Abstract
Genetic variants of cholesteryl ester transfer protein (CETP) and endothelial nitric oxide synthase (eNOS) influence high-density lipoprotein cholesterol (HDL-C) metabolism and nitric oxide (NO) synthesis, respectively, and might increase the risk of coronary artery disease (CAD). This study is to investigate the relationship between genetic polymorphisms and the risk of CAD and to evaluate their potential interactions. A total of 237 patients with CAD and 101 controls were genotyped. The association of the polymorphism with the risk of CAD varied among the ethnic groups. Moreover, the concomitant presence of both CETP B1 and eNOS 4a alleles significantly increased the risk of CAD in the Malay group (OR = 33.8, P < .001) and the Indian group (OR = 10.9, P = .031) but not in the Chinese group. This study has identified a novel ethnic-specific gene–gene interaction and suggested that the combination of CETP B1 allele and eNOS 4a allele significantly increases the risk of CAD in Malays and Indians.
Introduction
The pathogenesis of coronary artery disease (CAD) is multifactorial and results from the combined and interactive effects of genetic and environmental factors. These factors may vary depending on the ethnic group. 1 It is becoming increasingly evident that interactions of multiple candidate genes play an important role in the pathogenesis of complex diseases such as CAD. 2 The study by Kathiresan et al 3 found that multiple common alleles influence variation in quantitative traits like high-density lipoprotein cholesterol (HDL-C) level which may provide more information about cardiovascular risk beyond lipid levels alone. Therefore, better understanding of gene–gene interactions can provide insights into the novel pathological mechanisms of CAD.
Coronary artery disease is contributed by interplay of complex biochemical processes that include lipid and apolipoprotein metabolism, inflammatory response, endothelial dysfunction, homocysteine metabolism, thrombosis, insulin sensitivity, and blood pressure regulation. 4 It has been shown in many population studies that the HDL-C levels are inversely associated with the risk of CAD. 5 –8 The protective effects of HDL-C have been related to its role in stimulating cholesterol efflux as well as to its antioxidant, anti-inflammatory, and antithrombotic properties. 9
Cholesteryl ester transfer protein (CETP) maintains the cholesterol homeostasis in the body by participating in reverse cholesterol transport. 10 It promotes the transfer of cholesteryl ester from HDL-C to apolipoprotein B-containing particles in exchange for triglycerides, thereby reducing the concentration of HDL-C and increasing non-HDL cholesterol. 11 Recent genome-wide association studies have reported that CETP is the most significantly associated locus for HDL-C. 12,13 One of the most widely studied polymorphic sites of the CETP gene is the Taq1B (rs 708272) in the 277th nucleotide of the first CETP intron. B2 allele at this polymorphism site is reported to be associated with decreased CETP activity and levels and increased HDL-C levels. 14
Endothelial dysfunction plays a critical role in the initiation and progression of the atherosclerotic process.
15
Endothelial function is regulated by vascular endothelial nitric oxide (NO). Endothelial nitric oxide synthase (eNOS) produces NO from
Since HDL-C can also affect the production of NO by stimulating the activity of eNOS to boost NO production 24 ; therefore, the aims of present study are to determine the association of CETP and eNOS polymorphisms with the risk of CAD and to find possible interaction between CETP and eNOS polymorphisms on the risk of CAD in Malays, Chinese, and Indians residing in Malaysia.
Material and Method
Study Design and Patients
This study was approved by National Medical Research Registry (NMRR) of Malaysian Ministry of Health with the registration number of NMRR-13-435-14568 and University Research Ethics Committee, Universiti Putra Malaysia, with the reference number FPSK_Ogos(13)05IFR. Informed written consent was obtained from each individual prior to the study.
This was a hospital-based case–control study. Participants were those undergoing coronary angiography in the Cardiology Department at Hospital Serdang between 2013 and 2014. A total of 338 patients were recruited, comprising 237 patients with CAD and 101 controls. Patients with CAD were those who had more than 50% diameter obstruction in at least 1 major coronary vessel. Controls were those who had normal or insignificant coronary angiographic findings (<50%). Coronary angiograms were interpreted by 2 or more independent, experienced cardiologists.
The medical records were collected to obtain baseline characteristics of the patients. Hypertension was defined as systolic pressure >140 mm Hg and/or diastolic pressure >90 mm Hg on more than 1 occasion, according to the Clinical Practice Guideline on Hypertension and/or currently taking hypertensive medication. Diabetes was defined as fasting plasma glucose >7.0 mmol/L and included patients with diabetic medication. Hypercholesterolemia was defined as a fasting total serum cholesterol level of >5.2 mmol/L and the intake of antihypercholesterolemic medications.
Lipid Measurement
Blood samples were drawn into plain tubes after the participants had fasted overnight. Serum total cholesterol, triglyceride, and HDL-C concentrations were determined enzymatically using commercially available kits (BioRad Laboratories Inc, Hercules, California) and Architect autoanalyzer (Abbott, Wiesbaden, Germany). Serum low-density lipoprotein cholesterol levels were calculated using the Friedewald formula. 25
DNA Extraction
Total genomic DNA was extracted from 3-mL EDTA anticoagulated blood specimens using a commercial kit (GENE ALL Blood SV mini, General Biosystems, Seoul, Korea) according to the manufacturer protocol.
Genotyping
Polymerase chain reaction (PCR) and PCR restriction fragment length polymorphism analysis was used for the genotyping analysis. The PCR comprised genomic DNA (20-60 ng), 1 U of Taq DNA polymerase (GenetBio, Chungcheongnam-do, Korea), 1 mmol/L dNTPs mixture (GenetBio), and 0.5 μmol/L of forward and reverse oligonucleotide primers (First Base Laboratories, Malaysia), in a volume of 20 μL.
For the detection of the CETP TaqIB polymorphism, the forward and reverse primers used were 5’ CAC TAG CCC AGA GAG AGG AGT GCC 3’ and 5’ CTG AGC CCA GCC GCA CAC TAA C 3’, respectively. 14 DNA was amplified for 30 cycles of denaturation (95°C, 30 seconds), annealing (65°C, 30 seconds), and extension (72°C, 45 seconds). The PCR products were detected by electrophoresis on 2% agarose gel after digestion with 5 units of restriction endonucleases TaqI (New England Biolabs, United Kingdom) in 15 μL of PCR sample at 65°C for 3 hours. Presence of TaqI site (B1 allele) gave 2 bands of 174 bp and 361 bp while absence (B2 allele) showed 1 band of 535 bp (Figure 1).
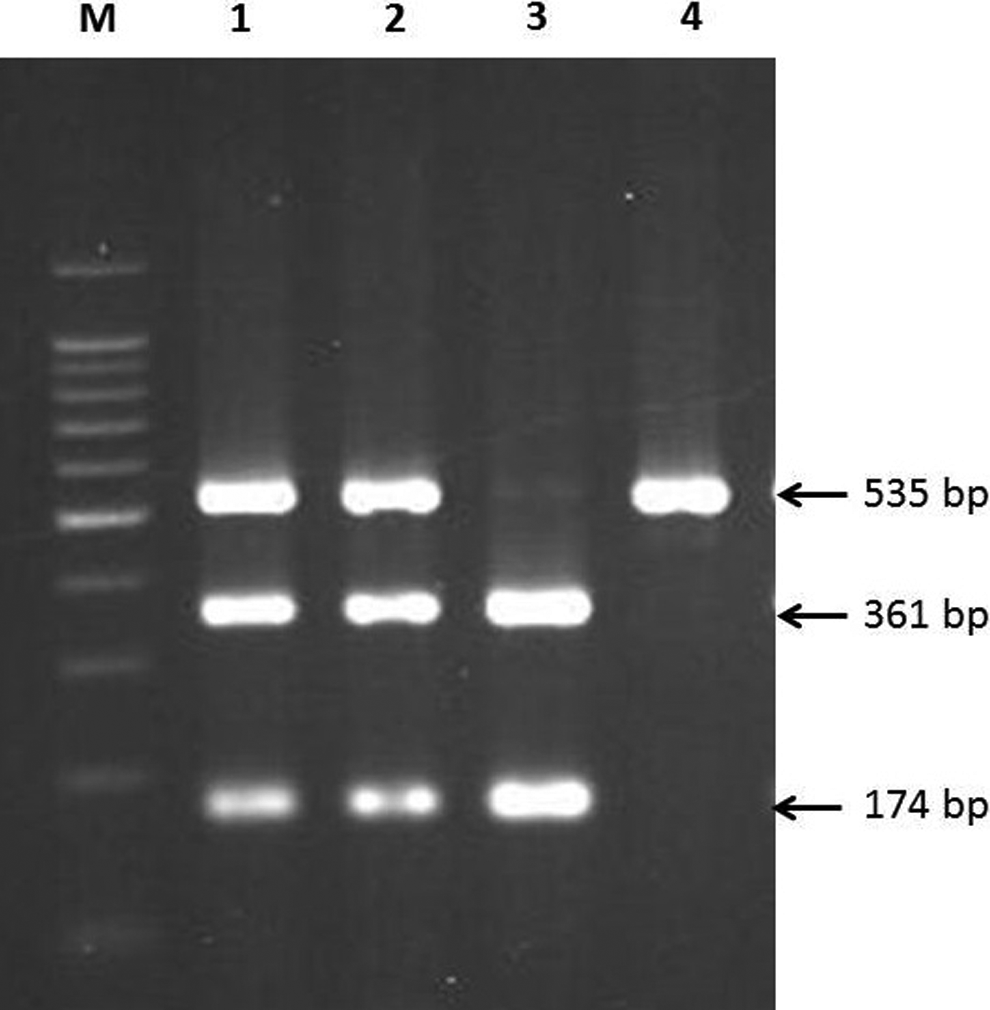
CETP TaqIB polymorphism. Gel picture shows B1B2 genotypes in lanes 1 and 2, B1B1 genotype in lane 3, and B2B2 genotype in lane 4. M is 100-bp DNA ladder.
For the detection of the eNOS 4a4b polymorphism, the forward and reverse primers used were 5ʹ AGG CCC TAT GGT AGT GCC TTT 3ʹ and 5ʹ TCT CTT TAG TGC TGT GGT CAC 3ʹ, respectively. 26 DNA was amplified for 35 cycles of denaturation (95°C, 30 seconds), annealing (63°C, 30 seconds), and extension (72°C, 45 seconds). The fragments of 393 and 420 bp that correspond with the eNOS alleles 4a and 4b were detected by electrophoresis on 3.5% agarose gel (Figure 2).
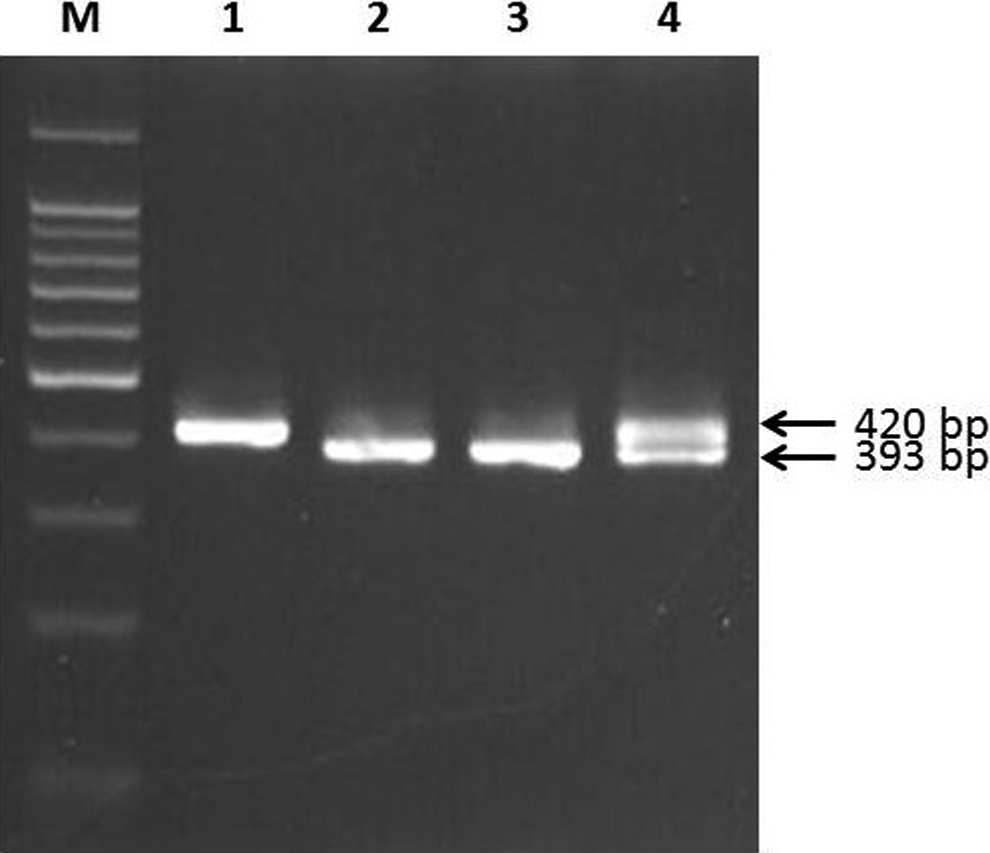
eNOS G894T polymorphism. Gel picture shows GG genotype in lane 1, TT genotype in lane 2, and GT genotype in lane 3. M is 100-bp DNA ladder.
For the detection of the eNOS G894 T polymorphism, the forward and reverse primers used were 5ʹ CAT GAG GCT CAG CCC CAG AAC 3ʹ and 5ʹ AGT CAA TCC CTT TGG TGC TCA C 3ʹ, respectively. 20 DNA was amplified for 35 cycles of denaturation (95°C, 30 seconds), annealing (68°C, 30 seconds), and extension (72°C, 30 seconds). The PCR products were detected by electrophoresis on 2.5% agarose gel after digestion with 2 units of restriction endonucleases MboI (New England Biolabs) in 4 μL of PCR sample at 37°C for 2 hours. The presence of a T at nucleotide 894 gave 2 bands of 119 bp and 87 bp while absence showed 1 band of 206 bp (Figure 3). Positive and negative mutation controls were always included during the genotyping.
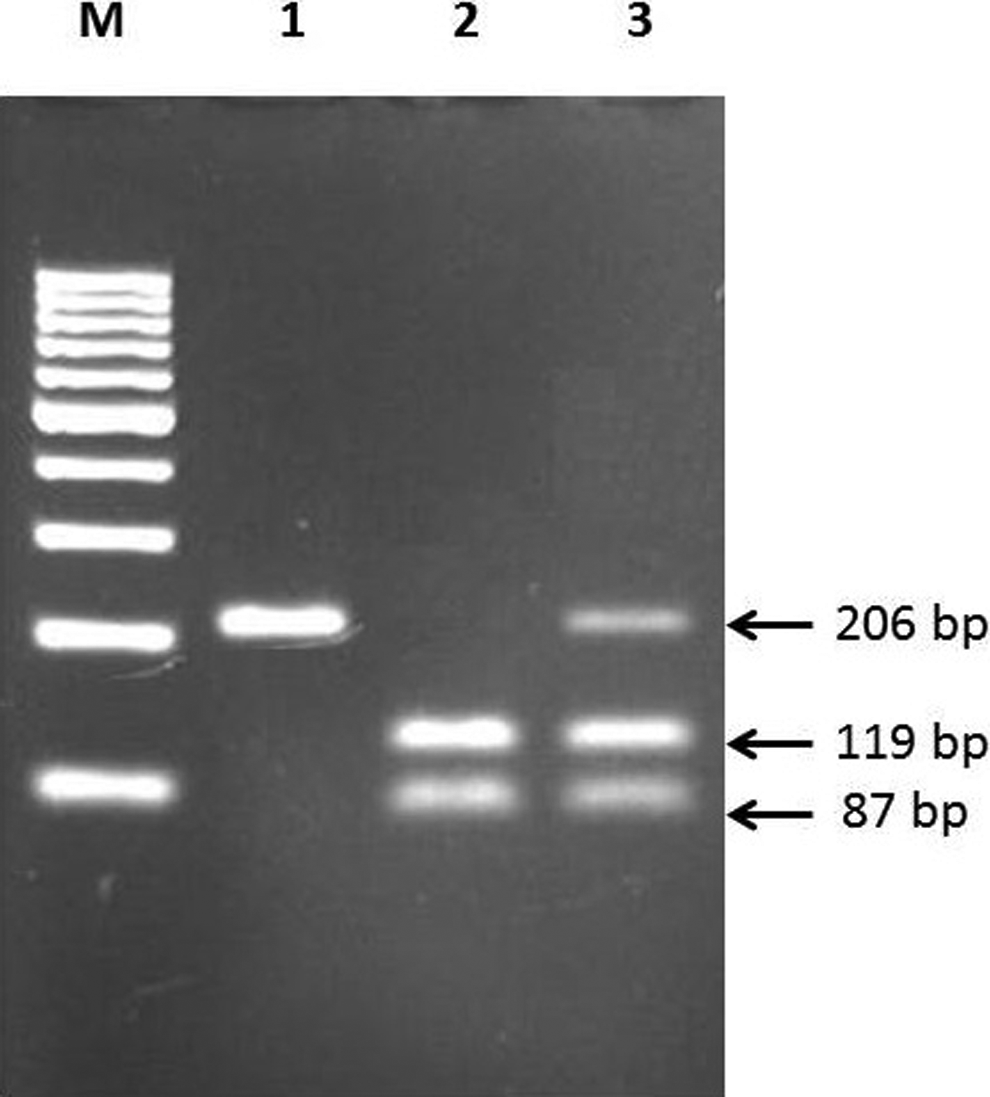
eNOS 4a4b polymorphism. Gel picture shows 4b4b genotype in lane 1, 4a4a genotypes in lanes 2 and 3, and 4a4b genotype in lane 4. M is 100-bp DNA ladder.
Statistical Analysis
All computations were done using the SPSS statistical software package version 21.0 (SPSS Inc, Chicago, Illinois). Data are presented as mean ± standard deviation or number (percentages). Continuous and categorical data were tested using independent t test or Mann-Whitney U test and chi-square (χ2) test or Fisher exact test, respectively. Differences between noncontinuous variables, genotype distribution, allele frequency, and Hardy-Weinberg equilibrium were tested by χ2 analysis. Statistical significance refers to a 2-tailed analysis where P < .05.
Result
Demographic and biochemical characteristics of the patients with CAD and controls are given in Table 1. Patients with CAD and controls were homogenous for age, gender, body mass index, prevalence of hypertension, dyslipidemia, family history of CAD, and lipid profiles. However, higher prevalence of diabetes was found in the Malay group.
Demographics and Biochemical Characteristics of the Study PAtients in 3 Ethnic Groups.
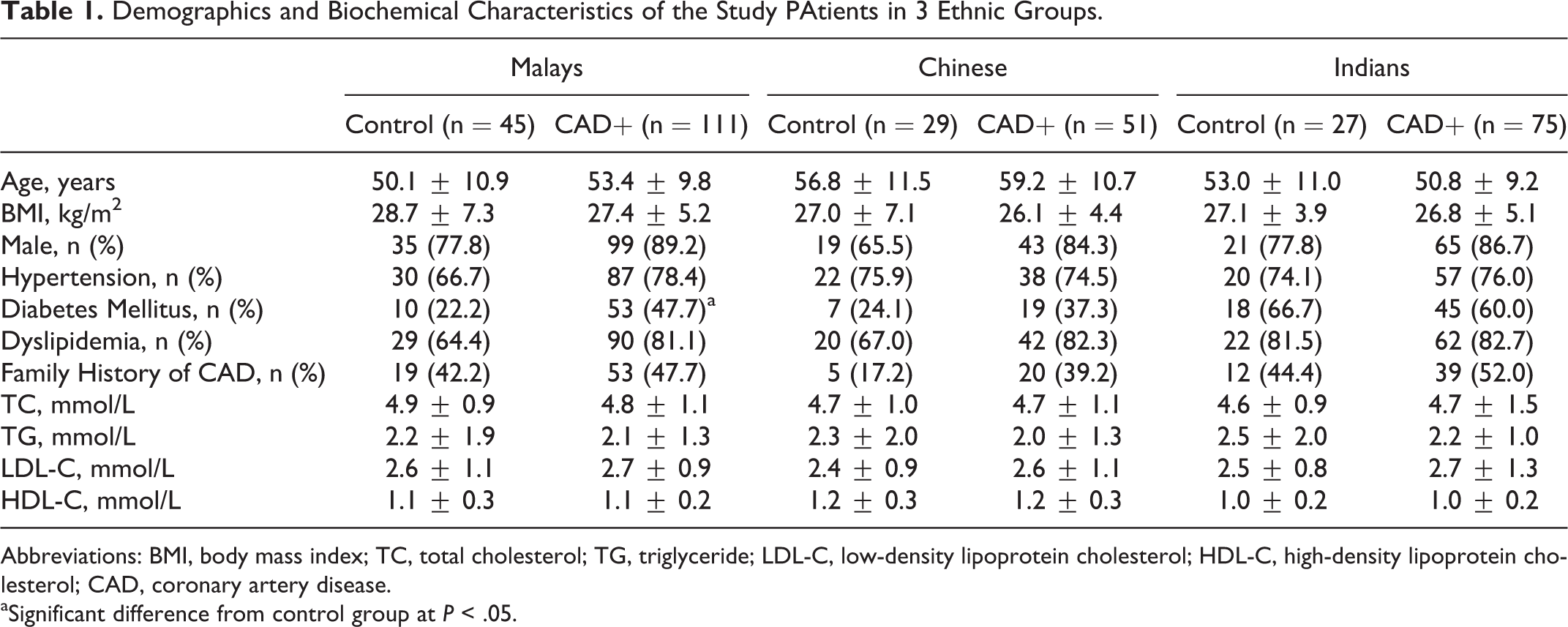
Abbreviations: BMI, body mass index; TC, total cholesterol; TG, triglyceride; LDL-C, low-density lipoprotein cholesterol; HDL-C, high-density lipoprotein cholesterol; CAD, coronary artery disease.
aSignificant difference from control group at P < .05.
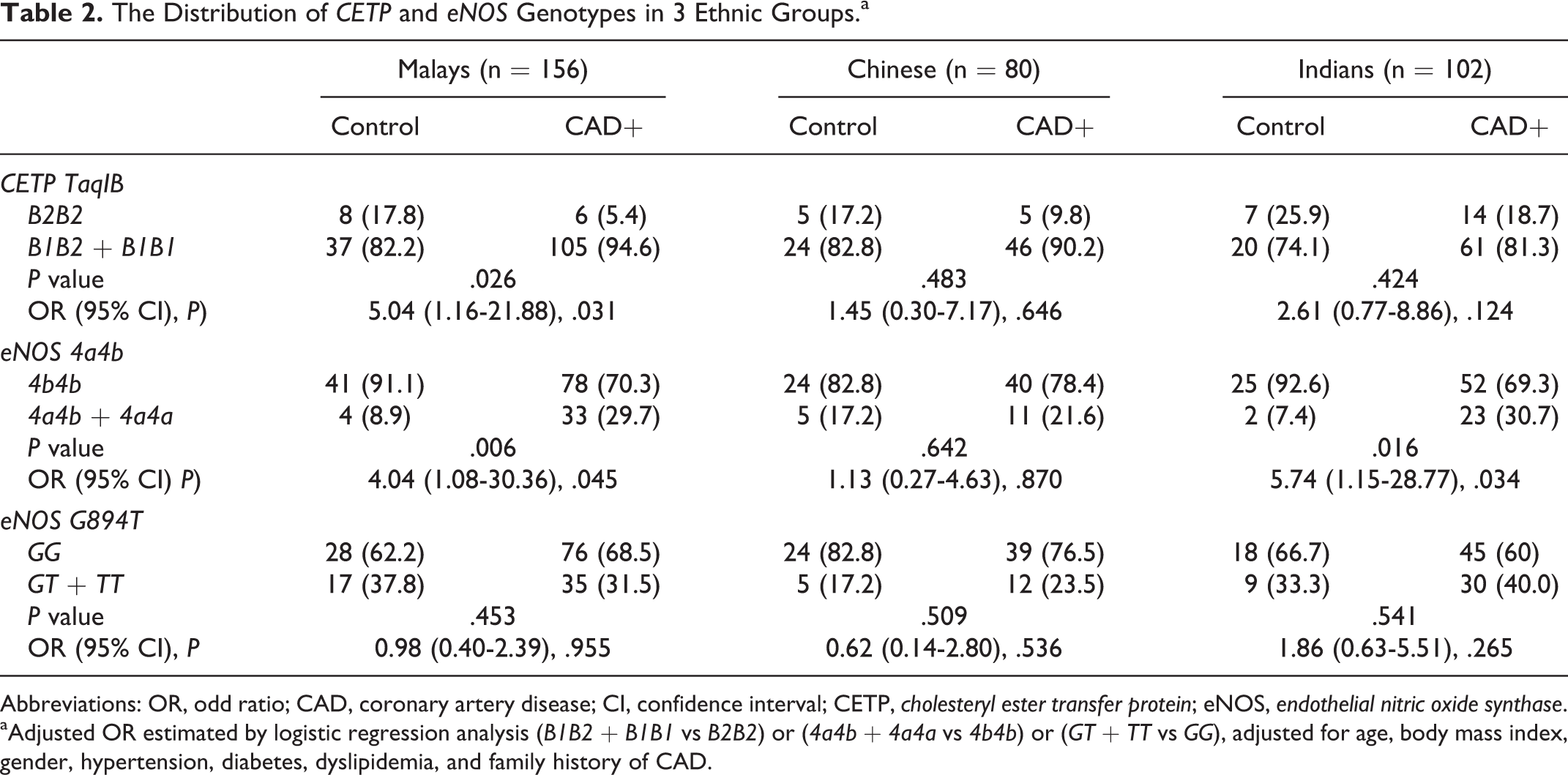
The distributions of CETP and eNOS genotypes among the 3 ethnic groups are demonstrated in Table 2. The distributions of genotypes for each polymorphism met Hardy-Weinberg equilibrium in all ethnic groups (all P > .05; data not shown). In the recessive genetic model to evaluate the risk effect of CETP TaqIB on the risk of CAD, B1B1 and B1B2 genotypes were combined and compared with B2B2 genotype, according to Tsujita et al. 27 The frequency of B1B2 + B1B1 genotype was significantly higher in patients with CAD than in the controls(94.6% vs 82.2%) in Malay group. The frequency of the CETP genotypes was similar in cases and controls in both Chinese and Indian groups.
The Distribution of CETP and eNOS Genotypes in 3 Ethnic Groups.a
Abbreviations: OR, odd ratio; CAD, coronary artery disease; CI, confidence interval; CETP, cholesteryl ester transfer protein; eNOS, endothelial nitric oxide synthase.
aAdjusted OR estimated by logistic regression analysis (B1B2 + B1B1 vs B2B2) or (4a4b + 4a4a vs 4b4b) or (GT + TT vs GG), adjusted for age, body mass index, gender, hypertension, diabetes, dyslipidemia, and family history of CAD.
The eNOS 894 T and 4a were considered as rare alleles, and their relationship with the risk of CAD was analyzed by dominant models. 28 The frequency of 4a4b + 4a4a was significantly higher in patients with CAD than in control patients 29.7% versus 8.9% and 30.7% versus 7.4% in Malay and Indian groups, respectively. However, the frequency of eNOS 4a4b genotype did not differ significantly between patients with CAD and controls in the Chinese group. No significant difference in genotype distribution of eNOS G894 T polymorphism was found between the 2 groups in the 3 ethnic groups.
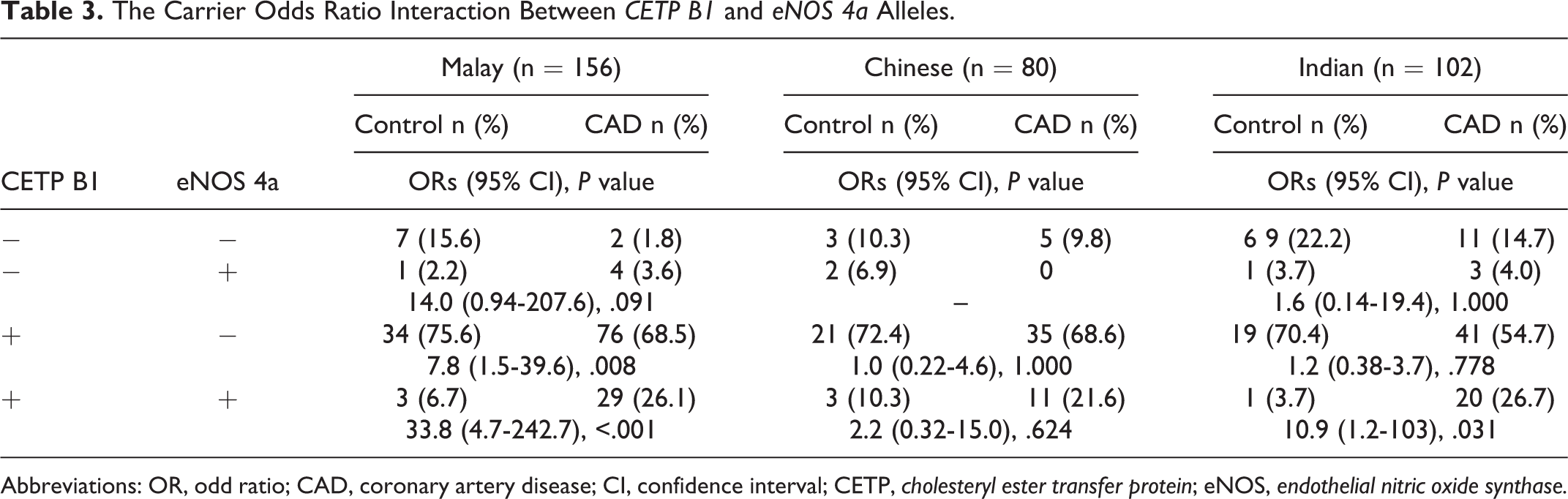
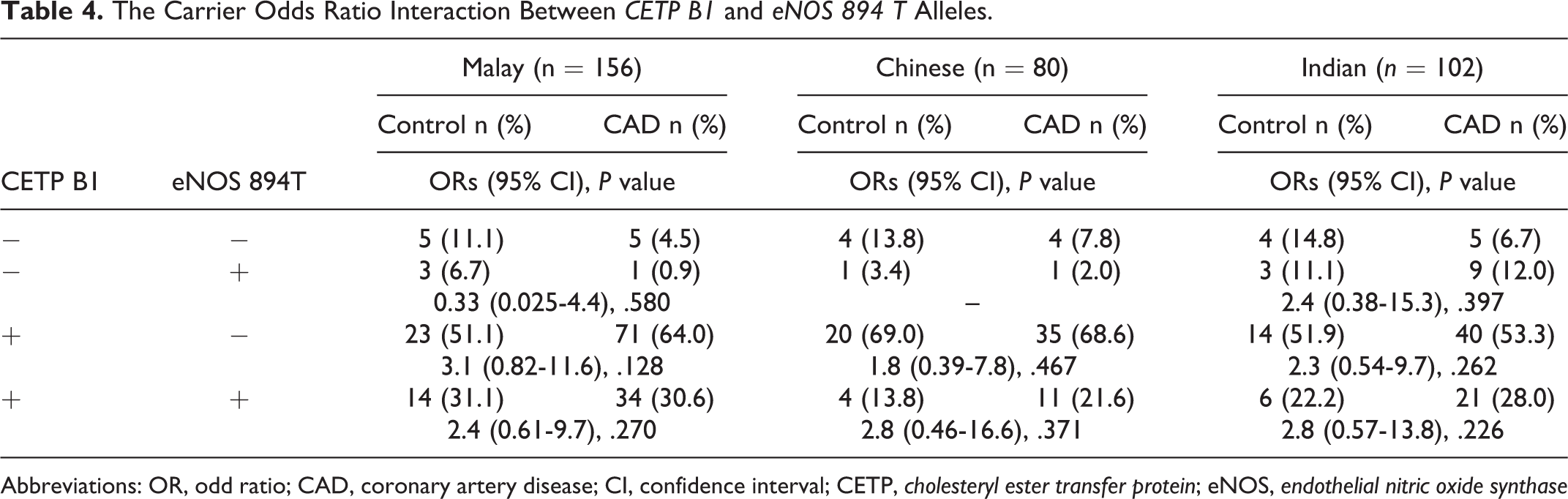
Table 3 indicated the interaction of both CETP B1 allele and eNOS 4a allele in the risk of CAD in the 3 ethnic groups. The concomitant presence of both alleles increased the risk of CAD significantly by 33.8-fold and 10.9-fold in the Malay and Indian groups, respectively. However, there was no significant interaction between the CETP B1 allele and eNOS 894 T allele in the risk of CAD in the 3 ethnic groups (Table 4).
The Carrier Odds Ratio Interaction Between CETP B1 and eNOS 4a Alleles.
Abbreviations: OR, odd ratio; CAD, coronary artery disease; CI, confidence interval; CETP, cholesteryl ester transfer protein; eNOS, endothelial nitric oxide synthase.
The Carrier Odds Ratio Interaction Between CETP B1 and eNOS 894 T Alleles.
Abbreviations: OR, odd ratio; CAD, coronary artery disease; CI, confidence interval; CETP, cholesteryl ester transfer protein; eNOS, endothelial nitric oxide synthase
Discussion
Coronary artery disease is a multifactorial disease that may differ in each ethnic group. This study provided a unique opportunity to examine the relationships of polymorphisms with CAD in multi-ethnic population in Malaysia.
A meta-analysis that involves a total of more than 147 000 individuals demonstrated that B2 allele of CETP TaqIB polymorphism decreases CETP activity and increases HDL-C by approximately 5% to 10%, respectively, and shows weak inverse associations with coronary risk. 29 Among the 3 ethnic groups in Malaysia, Malay group is the only group that shows a statistically significant association of CETP TaqIB polymorphism with the risk of CAD. Significant association of CETP TaqIB polymorphism with CAD was also reported for the Framingham 14 population, Scottish, 30 Tunisian, 31 and Tehran. 32 The Malaysian Indians migrated to Malaysia from Southern India. Interestingly, the frequency of B2B2 in Malaysian Indians was similar to that reported for North Indians 33 and Tamilian population from South Indian. 34 On the other hand, the Malaysian Chinese are from Southern China. The frequency of B2B2 in Malaysian Chinese was also similar to that in the Taiwanese. 35 In this study, lack of association of CETP TaqIB polymorphism with the risk of CAD was found in both Chinese and Indian groups. This finding is in agreement with the results of other studies of Chinese living in China 36,37 and Indians living in India. 38 These findings strongly suggest that the association of CETP TaqIB polymorphism with the risk of CAD is ethnic specific.
The eNOS 4a4b polymorphism is associated with reduced plasma NO levels. 39 Tsukada et al 40 found that the plasma NOx level of the patients with 4a allele was significantly lower than those without the 4a allele. Arecent meta-analysis of Yang et al, 41 including 37 studies indicated eNOS 4a4b is significantly associated with the risk of CAD among African and Asian populations but not caucasians. In the present study, eNOS 4a4b polymorphism was associated with the risk of CAD in Malays and Indians but not Chinese. The lack of correlation between eNOS 4a4b polymorphism with the risk of CAD has been demonstrated by Hwang et al 42 in the Taiwanese population.
The mutation of eNOS G894 T may alter eNOS activity by conformational change or by protease-mediated cleavage. Several studies have shown a possible association between genetic polymorphism of eNOS G894 T and the risk of CAD. 20,22,43 –46 Contrary to these studies, eNOS G894 T is not associated with the risk of CAD in all the 3 ethnic groups in Malaysia. There are also evidences that indicated a lack of association between eNOS G894 T and the risk of CAD among Taiwanense, 47 Polish, 48 Korean, 49 caucasian, 50 and South Indians. 51,52
In the present study, concomitant presence of the CETP B1 allele and eNOS 4a allele significantly increases the risk of CAD in Malays and Indians but not in the Chinese. The exact mechanism of the interaction of CETP TaqIB polymorphism with eNOS 4a4b polymorphism on the risk of CAD is unclear. CETP TaqIB polymorphism has been associated with the risk of CAD through its influence on HDL-C levels. 29 The HDL has been found to exert important antiatherogenic effects by stimulation of endothelial cell NO production and endothelial repair. Experimental studies have consistently demonstrated the ability of HDL to active eNOS expression and generate NO production. 53,54 Kuvin et al 55 found that endothelial function was abnormal in patients with low HDL-C and well preserved in those with high HDL-C. On the other hand, eNOS 4a4b polymorphism alters eNOS gene expression and has been associated with lower concentrations of endothelial NO metabolites, resulting in endothelial dysfunction. 39,40 Therefore, the concomitant presence of the CETP B1 allele and eNOS 4a allele might enhance the endothelial dysfunction and lead to an increased risk of CAD.
A few experimental studies have suggested an interaction of eNOS polymorphisms with other gene. For example, eNOS 4a4b polymorphism was highly associated with the genetic variants of the gene encoding platelet glycoprotein IIb/IIIa in patients with myocardial infarction. 56 This is the first study, to the best of our knowledge, which has shown the evidence of interaction between CETP B1 allele and eNOS 4a allele with the increased risk of CAD. Previous studies have suggested an interaction of eNOS 894 T allele with methylenetetrahydrofolate reductase (MTHFR) 1298C allele increases the risk of microalbuminuria 57 and CETP B1 allele with eNOS 894 T allele increases the risk of CAD and type 2 diabetes mellitus in Western Iran. 46 However, this study fails to show the interaction between CETP B1 allele and eNOS 894 T allele with the risk of CAD in all 3 ethnic groups. These findings suggest that the role of gene–gene interaction in the pathogenesis of CAD might be ethnic specific.
This study has several limitations. The first is the lack of functional studies. We did not actually measure NO concentrations to prove the biological effect. Second is a limitation of sample size, but these preliminary findings could serve as a reference point for further studies with larger sample sizes.
This is the first study to examine the association between these polymorphisms and CAD in Malaysian population. Our results have suggested that the increased CAD risk associated with the CETP TaqIB polymorphism is specific to Malay population, while the eNOS 4a4b polymorphism is associated with CAD in Malay and Indian population. Moreover, this study also indicated that CETP B1 allele in combination with eNOS 4a allele significantly increases the risk of CAD in Malays and Indians residing in Malaysia but not in Chinese. However, studies with larger sample sizes are needed to confirm these findings.
Footnotes
Acknowledgments
We gratefully acknowledge all the volunteers who kindly took part in this study and our clinical colleagues at Invasive Cardiac Laboratory of Hospital Serdang for obtaining the blood specimens. We also would like to thank Dr Kamaraj, Dr Norzian, Dr Foo, Dr Gary, Dr Ahmad, Dr Nazrul, and Dr Muizz for their generous support of obtaining the blood specimens from the patients. The authors would like to thank the Department of Biomedical Science, Faculty of Medicine and Health Sciences, Universiti Putra Malaysia for the facilities.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by Universiti Putra Malaysia Grant (GP-I/2014/9442200).
