Abstract
Introduction
Few machine learning (ML) studies have investigated the prediction of distant metastasis in patients with renal cell carcinoma (RCC). This retrospective study aimed to develop and validate predictive models based on ML algorithms for RCC patients with distant metastasis.
Methods
We extracted RCC data from the SEER database between 2004 and 2015 (n=192,912) and from the Chinese National Cancer Center (CNCC) database between 2010 and 2020 (n=3,034). Seven different algorithms were applied to predict distant metastasis in RCC. Fivefold cross-validation was employed for model construction. The data were analyzed by using Python based on incomplete data, complete data, upsampling data and downsampling data.
Results
After data cleaning and screening, 121,741 cases from the SEER dataset and 2803 cases from the CNCC external test set were retained. For incomplete data, the neural network model [area under the curve (AUC) 95% confidence interval (CI) of the external data: 0.7467±0.0573] achieved the highest accuracy. For the complete data, the support vector machine (SVM) model achieved the highest accuracy, with an AUC 95% CI of 0.8221±0.0485. The disparity between positive and negative samples significantly varied across the different datasets. Upsampling and downsampling analyses were also conducted. For the upsampling data, the extreme gradient boosting (XGBoost) model demonstrated the highest accuracy, with an AUC 95% CI for the external data of 0.8162±0.0558. For the downsampling data, the SVM model achieved the highest accuracy, with an AUC 95% CI of 0.8274±0.0546 for the external data.
Conclusions
Our study revealed that ML algorithms can effectively predict distant metastasis in patients with RCC. ML models exhibit favorable application prospects in clinical practice.
Introduction
Renal cell carcinoma (RCC) is among the most common malignant tumors affecting the urinary system. The incidence of renal cancer in China has demonstrated a continuous upward trend in recent years, with approximately 73,700 new cases being reported in 2022 and a standardized incidence rate of approximately 6.7 per 100,000 being observed. 1 RCC primarily comprises three histopathological types: clear cell RCC, papillary RCC, and chromophobe RCC, with clear cell RCC accounting for 70–80% of cases, papillary RCC accounting for 10–15%, and chromophobe RCC accounting for 5%. 2 The survival of RCC patients has improved due to surgical management and the use of targeted and immune drugs.3,4 However, a substantial proportion of RCC patients present with high-stage disease, with a poor prognosis being determined because of distant metastases.5-7 Thus, tumor metastasis is a critical factor in determining the prognosis of RCC patients.
Seventy percent of patients are diagnosed with stage I RCC, and 11% of patients are diagnosed with stage IV RCC. 8 The lungs are the most commonly affected site of metastasis, followed by regional lymph nodes, the brain, bone, and soft tissue. Lee CH et al 9 analyzed the Korean Renal Cancer Study Group metastatic RCC (mRCC) database. The results revealed the most common sites of metastasis (>5%), and the median cancer-specific survival (CSS) ranged from 13.9 (liver) to 29.1 months (lungs). Through continuous efforts in RCC research, for advanced RCC or mRCC, combinations of immune checkpoint inhibitors or combinations of immune checkpoint inhibitors with tyrosine kinase inhibitors are associated with a tumor response of 42% to 71%, with a median overall survival of 46 to 56 months being observed. 8 These results indicate that the prognosis of mRCC patients is still poor, regardless of the presence of lymph node metastasis or other organ metastasis.9,10 The early detection of distant metastasis is the top priority of clinical work because the appropriate treatment options (such as surgery, immunotherapy, and targeted therapy) can be chosen correctly.
The identification of the clinicopathological risk factors that promote distant RCC metastasis is vital. In recent years, several clinical diagnostic tools have been built using different prognostic and prediction models. Although predictive factors and models for mRCC have been developed, the limited data commonly restrict individualized accurate prediction. These studies exhibit significant limitations, such as the use of older tools and limited data. The Surveillance, Epidemiology, and End Results (SEER) database represents 28% of the current U.S. population; moreover, it includes large, multi-institutional patients and could be an essential source of mRCC research to provide greater statistical power. Additionally, machine learning (ML), which is a major subbranch of artificial intelligence, is a promising statistical method that can handle large amounts of heterogeneous data. ML techniques have not been used in many studies, and some models lack interpretability to fully capture the complex relationships among variables and to provide clinically actionable explanations. Previously, we used ML algorithms as auxiliary tools to predict overall survival in RCC patients. 11 This technique could be used to quantify the possibility of recurrence in patients and to help with more individualized postoperative clinical management.
In our study, we aimed to develop and validate an explainable ML model for the prediction of RCC metastasis. We collected a large amount of data from the SEER database and the Chinese National Cancer Center (CNCC) dataset. The RCC cases were subjected to data standardization and included multiple clinical parameters, such as tumor grade, side, pathological type, T stage, and N stage. The RCC M0/M1 metastasis prediction algorithm was explored and visualized.
Materials and Methods
Data Collection
The data used in this study were consecutively collected from the SEER database by using SEER*Stat software (version 8.3.6) and the CNCC. A data agreement form was signed and submitted to the SEER administration. This trial was a retrospective study and was approved by the Institutional Review Committee of the CNCC (Institutional Review Board number: 21/405-3076; date: October 13, 2021). The requirement for obtaining informed consent was also waived by the Institutional Review Committee. The study was conducted in accordance with the Helsinki Declaration of 1975, as revised in 2024. All of the patient details have been de-identified. The reporting of this study conforms to the TRIPOD + AI statement. 12
The variables included race, sex, age, marital status, tumor location, tumor size, histological type, tumor grade, tumor-node-metastasis (TNM) stage information, and surgical treatment. The SEER dataset was utilized as the internal dataset for model construction and performance evaluation, whereas the dataset from the CNCC served as an external independent dataset to test the generalizability of the models.
Data Preparation
We extracted RCC data, including data from the SEER database between 2004 and 2015 (n=192,912) and from the CNCC database between 2010 and 2020 (n=3,034). After data cleaning and screening, 121,741 cases remained in the SEER dataset, and 2803 cases remained in the CNCC external test set. The flowchart is shown in Figure 1. Given the substantial volume of the SEER data and the presence of incomplete data for some features, the dataset was initially divided into training and testing sets at a 9:1 ratio, thereby resulting in approximately 109,567 cases for the training set and 12,174 cases for the testing set. From all of the cases with complete data, 12,174 cases were randomly selected as the testing set, with the remaining 109,567 cases being allocated to the training set. Among the 109,567 cases, 80,119 cases had complete data, and 29,448 cases had incomplete data. Flowchart for SEER and CNCC data screening
Data Preprocessing
To facilitate extraction and analysis, numerical characterization of the clinicopathological data of the RCC patients was initially conducted through different numbers of assignments.
Processing of Incomplete Data
The random forest interpolation method was used to supplement missing values. First, a random forest regression model was trained based on available complete data. The average values were subsequently used to supplement all of the missing features. Finally, the random forest regression model was used to regress and predict each missing feature one by one, along with replacing the missing values with the original average values.
Feature Discretization
To improve the generalizability of the model, continuous features such as age and tumor size were discretized and binarized. This study employed a feature discretization method grounded in information entropy, which is a supervised approach. Information entropy serves as a metric for quantifying the uncertainty of an event, and higher entropy values indicate greater uncertainty. The following formula was utilized for calculating information entropy:
80,119 Cases With Complete Training Set

109,567 Cases With Incomplete Dada in Training Set

Stability Assessment
The “leave-top1-out” approach was used to evaluate feature ranking stability: (i) selected the top-n features from the full dataset (Set1); (ii) removed the highest-ranked feature and retrained the model on the reduced dataset; (iii) extracted the new top-(n−1) features (Set2); (iv) systematically compared the ranking orders between Set 1 and Set 2 using Spearman’s rank correlation coefficient and the Jaccard similarity index.
Statistical Analysis
We trained all of the ML models by using Python version 3.7.1, NumPy version 1.20.1, Scikit-Learn version 1.0.2, SciPy version 1.7.3, and XGBoost version 1.6.2. The ML models were developed and evaluated by using 5-fold cross-validation. The receiver operating characteristic (ROC) curve, decision curve analysis (DCA), calibration curve, Delong test and SHapley Additive exPlanations (SHAP) images were used to evaluate the accuracy of the model. Both F1(1) and F1(0) scores were calculated to provide a comprehensive performance evaluation.
Results
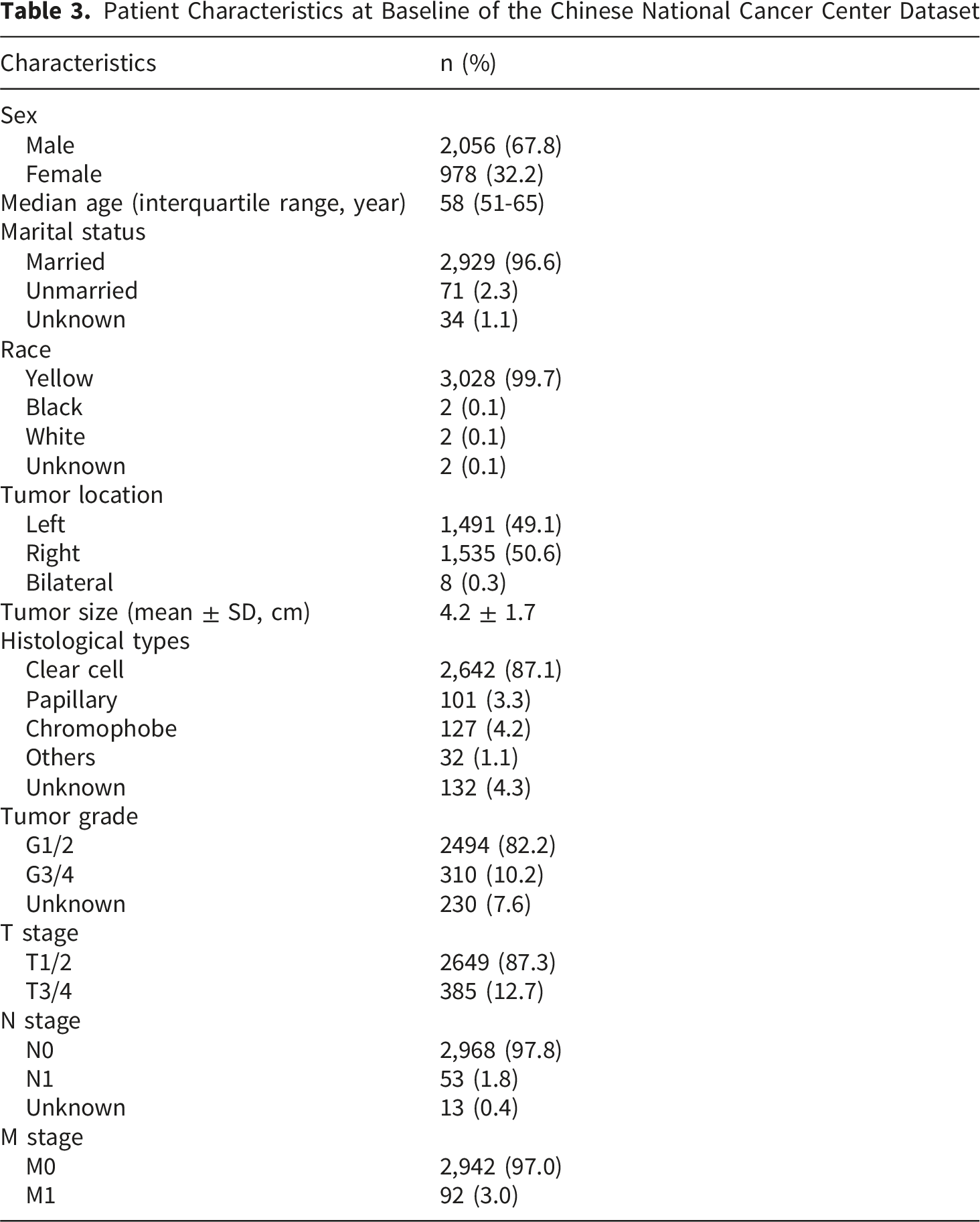
Clinical Characteristics of the Patients
Patient Characteristics at Baseline of the Chinese National Cancer Center Dataset

Model Construction
According to the aforementioned data partitioning table, it was necessary to investigate and compare the performance of the models that were constructed with either complete data or incomplete data. Additionally, as shown in Tables 1 and 2, there was a significant disparity observed between the numbers of positive and negative samples within each dataset. Consequently, the performance of the models was compared by using upsampling and downsampling methods. The research framework was divided into the following two sections.
Incomplete Data (n=109,567) vs. Complete Data (n=80,119)
The Integrated Model Prediction Results After Five-Fold Cross-Validation in the Incomplete Data
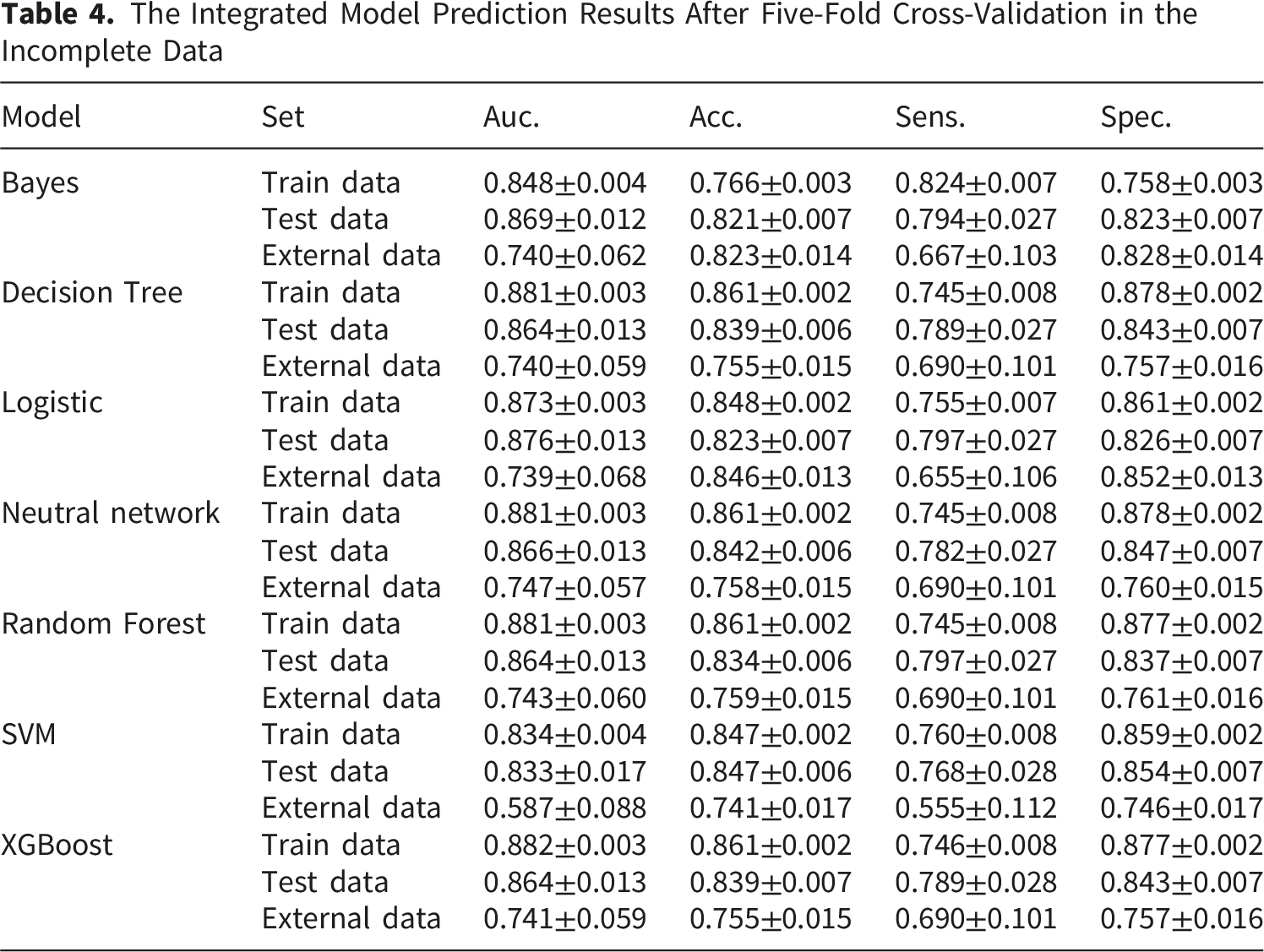

ROC curve analysis of different models based on the incomplete dataset (A) and complete dataset (B) for predicting metastasis in patients with RCC
The Integrated Model Prediction Results After Five-Fold Cross-Validation in the Complete Data

The model that was constructed with the complete training set outperformed the model that was constructed with the incomplete training set. Furthermore, the average prediction results of these two models based on the external test set were analyzed via the Delong test. The results revealed a p value of less than 0.0001, which demonstrated a significant difference in model performance.
Upsampling vs. Downsampling Analysis
Based on the analysis of positive and negative samples in each dataset, the disparity between the positive and negative samples significantly varied across the different datasets. Therefore, we investigated and compared the performances of the models that were constructed by using methods without sampling (the abovementioned results), along with methods involving upsampling and downsampling. We employed the fivefold cross-validation method to construct the models. First, downsampling was performed on the negative samples in the training set, and the model predictions after integrating the fivefold models were obtained, with the results being shown in Supplementary Table 1 and Figure 3A. The SVM model achieved the highest accuracy, with an AUC 95% CI of 0.8685±0.0125 for the test data and 0.8274±0.0546 for the external data being reported. Afterward, upsampling was performed on the positive samples in the training set, and the model predictions after integrating the fivefold models were obtained, with the results being shown in Supplementary Table 2 and Figure 3B. The XGBoost model demonstrated the highest accuracy, with an AUC 95% CI of 0.8819±0.0117 for the test data and 0.8162±0.0558 for the external data being reported. Both downsampling and upsampling led to relatively high AUC values and accuracy compared with those of the abovementioned nonsampling models. ROC curves based on the downsampling (A) and upsampling (B) analyses
To compare the performances of the three methods for constructing the models, a Delong test was also conducted on the average prediction results of these three models based on an external test set. The p value of the nonsampling model and the upsampling or downsampling model based on the external test set was less than 0.0001, thus indicating that the latter model performed better than the former model. The upsampling or downsampling method can significantly improve model performance. There was no significant difference observed in performance between the two models that were constructed by using the upsampling or downsampling methods (p=0.8734).
Based on the obtained results, we identified that the model constructed by using the upsampling method was the optimal model for this study. Furthermore, a visual analysis of the results is presented in Figure 4 (DCA curve) and Figure 5 (calibration curve). We also visualized each algorithm with a SHAP chart, as shown in Supplementary Figure 1. SHAP summary plots for both the test and external datasets—generated using the full feature set (10 features)—were presented in Supplementary Figure 2. Corresponding plots based on the reduced datasets (after removal of the top-ranked feature) were also provided for comparison (Supplementary Figure 3). Focusing on the full set of 10 features, we systematically evaluated the stability of feature rankings across top-n subsets (with n ranging from 3 to 8). As summarized in the Supplementary Table 3, the results showed that while a subset of top-ranked features remains relatively consistent, the overall ranking order is notably sensitive to the removal of the highest-ranked feature. This instability reflects the label-driven optimization dynamics inherent in supervised learning models and highlights the need for caution when interpreting feature importance rankings as robust biological signals. DCA curves based on the upsampling training set (A), test set (B), and external test set (C) Calibration curves based on the upsampling training set (A), test set (B), and external test set (C)

Based on the abovementioned feature visualization SHAP chart, it can be concluded that tumor size, T stage, N stage, and tumor grade demonstrate high feature importance in most models. Therefore, these four clinical characteristics can be considered as the main predictors of the metastasis parameters of RCC patients. Additionally, due to inherent imbalance of data, the model achieved low recall for the minority class, resulting in a substantially lower F1(1) compared to F1(0), as shown in Supplementary Table 4.
Discussion
With respect to model development, although nomograms are currently the most commonly used prediction models, ML models are favored by an increasing number of medical workers because of their practicality, innovation, and accuracy.12-14 In clinical practice, tumor-related models can accurately predict prognosis by combining multiple factors, such as tumor pathological subtype, tumor stage, tumor diameter, and molecular marker expression. However, few researchers have attempted to use ML methods to explore the prediction of metastasis in RCC patients. This study mainly tried to apply ML method to model construction and prediction. Firstly, it is to test its ability of dealing with large amount of data. Secondly, it is to test its ability of integrating data. Thirdly, the weights of each feature of RCC can be roughly described. The results indicated good sensitivity and specificity. These algorithms can be applied to accurately predict whether RCC patients have metastasis, thereby providing assistance for the determination of clinical metastasis.
Various statistical measures can facilitate the understanding and interpretation of data. However, the limitations and efficiency of the processing of big data limit computing power and accuracy. In recent years, ML has included algorithmic methods that enable machines to solve problems without the use of specific computer programming, thereby providing an avenue for predictive modeling tasks.15-17 The integration of big data with ML algorithms is becoming a clinical necessity. In RCC studies, ML models can be applied to analyze the risk factors associated with specific diseases based on patient information. Yin et al 18 integrated convolutional neural network models with Cox regression to identify potential prognostic biomarkers for overall survival. Terrematte P et al 19 created a novel ML 13-gene signature, which was able to improve risk analysis and survival prediction for RCC patients. Chen S et al 20 developed and validated an ML-based prognosis prediction model, which could contribute to clinical decision-making for patients with RCC. Similarly, in our previous study, we investigated and demonstrated that ML algorithms could be used as auxiliary tools to predict the overall survival of patients with RCC using SEER data. 11
Once distant metastasis occurs in RCC, patient prognosis becomes very poor. Therefore, the prediction of distant metastasis in RCC patients is extremely important. Many clinical studies have used clinicopathological factors to establish models to predict metastatic risk in patients with RCC. Fan Z et al 21 established nomograms using the SEER database to predict the risk of bone metastasis in patients with RCC. The calibration curve, ROC curve, and DCA confirmed good performance via diagnostic and prognostic nomograms. Wang J et al 22 developed and validated a nomogram to predict distant metastasis in elderly patients with RCC. The AUC values of the training and validation cohorts indicated excellent predictive ability. DCA indicated that the clinical application value of the nomogram was better than that of traditional TN staging. Some scholars have also explored the use of ML algorithm models to make predictions. Xu C et al 23 used data from 40,355 RCC patients in the SEER database to construct an ML model for predicting the risk of bone metastasis in RCC patients. Among the prediction models established by the six ML algorithms, the XGBoost model achieved the best prediction performance (AUC = 0.891). Dong J and colleagues also used the SEER database to predict distant metastasis in RCC patients based on interpretable ML models. 24 The calibration curve indicated that the predicted values were highly consistent with the actual observed values.
The abovementioned studies indicate that ML models for predicting RCC metastasis are feasible and highly accurate. However, most of these studies were based on SEER data and lacked external validation. In contrast, in this study, we collected RCC data from the SEER database (training set) and the CNCC database (external test set). First, all of the data were preprocessed. The preprocessing of the feature data consisted of two steps. In step 1, clinicopathological data, including sex, tumor grade, side, pathological type, N stage, T stage, surgery and marital status, were subjected to numerical characterization through different number assignments. In step 2, for the two features of age and tumor size, the features in the training set and in the internal and external test sets were separately discretized. Second, we compared the performances of the models that were constructed via different ML algorithms for incomplete data and complete data. The accuracy of the ML models with complete datasets was observed to be significantly greater than that with incomplete datasets. Finally, to overcome the problem of large differences between positive and negative samples in the dataset, we also used upsampling and downsampling methods to reconstruct the model and test its performance. The results revealed that the accuracy of the model was further improved. Our study is the first to combine the SEER database with an external dataset to predict the distant metastasis of RCC. The results demonstrated that after validation with the external dataset, our ML model achieved high accuracy, thus providing considerable guiding value for clinical decision-making. If future applications require reliable individual-level probability estimates, post hoc calibration methods (such as Platt scaling or isotonic regression) should be incorporated. However, the “important features” identified here in this study should be interpreted strictly as contributors to predictive performance under the current modeling framework, not as evidence of biological causality.
This study has several limitations. First, the data obtained from the external validation cohort and the SEER database were retrospectively collected, which may introduce some inherent bias. In addition, to validate our prediction model in the general population, prospective clinical studies with larger sample sizes are necessary. The limited number of external patients may also have affected the statistical significance of the results. The underlying data distribution remains highly skewed (approximately 1:10∼1:30 for positive: negative samples). Despite our best efforts, this extreme imbalance inherently limits the model’s ability to achieve high recall for the minority class, resulting in a substantially lower F1(1) compared to F1(0). This work represents an initial exploratory attempt, and future studies should incorporate more advanced imbalance-handling techniques or larger cohorts. Second, we were unable to obtain biomarkers, blood test results or time to the development of metastasis from the SEER database. In addition, feature importance reflects the predictive contribution within the model and should not be directly equated with the true clinical weight of risk factors. We expect that the addition of data from external data validation will result in a more sophisticated and effective adoption of ML models as supplementary tools for prediction research.
Conclusion
In summary, using data from the SEER database and CNCC dataset, this study explored the factors related to the prediction of RCC metastasis through multi-algorithm ML models. Relevant algorithms were established to predict the possibility of distant metastasis in RCC patients. The findings demonstrate that ML algorithms can effectively predict distant metastasis in patients with RCC and play a positive role in clinical applications.
Supplemental Material
Supplemental Material - Prediction of Distant Metastasis in Renal Cell Carcinoma Using Machine Learning Algorithms: A Multicenter Cohort Study
Supplemental Material for Prediction of Distant Metastasis in Renal Cell Carcinoma Using Machine Learning Algorithms: A Multicenter Cohort Study by Yajian Li, Xinwei Wang, Moxuan Wang, Jianzhong Shou, Cancan Chen and Li Wen in Cancer Control.
Supplemental Material
Supplemental Material - Prediction of Distant Metastasis in Renal Cell Carcinoma Using Machine Learning Algorithms: A Multicenter Cohort Study
Supplemental Material for Prediction of Distant Metastasis in Renal Cell Carcinoma Using Machine Learning Algorithms: A Multicenter Cohort Study by Yajian Li, Xinwei Wang, Moxuan Wang, Jianzhong Shou, Cancan Chen and Li Wen in Cancer Control.
Supplemental Material
Supplemental Material - Prediction of Distant Metastasis in Renal Cell Carcinoma Using Machine Learning Algorithms: A Multicenter Cohort Study
Supplemental Material for Prediction of Distant Metastasis in Renal Cell Carcinoma Using Machine Learning Algorithms: A Multicenter Cohort Study by Yajian Li, Xinwei Wang, Moxuan Wang, Jianzhong Shou, Cancan Chen and Li Wen in Cancer Control.
Supplemental Material
Supplemental Material - Prediction of Distant Metastasis in Renal Cell Carcinoma Using Machine Learning Algorithms: A Multicenter Cohort Study
Supplemental Material for Prediction of Distant Metastasis in Renal Cell Carcinoma Using Machine Learning Algorithms: A Multicenter Cohort Study by Yajian Li, Xinwei Wang, Moxuan Wang, Jianzhong Shou, Cancan Chen and Li Wen in Cancer Control.
Supplemental Material
Supplemental Material - Prediction of Distant Metastasis in Renal Cell Carcinoma Using Machine Learning Algorithms: A Multicenter Cohort Study
Supplemental Material for Prediction of Distant Metastasis in Renal Cell Carcinoma Using Machine Learning Algorithms: A Multicenter Cohort Study by Yajian Li, Xinwei Wang, Moxuan Wang, Jianzhong Shou, Cancan Chen and Li Wen in Cancer Control.
Supplemental Material
Supplemental Material - Prediction of Distant Metastasis in Renal Cell Carcinoma Using Machine Learning Algorithms: A Multicenter Cohort Study
Supplemental Material for Prediction of Distant Metastasis in Renal Cell Carcinoma Using Machine Learning Algorithms: A Multicenter Cohort Study by Yajian Li, Xinwei Wang, Moxuan Wang, Jianzhong Shou, Cancan Chen and Li Wen in Cancer Control.
Supplemental Material
Supplemental Material - Prediction of Distant Metastasis in Renal Cell Carcinoma Using Machine Learning Algorithms: A Multicenter Cohort Study
Supplemental Material for Prediction of Distant Metastasis in Renal Cell Carcinoma Using Machine Learning Algorithms: A Multicenter Cohort Study by Yajian Li, Xinwei Wang, Moxuan Wang, Jianzhong Shou, Cancan Chen and Li Wen in Cancer Control.
Footnotes
Acknowledgements
We are especially grateful to Moxuan Wang, who is a 14-year-old girl. She helped us with the layout and beautification of the figures. To encourage, she was listed one of authors.
Ethical Considerations
The authors state that they have followed the principles outlined in the Declaration of Helsinki for all human or animal experimental investigations. Our study was approved by Institutional Review committee of the National Cancer Center/Cancer Hospital, Chinese Academy of Medical Sciences (NCC/CHCAMS) (Institutional Review Board number: 21/405-3076, date: October 13, 2021).
Consent for Publication
Patient study consent was not required due to the study’s retrospective nature. The requirement for obtaining informed consent was waived by the Institutional Review committee.
Author Contributions
All authors listed in this manuscript contributed significantly to the study. Yajian Li and Xinwei Wang contributed to writing the manuscript. Moxuan Wang contributed to layout and beautification of the figures. Jianzhong Shou contributed to supervision. Li Wen and Cancan Chen contributed to reviewing the manuscript for critical revisions. All authors read and approved the final manuscript.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The datasets used and/or analyzed data in the current study are available from the corresponding author on reasonable request.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.

