Abstract
Objective:
Pregnane X receptor (PXR) is a transcriptional regulator of many drug-metabolizing enzymes including cytochrome P450 (CYP) 2C9. The objective of this study was to assess the possible association between PXR single-nucleotide polymorphisms (SNPs) and stable warfarin doses.
Methods:
A total of 201 patients with stable warfarin doses from the EwhA-Severance Treatment (EAST) Group of Warfarin were included in this study. The influence of genetic polymorphisms on stable warfarin doses was investigated by genotyping 11 SNPs, that is, vitamin K epoxide reductase complex 1 (VKORC1) rs9934438, CYP2C9 rs1057910, CYP4F2 rs2108622, constitutive androstane receptor (CAR) rs2501873, hepatocyte nuclear factor 4α (HNF4α) rs3212198, and PXR (rs3814055, rs1403526, rs3732357, rs3732360, rs2276707 and rs2472682). Subgroup analysis was conducted on CYP2C9 wild-type homozygote allele (AA) carriers.
Results:
One PXR SNP of rs2472682 (A>C) exhibited significant association with stable warfarin doses in study population and the subgroup; variant homozygote carriers required significantly lower daily doses of warfarin than those carrying wild allele by about 0.8 mg. Approximate 43.7% of overall interindividual variability in warfarin dose requirement was explained by multivariate regression model. VKORC1, CYP2C9, age, CYP4F2, PXR rs2472682, and CAR/HNF4α rs2501873/rs3212198 accounted for 29.6%, 5.9%, 3.7%, 2.3%, 1.3%, and 0.9% of the variability, respectively. PXR SNP of rs2472682 remained a significant factor in CYP2C9 wild-type homozygote carriers based on univariate and multivariate analyses. The combination of CAR/HNF4α/PXR SNPs of rs2501873/rs3212198/rs2472682 showed about 1 mg dose difference between grouped genotypes in study population and subgroup.
Conclusion:
Our results revealed that PXR could be a determinant of stable warfarin doses.
Introduction
Warfarin is the most frequently prescribed oral anticoagulant to prevent and treat thromboembolic disorders. Because warfarin has a narrow therapeutic window and large inter- and intraindividual variability, the effectiveness and safety of warfarin are critically dependent on maintaining therapeutic international normalized ratio (INR). Variations in individual warfarin dose requirement are largely attributable to genetic factor that accounts for 35% to 50% of the variability. 1 A number of studies have confirmed that genetic polymorphisms in cytochrome P450 (CYP) 2C9, vitamin K epoxide reductase complex (VKORC) 1, and CYP4F2 make significant contributions to variability in warfarin dose requirements. 2 –5
Genetic variants in CYP2C9 are closely associated with warfarin responses because warfarin undergoes metabolism by this polymorphic enzyme. CYP2C9 accounts for 80% to 85% of S-warfarin metabolism. S-warfarin is known to be 3 to 5 times more potent than R-warfarin. CYP2C9 is estimated to metabolize about 16% of clinical drugs. 6
Constitutive and altered expression of genes are largely governed by transcriptional control of certain nuclear receptors (NRs). 7 Recently, the significant role and genetic influence of NR superfamily have been extensively explored. 8 Androstane receptor (CAR/NR1I3) and pregnane X receptor (PXR/NR1I2) have emerged as key NRs that regulate drug-metabolizing enzymes (DMEs) including CYP2C9. Hepatocyte nuclear factor α (HNF4α/NR2A1) is another well-known transcriptional master regulator that affects complex cross talk between CAR and PXR. 6,9,10 These receptors are involved in transcriptional induction of warfarin metabolism enzyme. They could cause significant pharmacological consequence of warfarin. We have previously reported combined effects of CAR and HNF4α single-nucleotide polymorphisms (SNPs) on stable warfarin doses. 11
Based on our previous study, we examined the impact of PXR variants on stable warfarin doses in Korean patients with mechanical cardiac valves in this study. The PXR polymorphism effects on warfarin doses were further evaluated using a subgroup cohort comprising CYP2C9 wild-type homozygote carriers.
Methods
Study Population
A group of 229 patients (the EwhA-Severance Treatment [EAST] Group of Warfarin) who had undergone prosthetic mechanical cardiac valve replacements at Severance Cardiovascular Hospital of Yonsei University College of Medicine between January 1982 and December 2009 were recruited, of which 201 with stable warfarin doses were selected for this study. Subgroup analysis was conducted on 186 patients with CYP2C9 *1/*1 allele carriers. The study protocol was approved by the Ethics Committee of the Severance Hospital Institutional Review Board (IRB No. 2009-4-0283). Written informed consent was obtained from each patient before participation in this study.
Collection of Clinical Data
Data collection consisted of review of inpatient and outpatient medical records. Patients’ data included age, gender, age at time of the operation, body weight, body mass index, duration of warfarin therapy, position of valve prosthesis, valve types, comedications, comorbidity, and stable warfarin doses during warfarin therapy. Stable warfarin dose was defined as mean daily dose when an INR of 2 to 3 was maintained for at least 3 consecutive times.
Variant Selection and Genotyping
To select PXR SNPs, genetic information of the PXR gene was incorporated into the Haploview Program. 12 There were 22 SNPs in the PXR gene, with a minor allele frequency more than 20% in Japanese and Han Chinese populations. Linkage disequilibrium blocks were constructed following the D’-method in Haploview. 13 Tagger function within the Haploview was used to assign tag SNPs. Six tag SNPs (rs3814055, rs1403526, rs3732357, rs3732360, rs2276707, and rs2472682) in the PXR gene captured common variations within the gene and the surrounding area with a minimum r2 of 0.80. In addition to the selected PXR SNPs, VKORC1 rs9934438 (C>T), CYP2C9 rs1057910 (A>C), CYP4F2 rs2108622 (C>T), CAR rs2501873 (C>T), and HNF4α rs3212198 (C>T) polymorphisms were further included in this study. Therefore, a total of 11 SNPs were evaluated for this study.
Genomic DNA from EDTA blood samples of patients were extracted using QIAamp DNA Blood Mini Kit (QIAGEN GmbH, Hilden, Germany) according to the manufacturer’s protocol. Genotyping was conducted by single base primer extension assay using ABI PRISM SNaPShot Multiplex kits (ABI, Foster City, California) or TaqMan genotyping assay using RT-PCR system (ABI 7300, ABI, Foster City).
Statistical Analysis
Independent sample t test was used to compare 2 groups. One-way analysis of variance followed by post hoc Bonferroni test was used for multiple comparisons (mean doses among more than 2 different genotype groups). Multiple linear regression analysis was used to determine factors that independently affected interindividual variability of warfarin dose requirements. Stepwise selection of variables was entered into a regression method that met the default criteria of P < .05 for entry and P > .1 for removal. All analyses were performed with the SPSS Version 20.0 for Windows (SPSS 20.0K, SPSS INC, Chicago, Illinois). A P value less than .05 was considered statistically significant.
Results
Patients were followed up from June 1983 to May 2012 after their cardiac valve replacements. Follow-up periods ranged from 1.0 to 29.7 years. Interindividual warfarin dose variation among the study population ranged from 2.2 to 14.1 mg/d. Mean stable warfarin dose was 5.4 ± 1.9 mg/d. As reported in our previous article, atrial fibrillation was significantly associated with warfarin doses. 5
Observed genotype frequencies were consistent with Hardy-Weinberg equilibrium for all SNPs. The locations of the 6 selected PXR SNPs in the PXR gene were intron region (n = 4), 3′-untranslated region (UTR; n = 1), and upstream region (n = 1). Of the 6 PXR SNPs studied, rs2472682 (A>C) exhibited significant (P = .040) association with daily warfarin doses, with variant-homozygote (CC) carriers requiring significantly lower daily doses of warfarin than those carrying wild A allele (AA and AC; 5.38 ± 1.94 mg and 5.73 ± 1.86 mg vs 4.81 ± 1.65 mg, Table 1).
Differences in Stable Warfarin Doses Among Genotypes in Study Population.

Abbreviations: SD, standard deviation; VKORC, vitamin K epoxide reductase complex; CYP, cytochrome P450; CAR, constitutive androstane receptor; HNF4α, hepatocyte nuclear factor 4α; PXR, pregnane X receptor; UTR, untranslated region.
Multivariate analyses were performed with patient characteristics (age, gender, and atrial fibrillation) and SNP variables that had P value less than .2 in univariate analysis. The final multiple linear regression model explained 43.7% of overall interindividual variability in warfarin dose requirements among study patients. PXR rs2472682 was significantly (P = .034) associated with stable warfarin doses after adjusting potential confounding factors. Vitamin K epoxide reductase complex 1, CYP2C9, age, CYP4F2, PXR rs2472682, and CAR/HNF4α rs2501873/rs3212198 accounted for 29.6%, 5.9%, 3.7%, 2.3%, 1.3%, and 0.9% of the variability, respectively (Table 2).
Multiple Linear Regression Model for Stable Warfarin Doses in Study Population.a

Abbreviations: VKORC, vitamin K epoxide reductase complex; CYP, cytochrome P450; CAR, constitutive androstane receptor; HNF4α, hepatocyte nuclear factor 4α; PXR, pregnane X receptor.
aAdjusted for VKORC1 rs9934438, CYP2C9 rs1057910, CYP4F2 rs1057910, age, gender, atrial fibrillation, CAR/HNF4α rs2501873/rs3212198 combination, and PXR rs2472682.
Differences in stable warfarin doses in CYP2C9 wild-type homozygote carriers with gene polymorphisms are summarized in Table 3. Vitamin K epoxide reductase complex 1, CYP4F2, and PXR rs2472682 remained significant (P < .05) factors for stable warfarin doses in the subgroup (Table 3). Multiple linear analysis for stable warfarin doses in CYP2C9 wild-type homozygote carriers was summarized in Table 4. Vitamin K epoxide reductase complex 1, age, CYP4F2, PXR rs2472682, and CAR/HNF4α rs2501873/rs3212198 accounted for 30.1%, 3.5%, 3.0%, 1.4%, and 0.1% of the variability, respectively (Table 4).
Differences in Stable Warfarin Doses Among Genotypes in CYP2C9 Wild-Type Homozygote Carriers.
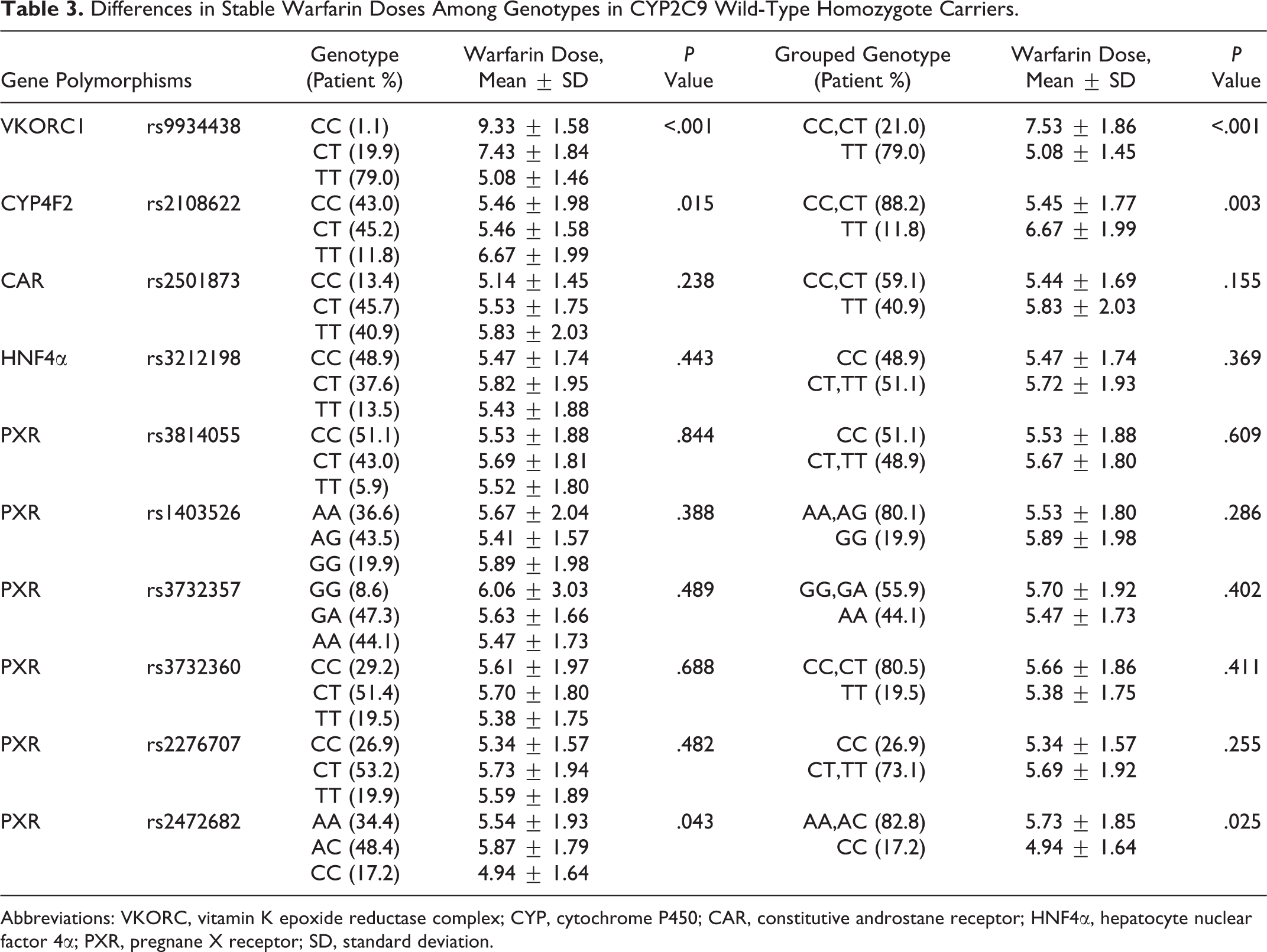
Abbreviations: VKORC, vitamin K epoxide reductase complex; CYP, cytochrome P450; CAR, constitutive androstane receptor; HNF4α, hepatocyte nuclear factor 4α; PXR, pregnane X receptor; SD, standard deviation.
Multiple Linear Regression Model for Stable Warfarin Doses in CYP2C9 Wild-Type Homozygote Carriers.a
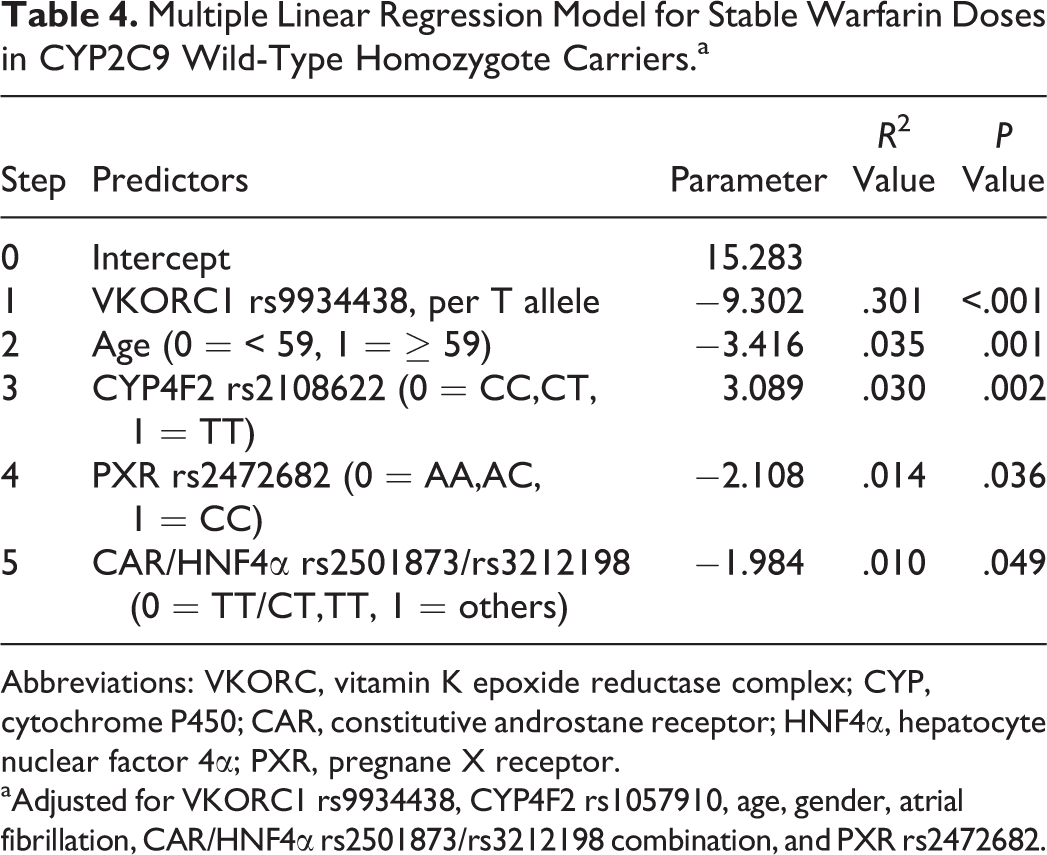
Abbreviations: VKORC, vitamin K epoxide reductase complex; CYP, cytochrome P450; CAR, constitutive androstane receptor; HNF4α, hepatocyte nuclear factor 4α; PXR, pregnane X receptor. aAdjusted for VKORC1 rs9934438, CYP4F2 rs1057910, age, gender, atrial fibrillation, CAR/HNF4α rs2501873/rs3212198 combination, and PXR rs2472682.
Effect of combined genotypes of CAR rs2501873/HNF4α rs3212198/PXR rs2472682 SNPs on stable warfarin doses in the study population is shown in Figure 1. The median warfarin daily dose was significantly different between the 2 groups as classified by genotype combination in the total study population (6.4 ± 2.5 mg vs 5.3 ± 1.7 mg; P = .018) and its subgroup (6.4 ± 2.5 mg vs 5.4 ± 1.6 mg; P = .041). Patients with variant-homozygote in CAR rs2501873 (TT), variant allele in HNF4α rs3212198 (T allele), and wild allele in PXR rs2472682 (A allele) required significantly (P < .05) higher warfarin doses than those with other combinations (Figure 1).

Effects of combined genotypes on stable warfarin doses in the total study population or cytochrome P450 (CYP) 2C9 wild-type homozygote allele (AA) carriers subpopulation. A, Effects of combined genotypes of constitutive androstane receptor (CAR) rs2501873/hepatocyte nuclear factor 4α (HNF4α) rs3212198/pregnane X receptor (PXR) rs2472682 single nucleotide polymorphisms (SNPs) on stable warfarin doses in total study population. B, Effects of combined genotypes of CAR rs2501873/HNF4α rs3212198/PXR rs2472682 SNPs on stable warfarin doses in CYP2C9 wild-type homozygote allele (AA) carriers. Box and whisker plots showing the distribution of stable warfarin doses by combined effect of CAR, HNF4α, and PXR polymorphic genotypes (CAR rs2501873/HNF4α rs3212198/PXR rs2472682 SNPs combination). The horizontal line indicates the median dose. Boxes cover the 25 to 75 percentiles. Whiskers are 5 to 95 percentiles. The numbers above whiskers represent mean values. Each outlier is shown by a point.
Discussion
The absorption, distribution, metabolism, and excretion (ADME) of drugs are performed by drug transporters as well as phase I (oxidation, reduction, or hydrolysis) or phase II (conjugation) DMEs. The expression of many important DMEs is largely under transcriptional control by specific NR family members. Of various DMEs, CYP3A4, CYP2B6, CYP2C9, and CYP2C19 are responsible for metabolizing more than 80% of clinically used drugs. Recent studies have shown that CAR and PXR (both belonging to the NR superfamily) are originally identified as xenobiotic receptors regulating CYP gene expression. 7,14 Both CAR and PXR are predominantly expressed in the liver, intestine, and tissues with high capacity for drug metabolism sharing unique characters such as ligand promiscuity, species differences, and regulation of the expression of a large number of overlapping sets of genes encoding phase I and phase II DMEs. 15
Although CAR and PXR overlap largely on their target genes, they differ in the regulation of transcriptional activities. Both CAR and PXR contain DNA binding domain (DBD) and ligand binding domain (LBD), respectively, and they bind to responsive elements and recruit coactivators to affect structure, therefore increasing the transcription of target genes after being activated by exposure to xenobiotics. There are now approximately 50 NRs working with more than 200 coactivators. 7,16,17 Therefore, the elucidation of CAR- and PXR-mediated signaling is very complex. They form a network of molecular cross-talk with other transcription factors, including HNF4α.
HNF4α is expressed at high levels in the liver, kidney, pancreas, and small intestine. HNF4α is involved in the regulation of expression and variety of CYP genes in human liver. 18 It has been shown that there is a cross-talk between the distal CAR/PXR binding sites and the 2 proximal HNF4α-binding sites in the CYP2C9 promoter. HNF4α synergizes with CAR/PXR to enhance the induction of CYP2C9 in HepG2 cell. 19 Compelling evidences have shown that the so-called “master regulator,” namely HNF4α, plays a dominant role in CAR- and PXR-mediated induction of CYP3A4 and expression of CYP2C9. 6,10 These findings suggest that the maximal and synergic transcriptional induction of CYP2C9 gene might be achieved through a cross talk between HNF 4α and CAR/PXR via cofactors or other hepatic factors. 16,19
To date, many splice variants of CAR and PXR have been established. These polymorphisms in the coding or noncoding regions of CAR and PXR are associated with functional change in CAR and PXR and further affecting drug metabolism. 14 Recently, many studies have been undertaken to identify whether genetic polymorphisms in regulatory NRs (PXR, CAR, and HNF4α) could make an implication on specific drug responses such as docetaxel and doxorubicin through changes in expression and/or activity of the CYP genes. 8,20
Based on the facts that S-warfarin is mainly metabolized by CYP2C9, we examined the effects of CYP2C9 regulating transcription factors, PXR, CAR, and HNF4a, on warfarin response. Of the 6 PXR SNPs examined in this study, 1 intronic SNP showed a statistically significant effect on warfarin maintenance doses. For the PXR rs2472682 A>C SNP, the CC genotype was associated with a lower warfarin maintenance doses compared to those with allele A (about 0.8 mg/d) in univariate analysis. It was found that PXR rs2472682 along with combined SNPs of CAR rs2501873 and HNF4α rs3212198 remained significant factors for stable warfarin doses after adjusting confounding factors. These results suggest that certain variants of CAR, HNF4α, and PXR could alter the expression of CYP2C9 gene, therefore affecting warfarin metabolism and sensitivity. Since these 3 transcription factors work together, and their polymorphisms have a small effect on stable warfarin doses, it may be that a combination of polymorphisms of transcription factors would be used to help guide warfarin therapy. The combination of CAR/HNF4α/PXR SNPs of rs2501873/rs3212198/rs2472682 showed about 1 mg dose difference between grouped genotypes in this study.
To investigate the effects of transcription factors on stable warfarin doses while ruling out the influence of CYP2C9, subgroup analysis was carried out using patients with CYP2C9 wild-type homozygote carriers. Results from subgroup were similar to those from whole study population. Combination of PXR SNP and CAR/HNF4α SNP showed significant effect on stable warfarin doses based on multivariate analysis, indicating that stable warfarin doses could be affected by variants in CYP2C9 transcription factors without any polymorphism effect of CYP2C9 itself. We were not able to conduct subgroup analysis with CYP2C9 variant carriers because the number of patients carrying CYP2C9 mutant-allele is 14, which was too small for statistical analysis.
Although PXR rs2472682, CAR rs2501873, and HNF4α rs3212198 are detected in the intron region, these noncoding region SNPs could also give rise to altered messenger RNA (mRNA) stability and degradation, gene expression, and alternative splicing, resulting in different protein isoforms. 14,21 These SNPs also could alleviate interaction with their partner microRNA (miRNA) and noncoding RNA molecules to regulate target genes by binding to complementary regions of transcripts, resulting in repression of their translation or mRNA degradation. 22 Recent data have indicated the possibility of miRNA-mediated PXR posttranscriptional regulation. It has been reported that miR-148a could recognize the complementary sequence in the 3′-UTR mRNA and lead to downregulation of PXR protein and its target genes. 23
Our algorithm could explain 43.7% of the variations in stable warfarin doses in Korean patients with cardiac mechanical valves. Previous studies from the United Kingdom and the United States reported that the CYP2C9 gene polymorphisms (*2 and *3) explain about 15% to 27% of the interindividual warfarin dose variation. 24,25 In an Asian study, the predictive contribution of CYP2C9 was 3.94%, 26 which was significantly lower compared with that of the Caucasian population. In our cohort, CYP2C9 variability accounted for about 5.9% of the overall warfarin dose variability. It is possible that the relatively low contributions of the CYP2C9 gene on the Asian population can explain lower R 2 in this study.
To our knowledge, this is the first report to investigate the effect of genetic variations of PXR genes on warfarin response in a clinical setting. There are several issues to consider in applying study results to clinical decision. First, study population was restricted to patients with cardiac valve replacements for securing homogeneity in this study. Because the association between genotypes and drug responses is not significantly affected by patient diseases, it is expected that study results could be applied to other populations such as patients with atrial fibrillation or venous thromboembolism. Second, PXR contributed to warfarin dose variability only about 1%. Nevertheless, result was meaningful in terms of dosage difference (5.58 mg vs 4.81 mg) because, in clinical practice, warfarin dose adjustments are made in 0.5 mg scale and doses are rounded to the nearest 0.5 mg.
In conclusion, as the role of NRs in the regulation of genes associated with warfarin metabolism becomes clearer, further studies on the correlation between other genetic variants of transcription factors and warfarin sensitivity are merited.
Footnotes
Author Contributions
Jung Yeon Moon contributed to acquisition, analysis, and interpretation; drafted the manuscript; gave final approval; and agreed to be accountable for all aspects of work ensuring itegrity and accuracy. Byung Chul Chang contributed to conception and design; critically revised the manuscript; gave final approval; and agreed to be accountable for all aspects of work ensuring itegrity and accuracy. Kyung Eun Lee contributed to acquisition and analysis; drafted the manuscript; gave final approval; and agreed to be accountable for all aspects of work ensuring itegrity and accuracy. Jun Seok Bang contributed to acquisition and analysis; drafted the manuscript; gave final approval; and agreed to be accountable for all aspects of work ensuring itegrity and accuracy. Hye Sun Gwak contributed to conception and design; contributed to interpretation; critically revised the manuscript; gave final approval; and agreed to be accountable for all aspects of work ensuring itegrity and accuracy.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
