h posttransduction), cells were treated with serial dilutions of small-molecule inhibitors for 2 h. The assay media were then removed via aspiration, and cells were lysed with LanthaScreen® cellular assay lysis buffer (20 µL per well) supplemented with Tb-anti-PRAS40 [T246] antibody (2 nM) and both protease and phosphatase inhibitor cocktails. For agonist activation assays (e.g., insulin dose-response in HEK 293E cells), the assay medium instead consisted of low-glucose DMEM plus bovine serum albumin (BSA, 0.1%), and LanthaScreen® experiments were performed as described previously. 3 Following lysis, assay plates were allowed to equilibrate for 2 h at room temperature, and TR-FRET emissions were then measured on a BMG LABTECH PHERAstar (Offenburg, Germany) or PerkinElmer EnVision® (Waltham, MA) fluorescence plate reader using recommended instrument settings for LanthaScreen®. Background-corrected emission ratio data (520 nm/490 nm) were determined by subtracting the ratios from Tb-labeled antibody alone (cell-free wells) from the TR-FRET ratios from each sample. To obtain a response ratio (RR), the background-corrected emission ratios were each divided by the average of background-corrected values from unstimulated/untreated samples. The Z′ factor was calculated as described previously. 3 Compound IC50 values were generated in GraphPad Prism 4 (La Jolla, CA) using the default settings for nonlinear regression analysis and curve fitting the data to the sigmoidal dose-response equation (variable slope) as shown below:
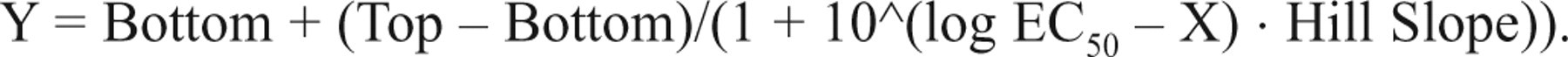
Results/Discussion
Gene delivery via BacMam is an established method for transient expression in mammalian cells and has been reviewed previously in the literature. 7,9 As illustrated schematically in Figure 1A , BacMam can be used to deliver GFP fusion proteins and can easily be incorporated into the LanthaScreen® cellular assay workflow for HTS and assay development. The general protocol begins by treatment of cells with the BacMam reagent (typically as a percentage of virus in growth media) followed by overnight incubation. Transduced cells can then be imaged the next day under a fluorescent microscope to confirm expression of the GFP fusion protein. Cells are then harvested and plated in a 384-well format for testing in the LanthaScreen® functional assay the next day (a total of 48 h posttransduction with BacMam). Alternatively, cells can be cryopreserved at this stage for use at a later date.
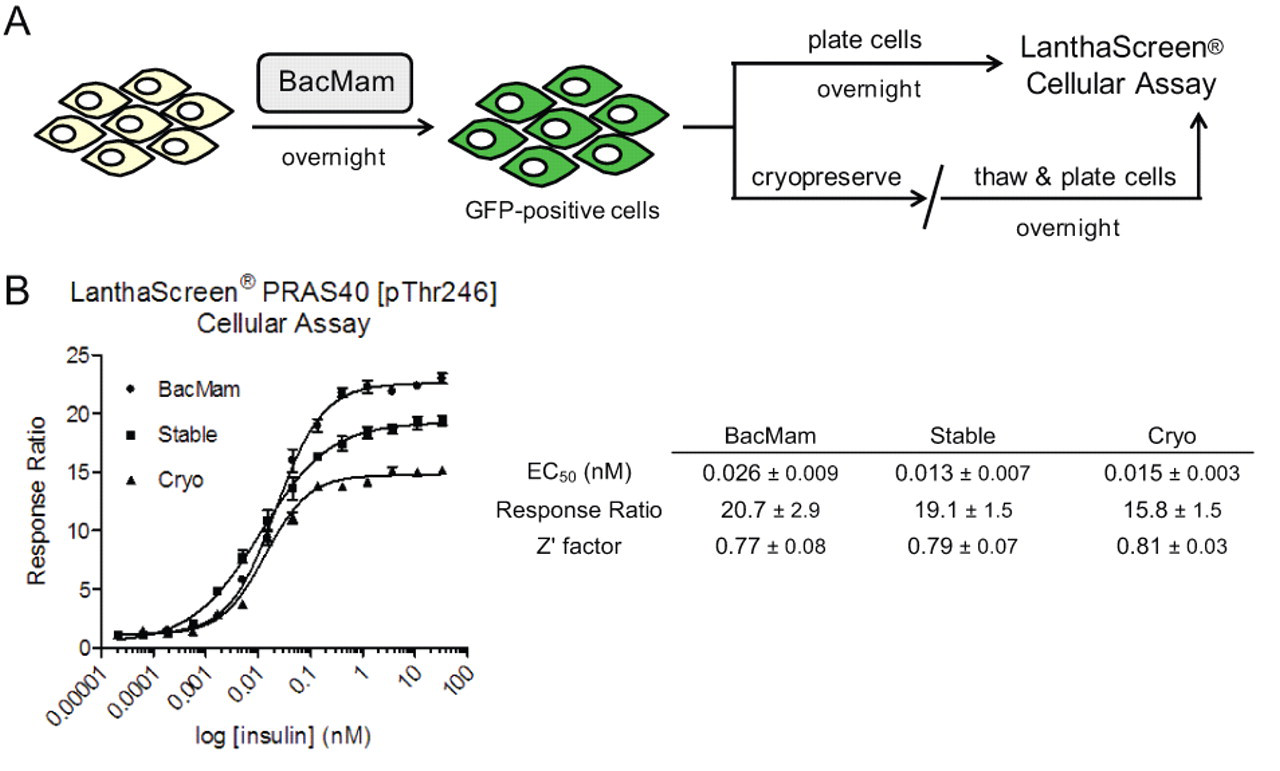
BacMam-mediated delivery of green fluorescent protein (GFP) fusion proteins for LanthaScreen® cellular assays. (
Following the workflow described above, we first wanted to compare the assay performance of cells transiently transduced with BacMam GFP-PRAS40 (including both dividing cells and also cells that were pretransduced and then cryopreserved) to a clonal cell line with the same target stably expressed. To this end, 3 different samples of HEK 293E cells expressing GFP-PRAS40 were serum starved overnight to reduce PI3K signaling and then stimulated with varying concentrations of insulin to activate the pathway. We observed comparable phosphorylation of PRAS40 at Thr246 in all 3 cases as indicated by a dose-dependent increase in TR-FRET signal relative to untreated cells ( Fig. 1B ). Moreover, the pharmacology for stimulation was reproducible (similar EC50s for insulin), the assay windows were good (RR > 10), and the data quality for a cell-based assay was excellent by HTS standards (Z′ factors above 0.7). Last, the fact that cells transduced with BacMam and then cryopreserved can be used without any significant loss in assay performance could greatly facilitate cell-based drug screening activities.
We next compared the transduction efficiency of the BacMam virus across a variety of cell types ( Table 1 ). We selected HEK 293E and HepG2 as cell lines with wild-type PI3K signaling, as well as other tumor types with relevance to the PI3K pathway. These include SKBr3 and BT474 (HER2 amplification), MCF7 and T47D (PIK3CA mutants), MDA-MB-231 (Ras mutation), and LNCaP, MDA-MB-468, and PC3 (PTEN deletion). Following a standard protocol (see Materials and Methods), cells were transduced overnight with BacMam virus, and the percentage of GFP-expressing cells was determined by flow cytometry the next day ( Table 1 ). Efficiencies of greater than 75% were achieved reproducibly for all BacMam transductions, thus emphasizing the impact of this technology as a robust method for the delivery of GFP fusion proteins. Of note, inhibitors of histone deacetylase (HDAC), such as trichostatin A (TSA), have been shown to enhance protein expression levels in cells treated with BacMam. 10 For all cell backgrounds except HepG2 and LNCaP, we included TSA during the transduction step.
Cell Lines and Pathway Mutations
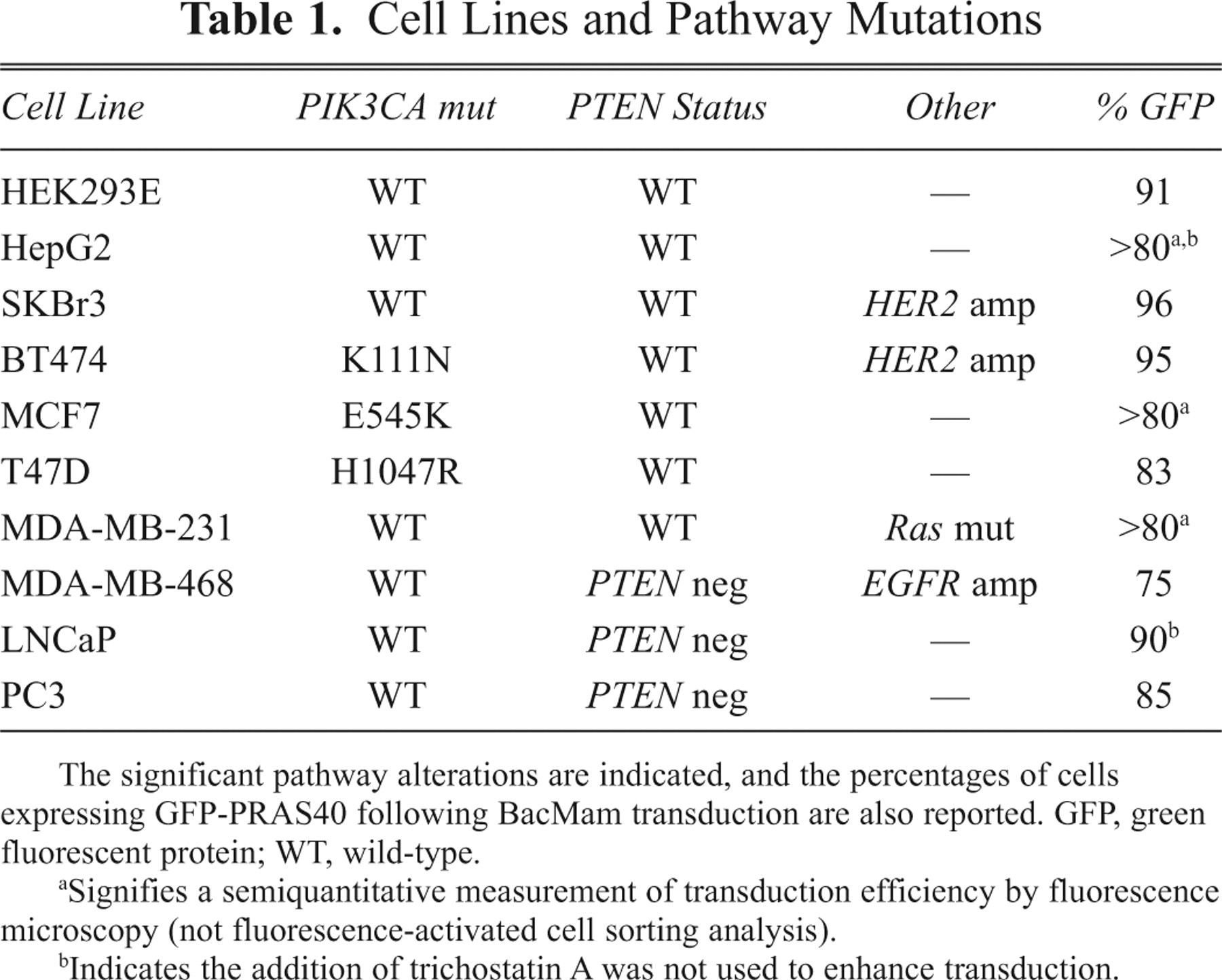
The significant pathway alterations are indicated, and the percentages of cells expressing GFP-PRAS40 following BacMam transduction are also reported. GFP, green fluorescent protein; WT, wild-type.
Signifies a semiquantitative measurement of transduction efficiency by fluorescence microscopy (not fluorescence-activated cell sorting analysis).
Indicates the addition of trichostatin A was not used to enhance transduction.
To demonstrate the utility of BacMam and LanthaScreen® cellular assay technologies together for pathway analysis, we then went on to profile a panel of 17 known inhibitors in dose-response format to generate IC50 values for each compound in the cell backgrounds mentioned above ( Table 2 ). To maintain consistency, we transduced each cell line with BacMam virus prior to running the LanthaScreen® assay according to the same protocol used for FACS analysis. The selected compounds, some of which are currently in clinical testing, cover a range of Akt and PI3K isoform specificities. Furthermore, many of these compounds were profiled previously using the HEK 293E clone stably expressing GFP-PRAS40, 3 and the data generated here in the same cell background using the transient BacMam system yielded matching IC50 values. Some remarkable disparities were uncovered, however, when certain inhibitors were assessed in the different cell backgrounds with altered pathway signaling.
Compound IC50 Values from BacMam-Enabled Profiling Experiments in Different Cellular Backgrounds
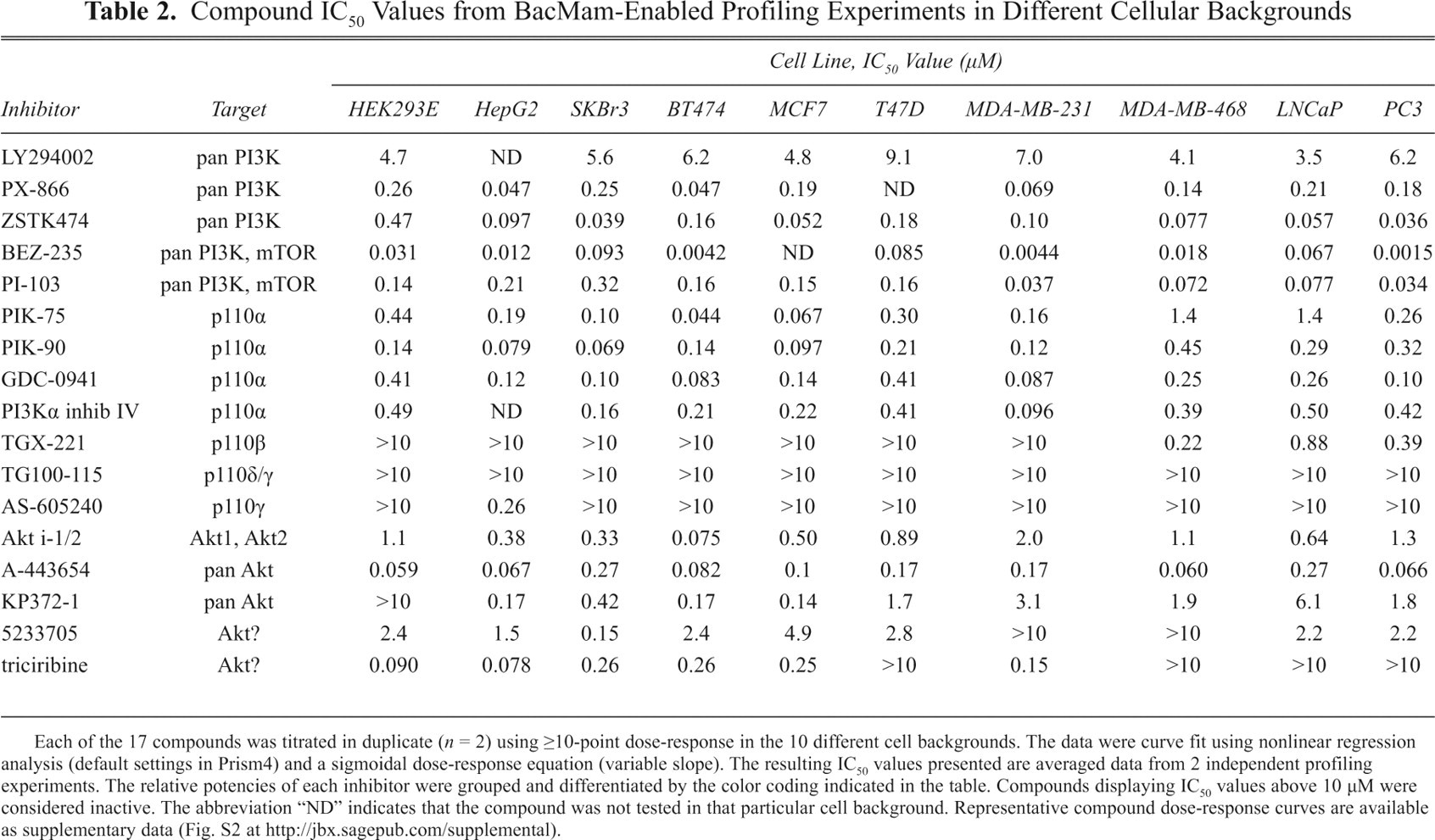
Each of the 17 compounds was titrated in duplicate (n = 2) using ≥10-point dose-response in the 10 different cell backgrounds. The data were curve fit using nonlinear regression analysis (default settings in Prism4) and a sigmoidal dose-response equation (variable slope). The resulting IC50 values presented are averaged data from 2 independent profiling experiments. The relative potencies of each inhibitor were grouped and differentiated by the color coding indicated in the table. Compounds displaying IC50 values above 10 µM were considered inactive. The abbreviation “ND” indicates that the compound was not tested in that particular cell background. Representative compound dose-response curves are available as supplementary data (Fig. S2 at http://jbx.sagepub.com/supplemental).
The PI3K family of lipid kinases includes 4 different class I isoforms: p110α, -β, -γ, and -δ. These enzymes are of particular interest in cancer drug discovery because the activating mutations that affect PI3K signaling mentioned previously are proving to be an essential factor in tumorigenesis. We selected 12 different PI3K inhibitors for this study that included the following: 5 compounds that target all isoforms (pan-PI3K inhibitors), 4 with reported p110α specificity, 1 p110β-selective compound, 1 putative inhibitor of p110γ, and 1 p110γ/δ-specific molecule. Each of these PI3K inhibitors was initially classified based on biochemical assay data, but a comprehensive collection of their cellular activity is lacking. Here, we have compiled data for each compound under a set of standardized conditions across a diversity of cell types as measured by the same cell-based assay readout (i.e., PRAS40 phosphorylation).
Because many cellular functions are regulated by PIK3CA, we expected the p110α-specific and pan-PI3K inhibitors to be effective in every cell line. Indeed, we observed a range of potencies, with the most active compound being the dual PI3K/mTOR inhibitor, BEZ-235 (IC50 values in the nanomolar range; Table 2 ). The classical broad-spectrum PI3K inhibitor, LY294002, also displayed complete inhibition of PRAS40 phosphorylation but was approximately 100-fold less potent (micromolar IC50 values) than BEZ-235. Overall, the data for these 9 compounds were in good agreement with previous reports in the literature. 1,11,12 The only visible trend based on pathway mutation status, although slight, is that PTEN-mutant cell types are less sensitive to p110α-specific than pan-PI3K inhibitors. For instance, the IC50 values for PIK-90 and PI-103 are relatively comparable in cells with wild-type PTEN status (e.g., HEK 293E, BT474, T47D) yet are >3-fold less potent for the p110α-targeted compound (PIK-90) in MDA-MB-468, LNCaP, and PC3 cells. The dependence of PTEN-deficient cancer cells on PIK3CB has been clearly demonstrated in the literature, and it follows that a compound able to inhibit all class I PI3K isoforms might be more efficacious.
In addition, with respect to the PTEN status of cells, it has been shown that loss of this lipid phosphatase also sensitizes cells to inhibitors of p110β. 12 TGX-221, a p110β-selective, adenosine triphosphate (ATP)–competitive PI3K inhibitor, was inactive (IC50 > 10 µM) in all cell lines with normal expression of PTEN. By contrast, addition of this compound attenuated PI3K signaling in each of the 3 PTEN-negative cell lines, with the most potent inhibition displayed in MDA-MB-468 cells (IC50 = 0.22 µM). Profiling of this compound in dose-response format in the LanthaScreen® cellular assay, however, revealed only partial inhibition by TGX-221 relative to the complete reduction in PRAS40 phosphorylation obtained using PI-103 in the same cell background ( Fig. 2A ). We did vary the compound incubation time in MDA-MB-468 cells to address any possible kinetic effects but observed similar results at each time point (data not shown). It is possible that the magnitude of inhibition may be more pronounced with other p110β-selective compounds or with a phospho-readout that is more proximal (e.g., Akt Ser473), but this has not been tested.
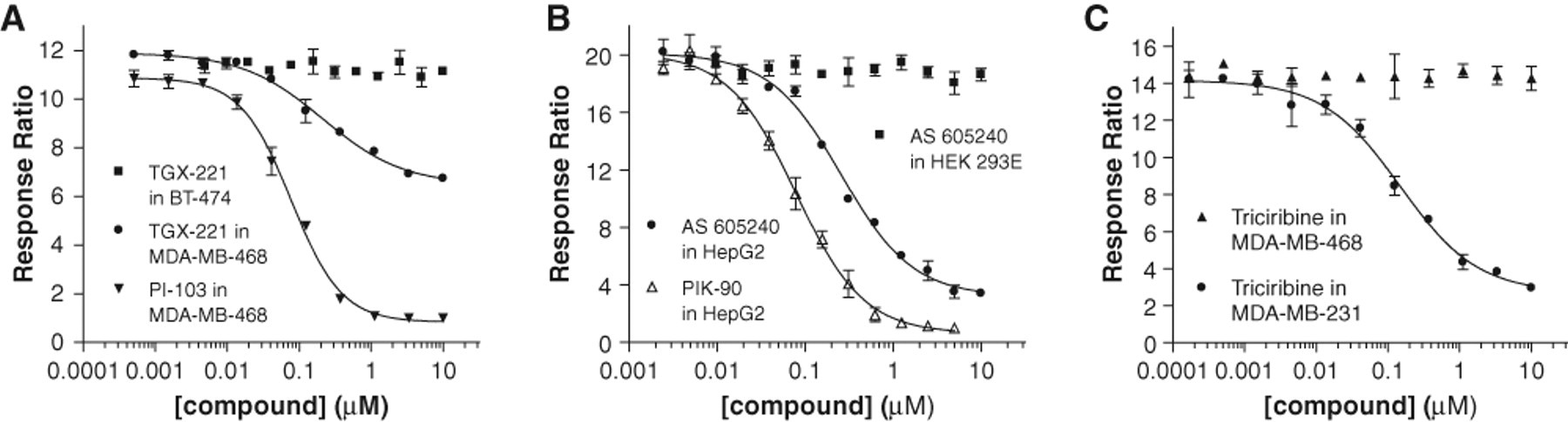
Selected compound dose-response curves from BacMam-enabled LanthaScreen® cellular assays performed in various cell types. (
The expression of PI3Kα and -β is rather ubiquitous in tumor cell lines, whereas PI3Kγ and -δ are mainly present in hematopoietic cells. Although some of the cell backgrounds used here have been shown to express significant levels of the latter 2 isoforms (e.g., p110δ in HepG2 hepatoma cells), 11 profiling of the p110γ/δ isoform-specific inhibitor, TG100-115, resulted in no inhibition in every cell line examined (IC50 values > 10 µM). Interestingly, the p110γ-selective compound, AS-605240, had a similarly inactive inhibition profile with one notable exception: potent activity was detected in HepG2 cells (IC50 = 0.29 µM) ( Fig. 2B ). Again for comparison, we have overlaid the data for the p110α-selective inhibitor, PIK-90, to illustrate the level of complete inhibition for this cell background. It is unclear why inhibition with AS-605240 was not detected in any other cell line tested, but it appears that the levels of enzymatic activity for a particular PI3K isoform must be different in HepG2 cells than in other backgrounds (under the conditions of our cell-based assay). On a related note, we have profiled this compound biochemically across the entire PI3K family and observe inhibition with kinase isoforms other than just p110γ (data not shown). Taken together, these data together suggest that AS-605240 may be less selective than previously described. 13
It is widely accepted that Akt is an important therapeutic target for human cancer. 14 Just as in lipid kinase drug discovery, however, the development of compounds specific for Akt has been complicated by the fact that Akt exists in 3 structurally similar isoforms (Akt1, Akt2, and Akt3), which differ not only in function but also in tissue distribution. In recent years, several new inhibitors—both ATP competitive and non–ATP competitive—have been described. 15 For our studies, we selected 5 different Akt pathway inhibitors for BacMam-enabled cellular profiling experiments.
The tricyclic quinoxaline compound AKTi-1/2 selectively inhibits Akt1 and Akt2 by allosterically targeting the pleckstrin homology (PH) domain of the kinase. Testing of this compound in HEK 293E (the same cell background that the stable GFP-PRAS40 cell line is built) yielded an IC50 of 1.1 µM, which is >10-fold less potent than seen in vitro. With respect to PTEN-deficient cell types, inhibition with AKTi-1/2 was also much less effective (IC50s ~ 1 µM). This result may be explained by higher expression levels of the Akt3 isoform in these cell backgrounds, 16,17 for which this inhibitor does not target as potently. In support of this idea, MDA-MB-231 also expresses Akt3, and inhibition in this cell type was poor (IC50 = 2 µM). By contrast, in a cell background that has both PIK3CA mutation and HER2 amplification but no Akt3 expression, such as BT474, AKTi-1/2 potently decreased the phosphorylation of PRAS40 (IC50 = 0.075 µM). This combination of pathway-activating abnormalities has been reported to hypersensitize tumor cells to Akt inhibition (specifically with this compound), as compared to cells with wild-type PI3K signaling. 17
Along these same lines, KP372-1, which has previously been shown to be a sub-micromolar inhibitor of Akt activity in both biochemical and cell-based assays, 18 surprisingly had no effect when we tested it in HEK 293E. Profiling of this compound in PTEN-negative cells, as well as in MDA-MB-231 and T47D, still only resulted in weak activity (IC50s > 1 µM). However, in other cell backgrounds tested, this compound was clearly inhibitory (IC50 values ranging from 0.14-0.42 µM). KP372-1 has not been reported to have any Akt isoform selectivity, but there is a clear cell background dependence for inhibition with this molecule.
Another well-characterized Akt inhibitor is the ATP-competitive indazole-pyridine called A-443654. In agreement with previously published data, 19 we found this compound to be an excellent inhibitor of Akt, reducing PRAS40 phosphorylation with IC50 values ranging from 0.06 to 0.34 µM across all 10 cell lines. On a side note, A-443654 inhibition of Akt substrates is paradoxically accompanied by the induction of Akt hyperphosphorylation at Ser473 in multiple cell lines, and we have reproduced that result in a LanthaScreen® assay with HEK 293E cells that stably express GFP-AKT (data not shown). More important to this study, however, A-443654 appears to be the one Akt inhibitor that is not influenced by any signaling pathway mutations.
The final 2 inhibitors analyzed yielded compound profiles that were the most clearly affected by mutational status of the pathway: the benzimidazole-based compound 5233705 (Akt inhibitor IV) and triciribine, a purine analog also known as API-2 or TCN. Both have been shown to inhibit the cellular activation of Akt in many examples, yet the exact molecular target of these compounds is not known. As such, they are commonly referred to as “Akt pathway inhibitors.” In the majority of cell lines tested, 5233705 was active with IC50 values in the single-digit micromolar range. The same compound showed no effect in MDA-MB-231 and MDA-MB-468 cells (IC50s >10 µM) but displayed potent inhibition in SKBr3 cells (IC50 = 0.15 µM). Equally disparate results were obtained with triciribine ( Fig. 2C ). This compound was inactive when tested in the 3 PTEN-negative cell lines (IC50s >10 µM), relatively effective in cells with wild-type PI3K signaling (IC50 ~ 1 µM in HEK 293E and HepG2), and quite inhibitory in cell backgrounds with PIK3CA mutations or HER2 overexpression (IC50 ~ 0.25 µM in BT474, MCF7, MDA-MB-231, and SKBr3). Curiously, T47D cells were an anomaly for this trend because the IC50 for triciribine was >10 µM. Given the dependence of certain tumors on Akt signaling, follow-up studies with these compounds is warranted.
There are several plausible explanations for each of the observed differences in compound profiles discussed above, including the varied expression levels of Akt isoforms among cell types or time-dependent kinetics of inhibition; however, elucidation of each of these reasons is unfortunately beyond the immediate scope of this study. Nevertheless, our results clearly underscore the importance of compound profiling in disease-relevant cell backgrounds (vs. the typical workhorse cell lines) and provide a valuable reference for the activity of many well-known inhibitors of the PI3K/Akt pathway.
We recognize that alternative cell-based methods exist to assess PI3K/Akt signaling, such as Western blot, phospho–enzyme-linked immunosorbent assay (ELISA), or cytoblot; however, the BacMam-enabled LanthaScreen® cellular assay described herein offers various advantages over these more traditional immunoassay formats. As addressed in our prior publications, 3-6 the LanthaScreen® cellular assay is homogeneous (no wash steps required), requires minimal hands-on time, and displays excellent assay performance in a 384-well format, which are all significant improvements. Moreover, the use of a GFP fusion protein simplifies assay development relative to other assay formats by eliminating the need for affinity-based capture reagents (i.e., a second antibody and/or beads) and allows for a ratiometric TR-FRET readout compatible with standard fluorescent plate readers (no image-based detection required). Last, the novel application of BacMam to deliver the GFP substrate for LanthaScreen® expands our assay technology into nearly any cell background and moves away from the dependence on stable cell line development. Taken together, the ability to measure target-specific phosphorylation by endogenous kinases in a cellular context of choice with a robust and flexible assay readout (particularly when cryopreserved “assay-ready” cells can be made available) is extremely valuable.
Summary
The use of BacMam as a vehicle for efficient gene delivery into mammalian cells has emerged as a powerful system for a variety of applications. The combination of BacMam technology to deliver substrates for the LanthaScreen® cellular assay platform enables the investigation of compound activity across a broad range of biologically relevant cell models. Consistent with our GFP expression results from FACS analysis, we detected a robust TR-FRET signal for phosphorylation of PRAS40 at Thr246 in every cell line tested. The profiling data with a panel of known PI3K and Akt inhibitors were in good agreement with expected compound pharmacology (if known for that cell type), yet we uncovered previously unreported differences in activity for several compounds in the cell lines with altered PI3K/Akt signaling. These results pose interesting questions for future investigation, and the development of additional assay readouts using these 2 technologies is currently ongoing in our laboratories.
Footnotes
Acknowledgements
The authors thank Thomas Machleidt, Elizabeth Frey, Jun Wang, Upinder Singh, Pamela Whitney, and Scott Brown for expert support throughout various stages of this work. Additional thanks to Michael Hancock, Matt Robers, and Kurt Vogel for critical evaluation of this manuscript and many valuable scientific discussions.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
