Abstract
Pulmonary embolism (PE) is difficult to diagnose due to nonspecific symptomatology and lack of accurate diagnostic tools. PE can be categorized into low-risk, submassive, and massive based on severity. Changes in several plasma and serum microRNAs (miRNA) are associated with PE, but most lack appropriate specificity for diagnostic utility. Given the emerging role of small extracellular vesicles (sEVs) in PE pathogenesis, we hypothesized that the microRNAs (miRs) circulating within these sEVs can be sampled to gain new information for biomarker studies and disease mechanisms. To address current technical limitations, we developed a novel high-yield size-exclusion chromatography (SEC) method for sEV isolation from small clinical plasma samples, and employed a non-biased method, RNA-Seq, to gain new information for RNA markers that have not been previously studied in the disease. Our goal was to establish the feasibility of the approach in a pilot study with pooled samples from a stratified (AHA criteria) patient population. We present information from sEV characterization studies of individual patient samples, and RNA-Seq data from the pools. PE patients had larger sEVs than healthy subjects. Small RNA sequencing of the sEV samples revealed differential miRNA expression patterns that correlated with PE severity and were predicted to be involved in PE pathology-related processes. Our data offer a first step toward identifying miRNA targets that may have utility in a miRNA biomarker panel for PE diagnosis, prediction of severity, treatment responses and adverse outcomes. This strategy can be extended to the detection of adverse effects and drug-drug interactions as well.
Keywords
Introduction
Venous thromboembolism (VTE) includes pulmonary embolism (PE) and deep-vein thrombosis (DVT). PE is the third leading cause of cardiovascular death worldwide and its prevalence has been steadily increasing over the past two decades.1,2 PE is caused by the blockage of the pulmonary artery by a thrombus travelling from the deep vein. This can severely impact the right ventricle and heart function in general, which is a major determinant of PE patient outcome. 3 Clinically, PE can be categorized into massive, submassive, and low-risk PE based on severity and hemodynamic compromise per the American Heart Association (AHA) guidelines. 4 The diagnosis of PE can be difficult due to nonspecific symptomatology. 3 A combination of tools, such as Wells score and Geneva score, clinical imaging, and laboratory tests is generally used for PE diagnosis. 3 The laboratory tests used include D-dimer, cardiac troponin and brain-type natriuretic peptide (BNP), however, these are not specific for PE. Currently, CT pulmonary angiography (CTPA) remains the gold standard for PE diagnosis. However, CTPA poses a significant risk, especially for patients with renal insufficiency, highlighting the need for more specific and less invasive diagnostic tools.
Circulating biomarkers have been studied since the 1800s, originally in the form of circulating tumor cells, and allow for non-invasive sampling and quantification. However, as different clinical manifestations and pathologies involve similar, and at times overlapping pathophysiological processes, no single marker is likely to meet all the criteria for a diagnostic PE biomarker.5,6 Biomarker panels, in contrast, offer the potential to reveal unique disease-specific biomarker signatures that may help diagnose, guide management, monitor therapy outcomes, and predict potential safety risks. 7
MicroRNAs (miRNAs) are 19-24-nucleotide-long small noncoding RNAs that mostly function in post-transcriptional degradation of mRNAs. 8 Circulating miRNAs have been extensively studied over the past decade, identifying them as key regulators of gene expression as well as biomarkers for different diseases, most notably malignancies.9,10 Many miRNAs are implicated in the regulation of pathophysiological processes that are often referred to as Virchow's triad, ie hypercoagulability, hemodynamic stasis, and endothelial dysfunction; some of these have been identified as potential biomarkers for PE, including miR-134, miR-28-3p, miR-27a/b, miR-1233, and miR-221.11–16
Compared with protein biomarkers, miRNAs are more stable and don’t require an antigen-antibody complex formation. 17 However, a common problem encountered for miRNA biomarkers in serum or plasma, the lack of specificity, derives in part from the fact that many cell types can secrete detectable levels of miRNAs into circulation and it is difficult to determine where the miRNAs come from or what stimulus elicits their increased production from particular cell types in patient population. However, Genschmer et al. demonstrated that neutrophil-derived extracellular vesicles (EVs) from chronic obstructive pulmonary disease (COPD) patients can be isolated by immunoselection for CD66b. 18 We envision that techniques will be developed to allow this approach in concert with biomarker development. Today, many studies use multiple miRNAs to form diagnostic panels to improve differentiation between diseases.19,20 Herein, we propose a new approach to investigate circulating miRNAs as biomarkers in PE, using circulating EVs and RNA sequencing.
Interestingly, reports suggest that some miRNAs are protected from degradation in the circulation when contained within EVs or bound to proteins.9,21 EVs are lipid vesicles secreted by cells into the extracellular space and the classifications relevant to this work include small EVs (sEVs), microvesicles and apoptotic EVs, based on their biogenesis, size, and function.22–25
We here investigated the sEV population, which includes exosomes derived from the endosomal reticulum and less than 200 nm in diameter. Exosomes are formed via the inward budding of the endosomal membrane in the multivesicular bodies (MVB) and later released into the extracellular space via membrane fusion of MVB and the plasma membrane.22,26 The sEVs are packed with lipids, proteins and RNA from the EV producer cells and can be taken up by other cells, delivering their cargo. The RNA cargo, including miRNAs, can regulate the gene expression in the recipient cells.26–28 Therefore, sEV contents represent the physiological state and gene expression network of the producer cells and carry miRNAs that characterize the cellular gene expression network in both normal physiology and pathophysiology and thus have the potential to affect recipient cells.
Increasing evidence supports that sEVs are involved in pathogenesis of diseases.29–31 Not only do they contain regulators of gene expression, but research also showed that sEVs can carry biologically active proteins to function in disease progression. Activated neutrophil (PMN)-derived sEVs have been identified as a pathogenic agent in COPD as they retain active neutrophil elastase (NE) on their surface. 18 Interestingly, pathogenic EVs isolated from human can confer COPD phenotypes to mice.18,32 Thus, EVs from injured lung cells can contribute to lung pathologies and may serve as biomarkers for these diseases.33,34 In 2019, FDA granted breakthrough designation for the first exosome-based liquid biopsy test. 35 The ExoDx Prostate IntelliScore (EPI) urine exosome test can detect three known mRNA biomarkers of prostate cancer, PCA-3, TMPRSS2:ERG, and SPDEF, from urine samples.36,37 This lends validity to sEV miRNA biomarker development and opens the door to further sEV biomarker discovery.
Herein, we tested the feasibility of utilizing miRNAs within circulating sEVs as potential biomarkers. Our hypothesis is that these sEV miRNAs contain expression patterns specific to PE severity of the patients, therefore can be used as biomarkers. The sEVs provide protection for the miRNA in circulation, prolonging their half-life, and allow determination of their cellular origin, and thus the miRNAs contained within them, increasing the specificity of the potential miRNA biomarkers. This study also pioneered a novel technique that increases the sEV and RNA yield from plasma that contributes to better investigating the circulating sEV content and can be applied to other cardiovascular diseases.
Materials and Methods
Patient Selection
Patients 18 years or older diagnosed with acute pulmonary embolism (PE) by computed tomographic angiography or ventilation/perfusion imaging were recruited to participate in this study through enrollment conducted in conjunction with an ongoing IRB-approved project by the Pulmonary Embolism Response Team (PERT) registry. Ethical approval for the collection of residual blood samples was obtained through the Loyola University Chicago Institutional Review Board (LU#2094572). Written informed consent was obtained from participants for the use of their anonymized information. All studies involving people, whether patients or research participants, must be evaluated by a qualified research ethics committee prior to undertaking the research in line with the Declaration of Helsinki.
All patient samples were obtained via the PERT registry between January 2020 and May 2022 at either Loyola University Medical Center or Gottlieb Memorial Hospital, upon admission. The patients were categorized into massive, submassive, and low-risk PE according to clinical presentation under the guidelines of the American Heart Association (AHA). 4 PE was confirmed by CT PE (or a V/Q scan in the small minority of patients who are not candidates for CT) and an echocardiogram. In addition, troponin and BNP are measured in all patients. Clinical data were collected and stored using REDCap (Research Electronic Data Capture, Vanderbilt University). Investigators were blinded to identifying data. Both clinical data and molecular biomarker analysis were identified exclusively by randomly generated numerical identifiers and investigators who did the molecular analyses were blinded to group assignment.
Clinical data obtained from the electronic medical record (EMR) included: age, gender, race, weight, body mass index (BMI), severity of PE, and past medical history (including history of hypertension, dyslipidemia, congestive heart failure, chronic kidney disease, end-stage renal disease, cancer, peripheral arterial disease, tobacco abuse, chronic obstructive pulmonary disease, diabetes, cerebrovascular disease, PE, DVT, or liver disease).
Additionally, presenting vital signs, labs, echocardiographic markers, presence of DVT on lower extremity duplex, CT parameters, mechanical ventilation, need for intensive care unit monitoring, acute kidney injury, mortality, need for pressors, bleeding, and treatment strategy were collected.
At our institution, PERT comprises a multidisciplinary team of experts from cardiology, vascular surgery, radiology, clinical pharmacy, pulmonary, and cardiothoracic surgery. When activated, typically by the emergency room or primary medical or surgical services, the on-site PERT fellow promptly assesses and staffs the patient with the on-call PERT attendings. A formal recommendation of either an initial conservative strategy or an escalation of treatment is made. This may involve mechanical thrombectomy, or a surgical procedure in the operating room. The PERT consult team continues to follow the patient throughout the hospital stay, with follow-up care arranged in the PE clinic at 1 month post discharge.
To limit the number of variables affecting miRNA expression in different severity groups, we excluded samples with the following criteria:
Patients that require cardiopulmonary resuscitation, vasopressor support and/or endotracheal intubation within 72 h of admission. Patients with previously known cardiac, pulmonary, hepatic and renal insufficiency before the date of diagnosis. Patients that have concomitant other ischemic diseases, including acute coronary syndrome, peripheral artery occlusion, and ischemic cerebrovascular disease, at the time of diagnosis. Patients with a history of venous thromboembolism or APE in the past 6 months and/or under anticoagulant therapy at the time of APE diagnosis. Patients with acute inflammatory disease state including active systemic infection and/or sepsis syndrome. Patients with previously known chronic inflammatory disease state including collagen vascular disease or inflammatory bowel disease before the date of diagnosis.
Blood Samples
Whole blood samples were drawn from patients within 24 h of acute PE diagnosis and collected under an IRB approved protocol. Samples were collected in 3.8% (0.109 mol/L) EDTA or sodium citrate tubes and processed into platelet-poor plasma (PPP) by centrifuging at 3000 rpm for 2 min within 2 h of collection and stored at −80 °C until sEV isolation. Healthy control plasma samples were collected from 14 consented adult volunteers, aged 33 to 65, at Loyola University Chicago Health Sciences Campus.
Extracellular Vesicle Isolation
Vesicles were isolated via size-exclusion chromatography (SEC) method, modified from Yang et al. 38 Depending on sample availability, 200-600 µL of PPP samples were used. Platelet-free plasma (PFP) was prepared by centrifugation at 2000×g for 15 min followed by 10,000×g for 30 min at 4 °C. The PFP was centrifuged at 2000×g for 10 min then the supernatant was filtered using 0.2 µm filters (ThermoFisher, Cat# 723-2520). The filtrate was added to the SEC column consists of Sepharose CL-4B (Sigma, Cat# CL4B200-100ML) in a 1:1 vol/vol ratio and centrifuged for 15 s (Corning, Cat# 6765/C1501 microcentrifuge, 2000×g). Multiple columns were used for individual samples if needed. The exclusion volume off the column should only contain particles sized between 30-200 nm and ranged between 150-500 µL in volume. The sEV samples were diluted 4000-fold in PBS and particle size distribution was analyzed with NanoSight NS300 (Malvern Panalytical, UK) by capturing 150 s of light scatter videos per sample.
Extracellular Vesicle Size Distribution Analysis
The sEV particle size (0-500 nm in 1 nm increment) and corresponding particle numbers were generated by NanoSight NS300 Software. The particle numbers of each size were converted to percentage of total particle number, then the percentage numbers of all technical repeats were averaged to generate a line graph of EV size distribution. Two intersections, at 65.5 nm and 108.5 nm, were selected as interval limits and percentage distribution in three subpopulations of sEV by size (0-65.5 nm, 66.5-108.5 nm, and greater than 108.5 nm) were determined (eg Figure 1C).

PE patients have more heterogeneous, lower concentration and larger sEVs compared to healthy controls. The sEV samples were isolated from platelet-free plasma (PFP) and analyzed with NanoSight NS300. The sEV mean size (A) and sEV concentration (B) were measured; data bars represent mean ± SEM (n = 6-8, samples excluded due to technical error). *p ≤ 0.05, **p ≤ 0.01 by Kruskal-Wallis test with Dunn's correction for multiple comparisons. The sEV size distribution (C) was evaluated by setting the range for small to large particles, and the distribution quantified as a percentage of each subpopulation.
RNA Isolation
Total RNA was isolated from up to 500 µL sEV samples. The sEVs were lysed and homogenized in Qiazol (Qiagen, Cat# 79306) in 1:3 ratio (vol/vol), and total RNA was isolated by phenol/chloroform phase separation and precipitation in ice-cold isopropanol per the manufacturer's protocol. The pellet was washed with ice-cold 75% ethanol in RNase free water, air dried, and resuspended in 20-30 µL of RNase/DNase free water. The concentration of RNA was measured with Nanodrop 8000 (ThermoFisher, Waltham, MA).
Small RNA Sequencing
RNA samples from (100 ng each, measured by NanoDrop) from eight individual donors for each PE severity group and healthy controls were pooled for small RNA sequencing at Northwestern University Sequencing Core. Small RNA library prep and sequencing were conducted at the Northwestern University NUSeq Core Facility. To start, 1 ng of input RNA was used (Agilent 2100 Bioanalyzer and quantity determined by Qubit). The NEXTFLEX Small RNA-Seq Kit from PerkinElmer (now Revvity) was used to build sequencing libraries. First, 3’ adapter and then 5’ adapter was ligated to small RNAs in the samples. After ligation, a reverse transcription step was carried out to generate single-stranded cDNA. A PCR step is then conducted to amplify the cDNA and incorporate Illumina adapter sequences including index sequences. The amplified cDNA constructs were then bead purified prior to sequencing. The Illumina HiSeq 4000 NGS System was used to generate single end 50 bp reads. DESeq2 approach as established by Anders and Huber was used to normalize the read counts. 39 The read depths were 1 million (submassive and healthy controls), 3 million (massive), and 8 million (low-risk).
Pathway and Network Analysis
For pathway analysis, the miRNAs of interest were input into MiRDB Database to generate a list of validated and predicted targets of the miRNA.40,41 Targets with a Target Score of 80 or higher were analyzed for their function and possible pathways involved in KEGG pathway database. 42 Pathways relevant to PE pathology or possible comorbidity pathologies were selected and displayed in Tables 2-4. For network analysis, a list of miRNAs was input into Mienturnet and network miRNA/gene analysis performed based on miRTarBase. 43 Number of interactions was set at 1 or 2, p value was set at 0.01 or 1, both strong and weak evidence were selected.
Statistical Analysis
Statistical analyses were conducted with GraphPad Prism 9.0 software. For data not normally distributed, Kruskal-Wallis test with Dunn's correction for multiple comparison was used to compare between two or more conditions. Values were expressed as mean plus or minus standard error of the mean (SEM). Statistical significance between groups was set at p ≤ 0.05. Specific assessment for small RNA Sequencing analyses is described in the appropriate sections.
Results
Previous pilot studies in our lab demonstrated that ultracentrifugation method for sEV isolation of small clinical samples (<2 mL) cannot provide sufficient sEV yield to study their cargo or function. This was mainly due to the large g-force that can damage the vesicles and the viscosity of clinical samples (plasma or serum) that makes it difficult to pellet the vesicles. To overcome these limitations, we developed the size-exclusion chromatography (SEC) spin column method of sEV isolation, which preserves the volume of the sample and causes less damage to the vesicles due to smaller g force applied. Indeed, as shown in Supplementary Figure 1, SEC generates almost 10 times higher sEV yield and ∼32-fold increase in sEV miR-146 from the same amount of plasma sample compared to ultracentrifugation.
However, this SEC spin column method is only suited for small sample volumes. Further experiments comparing the sEV pellet and supernatant from ultracentrifugation and size-exclusion methods are needed.
Samples from 67 pulmonary embolism patients and 14 healthy controls were collected in this study. Following sample exclusion, eight samples from each group were used for small RNA sequencing analysis. Table 1 depicts the demographic information of the PE patients, collected through the review of patient EMR. PE patients were categorized based on the AHA guidelines 4 and classified as low-risk (n = 24; 35.8%), submassive (n = 34; 50.7%), and massive PE (n = 9; 13.4%). NanoSight analysis of the 81 sEV samples showed that PE patients have lower particle concentration (PE: mean 5.50 × 1011 particles/mL, interquartile range (IQR) 3.50-6.61 × 1011 particles/mL; control: mean 8.40 × 1011 particles/mL, IQR 5.20-11.43 × 1011 particles/mL) and larger size (PE: mean 93.65 nm, IQR 85.60-98.00 nm; control: mean 73.19 nm, IQR 66.75-78.83 nm) compared to healthy controls.
Representative Distribution of Co-Morbidities in Pulmonary Embolism Patients.a

Abbreviations: PESI, pulmonary embolism severity index 61 ; COPD, chronic obstructive pulmonary disease; DVT, deep vein thrombosis; IQR, interquartile range.
The above analysis of the co-morbidities and demographics is based on the available data from 67 patients representing the cohort analyzed.
The RNA studies were conducted in a single-blinded manner using 32 samples from the four groups. Eight samples from each PE severity group and healthy controls were randomly selected for further analysis (Since there were only 9 samples available in the massive group, to match these, similar numbers from other groups were used). Due to the small sample size for each severity group, we were not able to draw any conclusions on potential sex or ethnicity difference among the severity groups. However, we did notice a trend in increasing age, body mass index and some other thrombosis-related parameters as well as co-morbidities with increasing PE severity (Supplementary Table 1). As shown in Figure 1, sEVs from all of these PE subgroups (n = 8) have larger size than the healthy control, and sEVs from the massive PE group have lower sEV concentrations relative to the healthy controls. These results are consistent with those from the entire population (see above). Furthermore, when analyzing the size distribution of sEV samples from the four groups, we saw stark differences between the healthy control sEVs and PE patient sEVs in that the sEVs from healthy controls are much more homogenous in size (Figure 1).
We then isolated RNA from all 32 sEV samples and pooled 100 ng RNA per sample (quantified by Nanodrop) from the randomly selected eight samples from each group. A total of 1 ng RNA by Bioanalyzer from each pool was used for small RNA sequencing. In total, the RNA sequencing results provided data on 2653 miRNAs with read counts ranging from 0 to 981,037 reads (healthy control: 0-80,970, low-risk PE: 0-981,037, submassive PE: 0-441,495, massive PE: 0-253,653). There were 220 miRNAs with more than 2000 normalized read counts in at least one group. Out of the 220 miRNAs, we elected to focus on 73 miRNAs with more than 10-fold differences between two groups, and we looked for patterns among PE severity groups (Table 2-4). Figure 2 and Table 2-4 show the most differentially expressed miRNAs between massive PE versus low-risk PE, low-risk PE versus healthy control and submassive PE versus healthy control, respectively. There were 34 miRNAs meeting the criteria between massive PE and low-risk PE groups listed in Table 2. Interestingly, not all miRNAs have significantly increased expression with higher PE severity. We observed two other patterns of expression (Tables 3-4), a class of miRNAs that increase in expression level between healthy controls and low-risk PE, but then decrease in submassive and massive PE (Figure 2B), and miRNAs that peak in submassive samples, decreasing again in massive PE (Figure 2C). A few miRNAs showed sequential increase from healthy to massive PE and others showed higher levels in 2 or all 3 PE severity groups compared to healthy controls (Data not shown). Many of the identified miRNAs are known or predicted to be involved in pathways relevant to pulmonary embolism pathophysiology, including platelet aggregation, coagulation, immune response, ischemia, and cell death, which warrants further investigation (Figure 3). Network analysis on the top 34 and top 10 miRNAs that increase with disease severity showed many target genes involved in immune and cell death pathways regulated by more than one miRNA (Figure 3 and Tables 5-6). Analysis of miRNAs from Tables 3 and 4 generated similar results (Figure 4).
Most Differentially and Highly Expressed Relevant miRNAs Between Massive PE sEV and low-Risk PE sEV.

Most Differentially and Highly Expressed Relevant miRNAs Between low-Risk PE sEV and Healthy Control sEV.

Most Differentially and Highly Expressed Relevant miRNAs Between Submassive PE sEV and Healthy Control sEV.

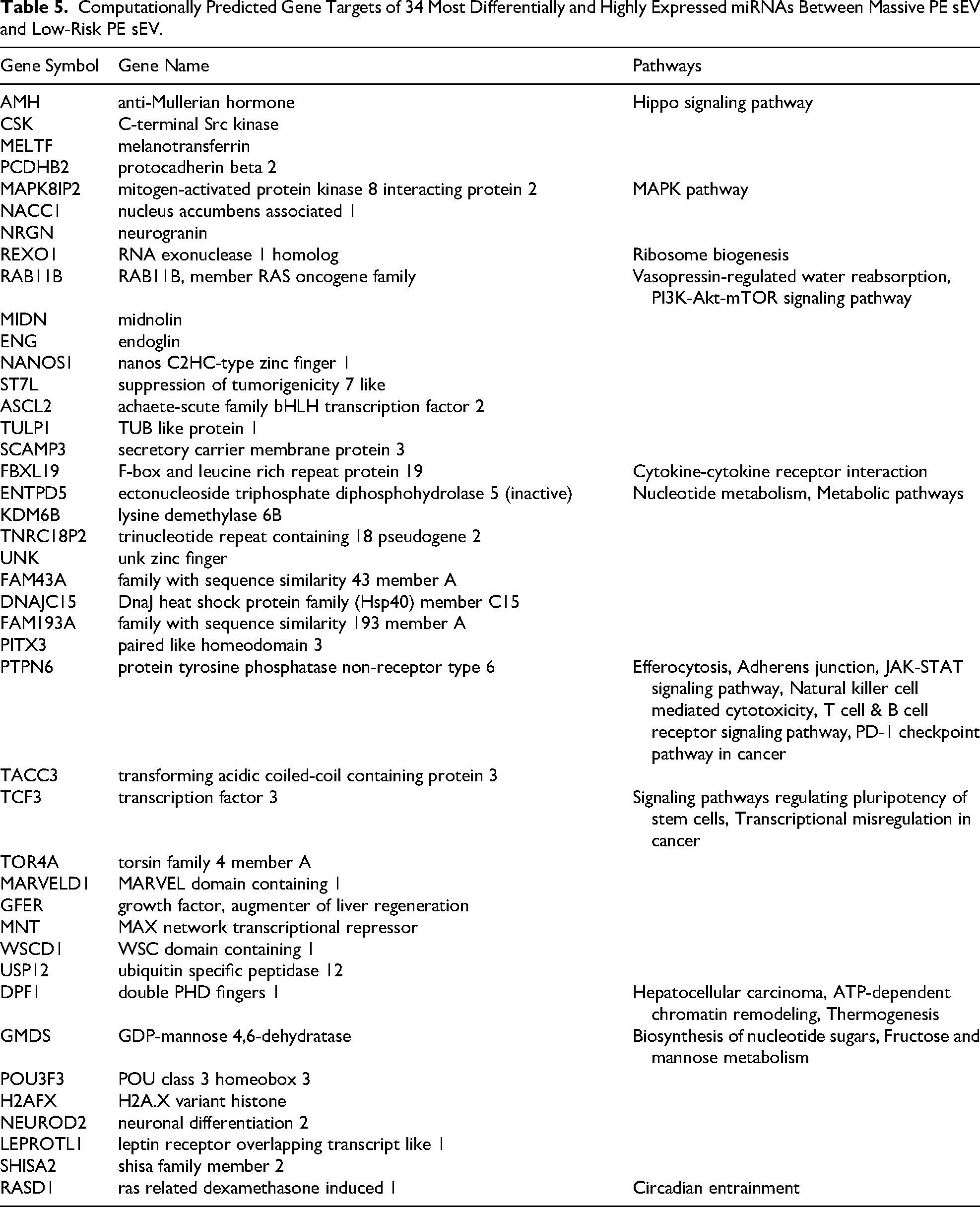
Computationally Predicted Gene Targets of 34 Most Differentially and Highly Expressed miRNAs Between Massive PE sEV and Low-Risk PE sEV.
Computationally Predicted Gene Targets of 10 Most Differentially and Highly Expressed miRNAs Between Massive PE sEV and Low-Risk PE sEV.


Three different patterns of miRNA levels observed across healthy and PE severity groups. (A) 34 most differentially expressed miRNAs that increase with disease severity as potential PE progression biomarkers. (B) Top 5 miRNAs differentially expressed between low-risk (increased in low-risk) and healthy control groups that decrease levels with increasing disease severity to submassive or massive. (C) Top 3 miRNAs differentially expressed between submassive (increased in submassive) and healthy control groups.

Gene network analysis of most differentially expressed miRNAs between massive PE and low-risk PE. Mienturnet analysis on (A) Top 34 most differentially expressed miRNAs between massive PE and low-risk PE with minimum of 1 interaction between miRNAs and target genes and p = 0.01, (B) Top 10 most differentially expressed miRNAs between massive PE and low-risk PE with minimum of 2 interactions between miRNAs and target genes and p = 0.4, (C) Top 3 most differentially expressed miRNAs between massive PE and low-risk PE with minimum of 2 interactions between miRNAs and target genes and p = 1.

Gene network analysis of most differentially expressed miRNAs between submassive PE, low-risk PE and healthy control. Mienturnet analysis on (A) Top 5 most differentially expressed miRNAs between low-risk PE and healthy control with minimum of 2 interactions between miRNAs and target genes and p = 0.1, (B) Top 3 most differentially expressed miRNAs between submassive PE and healthy control with minimum of 2 interactions between miRNAs and target genes and p = 1.
Discussion
PE is a complex disease characterized by disruption in fibrinolysis, coagulation, inflammation, hemodynamic insults, and vascular damage. 3 Given the challenges associated with PE diagnosis and risk stratification, it is imperative to identify biomarkers involved in PE pathophysiology. Biomarkers that are directly related to the underlying mechanisms or pathogenesis are more likely to serve as predictors. Further, identification of such can provide insight into mechanisms and therapeutic targets. Currently employed laboratory markers such as D-dimer, troponin, and BNP, all lack specificity for PE. Elevation of these markers may be caused by other pathophysiologic states such as DVT, end-stage renal disease (ESRD), myocardial infarction (MI), and heart failure. Despite the effort in miRNA biomarker discovery, most miRNAs identified in serum or plasma have a similar problem. Furthermore, circulating RNAs can be degraded which affects detection and quantitative analysis. Therefore, we proposed an alternative approach of assessing the miRNA content of circulating sEV miRNAs as more stable and specific biomarker candidates. The vesicles protect endogenous miRNAs against degradation in circulation and subpopulations of sEVs can be fractionated via surface markers to improve the identification of origin of these sEVs as well as their miRNA cargo. 18
In this study, we piloted the methods for survey analysis of up to 2653 miRNAs, using RNA-Seq as an unbiased method of looking at a maximum number of miRNAs, and the results provide the impetus to further study select miRNAs as candidate biomarkers.
This is the first study of sEV-associated miRNAs in samples from PE patients after stratification for PE severity for miRNA biomarker discovery. A previous study also used small RNA sequencing to study plasma miRNAs from acute and chronic PE patients. 44 They obtained data for 299 known mature miRNAs, among which, let-7i-5p and miR-320a have shown association with acute PE and chronic PE, respectively. Interestingly, of the top 4 miRNAs identified in that study, only miR-320a-5p was detected above 2000 normalized counts in our study, while let-7i-5p, miR-320b and miR-1291 were all below 200 normalized counts. We believe the mixture of samples from patients of differing PE severity (samples not stratified) in previous studies contributed to variability and confounded specificity analysis of the potential biomarkers identified. We found different patterns of miRNA expression across different PE severity groups (Figure 2). These patterns can be useful to survey disease stage and progression but would be masked when patients having different PE severities are grouped together. Future sequencing studies can compare the differences between stratified and unstratified PE severity groups to determine the extent of the masking effect from pooling different PE severity groups.
We used pooled RNA samples for our RNA sequencing to reduce RNA sequencing cost and gain a macroscopic view of the differential miRNA expression patterns within and across the PE severity groups. We noted that quantitation of RNA was different between Nanodrop 8000 and bioanalyzer (Agilent 2100). The bioanalyzer results showed that samples of different PE severity contained different percentages of small RNAs and large non-coding RNAs with lower proportions of miRNAs in the total RNA population as PE severity increases (Supplementary Figure 2). This is associated with a higher proportion of larger RNAs and can contribute to variation of miRNA detection across samples. We thus calculated RNA concentrations for the RNA sequencing based upon the small RNA measurements from the RNA-Seq core Bioanalyzer and employed small RNA sequencing methodology (Methods). The sEV analysis showed RNA/sEV particle (PE: mean 4.66 × 10−9 ng/particle, IQR 1.40-3.22 × 10−9 ng/particle; control: mean 1.26 × 10−9 ng/particle, IQR 0.74-1.76 × 10−9 ng/particle, p = 0.03) as well as estimated average sEV particle volume (PE: mean 4.56 × 106 nm3, IQR 3.57-6.08 × 106 nm3; control: mean 2.61 × 106 nm3, IQR 1.94-3.17 × 106 nm3, p = 0.0006) are significantly higher in PE patients compared to healthy controls. We speculate that the higher extent of cellular stress and/or damage in the more severe disease stages leads to more RNA molecules being packaged into sEVs and secreted into circulation. This is consistent with the literature.45–47 Increased RNA packing could also explain why sEVs from PE patients have larger size than healthy volunteers, although when also taking RNA analysis into account, the proportion of miRNAs in the RNA population may not have a significant role in different expression levels across PE severity groups (ie as RNA increases, the proportion of miRNAs decrease).
As mentioned, several miRNAs have been identified as potential PE biomarkers. The first miRNA reported as potential PE biomarker was plasma miR-134 in 2011 by Xiao et al. 12 The study compared plasma samples from 32 acute PE patients, 32 healthy controls and 22 non-PE patients. Using microarrays and qRT-PCR, they found miR-134 expression is significantly higher in PE patients than both healthy and non-PE subjects. ROC curve analysis showed miR-134 can be used to distinguish between acute PE and healthy and non-PE patients. However, others report that miR-134 is increased in the peripheral blood mononuclear cells of unstable angina pectoris patients. 48 They pointed out the discrepancy could be due to the different source of the samples (plasma vs peripheral blood mononuclear cells). Further studies were needed for confirmation, and the concern for translating miRNA analysis into clinical setting remains. Later, plasma miR-28-3p, miR-27a/b and miR-221 were also identified as potential PE biomarkers, but they all lack specificity to PE. The level of miR-28-3p also increases in Type 2 diabetes mellitus and gastric cancer,49,50 miR-27 levels can be affected by ventricular hypertrophy, 51 and miR-221 levels increase in acute myocardial infarction and pulmonary arterial hypertension.52,53 Interestingly, Kessler et al used similar method of microarray screening and qRT-PCR validation, and they reported that serum miR-1233-3p levels can be used to differentiate between PE and several other similar cardiovascular diseases, including acute DVT, acute non-STEMI, chronic non-thromboembolic pulmonary hypertension. 15 Thus, to a first approximation, miR-1233-3p remains a candidate biomarker, possibly with specificity for PE. Interestingly, our RNA sequencing result of the sEV RNAs show very low levels of miR-1233-3p in all severity groups. We speculate that this is mainly due to the different sources of the samples, that serum levels of miR-1233-3p are very different from levels of miR-1233-3p in plasma EVs. This may be related to the sources of the miR-1233-3p (for example, from cells that do not secrete sEVs, or dying cells, apoptotic bodies, etc), and further study is needed to address this.
Patients included in our study were recruited by the institutional PERT team. None of the subjects included in the study had active COVID-19 at the time of recruitment. Despite our careful sample selection process, aimed to exclude factors potentially influencing miRNA expression (details provided in the Methods section), it remains plausible that some subjects had preexisting changes in various miRNAs. Follow-up studies are planned to validate these specific patterns of expression across individuals in a broader patient population. Furthermore, these patient samples offer opportunities for stratification to explore miRNA expressions associated with comorbidities, such as cancer and COVID-19, as well as to investigate treatment responses and potential adverse effects.
As mentioned, RNA biomarkers from sEVs have been FDA-approved and found useful in predicting prostate cancer. 36 More recently, a clinical study analyzing levels of three RNAs, TMPRSS2:ERG, PCA-3 and SPDEF, from urine-derived sEVs, found these to provide predictive information about patients suspected of prostate cancer, and the EPI-LDT test was approved by FDA in 2019.37,54 It has also been shown that circulating sEV miRNAs can be predictive of changes in toxic effects in hepatotoxicity and after chemotherapy, and this relates to miRNA-target genes that affect drug metabolism.8,55
Our study found that many differentially expressed miRNAs have increased levels with higher PE severity, while some others have the opposite expression pattern, suggesting different functions of the miRNAs play in different stages of PE pathology. We performed signaling network analysis using Mienturnet and showed that the top miRNAs that we identified as increased in severe disease stages, form a gene regulatory network that includes target genes that are involved in cell death, immune cell activation, cytokine-cytokine receptor interaction, MAPK signaling, apoptosis, as well as solute transport, endocytosis, and metabolism 43 (Figure 3-4 and Table 5-6). Some of these targets are regulated by two or more of the miRNAs, indicative of network regulatory effects, and the presence of uptake and metabolic genes in the network supports a conjecture that miRNA gene networks may modulate the patient response to therapeutics in the clinical setting. The miRNAs with increasing expression with increasing PE severity (34) can be indicators of high levels of cellular dysfunction and inflammation associated with higher PE severity and overall systemic stress. Other processes such as fibrinolysis and coagulation, endothelial dysfunction and hemodynamic changes can be activated by inflammation and play a role in the progression of this pathology. The miRNAs with highest expression levels in low-risk PE and decreased expression with increased PE severity (5) could be involved in the initial thromboembolic injury and immune response and later subside. These miRNAs in particular can play an important role in early PE detection. The miRNAs that increase expression from healthy to submassive PE then decrease in massive PE (3) could be involved in hemodynamic changes. When specific miRNAs in the classes are better studied, we may determine a panel that when combined with other biomarkers can form a comprehensive diagnostic panel for PE, which warrants further investigation. If supported by clinical results, this could lead the unbiased methodology used herein to discovery of miRNA biomarkers for disease progression, drug efficacy and adverse effects in patient populations. Research has shown that miRNA profiles change with acetaminophen treatment and liver toxicity as well as chemotherapy and can be utilized as predictive tools for patient response and adverse effects.55–58 Thus, miRNA biomarkers may also be a significant advance for drug development and pharmacovigilance.
In summary, we show here that sEV isolation can be made feasible using our technique, that sEV-miRNA can be identified by differential expression across stratified PE patient samples, and that specific miRNAs identified to have differential expression exhibit specific patterns of expression. Further, the miRNAs dysregulated in PE map to targets that make sense in the context of pathophysiological mechanisms known to be involved in PE. We expect that further research and testing with a larger number of clinical samples, along with RNA-Seq analysis of individual samples, will potentially identify a panel of circulating sEV-derived miRNAs that provide clinically useful information for detecting PE, predicting severity, following progression, treatment responses, and adverse effects. The methodology of circulating sEV-derived biomarker discovery will likely be useful for other diseases, including lung cancer, ESRD, and other cardiopulmonary diseases, and it is not limited to miRNAs. The use of RNA sequencing provides an unbiased and highly sensitive technology (can detect as low as 60 copies of thousands of RNAs) to measure miRNA biomarkers from plasma sEV samples. This is superior to both microarrays and targeted qRT-PCR screening in that it captures information pertaining to a large number of markers in parallel. Dedicated instruments with rapid turnaround, increased throughput, and onsite monitoring should be developed to improve the cost-effective detection and diagnosis of PE as well as other illnesses. Combining this with practical, high yield sEV isolation and stratified patient populations allowed us to identify unusual patterns of expression that might similarly be employed to predict disease stage and response to treatment in patients with diverse ailments.
Limitations
One major limitation of this study is that we pooled 8 individual samples in each severity group. Although this provided an overview of the miRNA profiles across the severity groups, individual variations can be masked by this approach and extreme cases could skew the pooled sample. Further, the sample size of this study is small, and thus very limited in patient population representation and studies that include more patients are needed to validate the miRNAs identified in our study. As the cost of RNA sequencing comes down and throughput increases, the methodology will not be such a hindrance, and future studies will include larger sample sizes as well as analysis of individual samples while taking into account the effects of patient demographics and comorbidities.
Some reports suggest that some circulating RNAs are stable and are probably protected by extracellular vesicles or protein binding.9,21 RNases present in circulation can degrade free-floating RNAs, 59 so it is unlikely that other circulating biomarker studies include free-floating miRNAs. However, quantitative studies are needed to discern the proportions of RNA in circulation that are protein-bound or packaged in vesicles, including larger vesicles, which we did not explore. Our study focuses on the miRNAs packaged into sEVs, which may yield different results from those analyzing plasma or serum RNA, but both approaches aim to identify miRNA biomarkers in circulation that can be developed clinically.
The network analysis is limited by the assumption that the miRNAs entered into the calculation are interacting with each other. However, in the clinical or physiological setting, this may not be true as the origin of the miRNAs are unknown for now, and they can be expressed in different cells and thus may have no direct interaction. Further, the p-value used for Mienturnet/miRTarBase is affected by the number of miRNAs entered in the calculation as well as the number of targets these miRNAs have, which can affect the p-values generated for miRNA/target interactions. These values are simply predictions of interactions in a specific situation, namely miRNAs and their target mRNAs being present together in cells and likely do not have the usual predictive power in this scenario. Future analysis will take into account the different cell types that the miRNAs are expressed in for better evaluation of interactions with gene targets. Tools for evaluating interactions in complicated diseases will have to be developed, fed with data from different cell types and the EVs derived from them, and data from quantitative measurements of target mRNA levels in order to give actionable predictions. We look forward to these developments.
Finally, the sEVs we isolated may contain platelet-derived sEVs that although generally larger in size, do overlap with the size fraction that we studied. Production of platelet-derived sEVs can be activated by thrombotic events. 60 The SEC spin column and filtration procedures we adopted technically exclude particles outside the range of 30-200 nm, but our protocol cannot separate platelet-derived EVs from non-platelet-derived sEVs. This may also explain the larger size of sEVs from PE patients compared to healthy subjects and further study is needed to determine the prevalence of these sEVs and their roles and utility.
Conclusion
In this study, we demonstrate that circulating sEVs contain distinct miRNA signatures that correlate with PE severity, supporting their potential as stable and specific biomarkers for diagnosis and disease monitoring. Using a novel high-yield SEC method optimized for clinical plasma samples, we achieved efficient isolation of sEVs and comprehensive profiling of their miRNA cargo via miRNA sequencing. Differential expression and network analyses revealed that these miRNAs are involved in key biological pathways underlying PE pathogenesis, including coagulation, inflammation, apoptosis, and vascular injury. These findings establish a foundation for developing sEV-derived miRNA panels to enhance diagnostic precision and risk stratification in PE. Future studies involving larger, stratified patient cohorts and individual sample validation are needed to confirm their clinical utility and may facilitate the broader application of EV-based biomarkers in thromboembolic and cardiopulmonary diseases.
Supplemental Material
sj-docx-1-cat-10.1177_10760296251407689 - Supplemental material for Circulating Exosomal MicroRNAs as Potential Biomarkers for Pulmonary Embolism
Supplemental material, sj-docx-1-cat-10.1177_10760296251407689 for Circulating Exosomal MicroRNAs as Potential Biomarkers for Pulmonary Embolism by Chongyu Zhang, Bulent Kantarcioglu, Prakasha Kempaiah, Srdjan Nikolovski, Fakiha Siddiqui, Ahmed Kouta, Fabio Lievano, Thao Doan, Simon Kaja, Amir Darki, Jawed Fareed and W Keith Jones in Clinical and Applied Thrombosis/Hemostasis
Footnotes
Acknowledgments
The authors would like to acknowledge the numerous volunteers at Loyola University Chicago for their blood donations to our ongoing research program. The authors would like to thank Northwestern University Sequencing Core for their services and contributions to the small RNA sequencing and Methods section.
ORCID iDs
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Funding for this work was provided by Loyola University Institutional funds, including from the Cardiovascular Research Center and the Department of Molecular Pharmacology and Neuroscience. Support by Dr John P. and Therese E. Mulcahy Endowed Professorship in Ophthalmology (S.K.) is gracefully acknowledged.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
