Abstract
Abnormal transcriptomic profiles of coagulation genes have been linked to coagulopathies in patients with coronavirus disease 2019 (COVID-19). The objective of the present study was to explore the frequency of genotypes and potential association of polymorphisms in genes encoding coagulation factors with the disease severity in COVID-19 patients.
The patients were clinically categorized into four groups of COVID-19 disease severity (asymptomatic, mild, moderate, and severe). Three variants of genes, involving the coagulation genes rs3136516 (F2 gene), rs6042 (F7 gene), and rs6123 (PROS1 gene), were studied. Polymorphisms were genotyped by Sanger DNA sequencing.
Most of the subjects had moderate COVID-19 infection (n = 53, 62.4%), followed by mild (n = 16, 18.8%), and severe infections (n = 15, 17.6%). The frequency of the rs3136516 AG genotype was considerably higher in non-ICU patients compared to ICU patients (51.3% vs 34.1%, OR 3.167, 95% CI 1.094-9.170, P = .031). Furthermore, the dominant genetic model (AA + AG vs GG) was significantly associated with a decreased probability of admission to the ICU in COVID-19 patients (OR 0.340, 95% CI 0.127 - 0.905, and P = .028). No other variants of the coagulation genes studied were found to be associated with the severity of COVID-19 disease, admission to the ICU, and mortality (P > .05).
The rs3136516 AG genotype could predispose COVID-19 patients to increased disease severity and therefore admission to the ICU, while the dominant genetic model (AA + AG vs GG) of rs3136516 exerts a protective role.
Introduction
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is the causal agent of novel coronavirus disease 2019 (COVID-19). 1 COVID-19 symptomatology can take the form of asymptomatic to severe infections, often requiring hospitalization and admittance to intensive care units (ICUs) with a substantial increase in mortality. 2 The prevalence of severe COVID-19 is approximately 23% according to a systematic review and meta-analysis by Li et al (2021). 3
Among the classic characteristics of COVID-19 is the strong inflammatory response observed in a number of patients, particularly those with severe disease. 4 This is followed by systemic and potentially fatal clinical complications such as acute respiratory distress syndrome, arrhythmia, acute cardiac injury, heart failure, acute kidney injury, thrombosis, coagulopathy, and multi-organ failure, which is the reason why COVID-19 is regarded as a multi-system disorder and demands multi-disciplinary intervention.5–7
SARS-CoV-2 directly affects hemostasis, or can indirectly induce a pathobiological milieu of hypoxemia and hyperinflammation, causing dysfunctional coagulation and thromboembolic manifestations in COVID-19. These complications occur both in microvasculature and microvasculature owing to development of a systemic hypercoagulable state. This state is often known as “COVID-19-associated coagulopathy” which marks the derivation of multi-organ injury from COVID-19.1,8–10 Several research findings have indicated hemostatic imbalances such as COVID-19-associated coagulopathy11,12 and is reflected by elevated D-dimer and fibrinogen levels. The pathogenic mechanisms of COVID-19-associated coagulopathy are complicated and influenced by interactions between inflammatory responses and coagulation cascades ie thromboinflammation or immunothrombosis. Importantly, the overactivated systemic inflammation stimulating cytokine storm, critical illness, and other significant underlying predisposing factors might all increase the risk of developing such thromboembolic or other cardiovascular complication in COVID-19 patients. 13
Few studies have explored the realm of genetic polymorphism in coagulation genes and risk of severe forms of COVID-19. 14 An investigation by Lapić et al (2022) reported an association between polymorphism in the integrin beta-3 (ITGB3) gene and severe COVID-19. They further found a promoter polymorphism in the fibrinogen gene (ß-Fbg: 455G > A homozygous form) to be present in severe COVID-19 patients. 14 To be specific, the findings revealed 3.5-fold higher risk of severe COVID-19 disease in those who harbored at least one allele of integrin beta-3 PIA2/A2. Integrin beta-3 functions to produce platelet integrin glycoprotein IIb/IIIa (GPIIb/IIIa) complexes which assist in adhesion and stimulation of platelets. The integrin beta-3 PIA1/A2 polymorphism results in overactivation of platelet adhesion and activation via conformational changes involving the platelet integrin glycoprotein IIb/IIIa (GPIIb/IIIa) complex, leading to high thrombogenicity.15,16 This confirms the deleterious impact of such polymorphisms in COVID-19 patients, particularly by destabilizing hemostasis and thereby instigating disease severity. Notably, complications in severe COVID-19 are multifaceted and a plethora of genetic factors may be involved. Hence, it is emphasized to dissect genetic dynamics of proteins linked with coagulopathy complications in COVID-19 cases and thereby disease severity. Insight into underlying role of aforementioned genetic factors might assist in improving clinical management of severe COVID-19 cases and development of individualized therapeutic strategies.
This study was designed and conducted to achieve two main objectives: 1) to capture the genotype frequency of variants of genes encoding coagulation factors (F2 gene: rs3136516, F7 gene: rs6042, and PROS1 gene: rs6123) in patients with diverse forms of COVID-19 severity, and 2) to determine the association of genetic variants of coagulation factor genes with COVID-19 disease severity. Secondary objectives of the investigation were to compare genotypic differences of the aforementioned polymorphisms based on ICU admittance and variations in disease outcomes (ie survival vs death).
Materials and Methods
Ethical Considerations
The study protocol was approved by the Ethics Review Committee (ERC) of the King Faisal Specialist Hospital & Research Centre (KFSH&RC), Riyadh, Saudi Arabia (project number: 2201086). The study was conducted in accordance with the Declaration of Helsinki and its subsequent revisions.17,18 Written informed consent was gathered from all subjects. De-identified patient data and biological samples were utilized to preserve the privacy of research participants.
Study Design and Participants
This cross-sectional research study enrolled in-hospital COVID-19 patients at the KFSH&RC, Riyadh, Saudi Arabia between January 2021 and December 2022. Using World Health Organization (WHO) guidelines, infection with SARS-CoV-2 was confirmed from nasopharyngeal and oropharyngeal swab samples using reverse transcriptase real-time polymerase chain reaction (RT-PCR). Patients were clinically categorized into four groups of disease severity (asymptomatic, mild, moderate, and severe infection). COVID-19 cases who were discharged within 24 h of admission or those unable to provide blood samples for the evaluation of coagulation gene polymorphisms were excluded. Medical records were also reviewed to obtain information regarding patient demographics and clinical data, in particular, disease severity, ICU admission, need for mechanical ventilation, and disease outcomes (survival vs death).
Single Nucleotide Polymorphisms (SNPs)
Three SNPs rs3136516, rs6042, and rs6123 involving the genes F2 (Protein Product: Coagulation Factor II, Prothrombin), F7 (Protein Product: Coagulation Factor VII), and PROS1 (Protein Product: Protein S), respectively, were selected for genotyping. The basis for selection of the aforementioned coagulation factor genes was based on a report by Mast et al (2021), which confirmed the abnormal mRNA transcriptional expression profile of several coagulation genes in COVID-19 cases in contrast to controls, F2, F7 and PROS1 genes, in bronchoalveolar lavage samples and its implication to coagulation-associated pathologies. 19
Blood Sample Collection and Genotyping
Blood samples were obtained from COVID-19-infected hospitalized patients. Blood was drawn into 10 ml tubes containing ethylenediaminetetraacetic acid (EDTA). Genomic DNA from samples was extracted using commercial DNeasy kits (Qiagen, Valencia, CA, USA) as per the manufacturer's instructions. For identification of SNPs in genomic DNA samples, flanking PCR primers were designed using Primer3 software v.0.4.0 (available at: http://bioinfo.ut.ee/primer3-0.4.0/). Genotyping of three SNPs, rs3136516 (F2 gene), rs6042 (F7 gene), and rs6123 (PROS1 gene), was performed by Sanger DNA sequencing (Thermo Fisher Scientific, Carlsbad, CA). The Sanger dideoxy chain termination DNA sequencing method was employed using PCR products as templates, ABI Big Dye terminators, reverse and forward primers (10 pM). After sequencing, ABI, FASTA, and PDF files were assembled, and chromatograms were evaluated to identify the targeted SNPs (rs3136516 (F2 gene), rs6042 (F7 gene), and rs6123 (PROS1 gene)).
Statistical Analysis
Data normality distribution was evaluated using the Shapiro-Wilk test. Categorical data are presented as frequencies and percentages, while continuous data are shown as means and standard deviations (SD) or medians and interquartile ranges (IQRs), depending on data distribution. Comparison of frequency of genotypes among disease severity, ICU, and outcome groups of COVID-19 patients was conducted using Chi-squared (Χ2) tests or Fisher's exact test, where appropriate. Mann-Whitney U-test was utilized for skewed data, while normally distributed data were assessed using the independent samples Student's t-test. One-way analysis of variance (ANOVA) or Kruskal-Wallis H test was utilized for comparing three or more groups. Univariate and multivariate logistic regression analysis was performed to evaluate the association of polymorphisms with COVID-19 severity and outcomes. Results are presented as odds ratios (OR) with 95% confidence intervals (95% CI). P-values < .05 were regarded as statistically significant in two-tailed tests. To identify any deviation in genetic marker distribution, the Hardy-Weinberg equilibrium (HWE) test was also conducted. Statistical analysis was conducted using GraphPad Prism v.9.
Results
Eighty-five COVID-19 patients were studied in this investigation. The mean age of the subjects was 53.69 ± 17.26 years (Range: 18 to 96 years), and 56.5% (n = 48) of them were males. A majority of patients had moderate COVID-19 infection (n = 53, 62.4%), followed by mild (n = 16, 18.8%) and severe infections (n = 15, 17.6%). Forty-five (52.9%) patients were admitted to ICU and nine patients died (10.6%). Demographic and clinical data of COVID-19 patients are presented in Table 1.
Clinico-Demographic Data of Study Patients.
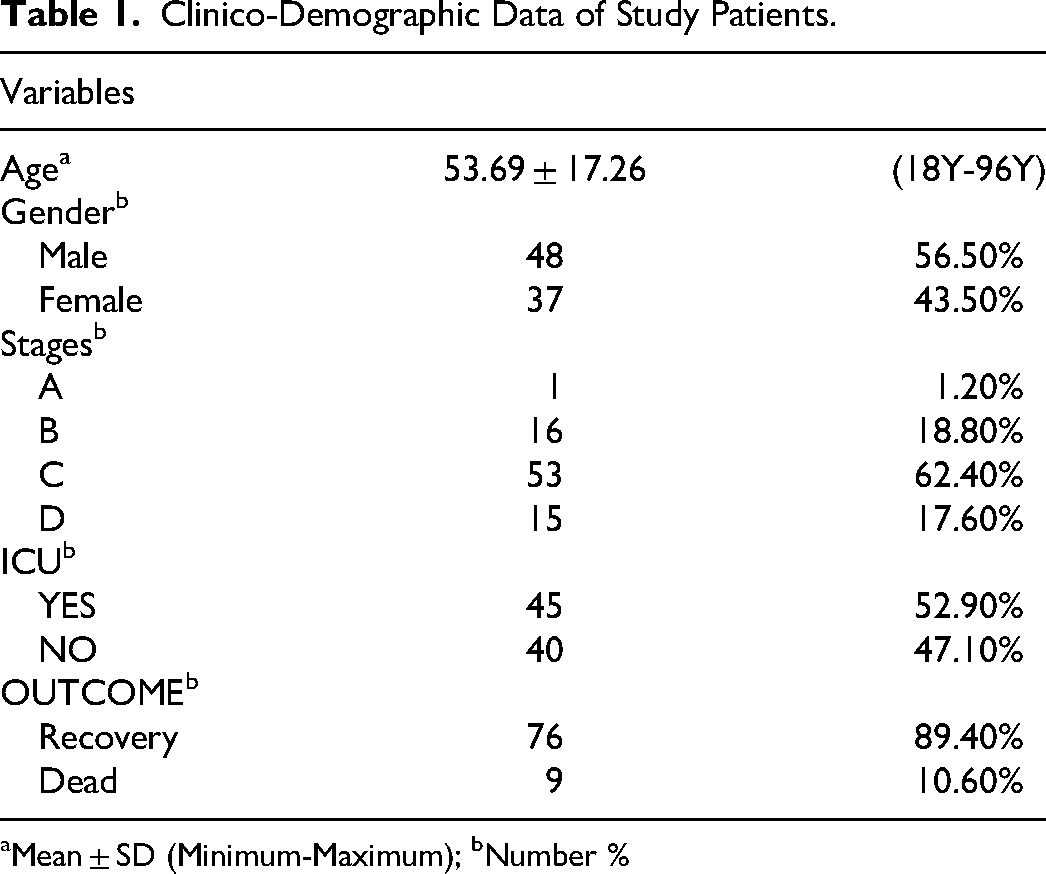
Mean ± SD (Minimum-Maximum); bNumber %
Genotypic Distribution of Coagulation Gene Polymorphisms in ICU and non-ICU COVID-19 Patients
The genotype distribution of the coagulation gene polymorphisms rs3136516, rs6042, and rs6123 in the ICU (n = 44) and non-ICU (n = 39) COVID-19 patients was investigated. The frequency of the heterozygous AG genotype of rs3136516 was significantly higher in non-ICU patients compared to that of ICU patients (genotype frequency 51.3% vs 34.1%, OR 3.167, 95% CI 1.094-9.170, and P = .031). In addition, the frequency of the A allele of variant rs3136516 was higher in non-ICU patients compared to ICU patients (genotype frequency 53.8% vs 39.8%, OR 1.767, 95% CI 0.953-3.274, and P = .070); however, no statistical significance was observed. Also, we observed that the dominant genetic model (AA + AG vs GG) was suggestively associated with diminished risk of susceptibility to ICU admission in COVID-19 patients (OR 0.340, 95% CI 0.127-0.905, and P = .028).
For rs6042, we found that the heterozygous CT genotype represented 15 (34.1%) of ICU cases and compared to 11 (28.2%) of non-ICU COVID-19 patients with no significant difference (OR 0.792, 95% CI 0.306-2.047, P = .630). Moreover, the T allele of rs6042 was identified in 15 (17.0%) COVID-19 cases admitted to ICU in contrast to 13 (16.7%) of non-ICU patients. Regarding rs6123, we did not observe significant genotypic or allelic differences between ICU and non-ICU COVID-19 patients (P > .05). A comparison of genotypic distributions of polymorphisms in coagulation genes in ICU and non-ICU COVID-19 patients is presented in Table 2.
Genotypic Distributions of COVID-19 SNPs Were Compared in the ICU with Those of non-ICU Patients.
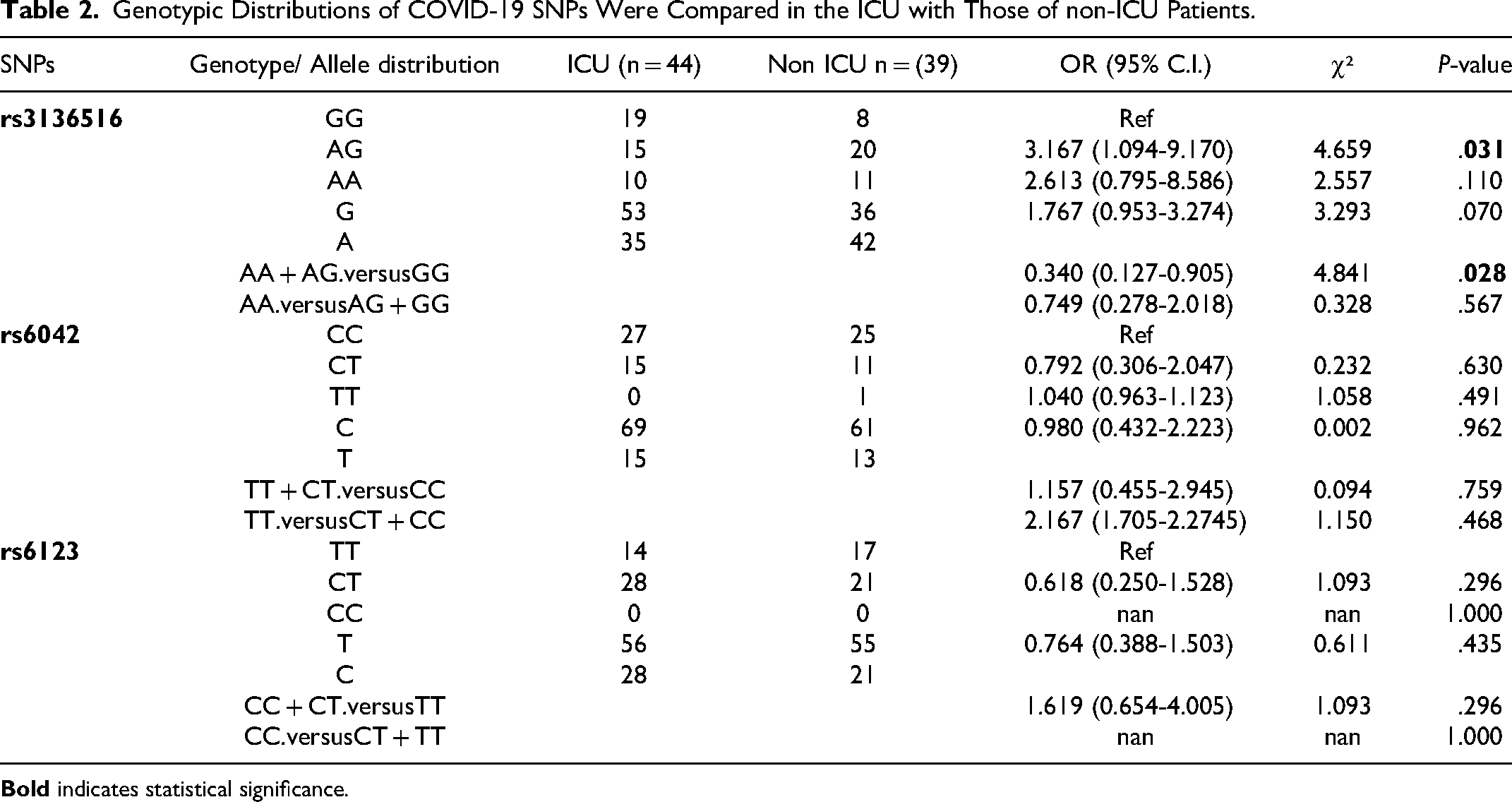
Genotypic Distribution of Coagulation Gene Polymorphisms in COVID-19 Patients with Recovery or Death
Table 3 illustrates the roles of the investigated coagulation gene polymorphisms in terms of survival outcomes (recovery [n = 74] and death [n = 9]) in COVID-19. The homozygous genotypes of the major allele were set as the reference genotype for each polymorphism. Our findings demonstrated that heterozygous genotypes of two polymorphisms (rs3136516 and rs6123) were higher in frequency in COVID-19 cases with outcome of death in comparison to those who recovered; rs3136516 (genotype frequency 44.4% vs 41.9%, OR 0.742, 95% CI 0.168-3.283, and P = .719) and rs6123 (genotype frequency 77.8% vs 56.8%, OR 2.417, 95% CI 0.468-12.473, and P = .470). However, we did not observe statistically significant differences. Conversely, the heterozygous CT genotype of rs6042 was detected in higher numbers in COVID-19 recovered patients compared to those who died, but with no statistical significance (genotype frequency 32.4% vs 22.2%, OR 0.536, 95% CI 0.103-2.783, and P = .710).
Genotype Distributions of COVID-19 SNPs Were Compared in Recovered Patients with Those of Deceased Patients.
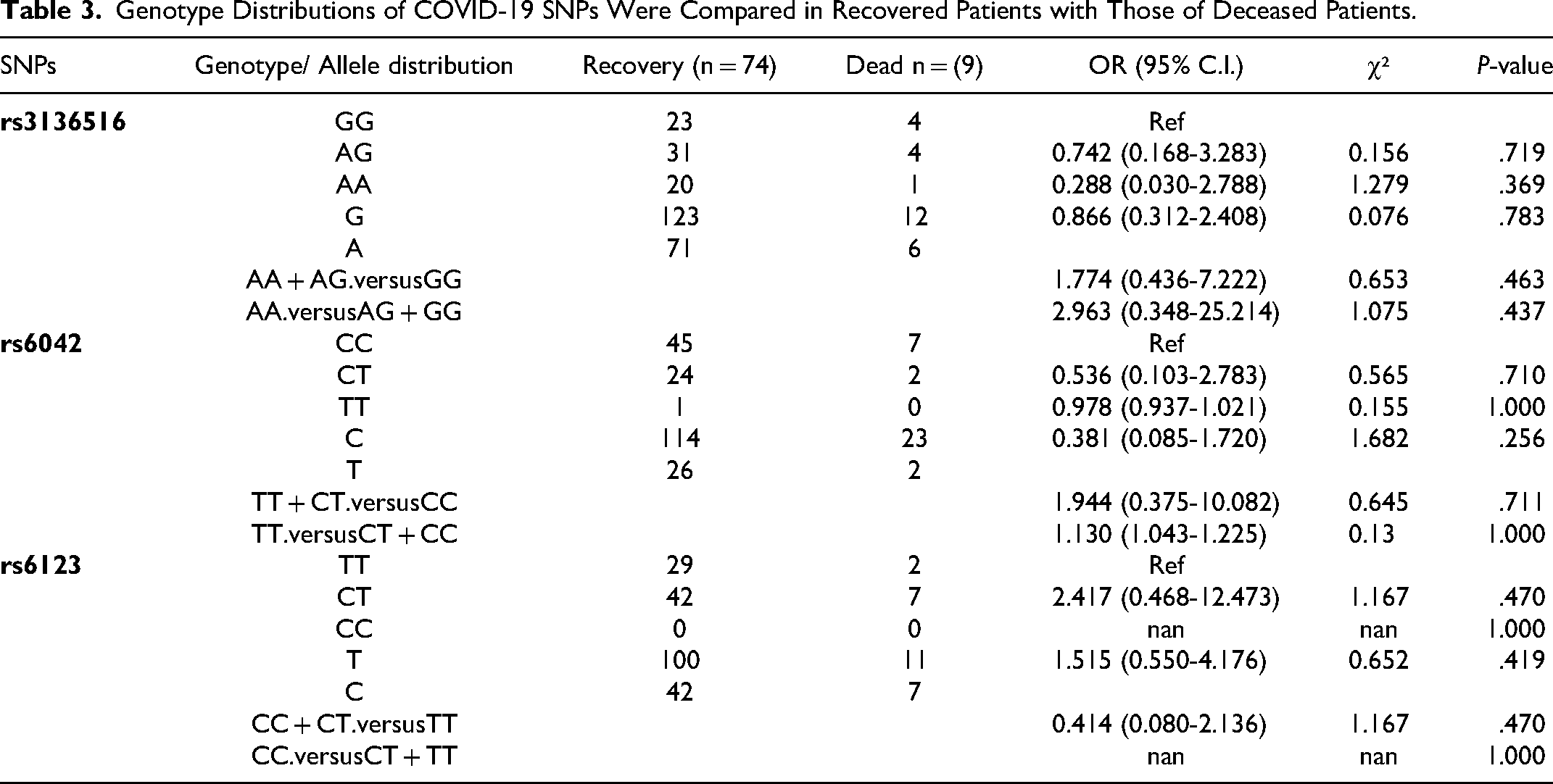
Genetic Variants in Coagulation Genes Across COVID-19 Severity Levels
We combined and compared two groups of COVID-19 patients with varied disease severity. Group 1 comprised of Stage A (asymptomatic infection) and B (mild infection) patients (n = 17), while Group 2 included Stage C (moderate infection) and D (severe infection) patients (n = 66). The results suggested that heterozygous genotypes of all three polymorphisms (rs3136516, rs6042, and rs6123) were considerably higher in number in COVID-19 patients with Stage C and D disease (moderate to severe infection) compared to those with Stage A + B disease (asymptomatic to mild infection); rs3136516 (genotype frequency 42.4% vs 41.2%, OR 0.696, 95% CI 0.181-2.674, and P = .742), rs6042 (genotype frequency 34.8% vs 17.6%, OR 2.30, 95% CI 0.587-9.008, and P = .223) and rs6123 (genotype frequency 59.1% vs 58.8%, OR 0.750, 95% CI 0.230-2.447, and P = .633). The analysis did not reveal statistically significant differences (Table 4).
Genotype Distributions of COVID-19 SNPs Were Compared among Patients with Stages A + B with That of Patients with Stages C + D.
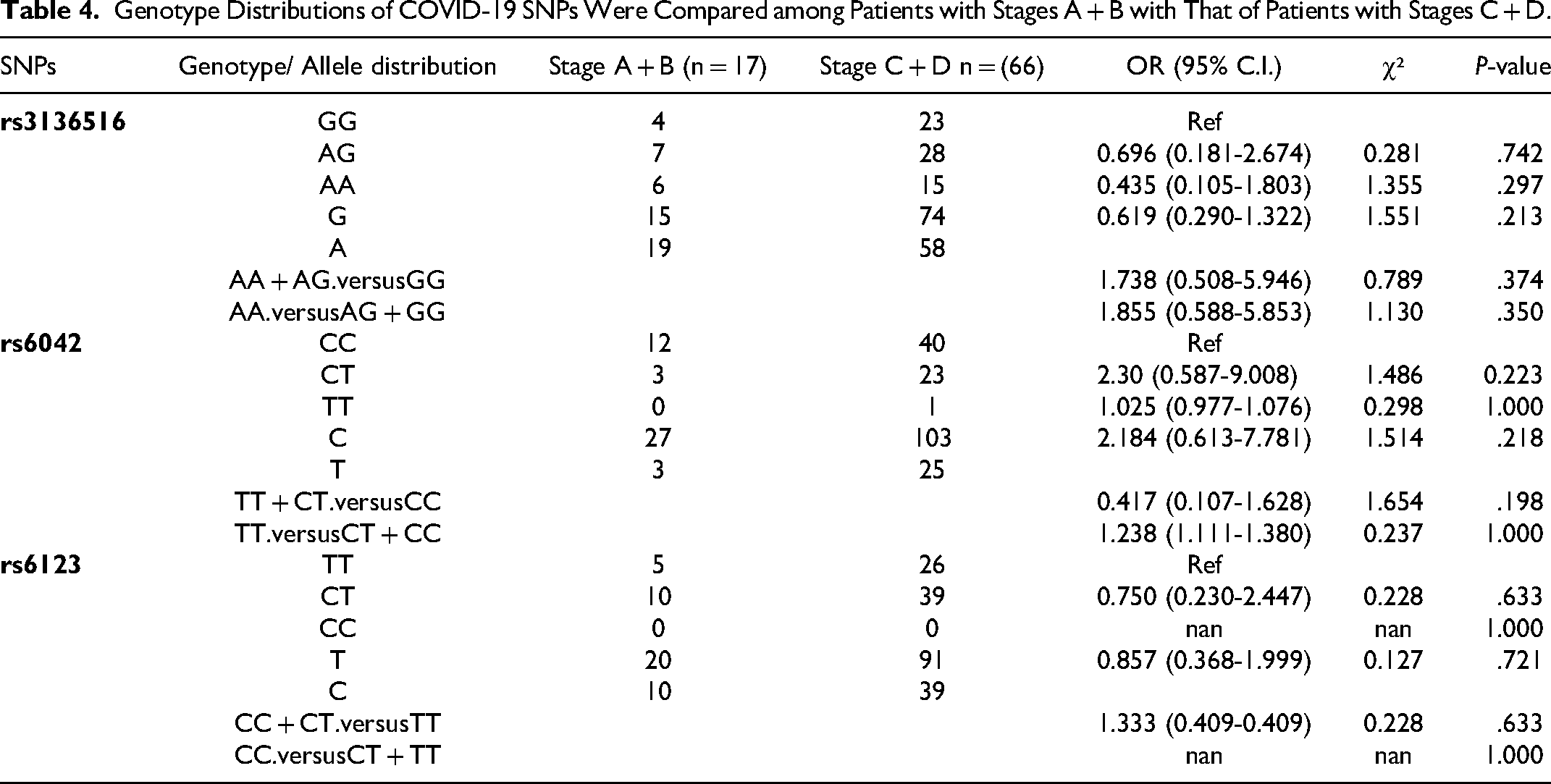
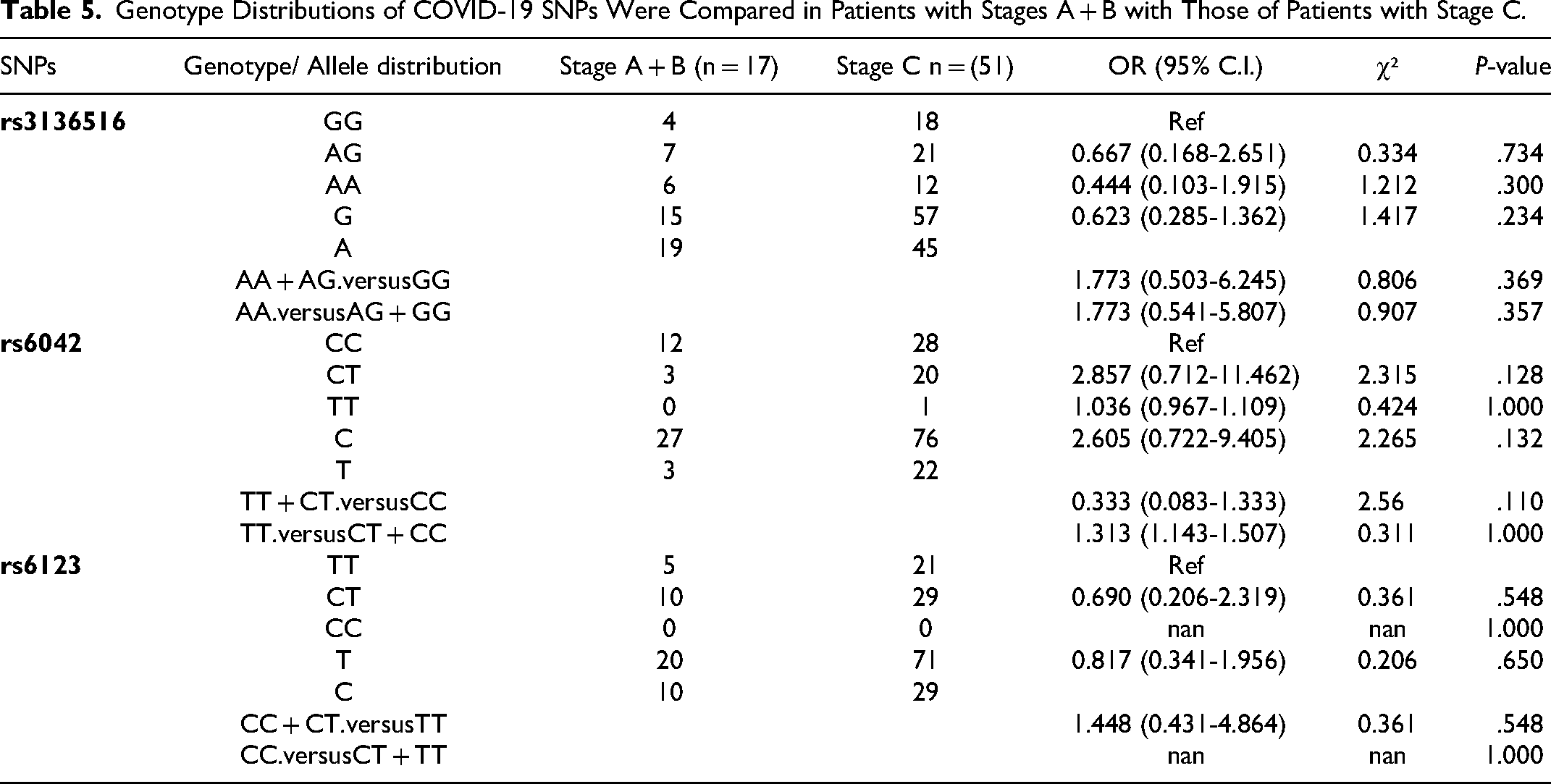
Table 5 displays the frequency distribution of genotypes, alleles, and dominant and recessive genetic models of the analyzed rs3136516, rs6042, and rs6123 polymorphisms between Group 1 which included Stage A + B disease (asymptomatic to mild infection) patients (n = 17) and Group 2, which included Stage C disease (moderate infection) in COVID-19 patients (n = 51). Importantly, for rs6042, we noticed that COVID-19 patients with Stage C disease (moderate infection) were frequent carriers of the CT heterozygous variant genotype compared to those with Stage A + B disease (asymptomatic to mild infection) (genotype frequency 39.2% vs 17.6%, OR 2.857, 95% CI 0.712-11.462, and P = .128). On the other hand, Stage A + B disease (asymptomatic to mild infection) patients had a higher frequency of the rs6123 CT genotype compared to Stage C disease (moderate infection) patients (genotype frequency 58.8% vs 56.9%, OR 0.690, 95% CI 0.206-2.319, and P = .548).
Genotype Distributions of COVID-19 SNPs Were Compared in Patients with Stages A + B with Those of Patients with Stage C.
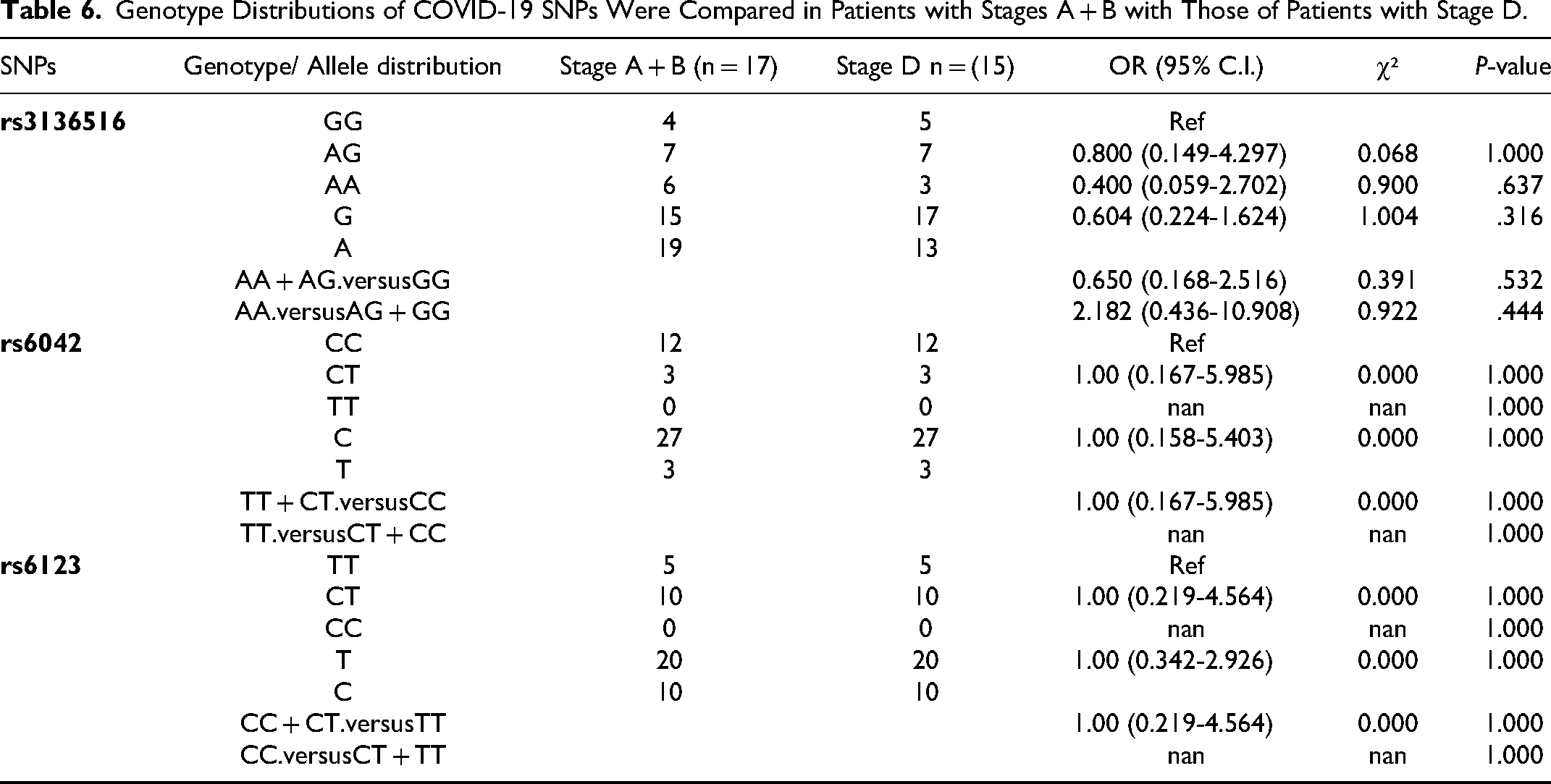
There was no statistical disparity in genotypes of rs3136516, rs6042, and rs6123 when Group 1 which included Stage A + B disease (asymptomatic to mild infection) patients (n = 17) and Group 2 which included Stage D disease (severe infection) patients (n = 15) were compared (Table 6). The findings were: AG genotype rs3136516 (genotype frequency 41.2% vs 46.7%, OR 0.800, 95% CI 0.149-4.297, and P = 1.000), CT genotype rs6042 (genotype frequency 17.6% vs 20.0%, OR 1.00, 95% CI 0.167-5.985, and P = 1.000), and CT genotype rs6123 (genotype frequency 58.8% vs 66.7%, OR 1.00, 95% CI 0.219-4.564), and P = 1.000).
Genotype Distributions of COVID-19 SNPs Were Compared in Patients with Stages A + B with Those of Patients with Stage D.
Discussion
COVID-19 possesses a broad symptomatology spectrum ranging from asymptomatic or mild infection to severe pneumonia evidenced by critical lung parenchymal injury leading to multi-organ failure and mortality. 20 Virus-, host-, and environment-related determinants have been suggested to increase predisposition to severe COVID-19.21,22 Genetics is one such host determinant that has complex relationship with health and disease. 23 The literature implies that genetic architecture is responsible for approximately half of the variation in predicted clinical COVID-19 phenotype. 24 Genetic diversity greatly determines systemic responses to COVID-19 infection and subsequent disease outcomes. Polymorphisms in the ACE2 gene have been linked with COVID-19 disease severity.22,25,26 A number of research studies have specifically demonstrated a role of genetic variants in COVID-19 disease severity phenotypes such as respiratory failure, hospitalization, or mortality.27,28 Hemostatic imbalance is another key aspect of COVID-19 disease severity, resulting in COVID-19 associated coagulopathy. Of late, studies have tapped associations between polymorphisms in coagulation genes and the spectrum of COVID-19 clinical phenotypes.14,29 Nevertheless, the genetic landscape of proteins associated with coagulopathy complications and thereby COVID-19 severity has been less explored.
In the present research, we studied rs3136516 (F2 gene), rs6042 (F7 gene), and rs6123 (PROS1 gene) polymorphisms in COVID-19 patients. The study aimed to determine the genotype frequency of the aforementioned polymorphisms in genes encoding coagulation factors in COVID-19 patients with diverse clinical phenotypes and to establish the association of these polymorphisms with COVID-19 disease severity. In the current investigation, we found a probable connection of the F2 gene (Protein Product: Coagulation Factor II, Prothrombin) rs3136516 heterozygous AG genotype with increased likelihood of ICU admittance in COVID-19 patients. Interestingly, the homozygous rs3136516 GG genotype, in a dominant mode of inheritance, was found to be associated with lesser susceptibility to admission to ICU and could be a protective factor in Saudi Arabian COVID-19 patients. F2 encodes coagulation factor II (prothrombin). The F2 gene rs3136516 polymorphism is reportedly linked with high levels of prothrombin activity and therefore increased risk of thrombosis.30,31
F2 gene polymorphism genotyping could be a sensitive genetic biomarker for thromboembolism and may aid in stratifying COVID-19 cases vulnerable to thromboembolic complications. 32 High mortality rates have been determined in severe forms of COVID-19 cases exhibiting clinically proven (high D-dimer) infection-triggered systemic coagulopathies, such as thrombotic microangiopathy or disseminated intravascular coagulation. To support this finding, anti-coagulant therapy, such as administration of low-molecular-weight heparin, has been indicated to decrease the probability of mortality due to COVID-19, thereby improving prognosis. 8 A research study by Zhou et al (2021) established that anti-platelet and anti-coagulant therapy reduces the odds of critical COVID-19. 33 Mast et al (2021) studied transcriptional profiles in COVID-19 in bronchoalveolar lavage samples to determine molecular mechanisms of COVID-19-associated coagulopathies and found abnormal expression of several coagulation gene mRNAs, including F2 (0.9 vs 0.0), in COVID-19 cases in relation to controls. 19 With evidence from the present study and apparent coagulation abnormalities in COVID-19, it is plausible to contemplate coagulation gene variants such as F2 gene rs3136516 as a risk factor for COVID-19 severity.
Coagulation factor VII is encoded by the F7 gene. 19 Factor VII is a protease that functions to instigate a coagulation cascade by proteolysis culminating in production of thrombin, deposition of fibrin, and stimulation of platelets. 34 High levels of activated factor VII and anti-thrombin complexes have been revealed in COVID-19 cases, highlighting their relationship with COVID-19-associated endotheliopathy and prothrombotic diathesis. 35 Several studies have also delineated the importance of F7 gene variants in identifying large plasma deposits of factor VII, with their role in susceptibility to thrombosis.36–39 Mast et al (2021) documented significant elevation in levels of transcripts encoding factor VII (10.1 vs 0.0, 363.5-fold change) in bronchioalveolar lavage samples from COVID-19 cases in contrast to controls. 19 Such evidence shows the importance of genetic polymorphism, rs6042, in the F7 gene in the regulation of transcriptional patterns related to increases in the procoagulant physiological environment in patients with COVID-19 patients. However, we did not find any association between the F7 gene rs6042 polymorphism and ICU admission, severe infection, or mortality in COVID-19 patients.
Healthy endothelium in lung bears membrane-bound anti-coagulant peptides which function to antagonize thrombin production at several points in the pathway. This encompasses tissue factor pathway inhibitor, thrombomodulin, and endothelial protein C receptor. Thrombomodulin converts thrombin to activated protein C from a pro-coagulant enzyme. Protein S (encoded by PROS1) is a co-factor for activated protein C and helps to diminish thrombin production by inhibiting actors VIIIa and Va and induces an anti-inflammatory effect by interacting with endothelial protein C receptor.40,41 Protein S is mainly synthesized in liver cells; however, the significance of its local production is not yet clear. Panigada et al (2020) described slightly low protein S levels in COVID-19 patients. 42 This finding was confirmed later by Mast et al (2021) who reported a substantial decline in transcripts encoding protein S (3.2 vs 195.3, −54.2-fold change) in bronchioalveolar lavage fluid from COVID-19 patients in contrast to controls. 19 Taken together, these data implicate COVID-19 in downregulation of mRNA transcripts associated with the ability of endothelial anti-coagulant proteins to modulate local thrombin production in COVID-19 patients. In the present study, we did not find any association between the PROS1 rs6123 variant and COVID-19 disease severity features (ICU admission, severe infection, or mortality).
Our research study had certain limitations which should be discussed and which merit cautious interpretation. Firstly, our study comprised of single center data and had a small number of COVID-19 patients which were further categorized into subgroups based on disease severity, ICU admission, and survival outcomes for the purpose of statistical analysis and comparison. We believe that this might affect the generalizability of our findings to a larger cohort of similar patients in Saudi Arabia. Secondly, any causal relationships could not be determined due to the retrospective nature of the study design. Finally, we were unable to include all clinical data, such as comorbidities, coagulation-related biomarkers, and treatments given, that could have acted as confounders or, conversely, further strengthened our findings. Finally, our study did not explore the link between overexpression of coagulation genes and development of thrombosis in COVID-19 patients. This might overlook variability in clinical outcomes associated with thrombosis, resulting in inadequate understanding of how genetic factors influence disease progression. Notwithstanding these study limitations, the potential role of F2 gene variant rs3136516 in the present study could serve as a reference for potential subsequent investigations involving polymorphic coagulation-related genes and identification of additional genetic prognostic indicators of COVID-19 severity. This is of significance for identifying at-risk cohorts, for increasing therapeutic efficacy, and designing tailored treatments.
Conclusion
In conclusion, the findings of the present study suggest that F2 gene (Coagulation Factor II, Prothrombin) heterozygous rs3136516 AG genotype could predispose to COVID-19 severity and therefore ICU admission. Furthermore, the F2 rs3136516 homozygous GG genotype, in a dominant mode of inheritance, is associated with diminished risk of admission to ICU and could be a protective factor in Saudi Arabian COVID-19 patients. Our findings should be confirmed, and further investigations should be carried out involving a larger cohort and in the light of additional clinical data surrounding patients with COVID-19.
Footnotes
Acknowledgements
The authors extend their appreciation to the Deanship of Scientific Research at Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2024R465), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received financial support for this work, as stated in the acknowledgements section.
