Abstract
Introduction
Current cervical cancer screening guidelines recommend colposcopy referral for women co-positive for high-risk human papillomavirus (hrHPV) and abnormal cytology (≥ASC-US). However, cytology exhibits suboptimal sensitivity, and this strategy leads to high colposcopy burdens, especially in populations where non-HPV16/18 genotypes predominate. We evaluated a novel triage strategy using HPV16/18 genotyping and PAX1 methylation to optimize resource allocation.
Methods
This was a retrospective cohort study. In a cohort of 3,233 hrHPV-positive women who underwent HPV genotyping, liquid-based cytology (TCT), PAX1 methylation testing, colposcopy, and histopathological confirmation, we compared two strategies: (A) standard referral (hrHPV+ & TCT ≥ ASC-US); (B) novel referral (HPV16/18+ → immediate colposcopy; non-16/18 hrHPV+ → colposcopy if PAX1 methylation ΔCt ≤ 8.79). CIN3+ (CIN3 or cancer) served as the clinical endpoint.
Results
Non-16/18 hrHPV types (especially HPV52/58) accounted for over 30% of infections and 60% of CIN2+ lesions. PAX1 methylation was strongly associated with lesion severity (median ΔCt: chronic cervicitis 17.92 vs. CIN3 7.36 vs. cancer 5.97; P < 0.001) and predicted CIN3+ with high accuracy (AUC = 0.82 in non-16/18 group). Strategy B detected 40 additional CIN3+ cases (+30.5%), reduced colposcopy referrals by 853 cases (absolute −25%, relative −46%), and increased the positive predictive value (PPV) from 6.6% to 14.6% (2.2-fold improvement) compared to Strategy A.
Conclusion
A triage algorithm combining HPV16/18 genotyping with PAX1 methylation significantly enhances CIN3+ detection while substantially reducing immediate colposcopies. This strategy is particularly well-suited for Chinese and other Asian populations where non-16/18 hrHPV types are prevalent, offering a more precise, cost-effective approach toward WHO’s 2030 cervical cancer elimination goals.
Keywords
Introduction
Cervical cancer remains a major public health challenge, particularly in low- and middle-income countries (LMICs), where it is the fourth most common cancer among women.1,2 Despite being one of the most preventable malignancies, an estimated 604,000 new cases and 342,000 deaths occurred globally in 2020, with over 90% of the burden concentrated in LMICs. 1 The World Health Organization (WHO) has launched a global strategy to eliminate cervical cancer as a public health problem by 2030, with a core target of 70% of women screened by age 35 and again by 45 using a high-performance test. 3 Primary high-risk human papillomavirus (hrHPV) testing has been endorsed as the preferred screening modality due to its superior sensitivity and longer safe rescreening intervals compared to cytology alone.4,5
However, the high sensitivity of hrHPV testing comes at the cost of reduced specificity—particularly in younger women, where transient infections are common. 6 Without effective triage, widespread hrHPV screening would lead to an unsustainable surge in colposcopy referrals, overwhelming health systems and exposing women to unnecessary anxiety and invasive procedures. Consequently, the development of accurate, objective, and scalable triage strategies is now the central bottleneck in the implementation of WHO’s elimination agenda.
Current guidelines often rely on cytology (e.g., ≥ASC-US) or partial HPV genotyping (e.g., immediate referral for HPV16/18-positive women) for risk stratification.7-9 Yet cytology suffers from well-documented limitations: poor inter-observer reproducibility, substantial false-negative rates (up to 30–50% for high-grade lesions), and inadequate performance in resource-limited settings where quality control is challenging. Meanwhile, HPV16/18 genotyping alone is insufficient in many Asian populations, where non-16/18 hrHPV types—particularly HPV52 and HPV58—account for a disproportionate share of cervical precancers and cancers. 10 In China, for instance, HPV52 and HPV58 together surpass HPV18 in prevalence and collectively contribute to over 30% of CIN3+ cases, 11 rendering a 16/18-only triage strategy both insensitive and inequitable.
This epidemiological reality has spurred interest in molecular biomarkers that reflect the host cellular response to persistent oncogenic HPV infection. Among these, DNA methylation of tumor suppressor gene promoters has emerged as a robust indicator of neoplastic progression. 12 Epigenetic silencing via promoter hypermethylation is a hallmark of carcinogenesis, occurring early and accumulating with lesion severity.13,14 The paired box 1 (PAX1) gene, a developmental transcription factor normally silenced in adult cervical epithelium, becomes aberrantly methylated in HPV-transformed cells. 15 Numerous studies have demonstrated that PAX1 methylation levels correlate strongly with histological grade, offering high discriminatory power for CIN3+—especially in hrHPV-positive women.15-17
Critically, however, most validation studies of PAX1 have been conducted in populations where HPV16/18 predominate, or have treated all hrHPV types as a homogeneous group. The performance of PAX1 specifically among non-16/18 hrHPV carriers—a group constituting the majority of screen-positive women in East Asia—has not been rigorously evaluated in large, real-world cohorts with complete pathological follow-up. Moreover, no study has prospectively compared a genotype-stratified triage algorithm (HPV16/18 → immediate referral; non-16/18 → PAX1-based referral) against the current standard of HPV+cytology co-testing in terms of both clinical sensitivity and system-level efficiency.
We address these critical gaps using data from 3,233 Chinese women who underwent comprehensive evaluation including HPV genotyping, liquid-based cytology, PAX1 methylation quantification, colposcopy, and histopathological confirmation. We hypothesized that a dual-path triage strategy leveraging HPV16/18 status and PAX1 methylation would simultaneously improve CIN3+ detection and drastically reduce immediate colposcopy referrals compared to conventional cytology-based triage. Our findings not only validate PAX1 as a powerful biomarker in non-16/18 hrHPV carriers but also provide a clinically actionable, cost-effective framework for precision cervical cancer screening in high-burden, resource-constrained settings—directly supporting the operationalization of WHO’s 2030 elimination goals.
Methods
Study Design and Population: A Retrospective Cohort Study
This was a real-world, retrospective cohort study. This study initially identified 3,253 consecutive women who underwent colposcopy at our institution between January 2023 and December 2024. The predefined inclusion criterion required completion of hrHPV genotyping as part of the integrated screening protocol. Among these, 3,233 women (99.4%) underwent hrHPV genotyping and were included in the primary analytical cohort. The remaining 20 women (0.6%) presented with visible cervical lesions on clinical examination and proceeded directly to colposcopy and biopsy without prior HPV testing; although all were subsequently confirmed to have CIN2+ histopathology, they were excluded from the primary analyses due to non-compliance with the inclusion criterion of hrHPV testing. A sensitivity analysis confirmed that inclusion or exclusion of these 20 cases did not materially alter the study conclusions (data not shown).
HPV genotyping, liquid-based cytology (TCT), and PAX1 methylation analysis were performed on the same cervical specimen collected at initial screening. Patients were referred for colposcopy per Chinese national guidelines (hrHPV-positive with TCT ≥ ASC-US), either from our hospital’s screening program or from community health centers within our regional referral network. All cervical specimens, including those from community health centers, were transported to and tested centrally at our institution’s laboratory. Specimens were stored at −80°C until analysis. While HPV and TCT were performed as part of standard clinical care, PAX1 methylation testing was performed retrospectively on residual DNA after study enrollment.
Data Collection
The study was conducted in accordance with the Helsinki Declaration of 1975, as revised in 2024. All patient identifiers were removed prior to analysis. Participants were consecutively enrolled from the hospital’s cervical screening database between January 2023 and December 2024 based on predefined eligibility criteria. The reporting of this study conforms to the STROBE guidelines. 18
Information was retrieved from the hospital’s electronic medical records system, including:
Demographics: Age.
HPV genotyping was performed using the Hybribio HPV GenoArray assay (Hybribio Biotech, China), a flow-through hybridization-based method that simultaneously detects 21 HPV genotypes, including:
15 High-Risk Types: HPV 16, 18, 31, 33, 35, 39, 45, 51, 52, 53, 56, 58, 59, 66, 68.
6 low-risk types: HPV 6, 11, 42, 43, 44, 81.
Cytology (TCT) Results
Interpreted according to the 2014 Bethesda System and categorized as: negative, ASC-US, ASC-H, LSIL, HSIL, or carcinoma. For diagnostic performance analyses, a threshold of ≥ASC-H was applied, defining ASC-H, LSIL, HSIL, and carcinoma as positive results.
NILM (cytology) and chronic cervicitis (histology) represent equivalent benign states at different diagnostic levels; NILM cases with biopsy confirmation were classified as chronic cervicitis.
Colposcopy Findings
Performed by experienced gynecologists following standardized international protocols. Colposcopic impressions were categorized based on the presence of acetowhite epithelium, mosaicism, punctuation, and/or atypical vessels, and classified as: normal, low-grade colposcopic impression, high-grade colposcopic impression, or suspicious for invasive cancer. Targeted biopsies were obtained from areas with abnormal colposcopic features. Final diagnosis was based exclusively on histopathological examination; colposcopic impressions were not used for diagnostic classification.
Clinical Decision Protocol
(a) Colposcopy referral strictly followed Chinese National Guidelines (2021): hrHPV-positive AND cytology ≥ ASC-US;(b) Biopsy indications: acetowhite lesions, abnormal vessels, or any visible abnormality;(c) Histopathological review: two independent senior pathologists blinded to HPV/PAX1 results; discordant cases adjudicated by a third pathologist.
Histopathological Diagnosis
All women who underwent colposcopy received cervical biopsy or multi-site biopsies. Histological diagnoses were independently and blindly reviewed by two senior pathologists, and final diagnoses were made according to WHO classification standards, categorized as: chronic cervicitis, CIN1, CIN2, CIN3, adenocarcinoma in situ (AIS), or invasive carcinoma (CA). For this study, CIN3, AIS, and CA were grouped into a single endpoint defined as CIN3+.
PAX1 Methylation Testing: PAX1 methylation testing was performed on the same cervical brush specimen collected for HPV genotyping at the initial screening visit, prior to colposcopy referral. PAX1 methylation was quantified using quantitative methylation-specific PCR (qMSP), and results were expressed as ΔCt (difference in cycle threshold between the target gene and an internal reference), where lower ΔCt values indicate higher methylation levels.
PAX1 methylation testing was performed on residual DNA from the cervical brush specimen originally collected for HPV genotyping at the initial screening visit. As described in the Study Design section, all specimens were transported to our central laboratory, stored at −80°C, and tested retrospectively after study enrollment. PAX1 results did not influence clinical management decisions.
Ethics Approval and Consent to Participate
This study was conducted in accordance with the ethical standards of the Declaration of Helsinki and approved by the Institutional Ethics Committee of Changsha Hospital for Maternal & Child Health Care (Approval Number: EC-20230726-02, dated July 26, 2023). The committee granted a waiver of informed consent for this retrospective study, as all data were collected from existing clinical records and residual clinical specimens were used for research purposes without affecting clinical management. All data were fully anonymized prior to analysis, and the use of residual specimens posed no additional risk to participants.
Statistical Analysis
All statistical analyses were performed using Python (pandas, numpy, scipy, scikit-learn, matplotlib, seaborn) and R.
Descriptive Statistics
Continuous variables (e.g., age, PAX1 methylation level) were presented as mean ± standard deviation or median (interquartile range), as appropriate. Categorical variables (e.g., HPV genotype distribution, TCT results, histological grade) were summarized as frequencies and percentages.
Group Comparisons
Continuous variables were compared using the independent-samples t-test or Mann–Whitney U test (depending on normality). Categorical variables were compared using the chi-square test or Fisher’s exact test.
Diagnostic Performance Evaluation
Receiver operating characteristic (ROC) curves were plotted to evaluate the diagnostic accuracy of PAX1 methylation and TCT (using ≥ASC-H as the positivity threshold) for detecting prevalent CIN3+ at the time of colposcopy. The area under the ROC curve (AUC) was calculated. Given the retrospective, cross-sectional nature of this study design, all performance metrics (sensitivity, specificity, PPV, NPV, AUC) estimate immediate disease risk (i.e.,prevalent CIN3+ detection) and do not assess longitudinal progression risk. The optimal cutoff for PAX1 methylation was determined using the Youden index.
Model Comparison
DeLong’s test was used to assess whether the difference in AUC between PAX1 methylation and TCT was statistically significant. We determined separate optimal PAX1 methylation cut-offs for the full cohort and for the non-HPV16/18 subgroup using the Youden index.
Potential Clinical Impact Assessment
We simulated the hypothetical impact of replacing TCT with PAX1 methylation as the triage strategy, estimating the reduction in colposcopy referral rates and the increase in CIN3+ detection rates.
All statistical tests were two-sided, and a P-value < 0.05 was considered statistically significant.
Results
Baseline Characteristics of the Study Population
Among 3,233 enrolled women who underwent comprehensive evaluation—including HPV genotyping, liquid-based cytology (TCT), colposcopy, PAX1 methylation testing, and histopathological confirmation—the distribution of high-risk HPV (hrHPV) genotypes was as follows: HPV52 (20.59%), HPV16 (19.60%), HPV58 (9.80%), HPV53 (7.40%), HPV18 (7.30%), HPV51 (5.50%), HPV59 (3.80%), and HPV not tested (0.60%). The remaining 25.0% harbored other non-16/18 hrHPV types (Figure 1A). Notably, HPV16/18 accounted for 26.9% of all infections, whereas HPV52/58 alone constituted 30.39%, underscoring the substantial contribution of non-16/18 genotypes to the overall hrHPV burden in this Chinese cohort. Baseline characteristics and PAX1 methylation levels across cervical lesion severity in the cohort of 3,233 Chinese women.
Histopathological diagnoses were categorized into five groups: chronic cervicitis (51.70%), CIN1 (31.20%), CIN2 (9.77%), CIN3 (5.35%), and invasive cervical cancer (CA, 1.84%) (Figure 1B). Among women with high-grade lesions (CIN2+, n = 493), the majority of CIN2 (n = 97/176, 55.1%) and CIN3 (n = 207/317, 65.3%) cases were associated with non-16/18 hrHPV types. In contrast, among the 60 cervical cancer cases, 44 (73.3%) were HPV16/18-positive, 12 (20.0%) were positive for other hrHPV types(Figure 1C), and 4 (6.7%) tested HPV “not test”—suggesting possible alternative carcinogenic mechanisms or limitations in current HPV detection assays.
Inverse Association Between PAX1 Methylation Levels and Cervical Lesion Severity
PAX1 methylation was quantified as ΔCt (cycle threshold), where lower ΔCt values correspond to higher methylation levels. A clear inverse correlation was observed between PAX1 ΔCt and lesion severity (Figure 1D):
Chronic cervicitis: median ΔCt = 17.92.
CIN1: median ΔCt = 16.96.
CIN2: median ΔCt = 12.04.
CIN3: median ΔCt = 7.36.
Cervical cancer: median ΔCt = 5.97.
This stepwise decrease in ΔCt (i.e.,progressive hypermethylation) across the histological spectrum (from benign inflammation to invasive carcinoma) provides strong molecular evidence that PAX1 methylation is dynamically upregulated during cervical carcinogenesis and may serve as a sensitive biomarker of disease progression.
Diagnostic Performance of PAX1 Methylation for Detecting CIN3+ Lesions
To assess the clinical utility of PAX1 methylation in identifying CIN3+ (CIN3 and cancer), we performed ROC analysis in the full cohort. The area under the curve (AUC) was 0.855 (95% CI: 0.828–0.882; P < 0.0001) (Figure 2A), indicating excellent discriminatory accuracy. Diagnostic performance of PAX1 methylation for detecting CIN3+ lesions.
Diagnostic Performance of Cytology (TCT) and PAX1 Methylation for CIN3+ Detection

This table compares the sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV) of liquid-based cytology (TCT, using ≥ASC-H as the positivity threshold) and PAX1 methylation (at the optimal cut-off ΔCt = 9.57) in identifying CIN3+ (CIN3 or invasive cancer) among 3,233 hrHPV-positive women. TCT missed 39.6% of CIN3+ cases, while PAX1 methylation demonstrated superior accuracy (AUC 0.855 vs. 0.70 for TCT as an ordinal variable), establishing its value as a more objective and sensitive triage biomarker.
Sensitivity: 71.1% (95% CI: 58.0–82.0%)
Specificity: 85.7% (95% CI: 81.6–89.1%)
Youden Index: 0.568.
These results establish PAX1 methylation as a robust, non-invasive biomarker for triaging women at high risk of CIN3+.
Limitations of Cytology-Based Triage
Among the 171 women diagnosed with CIN3+, cytology (TCT) using a threshold of ≥ASC-H achieved only 60.4% sensitivity (95% CI: 56.5–64.2%) and 84.7% specificity (95% CI: 83.2–86.0%). Critically, 250 CIN3+ cases (39.6%) were missed by cytology (reported as ≤LSIL). When TCT results were modeled as an ordinal variable (NILM=1 to HSIL+=5), the AUC for CIN3+ detection was merely 0.70 (95% CI: 0.67–0.72), further highlighting the suboptimal performance of cytology alone in high-grade lesion detection.
To further characterize the pattern of false-negative cytology results, we performed a detailed subcategory analysis of the 250 CIN3+ cases that were missed by cytology (reported as ≤LSIL). As shown in Supplementary Table 1, among these missed cases, 145 (58.0%) were classified as LSIL, 80 (32.0%) as ASC-US, and only 25 (10.0%) as NILM. This distribution reveals a critical finding: the majority of missed CIN3+ cases were not reported as completely negative but rather as low-grade or equivocal abnormalities. This suggests that the primary limitation of cytology-based triage is not merely false-negative sampling, but also the inadequate positive predictive value of low-grade cytological abnormalities for detecting high-grade lesions. These findings reinforce the need for more objective molecular biomarkers, such as PAX1 methylation, to improve risk stratification in hrHPV-positive women with ≤LSIL cytology.
Superior Clinical Performance of a Genotype-Driven, Methylation-Based Triage Strategy
We compared two triage strategies in the full cohort:
Strategy A (current standard): Referral to colposcopy if hrHPV-positive and TCT ≥ ASC-US.
Strategy B (proposed): unnecessary colposcopy for HPV16/18-positive women; or non-16/18 hrHPV-positive women, referral if PAX1 ΔCt ≤ 8.79 (i.e.,hypermethylated).
Performance of PAX1 Methylation for CIN3+ Detection in the Non-HPV16/18 hrHPV-Positive Subgroup (n = 2,376)

The Optimal Triage Cut-Off (ΔCt ≤ 8.79) was Determined Using the Youden Index Specifically for This Subgroup and Differs From the Cut-Off Derived for the Full Cohort (ΔCt = 9.57). CIN3+ was Defined as Cervical Intraepithelial Neoplasia Grade 3, Adenocarcinoma in situ (AIS), or Invasive Carcinoma. PPV and NPV Were Calculated Based on the Observed CIN3+ Prevalence (3.1%) in This Subgroup.
Comparative Performance of Two Cervical Cancer Screening Triage Strategies
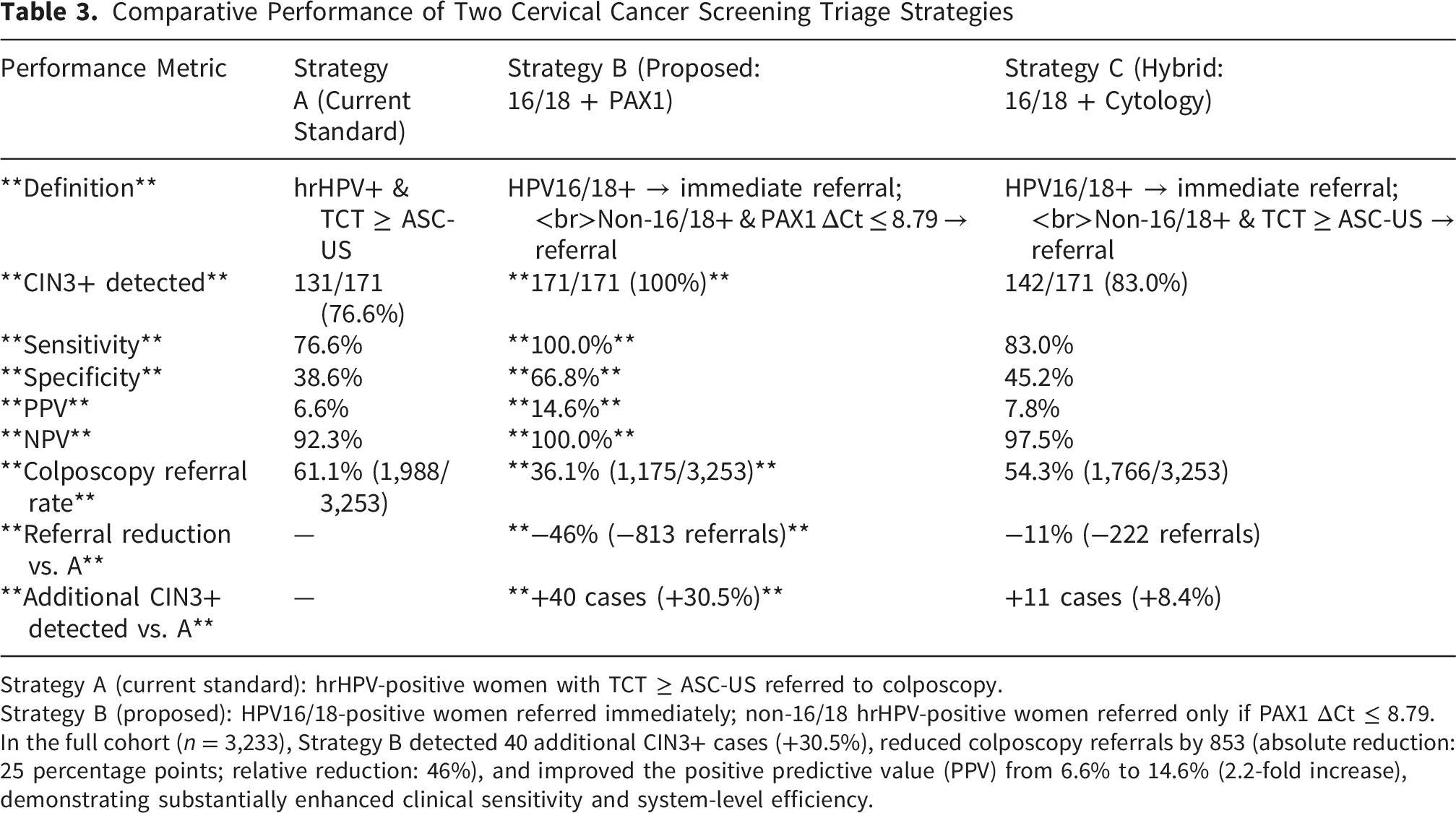
Strategy A (current standard): hrHPV-positive women with TCT ≥ ASC-US referred to colposcopy.
Strategy B (proposed): HPV16/18-positive women referred immediately; non-16/18 hrHPV-positive women referred only if PAX1 ΔCt ≤ 8.79.
In the full cohort (n = 3,233), Strategy B detected 40 additional CIN3+ cases (+30.5%), reduced colposcopy referrals by 853 (absolute reduction: 25 percentage points; relative reduction: 46%), and improved the positive predictive value (PPV) from 6.6% to 14.6% (2.2-fold increase), demonstrating substantially enhanced clinical sensitivity and system-level efficiency.
Increased CIN3+ detection: 40 additional CIN3+ cases identified (+30.5% vs. Strategy A);
Reduced colposcopy burden: 853 fewer referrals (absolute reduction of 25 percentage points; 46% relative decrease);
Higher diagnostic precision: Positive predictive value (PPV) improved from 6.6% to 14.6% (2.2-fold increase), indicating markedly improved resource efficiency.
Conclusion of Results
These findings demonstrate that a triage algorithm integrating HPV16/18 genotyping with PAX1 methylation (cut-off ΔCt ≤ 8.79) for non-16/18 hrHPV carriers simultaneously enhances sensitivity for CIN3+ detection and drastically reduces unnecessary colposcopy referrals. This strategy represents a clinically actionable, cost-effective refinement to current cervical cancer screening paradigms, particularly in populations where non-16/18 hrHPV types predominate.
Discussion
Our study demonstrates that a precision triage strategy integrating HPV16/18 genotyping with PAX1 methylation significantly outperforms conventional HPV+cytology co-testing in a large Chinese screening cohort. The key findings are threefold: (1) PAX1 methylation is a robust, inverse biomarker of CIN3+ risk in non-16/18 hrHPV+ women; (2) the proposed strategy increases CIN3+ detection by over 30%; and (3) it reduces immediate colposcopy referrals by nearly half, with a 2.2-fold improvement in PPV.
Our observed AUC of 0.82 for PAX1 methylation in non-16/18 hrHPV carriers aligns closely with validation studies conducted specifically in Asian cohorts. Huang et al (2010) first demonstrated that quantitative PAX1 methylation analysis achieved 100% sensitivity for invasive cervical cancer detection with 92.3% specificity in a Taiwanese hospital-based cohort. 16 Subsequent multicenter validation by the Taiwanese Gynecologic Oncology Group (TGOG) confirmed robust performance across histological grades, with PAX1 methylation showing 64% sensitivity and 91% specificity for CIN3+ detection. 19 Critically, Chang et al (2021) provided real-world evidence from a Taiwanese screening program demonstrating that PAX1 methylation achieved 91.6% accuracy and 92.5% specificity for CIN3+ detection—superior to HPV16/18 genotyping (76.9% accuracy)—in women aged 30–64 years. 20 However, these landmark studies evaluated PAX1 across all hrHPV types without genotype stratification. Our work is the first to rigorously demonstrate that PAX1 retains robust discriminatory power (AUC=0.82) specifically within the non-16/18 subgroup—a critical distinction given that non-16/18 types constitute >70% of hrHPV+ women in East Asia yet are systematically overlooked in triage algorithms focused exclusively on HPV16/18.
While PAX1 has emerged as a leading methylation biomarker in Asian populations, several alternatives have been evaluated globally. Kelly et al's systematic review and meta-analysis of 43 studies (n=16,336 women) established that human gene methylation markers collectively achieve 68.6% sensitivity and 70% specificity for CIN2+ detection at standardized thresholds. 21 Within this landscape, PAX1 demonstrates competitive performance: a recent Chinese study by Yang et al (2023) directly compared PAX1 methylation versus cytology in 387 non-16/18 hrHPV+ women, reporting 87% sensitivity and 75.5% specificity for CIN2+ detection—substantially outperforming cytology’s 24.8% specificity. 22 Notably, dual-marker panels incorporating PAX1 with complementary genes have shown incremental gains: Liang et al (2024) demonstrated that PAX1/JAM3 methylation (CISCER assay) achieved 92.6% sensitivity and 95.7% specificity for CIN2+ in 396 hrHPV+ Chinese women, with 100% sensitivity for cervical cancer detection—including two hrHPV-negative adenocarcinomas missed by conventional screening. 23 Nevertheless, our data suggest that single-marker PAX1 testing suffices for effective triage in non-16/18 carriers, avoiding the cost and complexity of multi-gene panels while maintaining high clinical utility.
More critically, HPV16/18-only triage strategies—endorsed in Western guidelines—perform suboptimally in Asian populations due to distinct HPV genotype distributions. Wang et al's systematic review of 156,803 Chinese women confirmed that HPV52/58 collectively surpass HPV18 in prevalence and contribute to 32.7% of CIN3+ cases—nearly double their contribution in European cohorts. 11 This epidemiological reality renders 16/18-only triage both insensitive and inequitable in Asia. Our genotype-stratified approach directly addresses this gap: by applying PAX1 triage precisely where it is most needed—in the non-16/18 majority (73.0% of hrHPV+ women)—we simultaneously improved CIN3+ detection (+30.5%) and reduced colposcopy referrals (−46%) compared to cytology-based triage. This dual benefit aligns with WHO’s precision prevention paradigm, which emphasizes maximizing clinical benefit while minimizing harm and resource utilization.
Notably, we observed that non-HPV16/18 genotypes—especially HPV52 and HPV58—accounted for 60% of CIN2+ lesions, underscoring a critical gap in strategies that triage solely on HPV16/18. This epidemiological pattern is consistent with prior reports from East Asia and highlights the need for region-specific algorithms. Importantly, even among cervical cancers, 26.7% were caused by non-16/18 types, and 6.7% were HPV “not test”—suggesting alternative oncogenic pathways and reinforcing the limitations of HPV-only risk stratification.
The inverse relationship between PAX1 ΔCt and lesion severity—where lower ΔCt (higher methylation) correlates with more advanced disease—is biologically plausible. PAX1, a developmental transcription factor, is epigenetically silenced during carcinogenesis, and its methylation serves as a surrogate for clonal expansion of transformed cells. The strong stepwise increase in methylation from chronic cervicitis to invasive cancer provides compelling evidence for its role as a “molecular histology” marker.
Our proposed cut-off (ΔCt ≤ 8.79) in non-16/18 women achieved a favorable balance between sensitivity (73%) and specificity (50.5%), but the true advantage lies in its system-level impact. By reclassifying low-methylation (low-risk) non-16/18 women as non-referral, we spared 853 women from invasive procedures and associated anxiety, without compromising—and indeed enhancing—CIN3+ detection. This aligns with the “precision prevention” paradigm advocated by WHO, which emphasizes maximizing benefit while minimizing harm. While our data support the clinical utility of PAX1 methylation, the optimal cut-off must be validated in independent, population-based screening cohorts before guideline adoption.
The PAX1 triage threshold (ΔCt ≤8.79) was determined post-hoc using the Youden index in this cohort and requires prospective validation before clinical implementation.” Added to Discussion: “While our data support the clinical utility of PAX1 methylation, the optimal cut-off must be validated in independent, population-based screening cohorts before guideline adoption.
Among 1,042 hrHPV+ women with NILM/chronic cervicitis, 28 (2.7%) harbored CIN2+ lesions. Of these, PAX1 methylation (ΔCt ≤8.79) detected 21 cases (75.0%), whereas cytology missed all 28. This highlights PAX1’s value in identifying high-grade lesions masked by benign cytology—a critical gap in current screening.
We observed HPV16/18 dominance in invasive cancer (73.3%) but predominance of HPV52/58 in CIN2/3 (65.3%). This pattern suggests differential oncogenic potential: HPV16/18 may drive more rapid progression from CIN3 to invasion, whereas HPV52/58-associated lesions may persist longer at precancerous stages or regress more frequently. This epidemiological nuance underscores the need for sensitive triage biomarkers like PAX1 methylation to capture high-risk non-16/18 lesions before progression.
Several limitations warrant mention. First, the cohort was enriched for hrHPV+ women referred for colposcopy, which may inflate prevalence estimates; however, this design reflects real-world clinical decision-making. Second, PAX1 testing used ΔCt from a methylation-specific qPCR platform; standardization across platforms will be needed for global implementation. Third, longitudinal data on progression of PAX1-low lesions are lacking, though the high NPV (89.4%) suggests safety.
First, our cohort comprised women already referred for colposcopy per national guidelines, which may overestimate CIN3+ prevalence compared to population-based screening. However, this design reflects actual clinical decision-making pathways and enables direct assessment of triage strategies within the high-risk population where resource optimization is most needed.
Third, as a retrospective cross-sectional study, our design only permits estimation of immediate (prevalent) CIN3+ risk at screening; we cannot assess whether PAX1-low lesions will progress, persist, or regress over time. However, the high NPV (89.4%) and biological plausibility of methylation as a marker of neoplastic transformation support its utility for prevalent disease triage. Prospective longitudinal studies are warranted to evaluate PAX1’s performance in predicting future progression.
While our retrospective data support PAX1’s utility, prospective validation in population-based screening cohorts remains essential before guideline adoption—consistent with WHO’s phased biomarker evaluation framework. 24 Encouragingly, emerging evidence supports PAX1’s safety profile: Chen et al (2025) recently demonstrated that among 989 HPV+ Chinese women, PAX1 methylation achieved 93.49% sensitivity and 93.24% specificity for CIN2+ detection, with negative predictive value exceeding 95%—suggesting that PAX1-negative women may safely defer colposcopy. 25 Furthermore, Su et al (2025) provided compelling evidence that PAX1/JAM3 methylation outperforms both cytology and HPV33/35 genotyping for risk stratification in non-16/18 hrHPV+ women, reducing colposcopy referrals by 61.9 percentage points while maintaining 39.1% CIN3+ risk in methylation-positive women. 26 Future implementation research should address three priorities: (1) standardization of ΔCt cut-offs across qMSP platforms; (2) cost-effectiveness analyses specific to resource-constrained Asian settings; and (3) integration with self-sampling approaches to improve screening coverage among underserved populations.
Conclusion
Our data support the adoption of a dual-path triage algorithm: HPV16/18 → immediate colposcopy; non-16/18 hrHPV → PAX1 methylation-based referral. This approach is not only more accurate and efficient but also more equitable for populations where non-16/18 types dominate. We recommend prospective validation in population-based screening programs and incorporation into national guidelines to accelerate cervical cancer elimination.
Supplemental Material
Supplemental material - HPV16/18 Genotyping Combined with PAX1 Methylation Triage Reduces immediate Colposcopy While Improving CIN3+ Detection: A Real-World Study of 3,233 Chinese Women
Supplemental material for HPV16/18 Genotyping Combined with PAX1 Methylation Triage Reduces immediate Colposcopy While Improving CIN3+ Detection: A Real-World Study of 3,233 Chinese Women by Xing Fan, Haichun Guo, Sifeng Wang and Xiangwen Peng in Cancer Control
Footnotes
Acknowledgements
We would like to express our sincere gratitude to all the participants who generously contributed their time and samples, making this study possible. We also thank the clinical staff at Changsha Hospital for Maternal & Child Health Care for their invaluable assistance with patient recruitment and sample collection. The authors acknowledge the use of Qwen (by Tongyi Lab) for language polishing to enhance the clarity and readability of the manuscript.
Ethical Considerations
This study was conducted in accordance with the ethical standards of the Declaration of Helsinki and approved by the Institutional Ethics Committee of Changsha Hospital for Maternal & Child Health Care (Approval Number: EC-20230726-02, dated July 26, 2023). The committee granted a waiver of informed consent for this retrospective study, as all data were collected from existing clinical records and residual clinical specimens were used for research purposes without affecting clinical management. All data were fully anonymized prior to analysis, and the use of residual specimens posed no additional risk to participants.
Consent to Participate
The committee granted a waiver of informed consent for this retrospective study, as all data were collected from existing clinical records and residual clinical specimens were used for research purposes without affecting clinical management. All data were fully anonymized prior to analysis, and the use of residual specimens posed no additional risk to participants.
Author Contributions
Xing Fan and Haichun Guo contributed equally to this work. Xiangwen Peng conceived and designed the study, acquired funding, and supervised the project. Xing Fan, Haichun Guo, and Sifeng Wang were involved in data curation, investigation, and formal analysis (including HPV genotyping, PAX1 methylation testing, and statistical analysis). Xing Fan and Haichun Guo wrote the original draft of the manuscript. Xiangwen Peng reviewed, edited, and finalized the manuscript. All authors have read and approved the final manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research supported by Clinical Medical Technology Demonstration Base for Genetic Research of Fetal Congenital Heart Disease in Hunan Province (2021SK4036), Hunan Province Children’s Safe Medication Clinical Medical Technology Demonstration Base(2023SK4083), Key research project of Hunan Provincial Department of Education(21A0043, Project of Changsha Science and Technology Bureau, ( KH2201045), Natural Science Foundation of Hunan Province (2023JJ30063); Changsha Science and Technology Bureau natural science surface project(kq2202030) and National Natural Science Foundation of China(32070817). Hunan Provincial Medical and Health Joint Fund (2026), granted to Xing Fan.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The data that support the findings of this study are not publicly available due to privacy and ethical restrictions. The dataset contains sensitive patient information collected as part of clinical care and was fully anonymized for research use under approval from the Institutional Ethics Committee of Changsha Hospital for Maternal & Child Health Care (Approval No. EC-20230726-02). Reasonable requests for access to de-identified data may be considered by the corresponding author upon formal request and in accordance with institutional and ethical guidelines.
Supplemental Material
Supplemental material for this article is available online.
