Abstract
Introduction
Methylenetetrahydrofolate reductase (MTHFR) polymorphisms modulate folate metabolism and carcinogenesis, yet their role in lung adenocarcinoma (LUAD) susceptibility and prognosis, particularly in underrepresented populations, remains unclear.
Methods
We conducted a retrospective case-control study of 306 lung cancer (LC) patients (predominantly LUAD) and 156 healthy controls of Guangxi Zhuang ethnicity, with prognostic follow-up of the cases. Genotyping (rs1801131/rs1801133), serum lactate dehydrogenase (LDH) levels, and tumor MTHFR protein expression were analyzed. Associations with susceptibility (logistic regression), survival (Cox models/Kaplan-Meier), and molecular mechanisms were evaluated.
Results
The MTHFR rs1801131 GG genotype conferred increased LUAD susceptibility (adjusted OR = 2.14, 95% CI: 0.99–4.65, P = 0.054) and predicted poor overall survival (OS, P = 0.043) and progression-free interval (PFI, P = 0.023). Among all LC subtypes, the GG genotype was also associated with worse prognosis (P = 0.033). Patients with both GG genotype and elevated LDH (eLDH) had higher mortality OS risk than those with TT or GG genotype with eLDH in the LC and LUAD cohorts (P = 0.034 and 0.042). Mechanistically, the GG genotype reduced tumor MTHFR protein expression (P < 0.05). The GG (rs1801131-G/rs1801133-G) haplotype elevated LUAD risk by 1.53-fold (P = 0.022).
Conclusion
In the Guangxi Zhuang cohort, the MTHFR rs1801131 GG genotype may represent a population-specific marker associated with LUAD susceptibility and poorer prognosis. Its combination with elevated LDH may provide supplementary information for risk stratification, although further validation in larger multicenter and multi-ethnic cohorts is required.
Plain Language Summary
Introduction
Lung adenocarcinoma (LUAD), the predominant histological subtype of lung cancer, remains the leading cause of global cancer mortality, with 5-year survival rates persistently below 20%. 1 In China, LUAD accounts for over 40% of all lung cancer cases, contributing significantly to the national burden of ∼870,000 new cases and ∼760,000 deaths annually. 2 While smoking is a major risk factor, LUAD frequently develops in non-smokers (particularly in Asian populations), underscoring the crucial role of germline genetic susceptibility.3,4 Furthermore, despite refinements in TNM staging and identification of targetable drivers (e.g., EGFR, ALK), approximately 35%-40% of patients experience unanticipated therapeutic failure, indicating there is unexplained clinical heterogeneity.5,6 This highlights an urgent need for novel biomarkers that elucidate subtype-specific mechanisms and refine risk stratification.
Dysregulated epigenetic control, particularly aberrant DNA methylation, and compromised genomic integrity are established as central drivers of lung cancer pathogenesis and disparate outcomes. 7 DNA methylation, essential for genomic stability and transcriptional regulation, is dependent on folate-mediated one-carbon metabolism for the supply of methyl groups via S-adenosylmethionine (SAM). 8 Methylenetetrahydrofolate reductase (MTHFR), a pivotal enzyme in this pathway, catalyzes the irreversible conversion of 5,10-methylenetetrahydrofolate to 5-methyltetrahydrofolate, directing folate pools towards homocysteine remethylation and SAM synthesis. 9 Functional polymorphisms in MTHFR (notably C677T [rs1801133] and A1298C [rs1801131]) significantly reduce enzymatic activity. 10 This impairment is mechanistically linked to genome-wide DNA hypomethylation which is a well-established precursor to chromosomal instability, oncogene activation, and tumor suppressor silencing due to altered cellular folate distribution and reduced SAM availability.11-13 Accumulating evidence suggests that these mechanisms contribute to the pathogenesis of LUAD. Nevertheless, epidemiological studies on the association between MTHFR variants and LUAD susceptibility or prognosis remain inconclusive, particularly among underrepresented populations such as the Zhuang ethnic group, the largest minority group in China, with a distinct genetic background.14-16 Existing studies suffer from ethnic bias, focusing predominantly on European or Han Chinese populations while neglecting the Zhuang ethnic group. There is a mechanistic gap: no prior LUAD research integrates germline MTHFR genotypes (e.g., rs1801131/rs1801133) with functional tumor phenotypes and key metabolic biomarkers like serum lactate dehydrogenase (LDH). Moreover, prognostic synergy between MTHFR variants and LDH which is a well-established marker of tumor glycolysis and LUAD progression,17,18 remains clinically unexplored, limiting biomarker-driven risk stratification.
Here, we address these questions through an integrated analysis of 306 Zhuang patients with lung cancer. We hypothesize that the MTHFR rs1801131 GG genotype contributes to LUAD susceptibility and adverse prognosis via reduced protein expression, with this effect potentially strengthened by elevated LDH levels. Our study integrates genotyping, tumor MTHFR protein quantification, and serum LDH measurements to investigate whether the rs1801131 GG genotype may serve as a population-specific marker for risk stratification in this underrepresented population.
Materials and Methods
Study Population and Ethics
This was a retrospective, single-center, hospital-based case-control study, with additional prognostic follow-up analyses among cases. The reporting of this study conforms to the STROBE guidelines. 19 A total of 700 consecutive patients with clinically suspected lung cancer (LC) were initially screened at the First Affiliated Hospital of Guilin Medical University (July 2021–March 2024). All lung cancer diagnoses adhered to the Chinese Society of Clinical Oncology (CSCO) Clinical Guidelines for Lung Cancer (2022) and were pathologically verified according to the WHO Classification of Thoracic Tumours (5th edition, 2021). Exclusion criteria included: (1) history of autoimmune disorders; (2) acute hepatic/renal dysfunction; (3) pregnancy; (4) familial relationships within the cohort; (5) histological classification failure (e.g., adenosquamous carcinoma or unclassifiable NSCLC); (6) non-primary lung malignancies; (7) genotyping failed; (8) Other groups of non-Zhuang nationality. The final analytical cohort comprised 306 histologically confirmed LC cases classified: 160 lung adenocarcinoma (LUAD) patients, 77 lung squamous cell carcinoma (LUSC) patients and 63 small cell lung cancer (SCLC) patients. A total of 156 age- and sex-frequency-matched Zhuang healthy controls (HC) were recruited from the health screening program of the same center. The study protocol was approved by the Institutional Review Board of the First Affiliated Hospital of Guilin Medical University (approval No. 2023QTLL-36; approved on December 15, 2023). Written informed consent was obtained from all participants. The study was conducted in accordance with the Declaration of Helsinki of 1975, as revised in 2024. All patient data were de-identified before analysis. Overall Survival (OS): defined as the time from diagnosis to death from any cause. Progression-Free Interval (PFI): defined as the duration from treatment initiation to radiological/clinical disease progression or cancer-related re-hospitalization or death from any cause (whichever occurred first). The median follow-up duration was estimated using the reverse Kaplan–Meier method.
SNP Selection and Genotyping
Functional polymorphisms were prioritized based on: (1) functional genomic regions: all SNPs localize to key regulatory or coding regions: MTHFR rs1801131 (T>G, p.E429A) and rs1801133 (G>A, p.A222V): exonic missense variants (alter enzymatic activity) (2) population frequency: minor allele frequency (MAF) >5% in East Asians (NCBI SNP database). (3) pathogenic evidence: established roles in lung carcinogenesis from the literature.
Genotyping was performed using the SNPscan™ platform with locus-specific probes (three per SNP: two allele-specific 5′-probes and one common 3′-probe) (Supplemental Table S1). DNA samples (30-50 ng/μL verified by agarose electrophoresis) underwent thermal lysis (4 μL DNA + 2.5 μL buffer at 98°C/5 min), followed by ligation reactions (10 μL lysate + 1 μL probe mix + 0.5 μL ligase; 94°C/1 min → 58°C/4 h → 72°C/2 min). Multiplex PCR amplified ligation products (1 μL) with touchdown cycling: 95°C/2 min; 9 cycles of 94°C/20 s → 62-57°C (-0.5°C/cycle)/40 s → 72°C/1.5 min; 25 cycles of 94°C/20 s → 57°C/40 s → 72°C/1.5 min. PCR products (1:10 diluted) were electrophoresed on ABI 3730xl with Liz600™ size standard after denaturation (95°C/5 min). Genotypes were called using GeneMapper 4.1, with quality controls including interplate negatives (n=3/96-well), 10% duplicate replicates (>99% concordance).
Immunohistochemical (IHC) Validation
Stratified analysis of representative formalin-fixed paraffin-embedded lung cancer tissues across histological subtypes was performed based on genotype. Tissue sections (4 μm) underwent antigen retrieval in citrate buffer (pH 6.0), followed by incubation with primary antibody against MTHFR (UpingBio, #YP-Ab-04024). Immunoreactivity was visualized using DAB chromogen and independently scored by two board-certified pathologists blinded to genotype. Staining intensity was quantified as follows: 0 (no staining), 1+ (faint/discernible staining in ≤10% cells), 2+ (moderate staining in >10% cells), and 3+ (strong/diffuse staining in >10% cells). Integrated optical density (IOD) per unit area was determined via digital image analysis (ImageJ v1.53).
Baseline Assessment and Laboratory Procedures
Hepatic and renal function indices for all participants were quantified in our ISO 15189-accredited laboratory (Certification No. ML00036). All assays were performed in strict compliance with manufacturer protocols for reagent kits and instrument operation manuals using Roche Cobas E701/E801 immunoassay analyzers. Based on the established clinical threshold, 19 values ≥180 U/L were classified as elevated lactate dehydrogenase (eLDH) and values <180 U/L as non-elevated LDH (neLDH). Blood specimens were collected contemporaneously with SNP genotyping to maintain batch-to-batch consistency.
Statistical Analysis
Statistical analyses were conducted using IBM SPSS 27.0 and R 4.2.1. Categorical data are presented as frequencies (n) or percentages [n (%)]. Pearson’s chi-square test assessed group differences in categorical variables, while the chi-square goodness-of-fit test evaluated Hardy-Weinberg equilibrium (HWE). Continuous data normality was assessed by the Kolmogorov-Smirnov test. Normally distributed data are expressed as mean ± standard deviation (SD) and analyzed using independent samples t-test (two groups) or one-way ANOVA (multiple groups). Non-normally distributed data are presented as median (interquartile range) [M (P25-P75)] and analyzed with the Mann-Whitney U test (two groups) or Kruskal-Wallis test (multiple groups). Binary logistic regression evaluated associations between biochemical markers, genetic SNPs, and LC susceptibility, reporting odds ratios (OR) with 95% confidence intervals (CI). Survival analyses were performed in R (survival package v3.3.1). Univariate Cox proportional hazards regression was first used to screen candidate variables, and only covariates with P < 0.10 in the corresponding univariate analysis were entered into the multivariate Cox regression model. This prespecified strategy was applied to reduce model complexity and limit potential overfitting. Kaplan–Meier survival curves were generated and visualized using survminer (v0.4.9) and ggplot2 (v3.3.6). Haplotype analysis was performed using SHEsis 20 (https://analysis.bio-x.cn/myAnalysis.php). Protein expression differences across genotypes were analyzed by ANOVA followed by Tukey’s post-hoc test. Statistical significance was defined as P < 0.05. No formal sample size calculation was performed; the sample size was determined by the number of eligible participants available during the study period.
Results
Comparative Analysis of Clinical and Biochemical Profiles
Clinical Characteristics of the Study Participants
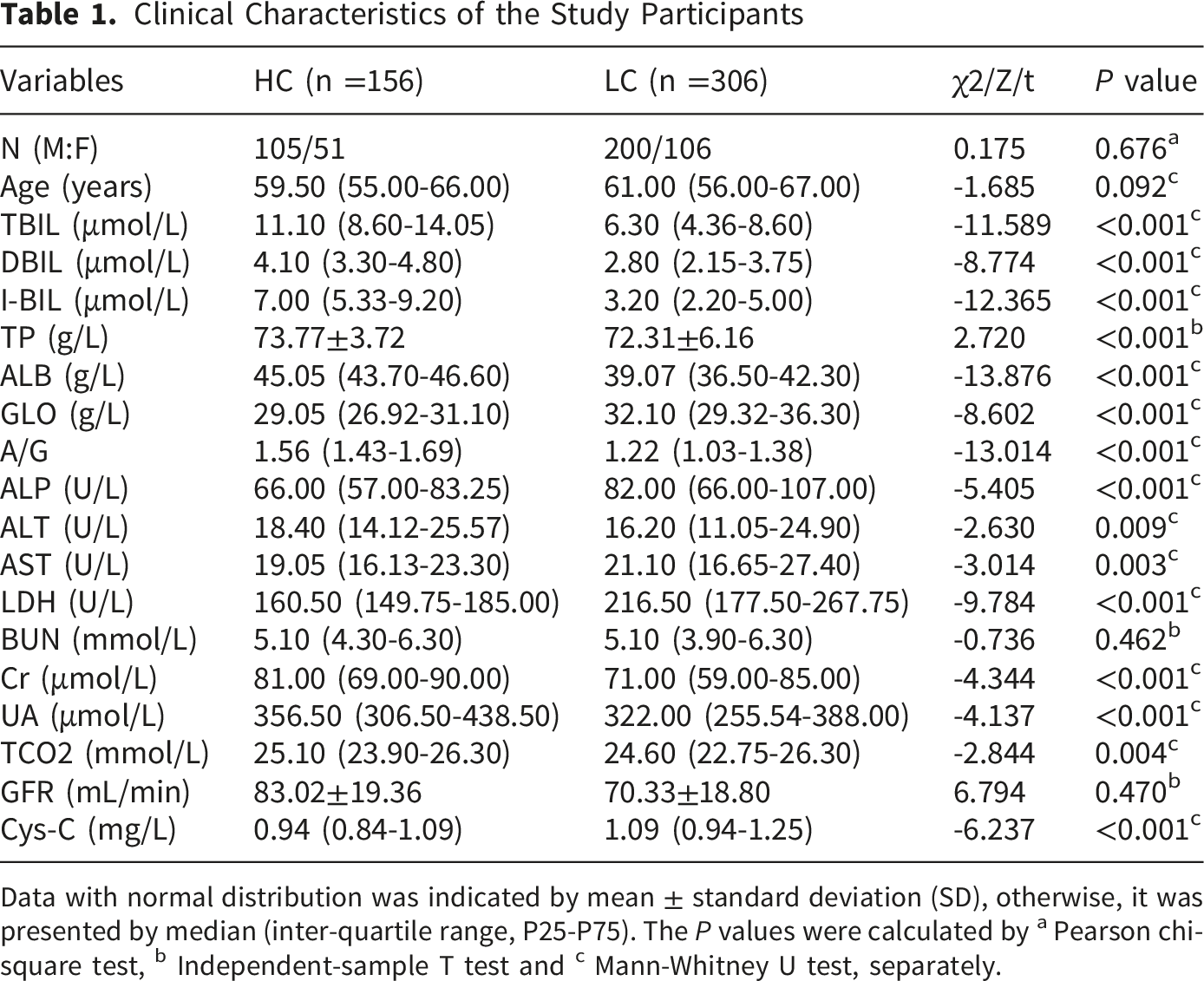
Data with normal distribution was indicated by mean ± standard deviation (SD), otherwise, it was presented by median (inter-quartile range, P25-P75). The P values were calculated by a Pearson chi-square test, b Independent-sample T test and c Mann-Whitney U test, separately.
MTHFR Genetic Polymorphisms and Lung Cancer Susceptibility
Genotyping of MTHFR variants (rs1801131, rs1801133) was performed using multiplex fluorescence PCR (Supplemental Figure S1). All polymorphisms conformed to Hardy-Weinberg equilibrium in controls, patients, and overall cohorts (P > 0.05), indicating no significant deviation from the expected genotype distribution..
Association Between Gene Polymorphisms and Healthy Individuals Versus LC Patients

HC, health control; LC, lung cancer. Data are presented as n (%). P values were calculated using the Pearson chi-square test.
Figure 1 summarizes statistically significant regression outcomes. In LC patients, (GG vs. TT) showed a significant association (adjusted OR = 2.614, 95% CI: 1.316–5.191, P = 0.006) (Supplemental Table S3). In the LUAD subgroup analysis, GG genotype carriers exhibited a trend toward increased LUAD risk relative to the TT genotype (unadjusted OR = 2.086, 95% CI: 0.976–4.456, P = 0.058) (Figure 1A). This association strengthened marginally after age and sex adjustment (adjusted OR = 2.143, 95% CI: 0.988–4.648, P = 0.054) (Figure 1B) (Supplemental Table S4). These findings suggest that the GG genotype may be associated with an increased risk of LUAD and warrant further investigation in larger cohorts. For LUSC, the GG genotype was infrequent, and the differences between patients and healthy controls were not statistically significant (P = 0.210) (Supplemental Table S2), although trends towards an association were observed (OR = 2.548, 95% CI: 0.830–7.824, P = 0.102) (Supplemental Table S5). For SCLC, the GG genotype was also associated with susceptibility in the subgroup analysis (TT vs GG, P = 0.022), with an OR of 6.596 (95% CI: 1.396–31.172, P = 0.017) (Figure 1B; Supplemental Table S6). However, given the extremely small number of GG genotype carriers in this subgroup, this finding should be interpreted with caution. The forest plots of positive results in the logistic regression analysis. * adjusted for age and gender
MTHFR Haplotype Associations in Lung Cancer Susceptibility
Analysis of haplotypes combining the rs1801131 and rs1801133 polymorphisms (order: rs1801131→ rs1801133) revealed a significant association with susceptibility specifically to LUAD (global P = 0.038). The GG haplotype (rs1801131-G/rs1801133-G) was significantly associated with an increased risk of developing LUAD (OR = 1.527, 95% CI: 1.062–2.195, P = 0.022). Conversely, the TG haplotype (rs1801131-T/rs1801133-G) was significantly associated with a decreased risk of LUAD (OR = 0.708, 95% CI: 0.518–0.967, P = 0.029). No significant global or individual haplotype associations were observed for overall LC (global P = 0.154), LUSC (global P = 0.842), or SCLC susceptibility (global P = 0.652) (Supplemental Table S7).
MTHFR rs1801131 GG Genotype Predicts Poor Survival in LUAD
Univariate and Multivariate Analyses of OS in LC Patients

CI, confidence interval; LUAD, lung adenocarcinoma; LUSC, lung squamous carcinoma; SCLC, small cell lung cancer; HR, hazard ratio; OS, overall survival; PR, partial response; SD, stable disease; PD, progressive disease.
Univariate and Multivariate Analyses of OS in LUAD Patients

CI, confidence interval; LUAD, lung adenocarcinoma; LUSC, lung squamous carcinoma; SCLC, small cell lung cancer; HR, hazard ratio; OS, overall survival; PR, partial response; SD, stable disease; PD, progressive disease.
Univariate and Multivariate Analyses of PFI in LUAD Patients
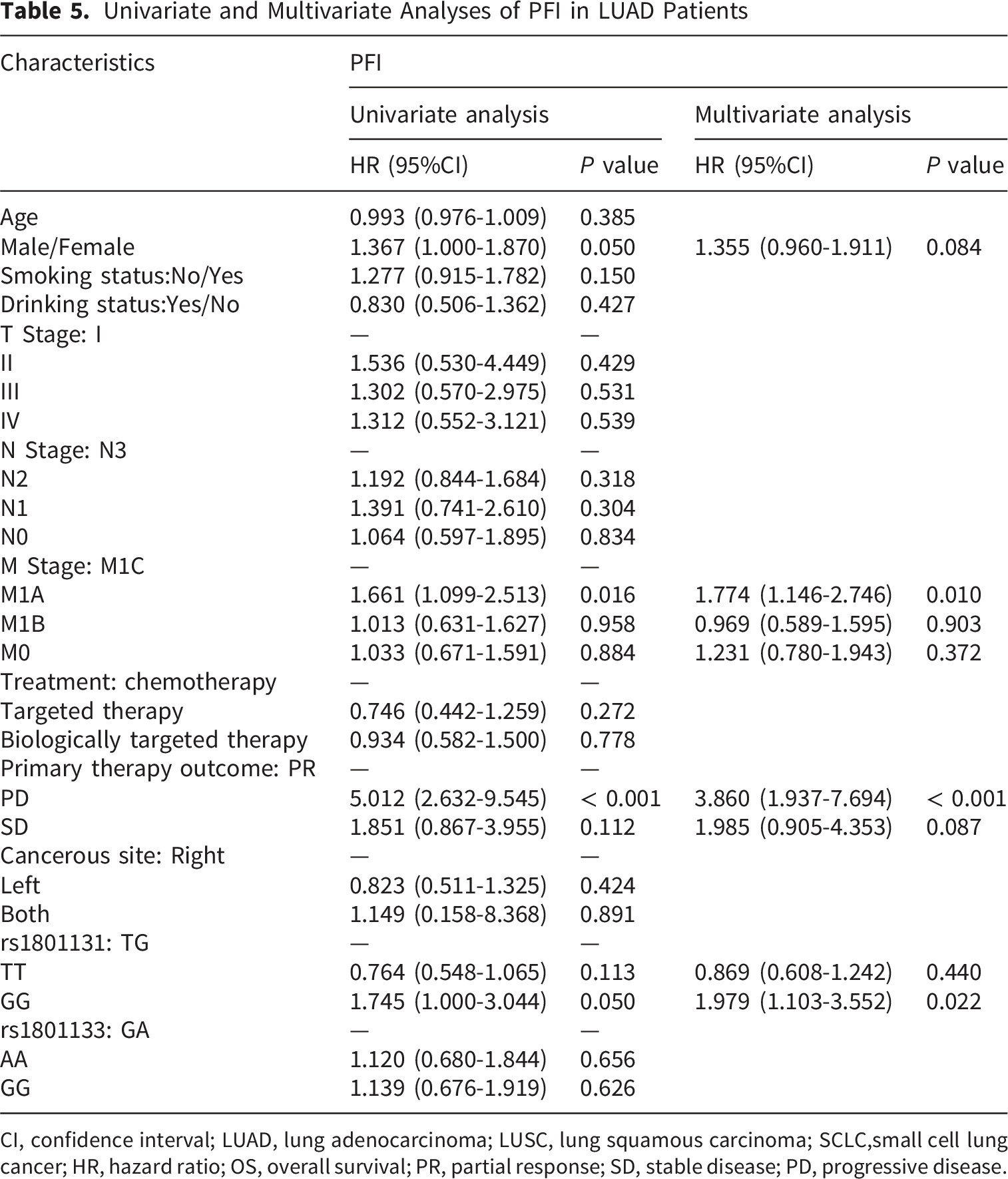
CI, confidence interval; LUAD, lung adenocarcinoma; LUSC, lung squamous carcinoma; SCLC,small cell lung cancer; HR, hazard ratio; OS, overall survival; PR, partial response; SD, stable disease; PD, progressive disease.

Kaplan–meier survival curves of overall survival (OS) and progression-free interval (PFI) for LC patients. (A) OS for all LC patients with rs1801131 polymorphism; (B) OS for LUAD patients with rs1801131 polymorphism; (C) PFI for LUAD patients with rs1801131 polymorphism; (D) OS for all LC patients with rs1801131 polymorphism and eLDH; (E) OS for LUADD patients with rs1801131 polymorphism and eLDH
Combined Analysis of MTHFR Genotype and LDH Expression Reveals Enhanced Prognostic Risk
To further elucidate the interplay between MTHFR genotype and tumor metabolism, patients were stratified by serum LDH expression levels (high: eLDH; normal: neLDH) and combined with rs1801131 genotypes. Within the entire lung cancer cohort, patients harboring the rs1801131 GG genotype and elevated LDH (GG-eLDH) exhibited significantly worse overall survival compared to those with TG/TT genotypes and high LDH expression (TG-eLDH/TT-eLDH) (P = 0.034) (Figure 2D). In LUAD, it was also found that patients with GG-eLDH had significantly lower overall survival rates compared to TG-eLDH and TT-eLDH (P = 0.042) (Figure 2E).
Association Between MTHFR Gene Polymorphism (rs1801131) and its Protein Levels in LUAD
Upon categorizing and examining the immunohistochemical outcomes by genotype in LUAD tissues, it was discovered that the rs1801131-GG genotype was associated with a lower protein expression level compared to both the TG and TT genotypes (Figure 3). In addition, no significant differences in protein levels were observed among the rs1801131 genotypes in other histological types of LC examined (LUSC and SCLC). Relationship between protein expression and rs1801131 genotype of MTHFR in LUAD patients. (A) rs1801131-TT; (B) rs1801131-TG; (C) rs1801131-GG; (D) differential statistics of MTHFR genotypes and protein levels in LUAD patients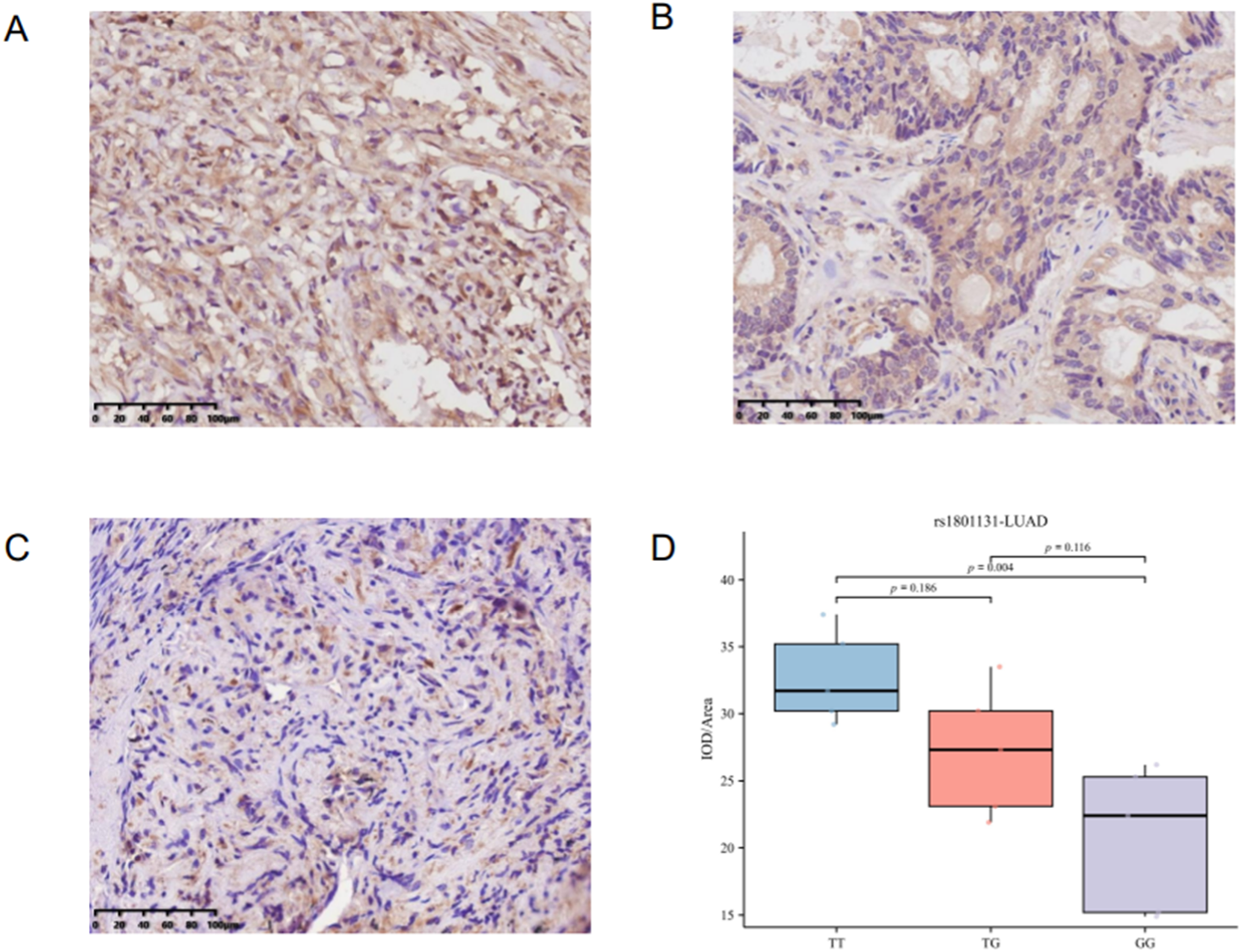
Discussion
In the present study, focusing on the Guangxi Zhuang population, we found that the MTHFR rs1801131 GG genotype was associated with poorer survival in patients with LUAD, as supported by multivariate Cox regression and Kaplan–Meier analyses. Patients carrying the GG genotype combined with elevated LDH also showed worse overall survival in both the overall lung cancer cohort and the LUAD subgroup. Given the limited data available for this ethnic group, our findings provide population-specific evidence and suggest that rs1801131, particularly in combination with LDH, may have prognostic value, although further validation in larger and multi-ethnic cohorts is needed.
Molecular genetic research has increasingly linked various genetic and epigenetic alterations to the pathogenesis of LC. Among these, polymorphisms in the MTHFR gene are significant, as they reduce enzymatic activity and expression, thereby disrupting folate metabolism and initiating a cascade of downstream events. 21 Currently, only two common MTHFR polymorphisms, rs1801133 (C677T) and rs1801131 (A1298C), have been conclusively shown to impact enzymatic function. 22 The rs1801133 variant involves a C>T substitution resulting in an alanine-to-valine replacement at position 222 (A222V) within the catalytic domain, which reduces enzymatic efficiency by approximately 35%. 23 In contrast to rs1801133 homozygosity, which is strongly associated with hyperhomocysteinemia, 24 the rs1801131 (A1298C) risk allele (C variant) does not typically elevate homocysteine levels. Nevertheless, this polymorphism similarly reduces MTHFR activity, potentially impairing folate cycling and associated metabolic processes. 25
Our findings suggest that the MTHFR rs1801131 GG genotype significantly elevates LUAD susceptibility and independently predicts adverse prognosis in the Guangxi Zhuang population—a contrast to studies reporting null associations in non-Asian cohorts.26,27 This discrepancy likely stems from ethnic-specific genetic architectures and histological stratification. Recent East Asian evidence further supports the possibility that ancestry-specific molecular contexts contribute to heterogeneity in LUAD susceptibility and may partly explain why certain loci show stronger effects in specific populations than in European cohort. 28 While rs1801131 showed no overall LC risk in European 29 or non-Hispanic White populations, 27 its effect was pronounced in LC or LUAD subgroups in Asians14-16 and Turkey. 30 Notably, the GG haplotype (rs1801131-G/rs1801133-G) conferred 1.53-fold increased LUAD risk (P = 0.022), aligning with reports of elevated DNA adducts in high-risk smokers carrying this variant. 31
The synergistic interaction between the rs1801131-GG genotype and eLDH underscores the established role of LDH as a biomarker of tumor metabolic reprogramming. Elevated serum LDH reflects enhanced glycolysis (Warburg effect) and correlates with aggressive phenotypes, metastasis, and poor survival in NSCLC.19,32 We showed that rs1801131-GG carriers exhibit both reduced tumor MTHFR protein and LDH-associated risk amplification, suggesting a model of metabolic dysregulation. The rs1801131 chronically impairs folate cycling.25,33 This disruption may alter NAD+/NADH homeostasis (33)—a key regulator of LDH activity—explaining the allele’s association with elevated serum LDH even in non-oncogenic contexts (e.g., high-intensity athletes 34 ). This study revealed that the combined effect of rs1801131 and LDH on the prognosis of LC and LUAD may account for the inconsistent findings regarding rs1801131’s association with chemotherapy toxicity, complications, and poor prognosis in LC patients reported in the current literature,35-38,39likely due to the lack of stratification for this factor. Taken together, these observations suggest that reduced MTHFR expression combined with elevated LDH may be associated with a more aggressive phenotype in LUAD; however, the underlying functional consequences and mechanistic basis still require further validation in dedicated mechanistic studies.
Several limitations of this study should be acknowledged. First, this was a single-center study conducted in a Zhuang cohort, which may limit the generalizability of the findings to other populations. Second, the sample sizes of certain subgroup analyses were relatively small, particularly in the SCLC subgroup, where the number of patients carrying the rs1801131 GG genotype was extremely limited. This restricted the statistical power of the subgroup analyses and may affect the robustness of the subgroup-specific observations. Therefore, these findings, especially those derived from the SCLC subgroup, should be interpreted cautiously and considered preliminary. Future studies should include larger multicenter and multi-ethnic cohorts, together with functional investigations and external validation, to further clarify the biological and clinical relevance of this association. Despite these limitations, our findings suggest that the combination of rs1801131 genotype and LDH may offer supplementary information for risk stratification in LUAD within the Guangxi Zhuang population.
Conclusion
In the Guangxi Zhuang cohort, the MTHFR rs1801131 GG genotype may represent a population-specific marker associated with LUAD susceptibility and poorer prognosis. Its combination with elevated LDH may provide supplementary information for risk stratification, although further validation in larger multicenter and multi-ethnic cohorts is required.
Supplemental Material
Supplemental Material - MTHFR rs1801131 GG Genotype Combined With Serum LDH Level Predicts Lung Adenocarcinoma Susceptibility and Poor Prognosis in Guangxi Zhuang Population
Supplemental Material for MTHFR rs1801131 GG Genotype Combined With Serum LDH Level Predicts Lung Adenocarcinoma Susceptibility and Poor Prognosis in Guangxi Zhuang Population by Chao Zuo, Yuanyuan Wang, Dongli Yang, Jing Cheng, Mengna Guo, Zhuo Yang, Yu Wang, Feng Wang and Yongchao Qiao in Cancer Control.
Supplemental Material
Supplemental Material - MTHFR rs1801131 GG Genotype Combined With Serum LDH Level Predicts Lung Adenocarcinoma Susceptibility and Poor Prognosis in Guangxi Zhuang Population
Supplemental Material for MTHFR rs1801131 GG Genotype Combined With Serum LDH Level Predicts Lung Adenocarcinoma Susceptibility and Poor Prognosis in Guangxi Zhuang Population by Chao Zuo, Yuanyuan Wang, Dongli Yang, Jing Cheng, Mengna Guo, Zhuo Yang, Yu Wang, Feng Wang and Yongchao Qiao in Cancer Control.
Footnotes
Acknowledgments
The authors of this article express gratitude for the collaborative efforts and contributions made by all members involved.
Ethical Considerations
This study was approved by the Institutional Review Board of the First Affiliated Hospital of Guilin Medical University (approval No. 2023QTLL-36; approved on December 15, 2023).
Consent to Participate
Written informed consent was obtained from all participants. The study was conducted in accordance with the Declaration of Helsinki of 1975, as revised in 2024. All patient data were de-identified before analysis. All patients provided written informed consent prior to their inclusion in the study.
Author contributions
Conceptualization: Y.Q., F.W.; Methodology: C.Z., Y.W., D.Y., J.C.; Formal analysis: M.G., Z.Y., Y.W.; Investigation: C.Z., Y.W., D.Y., J.C.; Writing - original draft: C.Z.; Writing - review & editing: Y.Q., F.W.; Supervision: Y.Q., F.W.; Funding acquisition: Y.Q., F.W.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Natural Science Foundation of Guangxi Zhuang Autonomous Region (2025GXNSFHA069198) and the Guangxi medical and health appropriate technology development and application project (S2024041).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
All data associated with this study are available in the main text or the supplementary materials. The original data analyzed in this study are available from the corresponding author upon reasonable request.
Use of Artificial Intelligence
Artificial intelligence was not used in the preparation of this manuscript.
Patient and Public Involvement
It was not appropriate or possible to involve patients or the public in the design, or conduct, or reporting, or dissemination plans of our research.
Supplemental Material
Supplemental material for this article is available online.
