Abstract
Background and Aim:
Chronic rheumatic heart disease (RHD) patients who undergo valve replacement with mechanical valves require lifelong anticoagulation. Acenocoumarol, a vitamin K antagonist has a narrow therapeutic range and wide inter-individual variability. Our aim was to investigate the influence of polymorphisms of VKORC1 and CYP2C9 genes on the mean daily dose requirement of acenocoumarol.
Methods:
205 chronic RHD patients, with mechanical heart valves and on acenocoumarol therapy, were recruited. Genotyping for VKORC1 (−1639G>A and 1173C>T) and CYP2C9 (*2 & *3 alleles) polymorphisms was done by PCR-RFLP. There was complete linkage disequilibrium between VKORC1 polymorphisms (r2 = 0.98, D′ = 1.0, LOD = 74.02). VKORC1 genotype distribution for GG/CC, GA/CT, and AA/TT were 57.6%, 36.1%, and 6.3%, respectively. CYP2C9 genotype distribution for *1/*1, *1/*3, *1/*2, *2/*2, and *2/*3 were 78.5%, 14.1%, 6.3%, 0.5%, and 0.5%, respectively. Patients with a wild type of both VKORC1 (−1639GG and 1173CC) and CYP2C9 gene variants required higher acenocoumarol dose compared to those with mutant genotype (P = 0.023 and P = 0.008 respectively). On combined genotype analysis, patients having a combination of wild type of VKORC1 with wild type of CYP2C9 (44.4%) required higher daily dose compared to patients bearing heterozygous VKORC1 (−1639GA & 1173CT) with wild type of CYP2C9 (30.2%, P = 0.008).
Conclusion:
Presence of a mutant allele of VKORC1 (−1639A & 1173T) and CYP2C9 genes increased the odds of requiring a lower mean dosage of acenocoumarol. Studying the combination of genotypes in RHD patients could predict acenocoumarol dose requirement more accurately.
Introduction
Rheumatic fever and chronic rheumatic heart disease (RHD), affecting mainly the young, continue to be a public health concern in rural India and in urban slums. Its prevalence varies anywhere between 0.12 per 1000 population in Cochin, Kerala, and 7.42 per 1000 population in rural Uttar Pradesh. 1 Severe cases of RHD are treated with valve replacement surgery, and patients with mechanical valves require lifelong oral anticoagulation to prevent thromboembolic complications. However, oral anticoagulant therapy is challenging because patients exhibit large variability in their dose–anticoagulant effect response. Chronic anticoagulation is associated with the risk of hemorrhage and requires close monitoring of prothrombin time and international normalized ratio (INR).
Acenocoumarol, an oral vitamin K antagonist, is a commonly prescribed anticoagulant following valve replacement surgery in India. This drug has a narrow therapeutic range, and small dose variations may result in hemorrhagic or thrombotic complications. The interindividual variability is known to depend on environmental factors, but genetic influence has also been demonstrated. Acenocoumarol acts by inhibiting the C1 subunit of vitamin K 2,3-epoxide reductase, the enzyme responsible for the regeneration of vitamin K, required for the synthesis of the vitamin K-dependent clotting factors. Single-nucleotide polymorphisms (SNPs) of VKORC1 gene, −1639G>A in the promoter region and 1173C>T in the intronic region, have been found to influence the anticoagulant dosage owing to reduced enzyme activity. 2,3
Cytochrome P450 2C9, the main enzyme responsible for the hepatic metabolism of acenocoumarol, has been demonstrated to be polymorphic. In humans, 3 major alleles have been found: (1) CYP2C9*1 (wild-type or reference allele) Arg144/Ile359, (2) CYP2C9*2, Cys144/Ile359, and (3) CYP2C9*3, Arg144/Leu359. 4 Those with CYP2C9*2 and CYP2C9*3 alleles have reduced enzyme activity compared to the wild type (CYP2C9*1), leading to slow metabolism of acenocoumarol and hence require a lower dosage. 5 There is only a limited data from India, in particular from the southern part, on the association of genetic variability with acenocoumarol maintenance dosage. The aim of this study was to determine the distribution of polymorphisms of VKORC1 (−1639G>A; 1173C>T) and CYP2C9 (*2 and *3 alleles) genes in the South Indian population of Bangalore and to elucidate its association with the maintenance dosage of acenocoumarol in patients who are on lifelong anticoagulation therapy following heart valve replacement with mechanical valves.
Materials and Methods
The ethics committee of Sri Jayadeva Institute of Cardiovascular Sciences and Research, Bangalore, approved the study. Clinical data and blood samples were collected after obtaining written informed consent from all the participants.
This is a prospective study, and 205 patients with RHD of South Indian origin who had undergone valve replacement with mechanical valves and were on acenocoumarol therapy with a stable therapeutic INR, between 2 and 3.5 for at least 3 months, were recruited. Of the 205 cases, 111 had mitral valve, 50 had aortic valve, and 44 had double valve replacements.
The exclusion criteria were age less than 18 years, the presence of renal or liver dysfunction, diabetes mellitus, malabsorption or chronic diarrheal diseases, and untreated thyroid disease. Patients on antiepileptics, including phenytoin and carbamazepine, or antituberculous treatment and those positive for HIV or hepatitis B or C viruses or reactive to Venereal Disease Research Laboratory test were also excluded.
A detailed history of diet, comorbidity, medication, episodes of bleeding, or thromboembolism, if any, was collected. Only those patients with a minimum of 3 consecutive INR readings in the therapeutic range were included in the study, and their INR readings were documented. The mean daily dose of acenocoumarol was calculated. Venous blood was collected in EDTA containers for genotyping. Laboratory investigations included complete hemogram and estimation of prothrombin time and INR.
Genotyping
Genomic DNA was extracted from peripheral blood using the conventional phenol–chloroform method 6 and quantified using NanoDrop 2000 (Thermo Fisher Scientific, Wilmington, Delaware).
Genotyping of VKORC1 Polymorphism
Genotyping for VKORC1 1173C>T (rs9934438) and −1639G>A (rs9923231) gene mutation was detected by the protocol described by Sconce et al. 7 A 200-base pair (bp) DNA fragment containing the C>T substitution at nucleotide position 1173 was amplified by polymerase chain reaction (PCR) and cleaved by restriction enzyme StyI (New England Biolabs, Massachusetts). The StyI-digested products were separated on 2% agarose gel. The band pattern obtained for wild homozygous (CC) was 144 and 56 bp. Heterozygous (CT) showed an intact band of 200 bp along with 2 cleaved bands of 144 and 56 bp, whereas homozygous mutant (TT) showed undigested intact band of 200 bp (Figure 1A). Similarly, amplicons of 290 bp containing the G>A substitution at nucleotide position −1639 in the promoter region of VKORC1 were cleaved by restriction enzyme MspI (New England Biolabs, Massachusetts). The individuals having homozygous wild type (GG) exhibited the band pattern of 168 and 122 bp. Heterozygous (GA) showed an intact band of 290 bp along with 2 cleaved bands of 168 and 122 bp. Homozygous (AA) showed an undigested single band of 290 bp (Figure 1B).
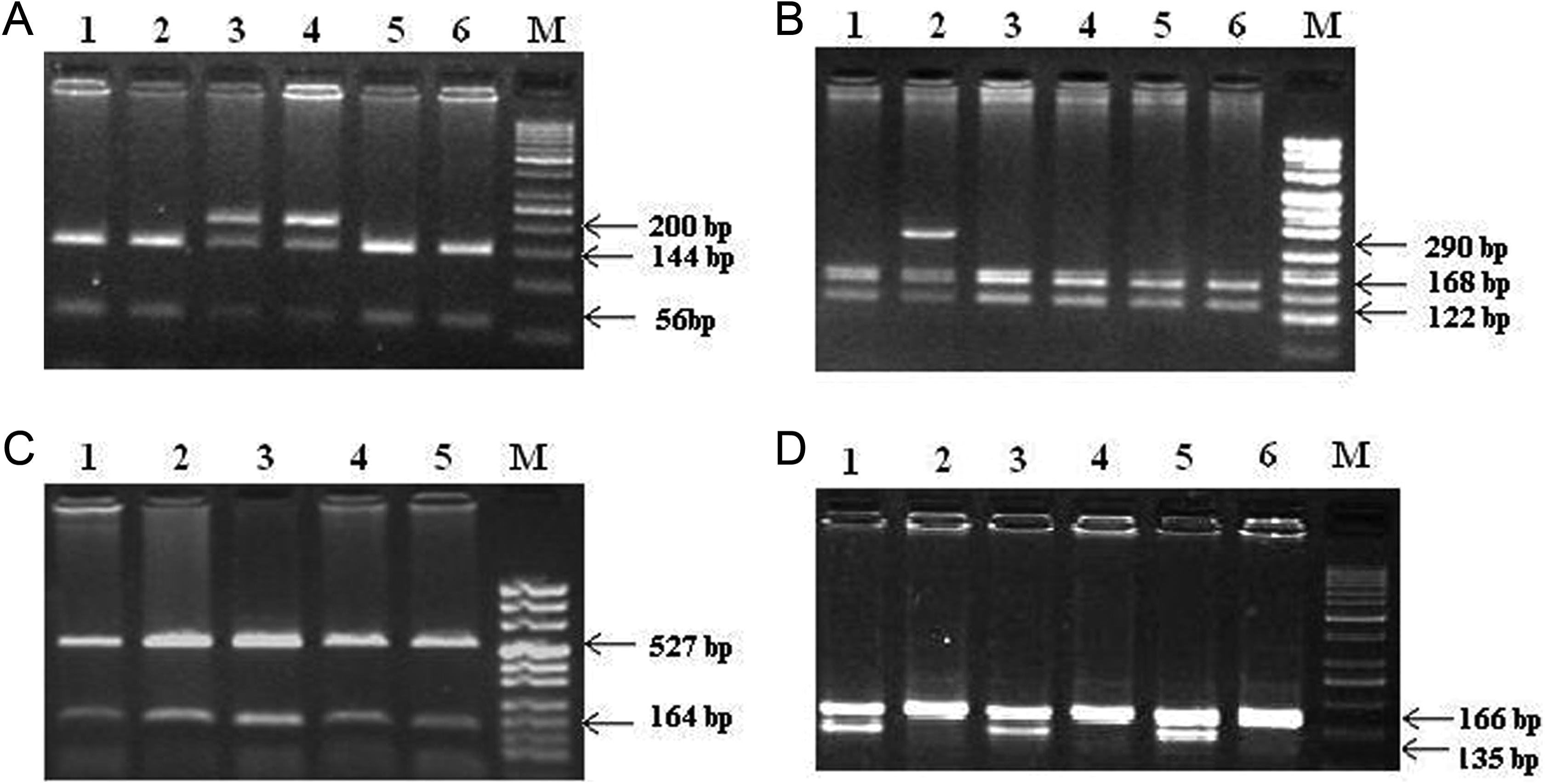
Restriction fragment length polymorphism (RFLP) analysis and representative band patterns of: (A) VKORC1 intron-1, 1173 C>T polymorphism samples digested with StyI. Lanes 1, 2, 5, and 6 are the samples of patients with homozygous (CC) genotype; lanes 3 and 4 are the samples of patients with heterozygous (CT) genotype. DNA marker: gene ruler 50 base pair (bp) ready to use. B, VKORC1 promoter, −1639 G>A polymorphism samples digested with MspI. Lanes 1, 3, 4, 5, and 6 are samples of patients with wild (GG) genotype; lane 2 is the sample of patient with heterozygous (GA) genotype. DNA marker: pUC 8 mix (C) CYP2C9 *2 polymorphism. Lanes 1, 2, 3, 4, and 5 are the samples of patients with homozygous genotype; DNA marker: pUC 8 mix. D, CYP2C9 *3 polymorphism. Lane 1, 3, and 5 are samples of patients digested with NsiI and lanes 2, 4, and 6 are samples of patients with KpnI. DNA marker: gene ruler 50 bp ready to use.
Genotyping of CYP2C9 Polymorphism
Genotyping of CYP2C9*2 430C>T (rs1799853) and CYP2C9*3 1075A>C (rs1057910) was detected by the protocol described by Sullivan-Klose et al. 8 For analysis of CYP2C9*1 and *2 (430C>T), the amplified PCR products of 691 bp were digested using restriction enzyme AvaII (New England Biolabs, Massachusetts). The digested products were separated on 1.5% agarose gel. The band pattern for wild type *1/*1 showed 527 and 164 bp, and heterozygous genotype *1/*2 showed an intact band of 691 bp along with 2 cleaved bands of 527 and 164 bp. Homozygous mutant *2/*2 showed an undigested single band of 691 bp (Figure 1C).
For detecting CYP2C9*3 (1075A>C) variant, the PCR products of 166 bp were amplified and digested using restriction enzymes NsiI and KpnI (New England Biolabs, Massachusetts) individually. The digested products were separated on 3% agarose gel. The wild type (*1/*1) was characterized by diagnostic NsiI restriction fragments of 135 and 31 bp. Individuals with *1/*3 and *2/*3 heterozygous genotype showed an intact PCR band of 166 bp and cleaved bands of 137 and 29 bp when digested with NsiI and also with KpnI. None of our patients showed the band pattern for *3/*3 (Figure 1D).
Statistical Analysis
Statistical analysis was performed using SPSS v.17.0 (IBM Corporation, New York). Genotype frequencies were determined using standard frequency analysis method. Pair-wise comparison between different genotypes was done using 1-way analysis of variance. The relation between the variable dose and genotype variants was established by univariate odds ratio (OR) at 95% confidence interval (CI). A multinomial logistic regression was used to assess the influence of certain parameters (age, sex, alcohol, smoking, and body mass index [BMI]) on the probability that a patient would receive a low or a high dose of acenocoumarol. A P value of .05 or less was considered statistically significant.
Results
Demographic Characteristics
Two hundred five patients with a stable therapeutic INR between 2 and 3.5 were recruited for the study (100 females and 105 males). Time in therapeutic range, calculated using the traditional method, 9 was >60% in all patients. The mean age of the study population was 39.9 ± 12.11 years and BMI (kg/m2) was 22.9 ± 5.2. Smoking (4.9%) and alcohol consumption (5.3%) were noted only in male patients. Two patients (0.9%) had history of prosthetic valve thrombosis, and 27 (13%) of them had atrial fibrillation. Concomitant medication included furosemide (5.8%), digoxin (16%), low-dose (75 mg) aspirin (32.7%), and amiodarone (1.5%).
Prevalence of Genotypes and Allele Frequency
The genotype distribution frequency of VKORC1 and CYP2C9 was determined for all 205 patients. The allelic frequencies for both VKORC1 −1639G>A and 1173C>T were in Hardy-Weinberg equilibrium. The distribution of VKORC1 −1639G>A (rs9923231) was as follows: wild type (GG) 57.6%, heterozygous (GA) 36.1%, and mutant homozygous (AA) 6.3%. The distribution of VKORC1 −1639G>A and 1173C>T was similar. There was complete linkage disequilibrium between these polymorphisms (r 2 = .98, D′ = 1.0, Log of Odds (LOD) = 74.02). Patients with heterozygous VKORC1 −1639GA genotype also had heterozygous VKORC1 1173CT genotype, and homozygous −1639AA was found to have homozygous 1173CC. The allele frequency for both G and C was 0.76 and 0.24 for A and T.
The genotype distribution of CYP2C9 was as follows: the wild type (*1/*1) 78.5%, mutants: *1/*3-14.1%, *1/*2-6.3%, *2/*2-0.5%, *2/*3-0.5%, and none detected for *3/*3. The most common allele frequency was 0.89 for *1, and the least common was 0.07 for *3 (Table 1).
Genotype, Allele Frequency, and Mean Dosage of Acenocoumarol in the Patients.a
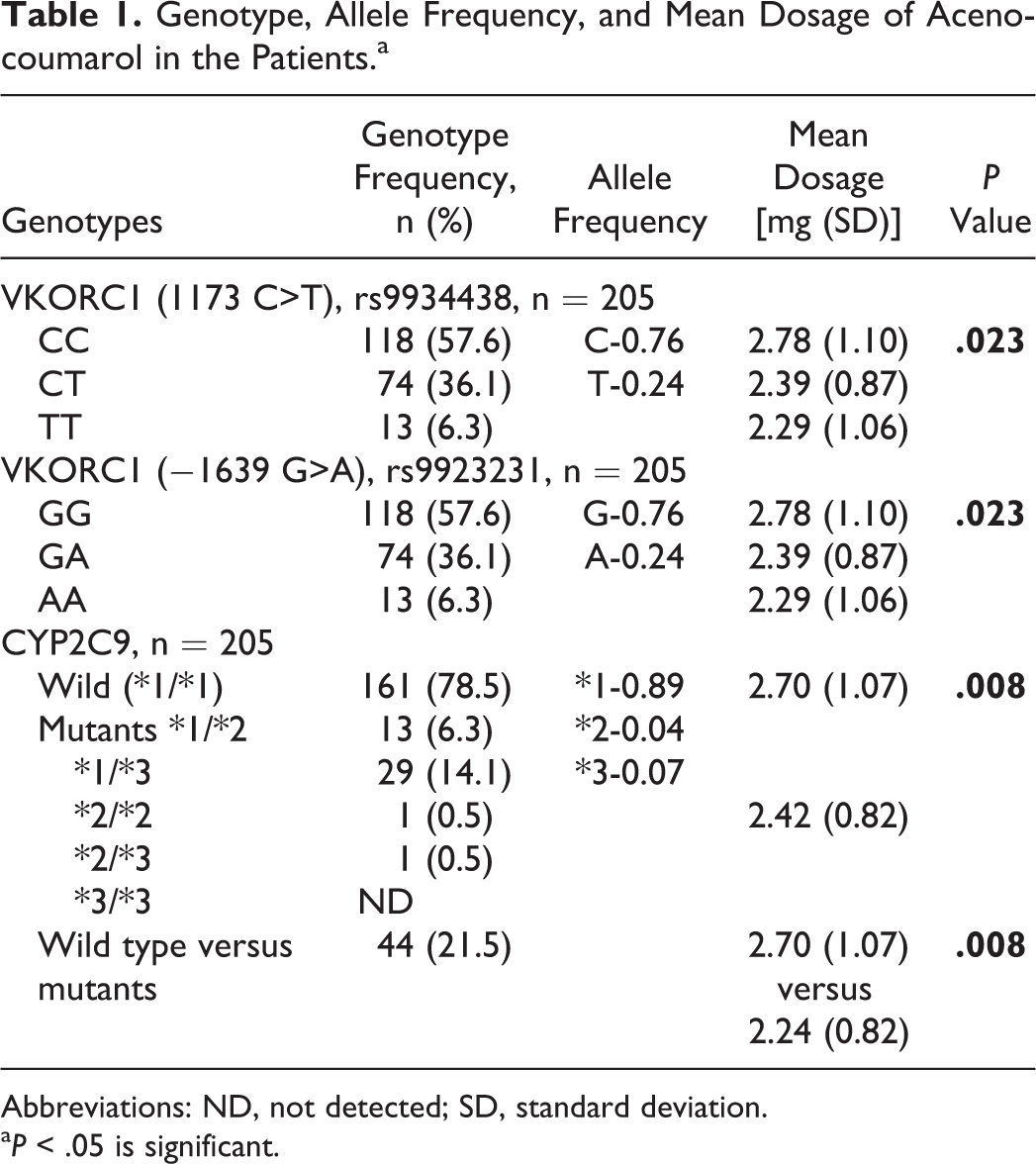
Abbreviations: ND, not detected; SD, standard deviation.
a P < .05 is significant.
Association of Genotype and Allele Frequency With Mean Acenocoumarol Dosage
Patients bearing wild genotype (GG) of VKORC1 −1639G>A required a mean acenocoumarol dose of 2.78 (1.10) mg/d to maintain the therapeutic INR, which was significantly higher (P = .023) as compared to those bearing the heterozygous VKORC1 −1639GA [2.39 (0.87) mg/d] and homozygous (−1639AA) genotype [2.29 (1.06) mg/d].
Patients with wild type of CYP2C9 (*1/*1) required a mean daily dose of 2.70 (1.07) mg as against 2.24 (0.82) mg for all patients with CYP2C9 mutant genotypes considered together (P = .008). Mean dosage for individual mutant genotype did not show statistical significance probably owing to the small sample size (Table 1).
In combined genotype analysis, patients bearing wild-type VKORC1 in combination with wild-type CYP2C9 genotype (−1639GG with *1/*1 allele) were most common (n = 91, 44.4%) in our study and required a mean dose of 2.90 (1.13) mg/d. A lower dose of 2.36 (0.90) mg/d was required by patients (n = 27, 13.2%) having wild type (−1639GG) of VKORC1 with mutants of CYP2C9 (P = .024), and heterozygous (−1639GA) patients (n = 62, 30.2%) carrying *1/*1 allele required a mean dose of 2.43 (0.93) mg/d (P = .008) (Table 2).
Mean Daily Dose in Subjects With Combined Genotype of Both VKORC1 and CYP2C9 Loci.a,b
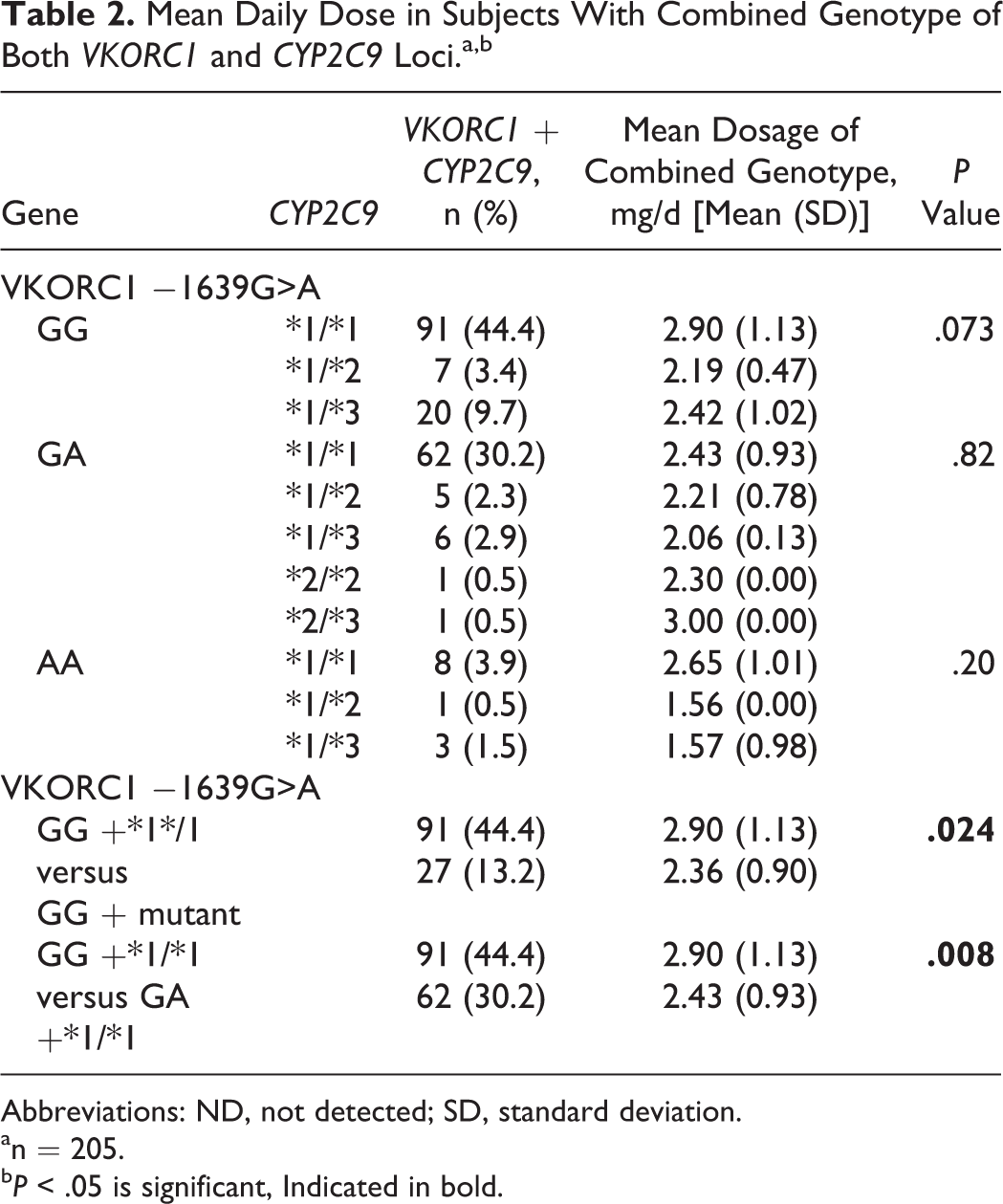
Abbreviations: ND, not detected; SD, standard deviation.
an = 205.
b P < .05 is significant, Indicated in bold.
Association of Gender With Mean Dosage
Only in male patients, presence of “A” allele of VKORC1 −1639 (GA and AA) and mutants of CYP2C9 (*2 and *3) independently increased the odds of requiring a lower dosage of acenocoumarol (adjusted OR: 1.65; 95% CI: 0.099-2.76; P = .05 and adjusted OR: 1.85; 95% CI: 1.0-3.43; P = .049, respectively).
Association of Concomitant Drugs With the Mean Dosage
On analyzing the effect of furosemide and digoxin on the mean acenocoumarol dose requirement, we found that patients who were taking these drugs concomitantly and had the same genotype (wild type and alleles of both VKORC1 and CYP2C9 genes) required a lower mean dosage when compared to others. However, the concomitant use of low-dose aspirin did not have any effect on the acenocoumarol dosage (Table 3). 10 Also, the presence of atrial fibrillation did not show any association on the maintenance dose requirement.
Association of Genotype Variants With Acenocoumarol Dosage in Patients With RHD and Odds of Requiring a Low Dosage.a,b
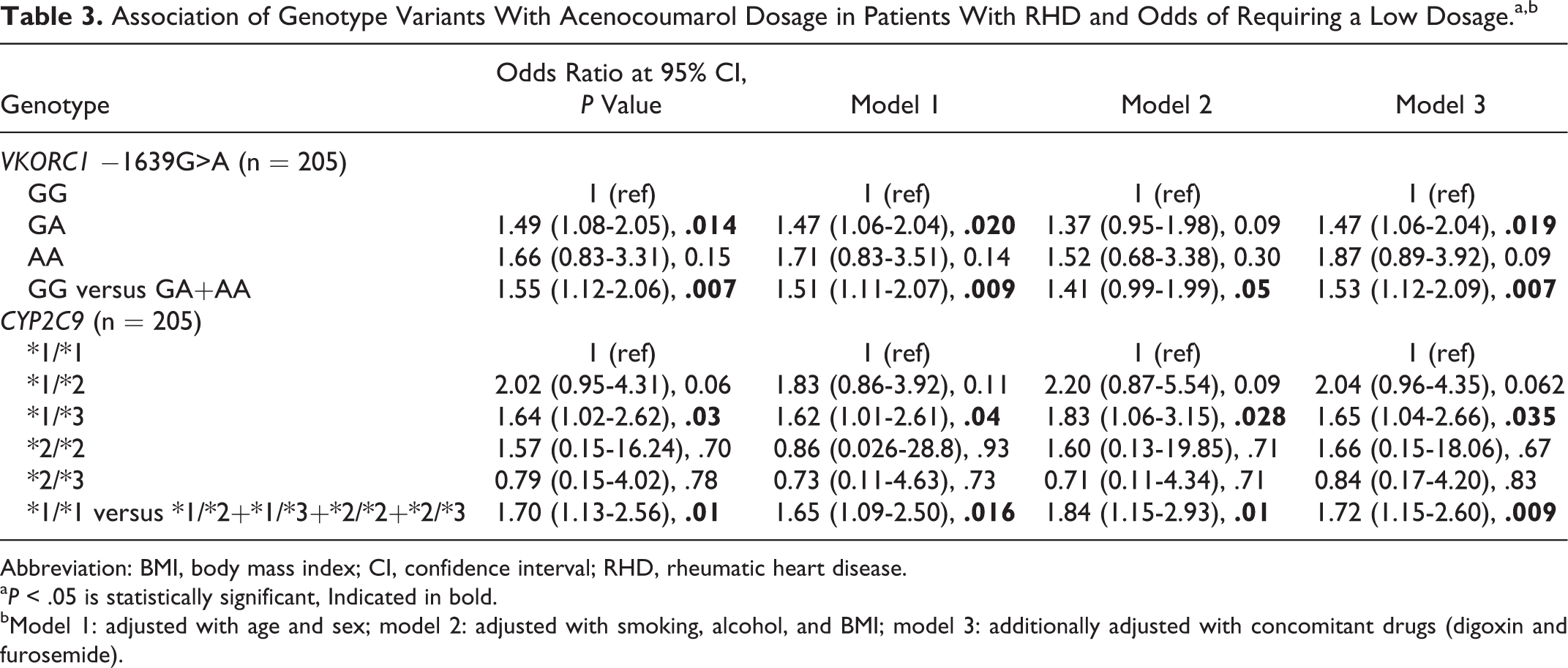
Abbreviation: BMI, body mass index; CI, confidence interval; RHD, rheumatic heart disease.
a P < .05 is statistically significant, Indicated in bold.
bModel 1: adjusted with age and sex; model 2: adjusted with smoking, alcohol, and BMI; model 3: additionally adjusted with concomitant drugs (digoxin and furosemide).
Discussion
This study aimed at determining the genotype frequency distribution in our patient population and evaluates the contribution of genetic variability of CYP2C9 and VKORC1 genes to the optimal maintenance dosage requirements of acenocoumarol, following cardiac valve replacement. We found that the wild genotype VKORC1 (−1639GG and 1173CC) as the most common (57.6%) in our study cohort, followed by heterozygous −1639GA genotype (36.1%). The majority of our patients (78.5%) had wild-type CYP2C9. Both wild types of VKORC1 and CYP2C9 required a higher mean daily dose compared to mutants. Of all, 44.4% of our patients had a combined genotype of VKORC1 −1639GG and 1173CC with CYP2C9 *1/*1 and required a higher mean daily dose.
An average of 2000 patients with RHD are treated annually at our center, a government-run tertiary cardiac care hospital situated at Bangalore, Southern India. About 600 valve replacement surgeries are carried out, and the majority of them are mechanical valves. Acenocoumarol, a coumarin derivative, is the anticoagulant of choice in our center on account of its affordability by the majority of our patients due to its lower cost compared to warfarin. Also, due to its longer duration of action and availability in all strengths compared to warfarin, our physicians find it easier to titrate the dose.
Acenocoumarol has a narrow therapeutic index with large pharmacokinetic and pharmacodynamic interindividual variability. 11 Polymorphisms in VKORC1 gene have been hypothesized to play a role in the variability of warfarin response based upon a seminal study, showing that an intronic 1173C>T polymorphism was associated with warfarin dose requirement. 12 A study from France has found that genotyping of the 2 SNPs, VKORC1 −1639G>A or 1173C>T could predict high risk of overdose before initiation of anticoagulation with acenocoumarol 4 and provide a safer and more individualized anticoagulant therapy. The CYP2C9 gene has also been demonstrated to be polymorphic, and its genetic variability is shown to be associated with variations in the levels of enzyme activity. 13
There are a plethora of studies showing a wide racial and multiethnic difference in the prevalence of these gene polymorphisms in various ethnic populations (Table 4). Indians are ethnically different not only from Caucasians and Africans but also differ considerably from their Asian counterparts, especially Southeast Asians. More than 80% of the East Asians have homozygous VKORC1 genotypes (AA and TT) and >90% of them have wild-type CYP2C9 (*1/*1), and they require a lower dosage than African and European patients. 27 A study on Han Chinese patients showed a prevalence of 81% homozygous AA and 18% heterozygous VKORC1genotype. 18
Global Genotype Frequency for VKORC1 (1173C>T and −1639G>A) and CYP2C9 Polymorphisms.
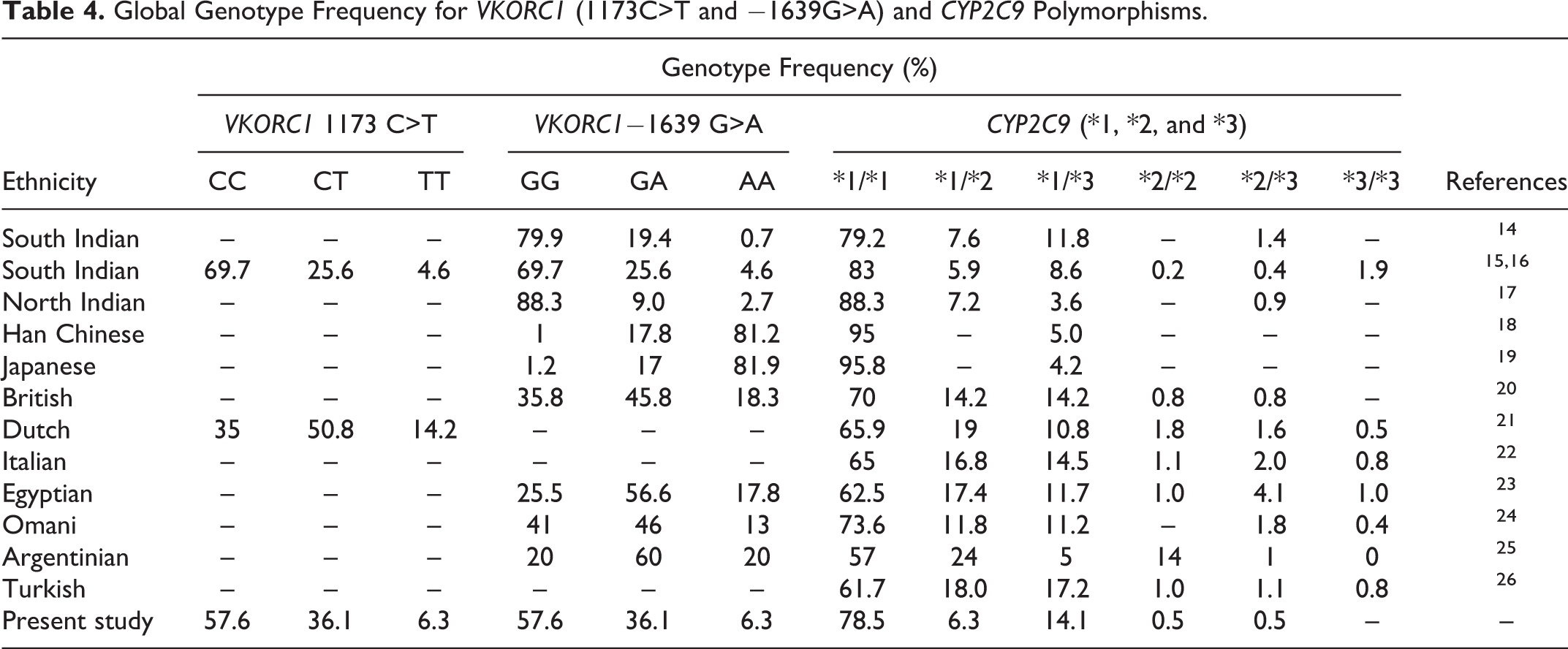
Although the wild type of VKORC1 (−1639 GG and 1173 CC) was the most prevalent genotype (57.6%) in our study, the frequency of heterozygous −1639GA and 1173CT (36.1%) was significantly higher compared to other studies from India. Our results were very similar to another study from Bangalore by De et al. 14,15,17,28,29 Whether this distribution is unique to the population of Bangalore needs to be confirmed with larger studies.
The mean daily dose required by our patients with VKORC1 mutant homozygous −1639AA and 1173TT and heterozygous −1639GA and 1173CT genotypes was much lower—2.29 and 2.39 mg, respectively, compared to patients with the wild type who required 2.78 mg (P = .023). The mean daily dose decreased by 18% for homozygous AA and 14% for heterozygous GA. This is in accordance with the seminal study by D’Andrea et al, who showed that patients carrying the “T” allele required a lower dose of warfarin compared with those carrying the “C” allele (the mean daily dose decreased by 43% for homozygous TT carriers and 22% for heterozygous). 12 Similar associations with mean dose requirement of warfarin were found in South Indian patients. 14,28 A study from North India found similar results with respect to wild and heterozygous types. However, patients with mutant homozygous genotype required a higher dose, which could probably be explained by a small sample size. 30 Only in male patients, the presence of heterozygous and mutant homozygous genotypes of VKORC1 −1639GG and GA gene independently increased the odds of requiring a lower dosage of acenocoumarol (adjusted OR: 1.65; 95% CI: 0.99-2.76; P = .05).
CYP2C9 *1 was the most prevalent (78.5%) genotype in our study, which is consistent with previous Indian studies on both North and South Indian patients where the prevalence varied from 70% to 90%. 14,17,29 Studies on other ethnic groups have also shown the wild type to be the most common genotype with the highest prevalence of 95% in Han Chinese patients. 7,18,24 The distribution of CYP2C9 wild and mutants in our study is very similar to previous studies on Indian patients 14,29 and also to a study by Biss et al on Caucasians. 20 The genotype *3/*3 was absent in our study, which is consistent with previous Indian studies. 14,17,28 The *2/*2 allele is almost absent in Asian population, and results of our study (0.5%) were similar to the previous study by De et al (0.2%). 5,16 North Indian patients have been reported to have a higher prevalence of *2 polymorphism at 4.1%. 14
In our study, the mean acenocoumarol dose requirement decreased by 17% in patients bearing alleles of CYP2C9 compared to patients with wild type [2.24 (0.82) mg/d vs 2.70 (1.07) mg/d].
It has been suggested that a combination VKORC1 and CYP2C9 gene polymorphisms is more accurate in predicting the anticoagulant dose requirement than when considered separately. 5 The most prevalent combined genotype in our patients was VKORC1 −1639GG with CYP2C9 *1/*1 (44%) followed by VKORC1 −1639GA with CYP2C9 *1/*1 (30.2%) and VKORC1 −1639GG with *1/*3 of CYP2C9 (9.7%). Patients with a combination of wild type (GG) of VKORC1 −1639 and wild type (*1/*1) of CYP2C9 required mean acenocoumarol dose of 2.90 (1.13) mg/d. The dose requirement decreased by 19% in patients with VKORC1 −1639GG with the mutant allele of CYP2C9 (P = .024) and 16% in patients with wild type CYP2C9 who have heterozygous (−1639GA) VKORC1 (P = .008). The least mean dose requirement was by patients with VKORC1 −1639AA + CYP2C9 *1/*2 genotype (1.56 mg/d)
When the patients were concomitantly using furosemide and digoxin, having a mutant allele of VKORC1 (−1639A and 1173T) and CYP2C9 (*2 and *3) increased the odds of requiring a lower mean acenocoumarol dosage by 1.53 (P = .007) and 1.72 (P = .009) folds, respectively (Table 3). These drugs are known to potentiate the effect of acenocoumarol by releasing the anticoagulant bound to plasma protein and increasing the concentration of free active form in the plasma. 31 However, coprescribing low-dose aspirin had no effect on the mean daily dose.
Conclusion
The South Indians in Bangalore showed a higher prevalence of heterozygous VKORC1 genotype. Bearing a mutant allele of VKORC1 (−1639A and 1173T) and CYP2C9 genes increased the odds of requiring a lower mean dosage of acenocoumarol, and studying the combination of genotypes could predict the dose requirement more accurately. Concomitant prescription of furosemide and digoxin increases the risk of bleeding, and hence, the patients, particularly those with mutant alleles, require a lower mean dosage.
Footnotes
Declaration of Conflicting Interests
The authors declare no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by the Indian Council of Medical Research, New Delhi, India.
