Abstract
Hemophilia A is an X-linked bleeding disorder resulting from heterogeneous mutations in the factor VIII (FVIII) gene. The disease is frequently caused by intron 22 inversion mutation in FVIII gene in approximately 50% cases and by intron 1 inversion (Inv1) in 2% to 5% cases with severe hemophilia A. Both inversion mutations occur due to intrachromosomal recombination between inverted repeats found in intronic sequences in upstream regions located telomeric to the FVIII gene and result in a severe phenotype. Frequency of Inv1 mutation varies in different populations. We have analyzed the frequency of the Inv1 in a group of 110 cases with Hemophilia A from North Indian Population and found 4 inversion-positive cases (3.6%). The Inv1 genotyping was further applied to analyze the hemophilia carrier status of the mother and proband's sister. Direct mutation detection strategy is more perfective in diagnosis of cases, carriers, and in prenatal diagnosis of hemophilia A.
Introduction
Hemophilia A (HA) caused by a deficiency of the coagulation factor VIII (FVIII), is a rare inherited bleeding disorder resulting from heterogeneous mutations in coagulation FVIII gene located on Xq28. The gene is 186 Kb in length and has 26 exons. Mutations described in the FVIII gene are mostly gene rearrangements, point mutations and large deletions and insertions (HAMSTeRS http://europium.mrc.rpms.ac.uk). 1 The only identified mutation hot spot of the FVIII gene is its largest intron (intron 22), with 40% to 50% of severe HA cases arising from intron 22 inversion mutation. After excluding a frequent intron 22 inversion, a complete screening of the large FVIII gene is necessary for direct genetic diagnosis. Intron 1 inversion (Inv1) is another large molecular defect resulting in severe HA. The pathogenic mechanism associated with this inversion involves homologous recombination between a 1041 bp region of intron 1 (int1h-1) of the FVIII gene and an extragenic copy (int1h-2) of region 5 approximately 140 kb telomeric to the FVIII gene. Both Inv1 and intron 22 prevent the formation of full-length FVIII messenger RNA (mRNA) and result in the absence of FVIII proteins leading to severe HA.2 –4 Intron 1 inversion has been reported to occur with a frequency of 5% among British cases with severe HA and may represent the second most common mutation. 2 Later studies indicate that the frequency is around 2% in the same population and also in others.2,5 –8 We have estimated the frequency of the Inv1 in a cohort of 110 unrelated cases with HA from North Indian population and also compared with frequency data from other populations.
Materials and Methods
Patients
The study group included 110 cases with severe HA. The cases were collected through the various city chapters of the Hemophilia Federation of India including Lucknow, Varanasi, Bareilly, Gujarat, Jammu, and Rajasthan. Ethical approval for the study was obtained from the Institutional Ethical Committee of CSM Medical University, India.
DNA Extraction
Peripheral blood, 5-10 mL, was collected in 0.5 M EDTA from patients with prior informed consent. DNA was extracted using Invitrogen Kit (Germany).
Polymerase Chain Reaction Amplification
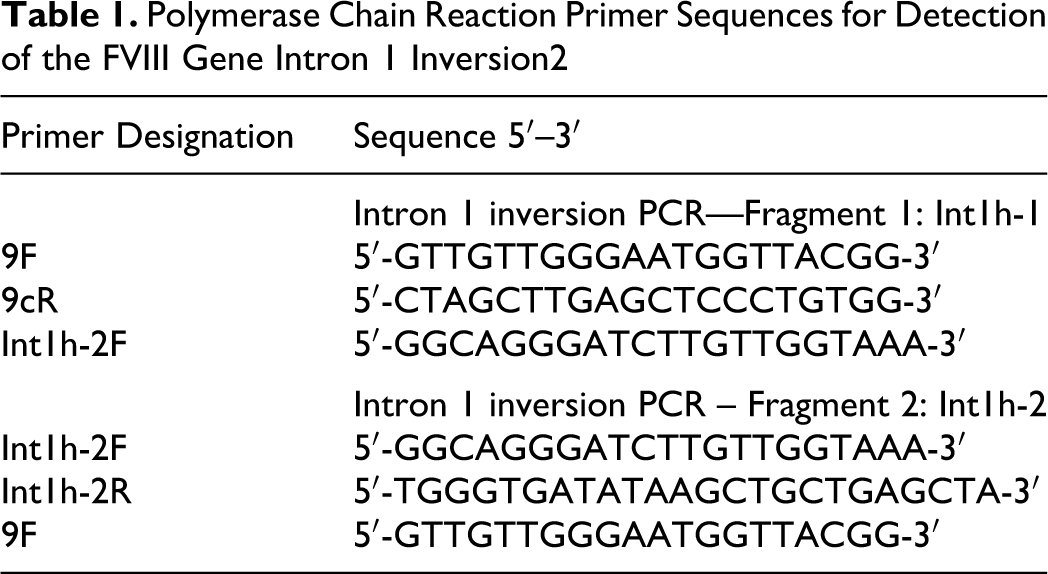
For Inv1 analysis, 2 independent polymerase chain reactions (PCRs) were performed on each genomic DNA sample using method and specific primers as designed by Bagnall et al 2 (Table 1 )
Polymerase Chain Reaction Primer Sequences for Detection of the FVIII Gene Intron 1 Inversion2
Polymerase chain reactions were run with 25 μL reaction mixture containing 100 ng of genomic DNA with 1× Mg2+ free reaction buffer, 2.5 mmol/L Mg2+, 10 mmol/L of each dinuclueotide triphosphate, 10 pmol of each oligonucleotide primer,1× reaction buffer, and 0.3 U (5 U/uL) Taq polymerase enzyme (Roche, Germany) in an Eppendorf Thermocycler ProS (Germany). After initial denaturation for 10 minutes at 94°C, 35 cycles of denaturation at 94°C for 0.30 seconds, annealing at 55.5°C for 0.30 seconds, and extension at 72°C for 2 minutes were carried out, followed by a final extension step at 72°C for 10 minutes. Polymerase chain reaction products were observed in 1% agarose gel electrophoresis.
In the first reaction, 3 different primers that is 9F and 9cR specific for int1h-1 region and int1h-2F primers were used that yielded a 1908 bp product from normal DNA and a 1323 bp product if the inversion was present (Figure 1 ). In the second reaction primer specific for int1h-2 (int1h-2F, int1h-2R) and 9F, which are specific for the telomeric side of the int1h-1 region were used in an amplification reaction that yielded a 1191 bp product from normal DNA and a 1776 bp product in the presence of inversion, whereas female carriers showed 2 bands in the both PCRs (Figure 2 ).
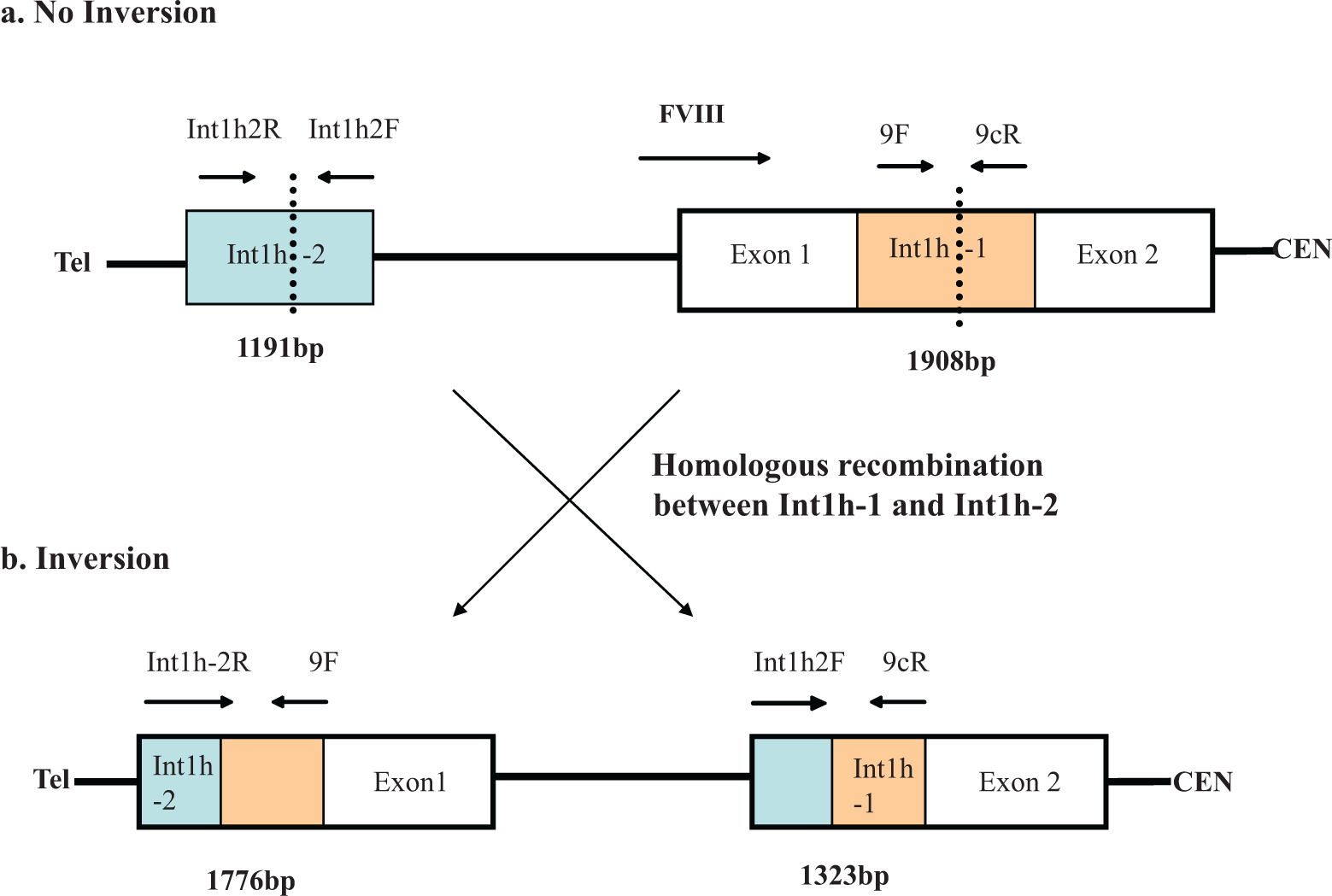
Illustration of (A) normal and (B) inverted intron 1 in factor VIII (FVIII) gene. The Int 1h-1(orange) and Int1h-2 region downstream (blue) are similar. A homologous recombination with inversion occurs. The polymerase chain reactions (PCRs) in normal cases amplify Int 1h-1 region of 1908 bp (primers 9F to 9cR) and int1h-2 region of 1191 bp (primers Int1h-2F to Int1h-2R). In cases with inversion exchange of sets of primers in PCR (Int1h-2R to 9F and Int1h-2F to 9cR.) results in 1776 and 1323 bp fragments.
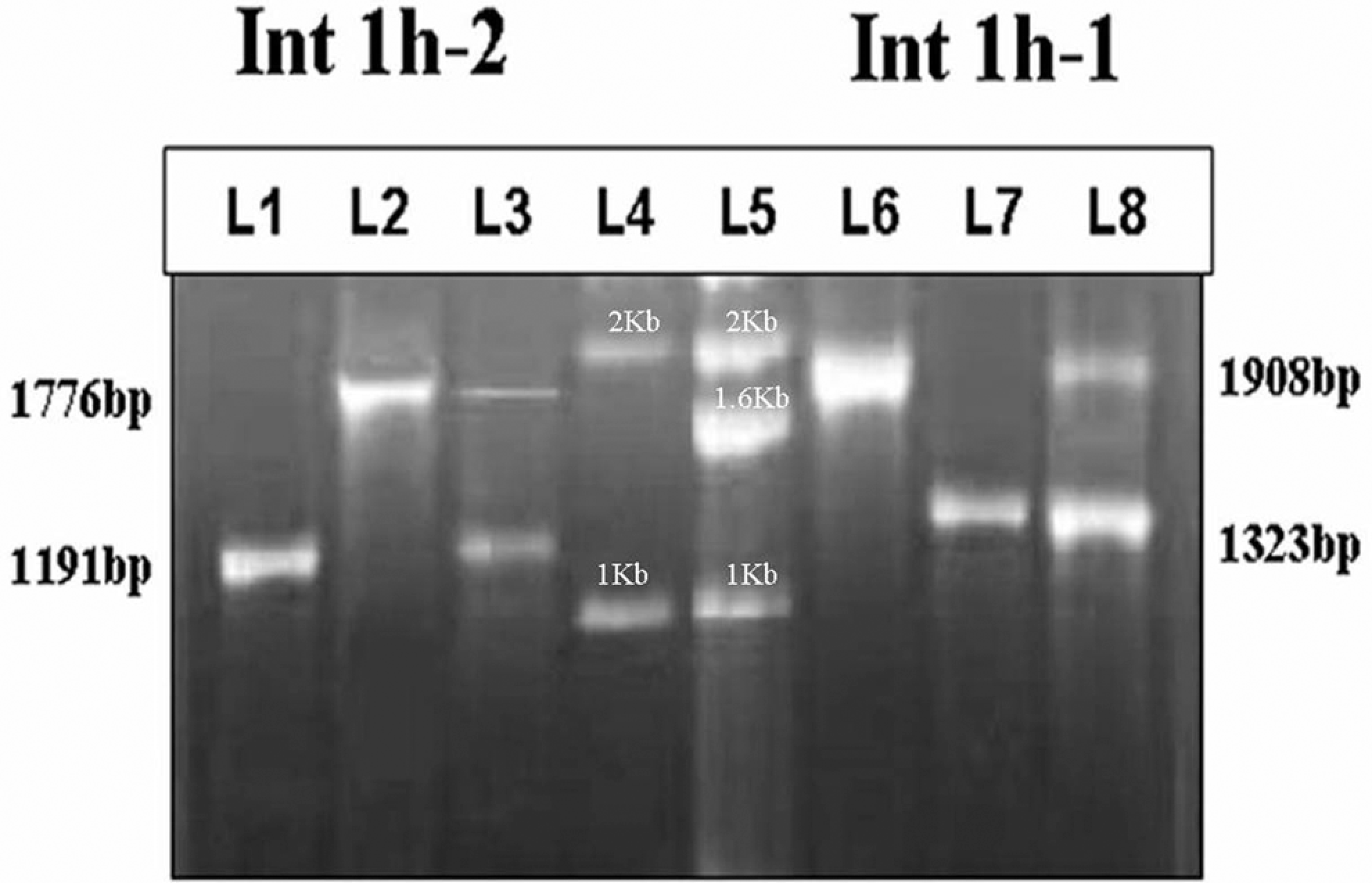
Polymerase chain reaction (PCR) amplification showing intron 1 inversion in Cases with Hemophilia A and carrier mother: A, Lane 1-3 shows PCR for Int1h-2 region; Lane 1: wild-type control, Lane 2: intron 1 inversion–positive CWH; Lane 3: carrier mother. B, Lane 6-8 shows PCR for Int1h-1 region; Lane 6: wild-type control, Lane 7: intron 1 inversion–positive CWH, Lane 8: carrier mother. C, Ladders: Lane 4: 1 kb DNA ladder marker (Bangalore Genei, Bangalore, India), Lane 5: Marker10 (Roche, Germany).
Results and Discussion
In 110 cases with HA studied, our results revealed 4 patients with Inv1 (Figure 1). Pedigree analysis of these cases revealed no family history of HA. In 3 cases, the mother also carried intron 1 mutation; while in 1 case, the mother tested negative, suggesting a sporadic mutation. All the remaining cases tested had the expected size of the normal band in the first and second PCRs. Overall frequency of Inv1 mutation in our population was 3.6%.
Intron 1 inversion mutation was first reported by Brinke et al. 9 They identified an inversion in intron 1 FVIII gene which resulted in the formation of 2 chimeric mRNA fragments of which 1 mRNA fragment was under control of the promoter region of FVIII gene contained the first exon of the FVIII gene and facultative exons with exons 2 to 6 of VBP1 gene. 10 The other mRNA fragment transcribed was under control of C6.1A promoter and contains the last exon of the C6.1A gene and exons 2 to 26 of the FVIII gene with facultative exons. 9 Bagnall et al 2 characterized the break point region, elucidated the mode of the inversion mutation, and also devised a PCR method for its detection.
The overall frequency of Inv1 in our study data was 3.6%. In other Indian studies Shetty et al 5 detected 3 (1.24%) inversion-positive mutations out of a large cohort of 241 severe cases of HA, while an incidence of 3.75% was reported by Ahmed et al in 80 samples from the North Indian population. 6 Reported frequencies of Inv1 in FVIII gene vary from 1.8% to 4.8% in the United Kingdom, 8 1.2% to 3.8% in Indian population,6,11,12 4.3% in the Czech Republic, 13 and 1.26% to 4.6% in China14,15 –17 (Table 2 ).
Prevalence of Intron 1 Inversion in Different Ethnic Populations
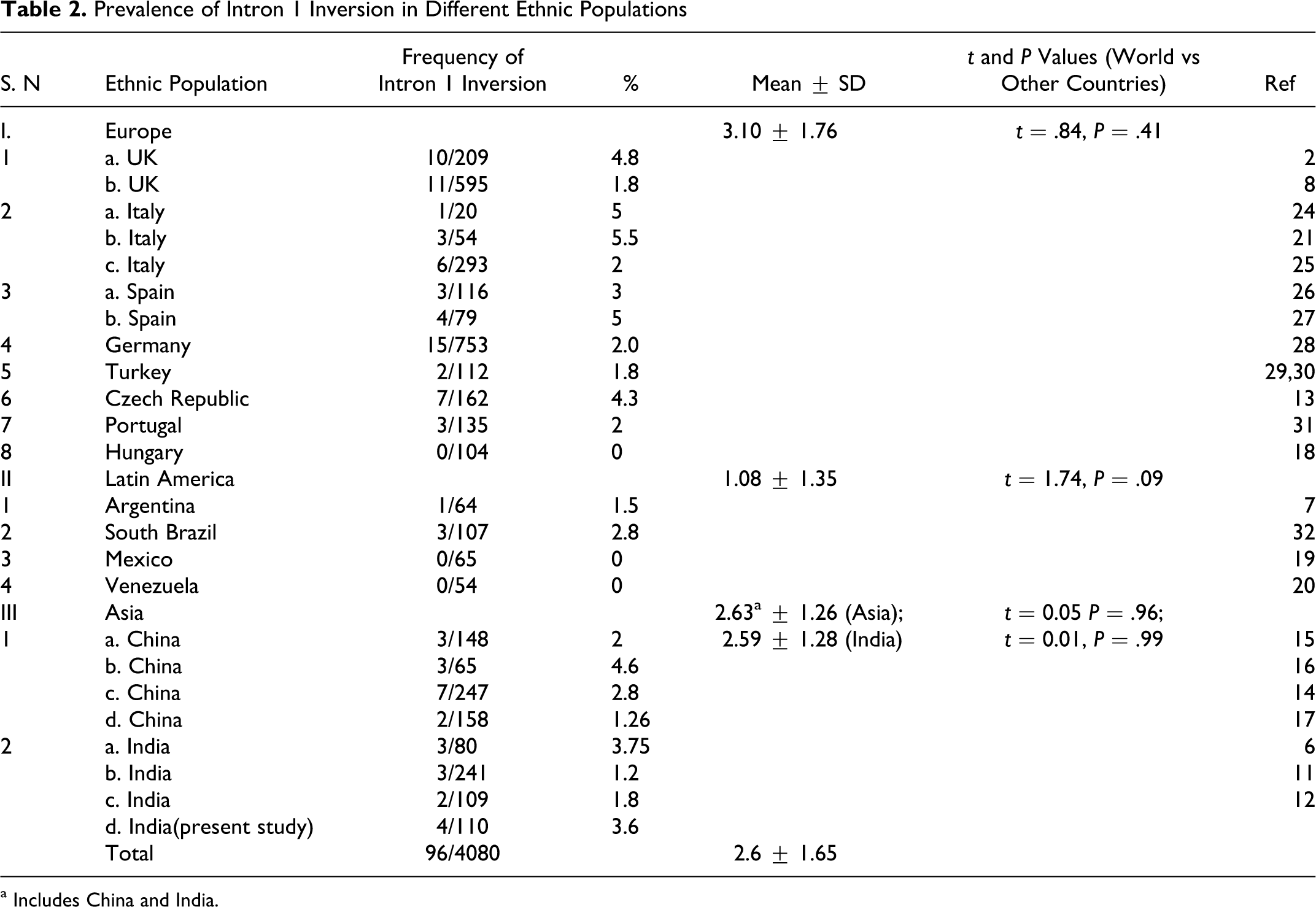
a Includes China and India.
Table 2 shows the results of Inv1 in our cases, comparing them with different series studied in 14 countries. All reviewed studies together include 4080 individuals of which 96 showed Inv1, giving a worldwide mean of 2.6. When grouped into continents, the mean prevalence Inv1 mutation for Europe was 3.10%, for Latin America 1.08% and 2.63% for Asians. The unpaired t test was used to compare the mean prevalence of Inv1 between regions and the overall world mean. Considering a P value <.05 as significant, no significant difference was observed. A minimum incidence of 0% was observed in some studies from Hungary, 18 Mexico, 19 and Venezuela. 20 Larger studies from these countries are required since related ethnic groups shown in the table show the presence of the mutation. Highest incidence of Inv 1 mutation was 5.5%, which was reported from Italy. 21
Germ line origin of Inv1 mutation has been traced to male germ cell meiosis in most cases. 22 Since the X chromosome in male cells is unpaired with a homologous chromosome, it tends to turn over inducing a crossover between homologous copies of FVIII. In female meiosis, the pairing of 2 X chromosome inhibits the gene rearrangements. On the basis of pedigree analysis, 1 out of 4 inversion-positive patients in our series was a sporadic mutation. Mothers in 3 of the 4 cases tested positive for Inv1. Fourth inversion-positive case had no family history of HA, and the mother tested negative for the Inv1 mutation, suggesting a de novo meiotic mutation in the male germ line in the father.
The commonest mutation in HA also occurs due to meiotic homologous recombination between a 9503-base pair sequence (int22h-1) in intron 22 of the F8 gene and one or other of the 2 inverted copies of this sequence (int22h-2 and int22h-3) located, respectively, 500 and 600 kb more telomeric.3,23 It appears therefore that Inv1 mutation is, after the intron 22 inversion, the most common of the FVIII gene mutation causing severe HA.
The results of our study show that frequency of Inv1 mutation in Indian patients with HA is 3.6%. Direct mutation detection is a resource-intensive but an effective method for carrier detection and prenatal diagnosis of HA. We recommend that screening for this mutation should be part of routine genetic diagnosis for this disorder. It is readily detectable by PCR and effective in diagnosing carriers at risk. Direct mutation detection strategy should be implemented in developing countries since definitive diagnosis can be achieved in cases and it can be applied where all family members are not available.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: financial support from Department of Science & Technology Govt. of India Grant no: SR/SO/HS/102/2007 and University Grant Commission (RGNF), New Delhi, India.
