Abstract
Background:
Antimicrobial resistance (AMR) is a growing global threat that complicates the treatment of infectious diseases, including plague. Yersinia pestis, the causative agent of plague, remains a serious public health concern in natural foci, such as those in Kazakhstan, where approximately 40% of the territory is plague-endemic. Despite the last reported human case in 2003, data on antibiotic resistance among Y. pestis isolates from these foci, especially historical ones, remain limited.
Materials and Methods:
A total of 75 Y. pestis strains were examined, including 61 isolates obtained from patients and deceased individuals during epidemic outbreaks (1926–2003) and 14 isolates from carriers and vectors in natural plague foci. Taxonomic identification was conducted using the Vitek 2 Compact 30 system. Antibiotic susceptibility was assessed by Kirby–Bauer disk diffusion and E-test methods. Extended-spectrum β-lactam (ESBL) activity was evaluated phenotypically, and resistance genes to glycopeptides and β-lactams were screened by real-time polymerase chain reaction (RT-PCR) using the BacResista GLA Detection Kit.
Results:
All isolates showed complete susceptibility (100%) to β-lactams, tetracyclines, aminoglycosides, amphenicols, glycopeptides, lincosamides, and quinolones. The overall susceptibility rate across antibiotic classes was 97.5%. Macrolides exhibited low activity (0.0–58.0%), consistent with known limitations against Gram-negative bacteria. No ESBL production was detected phenotypically, and RT-PCR screening found no resistance genes (vanA/B, mecA, tem, ctx-M-1, shv, oxa, imp, kpc, ndm, etc.).
Conclusions:
These findings confirm a lack of resistance to key antibiotic classes in historical Y. pestis isolates from Kazakhstan. Despite the absence of recent human cases, ongoing epizootics among wild animals highlight a persistent risk of transmission. This study, conducted for the first time in Kazakhstan, has important implications for public health preparedness and clinical management during plague outbreaks.
Introduction
Antimicrobial resistance (AMR) is a critical and growing global threat to public health and sustainable development. The World Health Organization (WHO) has identified AMR as one of the top 10 global public health threats facing humanity today.1,2 Although antibiotics prevent millions of deaths annually, inappropriate prescription practices and the widespread overuse of antibiotics have led to escalating resistance, culminating in a global health emergency. 3
According to WHO projections, by 2029, approximately 70% of known bacterial species may develop resistance to one or more antibiotics, potentially resulting in 1.27 million deaths annually. 4 In response, Kazakhstan, with support from the WHO and within the framework of international initiatives—the Global Antimicrobial Resistance Surveillance System (GLASS) and the European Antimicrobial Resistance Surveillance Network (EARS-Net)—has developed a National Surveillance Program on AMR, aligned with the “One Health” approach.
Plague is a highly infectious, natural-focal, quarantinable disease characterized by an acute onset, severe systemic illness, fever, and the involvement of the lymph nodes, lungs, and other internal organs, often progressing to sepsis. The causative agent, Yersinia pestis, 5 exhibits nearly 100% lethality and extremely high transmissibility. 6
Natural plague foci are distributed across a broad geographic belt encompassing tropical, subtropical, and temperate zones between approximately 55°N and 40°S. 7
At present, the global epidemiological situation regarding plague remains concerning, with no signs of decline. 8 Between 2014 and 2024, plague outbreaks were recorded in 10 countries, with 5,514 reported cases and 605 deaths (fatality rate: 11.1%).9–13 The most severe outbreaks during this period occurred in the Republic of Madagascar (3,457 cases, 373 deaths) and the Democratic Republic of the Congo (1,352 cases, 76 deaths), accounting for 87.2% of all cases globally.14,15
Kazakhstan hosts six natural plague foci, covering an area of approximately 1,083,900 km2—about 40% of the national territory—placing over 1.5 million people at risk of infection. 16 From 1920 to 2003, more than 2,000 human cases were recorded within the Central Asian natural plague focus, with a case fatality rate of 90.6%. More than 1,000 of these cases occurred in Kazakhstan, where the mortality rate has reached 80.0%.17–19 In recent years, many scientists from Kazakhstan and Russia have made significant contributions to the study of the properties of Y. pestis circulating in the natural foci of Kazakhstan.20–29
In 2024, our team published a meta-analysis summarizing human plague cases in Kazakhstan between 1926 and 2003, 30 which served as the foundation for further investigation. Prior to the introduction of antibiotics,31,32 large-scale plague outbreaks were reported, comprising 80.7% of all documented cases during that period. 30 Prior to the introduction of antibiotic therapy in 1947–1948, the region repeatedly experienced large-scale plague outbreaks, with the most significant occurring in 1926, 1929, 1945, 1947, and 1948. These outbreaks accounted for 80.7% of all recorded human transmission cases during the study period, reflecting the extremely high epidemic activity of Y. pestis in the absence of effective treatment options. The widespread transmission of the disease during these years was facilitated by a combination of poor sanitary and hygienic conditions, the lack of specific therapeutic agents, limited access to medical care, and frequent human interaction with natural plague foci. This contact occurred through trade, transportation and slaughter of camels, hunting of wild animals, and human migration into enzootic areas. These historical outbreaks underscore the population's vulnerability in the pre-antibiotic era and highlight the critical importance of the timely implementation of chemotherapeutic and preventive measures in controlling high-risk infectious diseases.
The aim of this study was to perform, for the first time, a comprehensive screening and identification of genes associated with resistance to β-lactam antibiotics and other classes of antimicrobials in clinical isolates of Y. pestis collected during epidemic outbreaks in the desert plague foci of Kazakhstan from 1926 to 2003 and additional isolates (n = 14) from other natural plague foci in Kazakhstan. The study also aimed to contribute to the development of a microbial biorepository and to enhance national biosafety infrastructure.
Materials and Methods
Bacterial strains and isolates
In this study, Y. pestis clinical isolates (n = 61) were obtained during plague outbreaks in natural foci across Kazakhstan between 1926 and 2003, and additional isolates (n = 14) from other natural plague foci in Kazakhstan. The strains were obtained from the National Working Collection and the Microorganism Depository of the M. Aikimbayev National Scientific Center for Especially Dangerous Infections. The strains were cultured on Mueller–Hinton agar (pH 7.3 ± 0.2) and Hottinger agar (pH 7.2 ± 0.1) at incubation temperatures ranging from 28°C to 37°C. 33 Taxonomic identification was performed using an automated microbial identification system (Biomerieux Vitek 2 Compact 30 Microbiology Analyzer, Marcy-l’Étoile, France). A summary of the strain characteristics is presented in Table 1.
Geographic Origin, Plague Natural Foci, Principal Hosts and Vectors, Years of Human Plague Case Registration, and Yersinia pestis Isolates Included in the Screening for Glycopeptide and β-Lactam Resistance Genes
Antimicrobial susceptibility testing
Antibiotic susceptibility analysis was performed using two complementary phenotypic methods: Kirby–Bauer disk diffusion and E-test gradient diffusion. The Kirby–Bauer method was selected because of its wide acceptance, reproducibility, and simplicity, which enables the rapid screening of susceptibility to multiple antibiotics simultaneously. The E-test was employed to precisely determine the minimum inhibitory concentrations (MICs), which are essential for constructing a detailed antibiotic susceptibility profile. Both assays were conducted in accordance with the guidelines of the Clinical and Laboratory Standards Institute (Malvern, PA, USA).33–36
A bacterial suspension was prepared from the Y. pestis culture in a sterile test tube and evenly spread across the surface of nutrient agar in Petri dishes. Testing was conducted on Mueller–Hinton or Hottinger agar using 18–24-hour cultures incubated at 28°C–30°C. Bacterial suspensions were standardized to 0.5 McFarland (∼1.5 × 108 CFU/mL). Ten to fifteen minutes after inoculation, antibiotic-impregnated disks were placed onto the seeded agar surface, ensuring adequate spacing between the disks to prevent overlapping of inhibition zones. The inoculated plates were then incubated at 28°C. Preliminary observation of inhibition zones was performed after 24 hours, with final measurements taken after 48 hours. To ensure the reliability of the results, reference strains were used alongside control plates containing sterile disks. The diameters of the inhibition zones were measured to the nearest 1 mm using a HiMedia zone scale (AG Medical Company Pvt. Ltd., New Delhi, India).33–36
The antibiotics tested were chosen based on their clinical relevance and included β-lactams, tetracyclines, aminoglycosides, amphenicols, glycopeptides, lincosamides, quinolones, macrolides, and several other classes. To ensure the accuracy and reliability of the results, standard reference strains were used, including Y. pestis EV, Escherichia coli ATCC 25922, Klebsiella pneumoniae ATCC 70060, Staphylococcus aureus ATCC 372, and Pseudomonas aeruginosa ATCC 377.33–36
Results were classified as sensitive (S), intermediate (I), or resistant (R). Resistant findings were confirmed by repeat testing and, when necessary, molecular methods (e.g., detection of resistance genes).
Biochemical testing
The biochemical characterization of Y. pestis strains was performed using a VITEK 2 System (Biomerieux), with a GN card 07.01 (Code: GN 00001B1B42DB, Lot: 19086). For antimicrobial susceptibility testing, bacterial suspensions were prepared from 24 hours agar cultures in 0.85% isotonic sodium chloride solution and standardized according to the McFarland turbidity standard (McFarland Standard Set R092-1NO, Lot No. 0000633797; valid until February 2026; HiMedia Laboratories Pvt. Ltd., Maharashtra, India), using indicators R092A and R092B.33–36
Molecular genetic screening
Real-time polymerase chain reaction (RT-PCR) was used to screen Y. pestis isolates for genes associated with resistance to glycopeptide and β-lactam antibiotics. This method was selected because of its high sensitivity, specificity, and rapid turnaround, enabling the precise identification of AMR genes. 37
Genomic DNA was extracted using commercial kits (Real-RealBest UniMag and “RealBest UniMag” and “RIBO-prep”) optimized for automated DNA/RNA extraction platforms, ensuring standardized procedures and minimizing the risk of cross-contamination. The genetic screening targeted a range of resistance determinants, including vanA/B, mecA, tem, ctx-M-1, shv, and multiple carbapenemase-associated genes (oxa-40-like, oxa-48-like, oxa-23-like, oxa-51-like, imp, kpc, ges, ndm, and vim), using the BacResista GLA Real-Time PCR Detection Kit (DNA-Technology LLC, Moscow, Russia).
For the detection of Y. pestis genetic targets by multiplex PCR, amplification curves were monitored across three fluorescence channels: 6-carboxyfluorescein (FAM), a red fluorophore used for labeling DNA probes, characterized by high fluorescence intensity and applied for the detection of the first genetic target; hexachlorofluorescein (HEX), a blue fluorophore used for labeling a second target, allowing differentiation of signals from multiple targets within the same sample; and cyanine-5 (CY5), a yellow fluorophore with excitation and emission wavelengths distinct from FAM and HEX, which enables the simultaneous detection of a third target, thereby increasing the sensitivity and specificity of the analysis.
Registration of fluorescence signals in these channels allows for the monitoring of PCR product accumulation across successive thermal cycles, where the fluorescence intensity is directly proportional to the quantity of target DNA present in the sample. The sigmoidal shape of the amplification curve confirms the efficiency of the reaction, and the cycle threshold (Ct) value is defined as the point at which the fluorescence signal exceeds the background level, providing a semi-quantitative measure of target abundance in the test sample.
Statistical analysis
MIC data were analyzed for central tendency and variability and were expressed as MIC50 and MIC90. The diameters of the inhibition zones were evaluated using descriptive statistics, including mean values and standard deviations. Statistical significance was assessed using non-parametric tests (Mann–Whitney U test) to compare groups of isolates, with significance defined at p < 0.05.
Results
The antimicrobial susceptibility of microbial cultures was studied simultaneously with a preliminary analysis of the phenotypic and molecular genetic properties of 75 Y. pestis strains. These included clinical isolates (n = 61) obtained during plague outbreaks from 1926 to 2003 from 10 autonomous natural foci, as well as Y. pestis strains (n = 14) from seven autonomous plague foci in the Republic of Kazakhstan, isolated over the past 10 years, where no human plague cases had been reported.
The expansion of the sample size to 75 strains complied with the methodological standards for enhancing the reliability and interpretation of the results. The taxonomic identification of all Y. pestis strains was performed using the Biomerieux VITEK 2 Compact 30 Microbiology Analyzer (Table 2).
Biochemical Characteristics of 75 Y. pestis Strains Tested Using VITEK 2 Systems (as of 07.01)
The biochemical characterization of the isolate demonstrated positive results for the following markers: PyrA (
Thus, the VITEK 2 GN system enabled the identification of Y. pestis strains with an accuracy of up to 96.0%, underscoring the high reliability of its biochemical profile for taxonomic classification.
Y. pestis isolates generally showed susceptibility to streptomycin, gentamicin, doxycycline, ciprofloxacin, and chloramphenicol.
The results of the phenotypic susceptibility testing to antibacterial agents, including the range of inhibition zone diameters for the 75 Y. pestis strains, are presented in Table 3.
Range of Growth Inhibition Zone Diameters (in Millimeter) for Groups of Antibacterial Agents Against 75 Y. pestis Strains on Mueller–Hinton Agar
MIC, minimum inhibitory concentration.
The results demonstrated that the strains were highly susceptible to major classes of antibiotics, including β-lactams, tetracyclines, aminoglycosides, amphenicols, glycopeptides, lincosamides, and quinolones (100%), as well as to antibiotics from other classes (97.5%). At the same time, all strains exhibited extremely low sensitivity to macrolides (0.0–58.0%), which confirms the well-known inefficacy of this antibiotic class against Gram-negative bacteria of the Yersiniaceae family.
Representative images illustrating the results of antimicrobial susceptibility testing are provided in Figure 1.
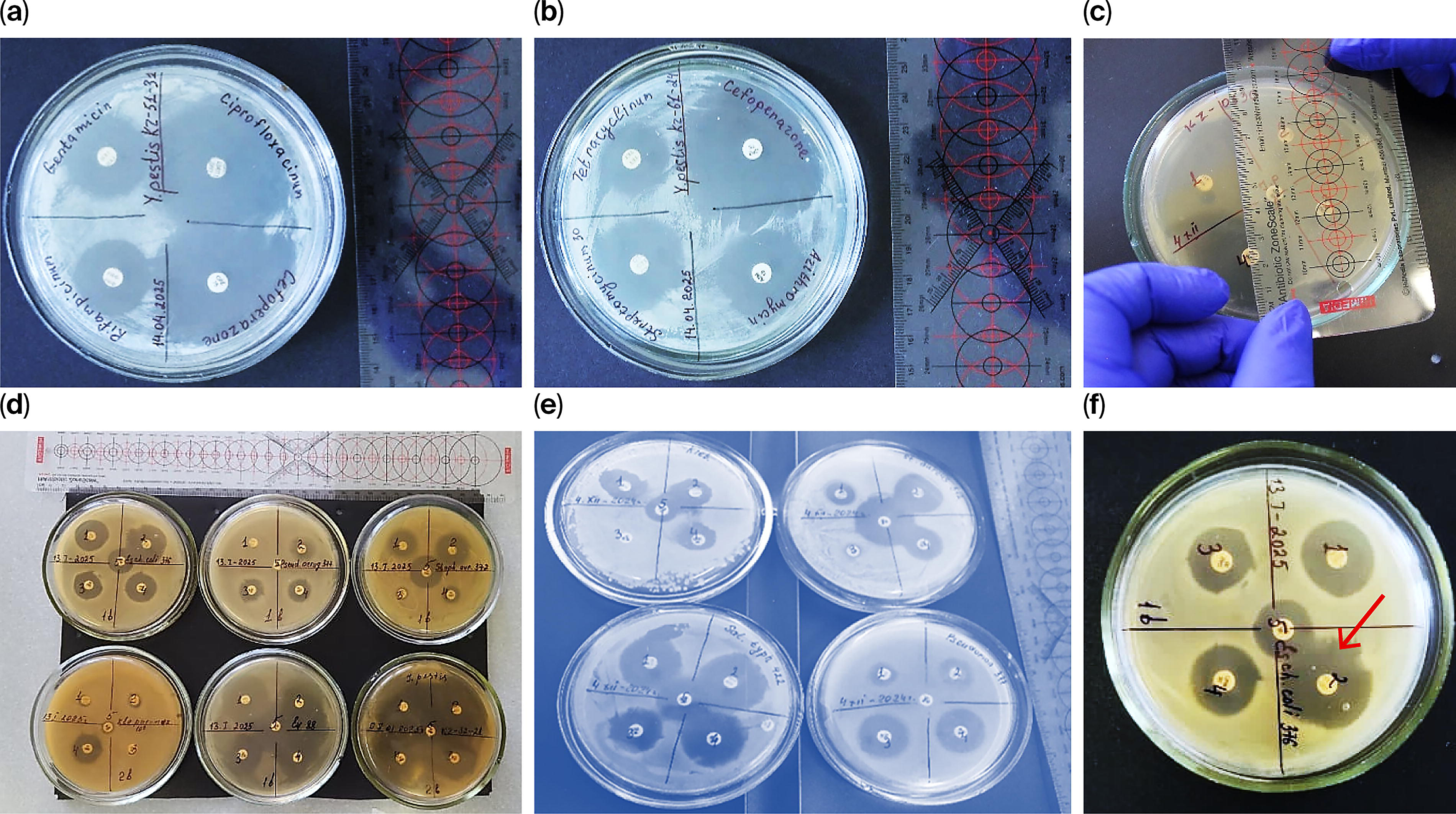
Representative photographs of selected inhibition zones obtained by the disk diffusion method and phenotypic tests for the detection of extended-spectrum β-lactam (ESBL) resistance genes:
These include clear examples of inhibition zones around antibiotic-impregnated disks, as well as phenotypic confirmatory tests for the detection of extended-spectrum β-lactam (ESBL) production. The selected photographs demonstrate both the presence and absence of inhibition, enabling visual comparison of resistant versus susceptible phenotypes. Such visual documentation supports the interpretation of phenotypic resistance profiles and complements the molecular findings obtained in this study.
The antimicrobial susceptibility studies were conducted using two types of growth media—Mueller–Hinton and Hottinger agars. However, the summary diagram presented (Fig. 2) reflects the results obtained on Hottinger agar only. This figure includes the specific Y. pestis strain numbers and the full list of antibiotics tested, allowing for a comparative analysis of inhibition zone diameters across antimicrobial classes, with particular focus on the bacterial response observed on this nutrient substrate.
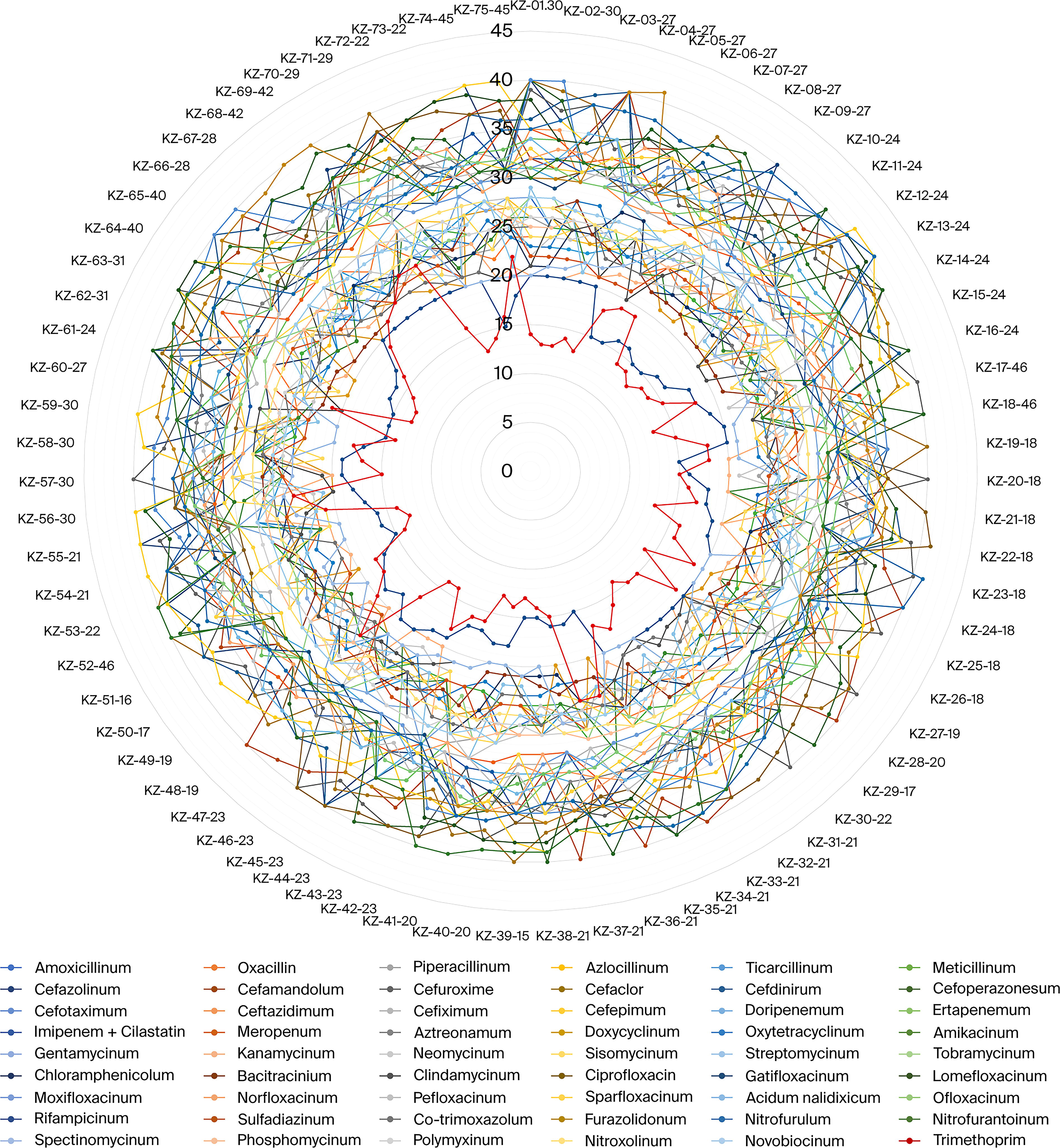
Range of minimum inhibitory concentrations (MICs) of antibacterial agents and growth inhibition zone diameters for 75 Y. pestis strains on Hottinger agar, in millimeters.
The radar chart illustrates the range of growth inhibition zone diameters (in mm) for 75 strains of Y. pestis tested against a wide panel of antibacterial drugs on Mueller–Hinton agar. Each radial line corresponds to an individual Y. pestis isolate (labeled KZ-01-75), while the concentric circles represent the diameter of the inhibition zones. The different colors of the curves represent different antibiotics, as indicated in the comprehensive legend below the figure.
The figure clearly demonstrates a consistent susceptibility pattern across most of the strains, with the majority showing inhibition zone diameters between 10 and 45 mm for most antibiotics. This indicates a high level of sensitivity among Y. pestis isolates to a broad range of antimicrobial agents.
The central red-colored polygon represents the range of MICs for the antibiotics tested, providing a comparative reference. The relatively smaller size of the MIC contour in comparison to the growth inhibition curves supports the phenotypic finding of high antimicrobial susceptibility.
Notably, no isolate falls significantly outside the general pattern, suggesting the absence of outliers with reduced susceptibility or resistance. The close clustering of lines across most of the antibiotics further reinforces the conclusion that the Y. pestis strains tested remain uniformly sensitive to multiple antibiotic classes, including β-lactams, aminoglycosides, fluoroquinolones, and tetracyclines.
This visualization supports and complements molecular findings (e.g., absence of resistance genes by PCR), underscoring the continued efficacy of commonly used antibiotics against plague agents in Kazakhstan.
Among the β-lactams, the most pronounced activity was observed with cefepime and ceftriaxone. Ciprofloxacin, a member of the fluoroquinolone group, demonstrated high activity, confirming its effectiveness in the treatment of plague. Traditional antibiotics used in the treatment of plague (streptomycin, tetracycline, and chloramphenicol) demonstrated moderate activity, whereas trimethoprim and rifampicin showed the least activity.
The results of the sensitivity and resistance studies were confirmed by the standard “E-test” method, using strips indicating the MIC. Representative photographs illustrating selected results obtained using the standard “U-test” method are presented in Figure 3. These images demonstrate typical patterns observed during the analysis and serve to visually support the interpretation of experimental outcomes.
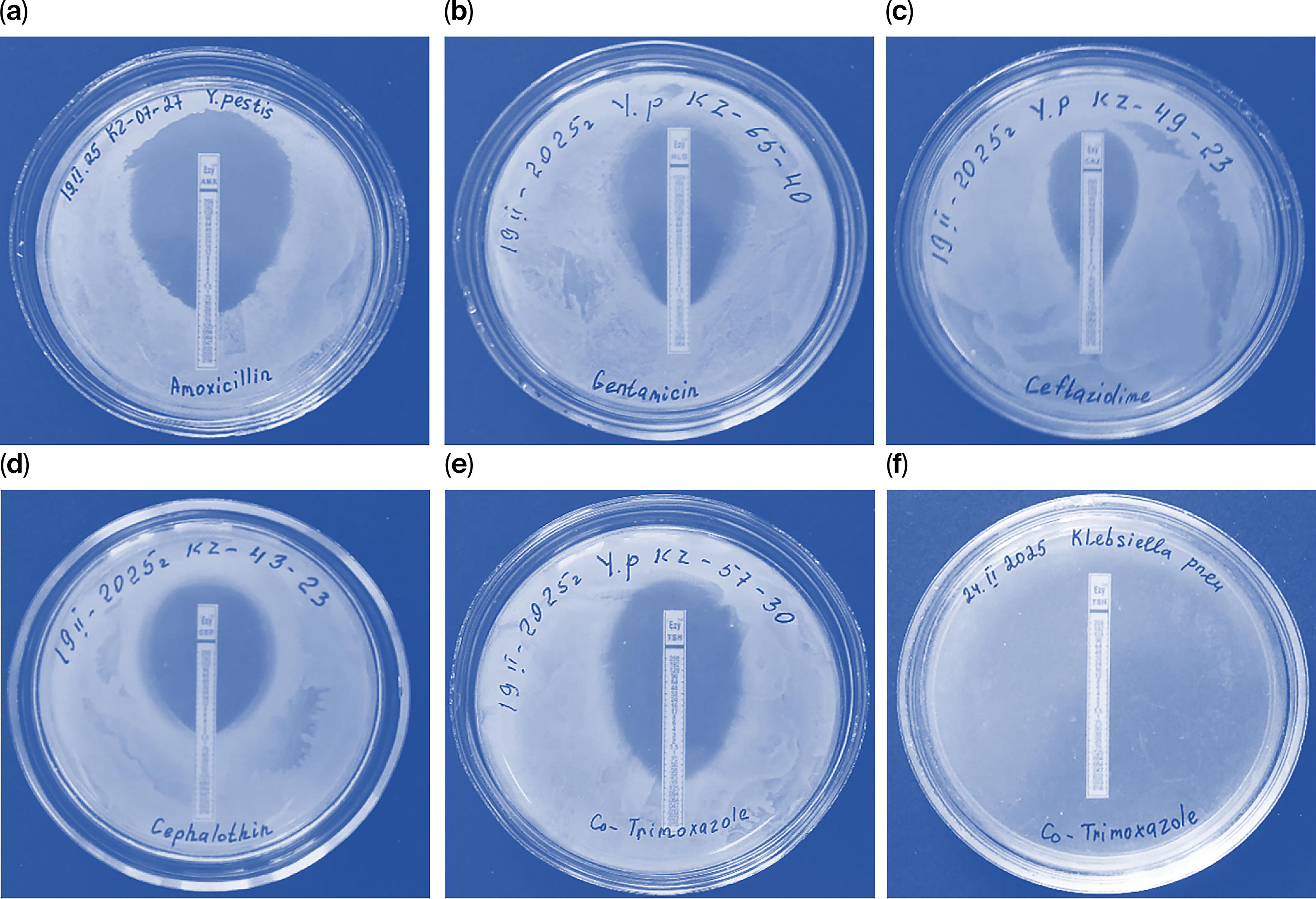
Results of the “E-test” method:
An assessment of antimicrobial susceptibility was conducted for both reference and field strains of Y. pestis using the E-test method. MICs were determined for a range of antibiotics, including representatives of tetracyclines, aminoglycosides, β-lactams, and other classes.
The reference strain Y. pestis EV 88 exhibited high susceptibility to aztreonam (0.064 µg/mL), while the MICs for gentamicin and tetracycline were 24 µg/mL. The average MIC for this strain was calculated to be 12.516 µg/mL. The strain Y. pestis pstbc 2841 showed lower MIC values: 0.047 µg/mL for aztreonam, 12 µg/mL for tetracycline, 16 µg/mL for gentamicin, and 1.5 µg/mL for cefdinir, with a mean MIC of 7.38 µg/mL.
Analysis of field isolates (KZ series) also demonstrated high susceptibility to the tested antibiotics. MIC values ranged from 0.023 µg/mL (moxifloxacin) to 4 µg/mL (amikacin), with an overall mean MIC of 1.065 µg/mL. These findings confirm the continued efficacy of key antibiotics against Y. pestis. The E-test method proved to be reliable and suitable for quantitative assessment of antimicrobial susceptibility in both laboratory and field-derived strains.
In the search for antibiotic resistance determinants in the genomes of lysates, genes conferring resistance to glycopeptide and β-lactam antibiotics were investigated in 75 Y. pestis strains using the “BacResista GLA Real-Time PCR Detection Kit” system through real-time PCR, searching for the resistance genes van A/B (vancomycin, teicoplanin); mec A (methicillin, oxacillin); tem, ctx-M-1, and shv (penicillins and cephalosporins); and oxa-40-like, oxa-48-like, oxa-23-like, oxa-51-like, imp, kpc, ges, ndm, and vim (carbapenems). The results of the real-time PCR are presented in Table 4 and Figure 4.
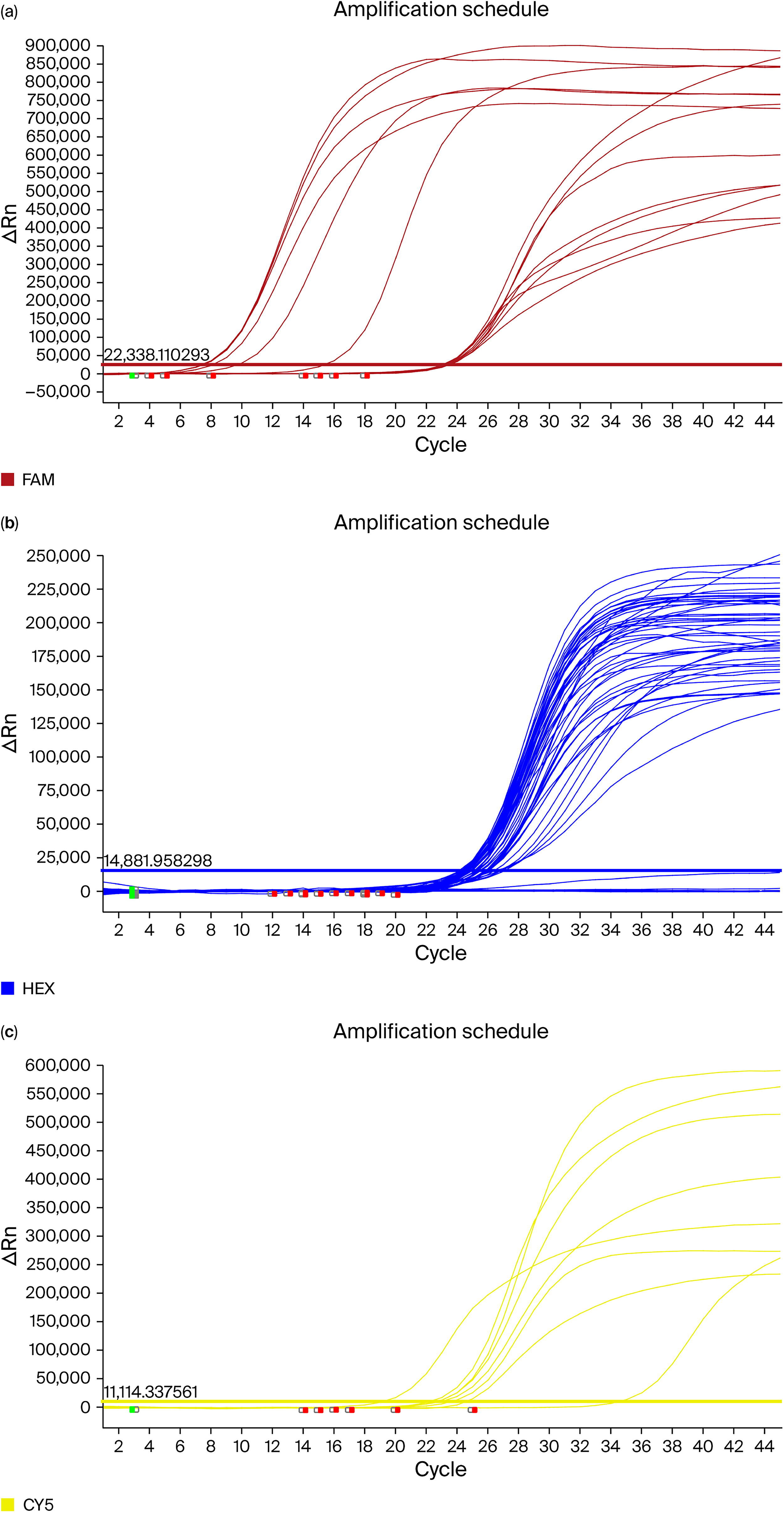
Amplification graph of fluorophore detection channels: real-time PCR amplification curves demonstrating the fluorescence signal increase in three detection channels:
Results from the Study of Genes Resistant to Antibacterial Drugs (ABP) in 75 Y. pestis Strains Using the “BacResista GLA Real-Time PCR Detection Kit” System Through Real-Time PCR
KZ61-24, average value based on results from 61 Yersinia pestis strains.
CY5, cyanine-5; FAM, 6-carboxyfluorescein; HEX, hexachlorofluorescein; IC, internal control; PCR, polymerase chain reaction; TBM, total bacterial mass.
The results presented in Table 4 provide important insights into the current genetic resistance landscape of Y. pestis. Using the “BacResista GLA Real-Time PCR Detection Kit,” we screened 64 strains, including the reference strain EV and 14 field isolates (grouped as KZ61-24), for the presence of key resistance genes associated with β-lactams, carbapenemases, glycopeptide, and methicillin resistance.
Strikingly, none of the Y. pestis strains tested positive for any of the targeted resistance determinants. Genes commonly associated with ESBL production, such as ctx-M-1, shv, and oxa-type variants, were absent. Similarly, no amplification signals were observed for carbapenemase genes (kpc, ndm, vim, oxa-48-like, oxa-23-like, oxa-40-like), nor for vanA/B and mecA, which are typically involved in glycopeptide and methicillin resistance, respectively.
The absence of amplification signals in the FAM and CY5 channels across all Y. pestis samples supports a lack of horizontally acquired resistance elements in these strains for the tested targets.
In contrast, the presence of target genes in the positive control strains validated the performance and specificity of the assay. For example, Klebsiella pneumoniae ATCC 70060 tested positive for shv, Escherichia coli ATCC 25922 for tem (Ct = 34.60), Pseudomonas aeruginosa ATCC 377 for tem (Ct = 24.85), and Yersinia pseudotuberculosis R-2841 for vanA/B. Internal control amplification (HEX channel) produced cycle threshold (Ct) values within the expected range (∼24–27) in all tested samples, confirming assay reliability and the presence of amplifiable DNA.
Collectively, these findings reinforce previous phenotypic observations that Y. pestis remains highly susceptible to a broad spectrum of antibiotics. Importantly, the molecular evidence from this study demonstrates that the organism does not currently carry genetic determinants commonly associated with acquired AMR in other Gram-negative and Gram-positive bacteria. Continuous monitoring remains essential, however, due to the global rise of multidrug-resistant pathogens and the known capacity of Yersinia species to acquire resistance genes under selective pressure. The results provide important evidence of the epidemiological stability of natural plague foci in Kazakhstan and substantiate the feasibility of using existing antibiotic therapy regimens.
Discussion
An integral part of molecular epidemiological monitoring is the tracking of both the phenotypic resistance profiles of pathogens associated with health care-related infections (HCRIs) to antibiotics (ABPs) and the mechanisms of resistance that have clinical and epidemiological significance. 38
Antimicrobial therapy is effective in alleviating symptoms in patients, especially when administered during the early stages of infection.39–41
Traditional antimicrobial agents used for the treatment and/or prevention of plague include aminoglycosides (streptomycin and gentamicin), tetracyclines (doxycycline and tetracycline), chloramphenicol, and co-trimoxazole.42–44 In 2011–2012, a large-scale study was conducted by American scientists to assess the resistance of 392 Y. pestis isolates from 17 countries (America, Africa, and Asia) to eight antibiotics used in the treatment or prevention of plague. 45
The sensitivity screening and molecular genetic analysis of Y. pestis strains isolated in Kazakhstan between 1926 and 2003 confirmed their high sensitivity to the main groups of antibiotics used for plague prevention and treatment.
It is important to note that in 1995, a multidrug-resistant (MDR) strain of Y. pestis was isolated from a human patient on the island of Madagascar. This strain exhibited resistance to multiple classes of antibiotics, including those used for treatment (streptomycin, chloramphenicol, tetracycline), prophylaxis (sulfonamides, tetracycline), as well as reserve drugs (kanamycin, ampicillin, spectinomycin).32,46,47
In 1996, another resistant strain of Y. pestis—designated S19960127 and classified as biovar Antiqua—was isolated during an outbreak of pneumonic plague in the Qinghai-Tibet Plateau region of Tibet, China. This strain demonstrated a high level of resistance to streptomycin due to a point mutation in the rpsL gene, which encodes the S12 ribosomal protein responsible for binding streptomycin. 48
Between 1995 and 2021, a total of seven Y. pestis isolates with varying mechanisms of antibiotic resistance were reported. These cases highlighted that Y. pestis can acquire resistance through both horizontal gene transfer—such as the conjugative uptake of plasmids bearing AMR genes from other bacterial species—and through spontaneous point mutations in chromosomal loci. 48
In 2022, researchers from China and Malaysia conducted a comprehensive review of AMR in Y. pestis, analyzing 1,499 publications, including 68 full-text articles. The majority of these works focused on characterizing the antibiotic resistance profiles of Y. pestis isolates and provided valuable insight into the global status of drug resistance in this pathogen. 49 The review presented the following data: in 2006, streptomycin-resistant strains were identified in China; in 2010, MDR strains were detected in North America; in 2011, strains resistant to gentamicin, streptomycin, tetracycline, doxycycline, ciprofloxacin, levofloxacin, chloramphenicol, and trimethoprim-sulfamethoxazole were reported in North and South America, Asia, and Africa; and in 2018, streptomycin-resistant, anti-β-lactam, and anti-sulfonamide Y. pestis strains were identified in Qinghai Province, China. 49
These observations underscore the importance of continuous surveillance and molecular monitoring of Y. pestis populations. Although resistance remains rare, its emergence through natural mechanisms reinforces the need for preparedness and early detection in both endemic and non-endemic regions.
These results are of great significance, as the resistance of Y. pestis to antibiotics has been reported in other regions. The absence of similar resistance mechanisms in the Kazakhstani Y. pestis isolates may be attributed to several factors. First, the limited use of modern antibiotics in the natural plague foci of Kazakhstan may have reduced selective pressure, which may have promoted the development of resistance. Second, the potential absence of the horizontal gene transfer of resistance plasmids from other pathogenic bacteria in the region's natural foci may have contributed to the maintenance of the sensitivity of the strains.
These results underscore the need for the continued monitoring of Y. pestis’s sensitivity to antibiotics in Kazakhstan. Regular phenotypic and genetic analyses of strains from natural plague foci are recommended for the timely detection of any changes in the resistance profile. It is also important to monitor antibiotic usage in endemic regions to minimize the risk of the emergence and spread of resistant strains.
Thus, the findings of this study have practical significance for Kazakhstan's health care and biosafety system. The results allow for the optimization of therapeutic strategies and the maintenance of preparedness for rapid response in the event of plague outbreaks, based on up-to-date data on pathogen sensitivity.
Conclusions
In this study, screening for antibiotic sensitivity and the detection of antibiotic resistance genes in 75 clinical and natural Y. pestis isolates from various natural foci in Kazakhstan (1926–2003) was conducted for the first time. The study found a complete absence of phenotypic and genetic resistance to key antibiotic groups (β-lactams, tetracyclines, aminoglycosides, amphenicols, glycopeptides, lincosamides, and quinolones) in the examined strains.
All of the studied strains exhibited low sensitivity to macrolides, which aligns with well-known data on the limited effectiveness of this group against Gram-negative bacteria, including Enterobacteriaceae.
In conclusion, the findings of this study hold significant practical value for Kazakhstan's public health and biosafety systems. The data obtained contribute to the optimization of treatment protocols and support evidence-based decision-making for outbreak preparedness. By providing current insights into the antimicrobial susceptibility of Y. pestis, this study strengthens the foundation for effective therapeutic planning and timely response in the event of potential plague reemergence.
The obtained data have significant epidemiological and clinical implications, confirming the continued high efficacy of the standard antibiotics recommended for the prevention and treatment of plague in Kazakhstan. The results of this study provide a foundation for the creation of a national biorepository and an information database on the antibiotic susceptibility of pathogens that cause particularly dangerous infections. This will enable the expansion of resistance monitoring using modern approaches such as whole-genome sequencing for a deeper understanding of the mechanisms underlying the formation and spread of antibiotic resistance in Y. pestis.
Authors’ Contributions
Conceptualization: Z.A., Z.Z., and R.S.M. Methodology: Z.A., R.M., and B.A. Software: N.S. Validation: Z.A., R.M., A.A., A.R., S.I., B.B., B.T., D.O., N.S., A.J., and B.A. Formal analysis: A.A., A.R., B.B., D.O., N.S., and B.B. Investigation: Z.A., Z.Z., and R.M. Resources: A.A., A.R., S.I., B.B., B.T., D.O., and A.J. Data curation: Z.A., R.M., D.O., and N.S. Writing—original draft preparation: Z.A. Writing—review and editing: Z.Z. and R.M. Visualization: Z.A. and N.S. Supervision: Z.Z., R.M., and A.A. Project administration: N.S. Funding acquisition: Z.A. All authors have read and agreed to the published version of the article.
Footnotes
Disclosure Statement
No competing financial interests exist.
Funding Information
This research was conducted as part of the Ministry of Science and Higher Education of the Republic of Kazakhstan's project titled “Study of Antibiotic Resistance Genes in Plague and Cholera Pathogens, Development of a PCR Test System,” project IRN—AP19679355, funded by the Committee of Science of the Ministry of Science and Higher Education of the Republic of Kazakhstan.
