Abstract
Objective
This study aimed to explore the relationship between CD276 and clear cell renal carcinoma (ccRCC) and assess the diagnostic value of CD276 in ccRCC.
Methods
Expression levels of CD276 in ccRCC and para-cancer tissues were compared and analyzed retrospectively using data obtained from TCGA and GEO databases. The clinical data was analyzed prospectively. Immunohistochemistry and RT-PCR analyses were used to analyze the expression of CD276 at the mRNA and protein levels. These analyses compared the expression between ccRCC tissues and para-cancer tissues obtained from 70 patients with ccRCC. Next, ELISA was used to analyze peripheral blood samples from 70 patients with ccRCC and 72 healthy individuals, facilitating the differentiation of ccRCC patients from normal controls. Finally, we utilized the Kaplan-Meier method to generate ROC curves for assessing the diagnostic value of CD276 for ccRCC.
Results
Analysis of TCGA and GEO data revealed that the mRNA expression of CD276 was higher in ccRCC tissues than in para-cancer tissues (
Conclusion
The expression of CD276 was higher in ccRCC tissues and positively associated with the pathological grade. Therefore, CD276 may serve as a molecular biomarker for ccRCC prediction.
Introduction
Renal cell carcinoma (RCC) originates from a malignant epithelial tumor of the renal tubules, accounting for 2-3% of all adult malignancies, and has the highest mortality rate among urological cancers. 1 Clear cell renal cell carcinoma (ccRCC) is the most prevalent form of renal cell carcinoma, accounting for approximately 75% of cases. 2 Statistics indicate an annual increase of nearly 2% in the incidence of kidney cancer.1,2 Currently, surgery remains the primary clinical treatment for kidney cancer. 3 However, studies have shown that approximately 25% of patients experience distant metastases postoperatively, 4 posing a significant threat to their lives. For such patients, adjunct targeted therapy or immunotherapy becomes necessary in addition to surgical resection of metastatic lesions. 5 Nevertheless, current molecular targeted therapy or immunotherapy offers limited improvement in progression-free survival (PFS) for patients with advanced kidney cancer, typically maintaining PFS at approximately 12 months or even less.6–8 Therefore, an urgent need exists to identify new molecular targets that can offer fresh perspectives on the diagnosis and treatment of kidney cancer.
The B7-Homologue 3 molecule (B7-H3, also known as cluster of differentiation 276 [CD276]) is a relatively new member of the immune co-stimulatory B7/CD28 family, discovered by Andrei et al in 2001. 9 B7-H3 shares 20-27% amino acid homology with other members of the B7 family and exists in two isoforms: 2IgB7-H3 and 4IgB7-H3. 10 Recent studies have implicated CD276 in the occurrence and development of a variety of cancers, including colorectal cancer, lung cancer, and prostate cancer.11–13 However, few reports on CD276 and RCC have been published.14–17 There exist several histological subtypes of RCC, such as ccRCC (constituting 75% of cases), papillary RCC (accounting for 10%), chromophobe RCC (comprising 5%), and other unclassified types, with ccRCC being the predominant variant.2,15 Roth TJ et al 18 have indicated that B7-H1 and B7-H4, members of the B7-family of T-cell costimulatory molecules, are associated with unfavorable pathologic characteristics and foreshadow poor outcomes for patients suffering from ccRCC. Nevertheless, the implications of CD276 in relation to ccRCC remain ambiguous. This study aimed to explore the relationship between CD276 and ccRCC and to investigate the diagnostic role of CD276 in ccRCC.
Materials and methods
The reporting of this study conforms to STARD guidelines. 19
TCGA Data Description
Gene expression data of 613 patients were obtained from The Cancer Genome Atlas (TCGA) -Kidney Renal Clear Cell Carcinoma (KIRC) project from the TCGA database (https://portal.gdc.cancer.gov) and analyzed retrospectively. A total of 541 tumor specimens and 72 para-cancer tissues were included in Tumor Group and Normal Group, respectively. Data were processed by spliced transcripts alignment to a reference (STAR) pipeline and extracted in Transcripts Per Million (TPM) format. This dataset presented messenger ribonucleic acid (mRNA)-level transcription estimates as log2(x+1) transformed RNA-Seq by expectation-maximization normalized counts. For the statistical analyses, false discovery rate (FDR) corrected q-values adjusted to .05 were employed. The Wilcoxon rank-sum test was then applied to compare the differences of CD276 expression between Tumor Group and Normal Group. A receiver operating characteristic (ROC) curve was generated using Kaplan-Meier analysis to assess the diagnostic value of CD276 in ccRCC.
GEO Data Description
The dataset GSE40435 was selected and obtained from the Gene Expression Omnibus (GEO) database (https://www.ncbi.nlm.nih.gov/geo/). It was analyzed retrospectively. The inclusion and exclusion criteria were as follows: (1) only human tissue studied were included; (2) only ccRCC studies were included, while studies about kidney renal papillary cell carcinoma and kidney chromophobe were excluded; (3) studies without a control group were excluded; (4) only mRNA-level arrays studied were included and dual-channel microarray studies were excluded; (5) studies with < 50 cases were excluded. Differentially expressed genes (DEGs), identified by comparing expressions in ccRCC and normal samples, were analyzed using R language software. Samples with both corrected
Clinical Data Description
A total of 70 cases of ccRCC and 72 cases of healthy individuals who vistied the First Affiliated Hospital of Soochow University between 1st Jan 2023 and 30th September 2023 were enrolled in this study and analyzed prospectively. The inclusion and exclusion criteria for Normal Group were as follows: (1) those with other malignant tumors or infectious diseases, such as acquired immunodeficiency syndrome (AIDS) were excluded; (2) those with recent diagnosis of acute or chronic inflammatory infections as well as those with immune system diseases, such as systemic lupus erythematosus and hematologic diseases were excluded; (3) those underwent nephron-sparing surgeries were excluded; and (4) those who agreed to participate in the study were included. The inclusion and exclusion criteria for Tumor Group were the same as those of Normal Group except that (1) patients with pathologically confirmed ccRCC using the World Health Organization (WHO) grading system were included; (2) patients with a history of radiotherapy or chemotherapy before surgery were excluded; and (3) patients with complete clinical information and tumor and adjacent cancer samples obtained were included. Peripheral blood samples taken before surgery from all 142 cases and 70 pairs of ccRCC and para-cancer tissues were further analyzed by immunohistochemistry (IHC), real-time polymerase chain reaction (RT-PCR), and enzyme-linked immunosorbent assay (ELISA). A ROC curve was generated to demonstrate the ability of CD276 to distinguish patients with ccRCC from healthy individuals. This study was aprroved by the Ethics Committee of the First Affiliated Hospital of Soochow University (Approval No. 257, 2023). Written informed consents were signed and obtained from all participants prior to their inclusion in the study. And we have de-identified all patient details.
Immunohistochemistry
Tumor tissue specimens were obtained from 70 patients with ccRCC and corresponding adjacent normal tissues (at least two centimeters away from the edge of the tumor and no invasion of tumor cells confirmed on post-surgical pathology). The specimens were preserved by fixation in 10% formaldehyde (formalin) solution, embedded in paraffin wax, and sectioned to ensure proper archiving for future reference. Paraffin-embedded tissue slices underwent a series of preparatory treatments to facilitate IHC. The process began with sequential dewaxing, rehydration with graded alcohols, and thorough washing with phosphate buffer saline (PBS). Subsequently, the sections were gently rinsed with citric acid-based buffer. Endogenous peroxidase activity was quenched by applying 3% hydrogen peroxide directly to the tissue slices, followed by a 10-minute incubation at room temperature and another PBS wash step. To block non-specific binding, the slides were treated with 5% bovine serum albumin (50 μL) and incubated at room temperature for 10 minutes. Primary antibodies specific to CD276, diluted to 1:200, were then administered to the slices, which were subsequently incubated overnight in a 4°C environment. The following day, the slices were washed with PBS rinse, and the secondary antibody was applied, followed by one hour incubation at room temperature. After another PBS rinse, the antibody-antigen complexes were developed using the diaminobenzidine chromogen for 5-minute period. Counterstaining was performed using hematoxylin to highlight the cellular and tissue structures. After staining, the sections were dehydrated using ascending alcohol concentrations, and the preserved tissues were sealed under a cover slip with neutral resin, thus optimizing them for detailed examination under a microscope (Nicolet™ iN™ 5 FTIR, ThermoFisher Scientific, USA). The acquisition software was ToupCam (E3ISPM06300KPB).
Given that CD276 predominantly localizes in the cytoplasm, pale yellow, brownish-yellow, or brown granular staining within the cytoplasm at ×200 magnification was indicative of a positive result. To determine the positivity of a section, ten microscopic fields were randomly selected and evaluated at ×400 magnification, with scoring based on both the percentage of stained cells and the intensity of the staining. The scoring system for the proportion of positive cells was as follows: a score of zero was assigned for < 5% positivity, one for 5-30% positivity, two for 31-60% positivity, and three for > 60% positivity. For the staining intensity, the scale ranged from zero for no coloration to one for light yellow, two for brownish-yellow, and three for brown staining. The final assessment was calculated by multiplying the proportion of positively stained cells with the staining intensity score. Results yielding a combined score of < two points were classified as negative (−), while scores equal to or exceeding two were considered positive (+). 20
Quantitative Real-Time PCR
DNA Sequences of the Primers Used for Rt-PCR.

DNA, deoxyribonucleic acid; rt-PCR, real-time polymerase chain reaction; GAPDH, glyceraldehyde-3-phosphate dehydrogenase; CD276, cluster of differentiation 276.
ELISA
We performed ELISA tests on peripheral blood from 70 preoperative ccRCC patients (taken before surgery) and 72 healthy volunteers. Serum CD276 levels were quantified using ELISA kits (R&D Systems, Minneapolis, MN, USA). This assay was performed in strict compliance with the manufacturer’s instructions as described in previous studies. 21
Statistical Analysis
R software (version 4.2.1) with related accessory packages including “ggplot2[3.3.6], stats[4.2.1], car[3.1-0] and pROC[1.18.0]” was utilized in this study. Qualitative data were compared between the two groups using the Wilcoxon rank-sum test, whereas the Kruskal-Wallis test was used to compare the three groups. Cumulative data between the two groups were examined the χ2 test. Statistical significance was set at
Results
CD276 Showed Elevated Expressed in ccRCC Tissues Compared to Para-Cancer Tissues in TCGA Database
We first analyzed the 541 ccRCC and 72 para-cancer tissue samples from the TCGA-KIRC project dataset. The median value of CD276 expression in tumor tissues was 5.394, while that in para-caner tissues was 4.253. Significant disparities were observed in the expression of CD276 between cancerous and adjacent noncancerous tissues in patients with ccRCC (Mann-Whitney U = 4132.5, CD276 expression in ccRCC tissues and para-cancer tissues in TCGA and GEO databases. (A) The difference of CD276 expression between 541 cases of ccRCC tissues and 72 cases of adjacent samples from TCGA database was calculated using Wilcoxon rank sum test. (B) The difference of CD276 expression between 72 cases of ccRCC tissues and its counterpart para-carcinoma tissues from TCGA database was evaluated using Wilcoxon rank sum test. (C) The efficacy of CD276 expression to distinguish ccRCC tissues from normal samples was verified from ROC curve in TCGA dataset. (D) The difference of CD276 expression between 101 cases of ccRCC tissues and 101 cases of adjacent samples from GEO database was calculated using Wilcoxon rank sum test. (E) The efficacy of CD276 expression to distinguish ccRCC tissues from normal samples was verified from ROC curve in GEO dataset. FPR, false positive rate, TPR, true positive rate. CD276, cluster of differentiation 276; ccRCC, clear cell renal cell carcinoma; TCGA, The Cancer Genome Atlas; GEO, Gene Expression Omnibus; ROC, receiver operating characteristic; AUC, area under the curve; TPM, Transcripts Per Million.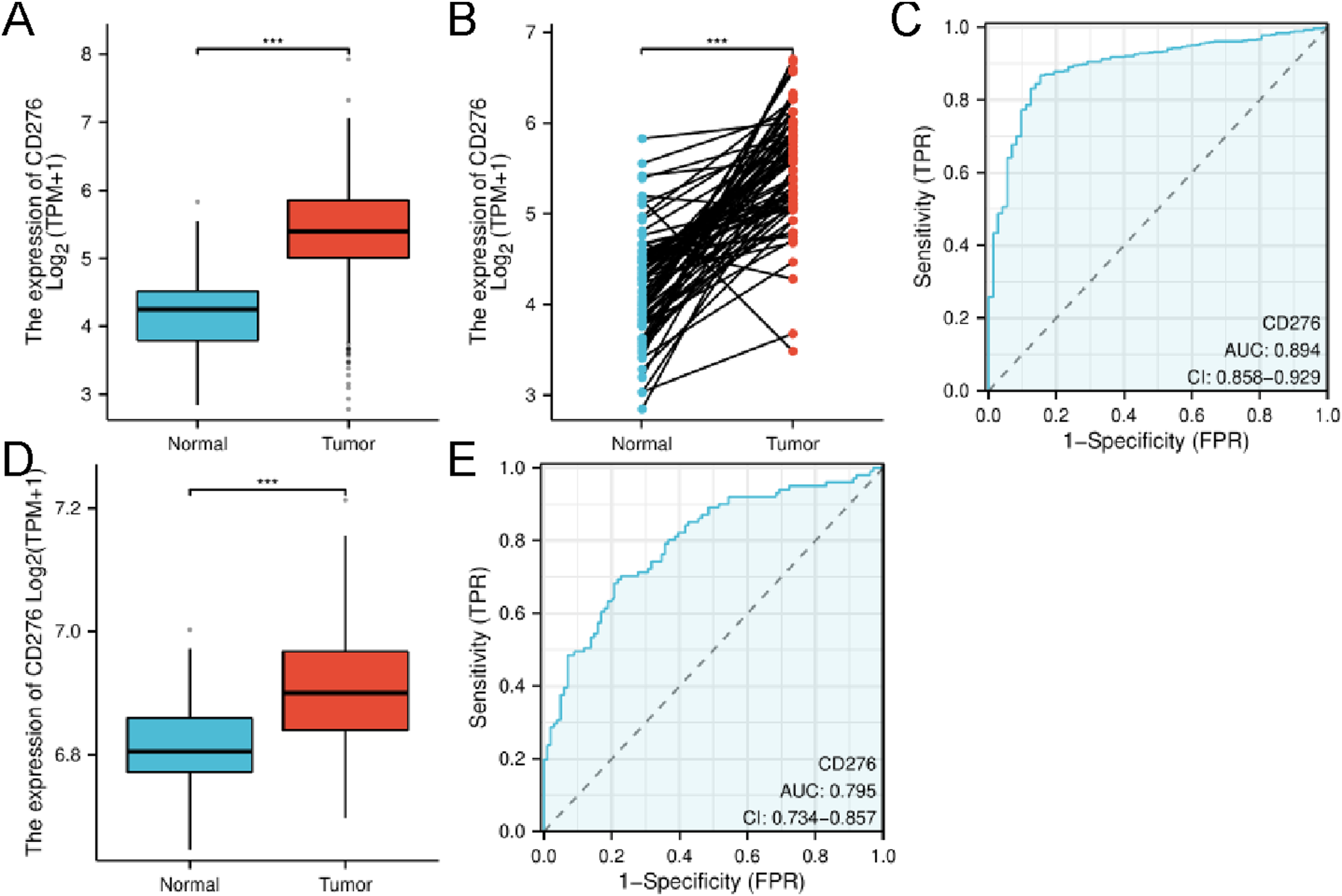
CD276 Showed Elevated Expression in ccRCC Tissues Compared to Para-Cancer Tissues in GEO Database
A total of 101 pairs of tumor samples and corresponding adjacent para-cancer samples were obtained from the GEO dataset GSE40435. The median value of CD276 expression in Tumor Group was 6.901, and that in Normal Group was 6.806 (t = 175.647,
CD276 Showed Elevated Expression in ccRCC Tissues Compared to in Para-Cancer Tissues in Clinical Dataset
Clinicopathologic Features of 70 Enrolled ccRCC Patients and 72 Healthy Volunteers.

IQR, interquartile range; SD, standard deviation; BMI, body mass index; HBP, high blood pressure; DM, diabetes mellitus; WHO, World Health Organization; N/A, not available.

The expression of CD276 in ccRCC and adjacent normal tissues. (A) Negative expression of CD276 in ccRCC (200x). (B) Negative expression of CD276 in ccRCC (400x). (C) Positive expression of CD276 in ccRCC (200x). (D) Positive expression of CD276 in CD276 (400x). (E) Negative expression of CD276 in para-cancer tissues (200x). (F) Negative expression of CD276 in para-cancer tissues (400x). (G) Positive expression of CD276 in para-cancer tissues (200x). (H) Positive expression of CD276 in para-cancer tissues (400x). ccRCC, clear cell renal cell carcinoma; CD276, cluster of differentiation 276.
CD276 Expression in ccRCC and Corresponding Adjacent Normal Tissues.

CD276, cluster of differentiation 276; ccRCC, clear cell renal cell carcinoma.

The expression and relationship of between CD276 and ccRCC in clinical dataset. (A) The expression of CD276 expression in 70 pairs of ccRCC tissues and adjacent samples was measured by RT-PCR and the difference between the two groups were calculated using Wilcoxon rank sum test. (B) The expression of CD276 expression in peripheral blood samples from 70 cases of ccRCC and 72 cases of normal individuals was measured by ELISA and the difference between the two groups were calculated using wilcoxon rank sum test. (C) The 142 cases were divided into three groups: Low Grade Group with 56 cases, High Grade Group with 14 cases and Normal Group with 72 cases. The differences of CD276 expression in peripheral blood samples were compared by kruskal-Wallis test. (D) The efficacy of CD276 expression to distinguish ccRCC patients from normal individuals was verified from ROC curve in clinical dataset. FPR, false positive rate, TPR, true positive rate. CD276, cluster of differentiation 276; ccRCC, clear cell renal cell carcinoma; ROC, receiver operating characteristic; AUC, area under the curve; rt-PCR, real-time polymerase chain reaction; ELISA, enzyme-linked immunosorbent assay.
CD276 Showed Elevated Expression in Patients With ccRCC Compared to Normal Individuals and was Positively Associated With WHO Grading System
Peripheral blood samples from 70 preoperative ccRCC patients and 72 healthy volunteers were analyzed using ELISA. The results showed that the median value of CD276 expression in Tumor Group was .185 pg/mL, whereas that in Normal Group was .126 pg/mL (U = 4729,
Discussion
Currently, treatment for advanced RCC mainly includes surgery, molecular targeted therapy, and immunotherapy. 22 Gene-targeted therapy is a promising therapeutic modality for advanced RCC. 23 Targeted drugs commonly used in clinical practice include sorafenib, sunitinib, everolimus, and nivolumab. However, these drugs are not ideal for extending the progression-free survival (PFS) of patients and only offer approximately one year of benefit.24–26 Therefore, an urgent need persists for identifying new therapeutic targets to provide new ideas for the treatment of advanced RCC.
CD276 is a relatively new member of the immune costimulatory molecule B7/CD28 family. 9 Subsequent research confirmed that CD276, a classic immune checkpoint molecule, could significantly inhibit the function of T cells.27,28 The earliest discovery was that programmed cell death-1 (PD-L1; also known as B7-H1) is expressed on antigen-presenting cells. Upon binding to PD-1 expressed on T cells, PD-L1 transmits inhibitory signals to suppress the immune response of T cells. Later, various malignant tumor cells were found to express PD-L1, and inhibit the immune response of T cells against tumors through the interaction between PD-L1 and PD-1 expressed on T cells, leading to immune evasion. 29 The role of CD276 is similar to that of PD-L1 in T cells. 30 However, as the receptor for human CD276 has not yet been discovered or confirmed, the exact immunological and biological functions and mechanisms of action of this molecule are not fully understood. Although the receptor for CD276 is not yet clear, studies suggest that CD276, as a “membrane molecule,” can mediate signals within the cells that express CD276. This phenomenon is known as “reverse signaling”.31,32 Suh et al found that the CD276Ig fusion protein can inhibit the mineralization and calcification of osteoblasts; using a CD276 gene knockout mouse model, they found that compared with wild-type mice, CD276 gene knockout mice had significantly lower cortical bone density, and their bone hardness and toughness were significantly reduced. 31 These results indicated that the absence of CD276 can lead to a disorder in bone differentiation and maturation. Thus, it is evident that the CD276Ig fusion protein does not operate by enhancing the inhibitory signal mediated by a supposed CD276 receptor (CD276R) in a “forward direction.” Instead, it operates by competitively inhibiting the “reverse signaling” mediated by CD276, which promotes bone development. This interference results in the hindrance of osteoblast differentiation and maturation. Another study 32 has provided evidence that the CD276 may reverse signaling. This research discovered that CD276 can be stably expressed at various differentiation and maturation stages of dendritic cells derived from monocytes (Mo-DCs). This result suggested that as a constitutively high-expressed membrane molecule on the surface of Mo-DCs, it might play an important role in the biological functions mediated by DCs. CD276 monoclonal antibody can upregulate the expression of costimulatory molecules CD80 and CD86 on Mo-DCs, enhancing their capacity to co-stimulate T cells; under the combined action of TNF-α, CD276 monoclonal antibody can significantly upregulate the level of CD83 on Mo-DCs, promoting their differentiation and maturation. This leads to the speculation that the expression of CD276 on the surface of DCs likely has reverse signaling effects on the biological functions of DCs.
Crispen PL et al conducted an immunohistochemical analysis on specimens from 743 patients who underwent radical nephrectomy for kidney cancer, discovering a heightened expression of CD276 in the ccRCC tissue samples. 14 This finding is supported by further research led by Saeednejad Zanjani L et al, which found a correlation between elevated CD276 levels and malignant biological behavior in ccRCC.15,16 Additionally, a study by Inamura K et al found that ccRCC patients with high CD276 expression showed an increase in FOXP3 + expression in the tumor microenvironment, thereby promoting a tumor developing response. 17
Despite the scarcity of research on the relationship between CD276 and ccRCC, our analysis of data from the TCGA and GEO databases verified that CD276 is expressed at significantly higher levels in cancer tissues than in adjacent non-cancerous tissues. To further validate our findings, we enrolled 70 ccRCC patient samples from the First Affiliated Hospital of Soochow University for analysis, and the experimental results further supported our conclusions. Furthermore, we collected peripheral blood samples from 70 patients with ccRCC and 72 normal controls to analyze CD276 expression. The results showed that the CD276 expression levels in the peripheral blood of ccRCC patients were higher than those in normal patients. Further subgroup analysis revealed that the expression level of CD276 was significantly higher in the High Grade Group compared to the Low Grade Group. The subsequent ROC diagnostic curve indicated that CD276 could serve as a predictive marker for renal clear cell carcinoma. Although so far, there have been no reports on the development of CD276 antibodies for treatment, as the mechanism between CD276 and the development of ccRCC becomes more clearly understood in the future, CD276 has the potential to be a biomarker for diagnosis, prognostic assessment, and even treatment of ccRCC.
However, our study has some limitations. First, this study involved a small sample size and is a single-center study; therefore, the conclusions need to be further confirmed by subsequent multi-center studies with larger samples. Second, this study only conducted research on the diagnosis of ccRCC using CD276, and because of the short follow-up period, we were unable to observe patient prognosis. We plan to further study the relationship between CD276 and ccRCC prognosis after accumulating data over an extended follow-up period. Third, our study’s audience was limited due to our lack of research into the relationship between the expression level of CD276 and different histological subtypes of RCC. In the future, we will consider including other types of RCC for further study. Furthermore, we did not investigate the mechanisms underlying the effect of CD276 on renal cancer. We will continue our in-depth research to explore the molecular mechanisms by which CD276 functions in ccRCC.
In summary, we found that CD276 is highly expressed in ccRCC and is positively correlated with pathological grade, demonstrating a strong capability to diagnose ccRCC.
Footnotes
Author Contributions
(I) Conception and design: Ouyang Jun, Lin Yuxin, Zhang Jianglei; (II) Administrative support: Ouyang Jun, Zhang Jianglei and Lin Yuxin; (III) Provision of study materials or patients: Zhang Zhiyu and Xu Jianhao; (IV) Collection and assembly of data: Zhiyu Zhang, Xu Jianhao and Song Zhen; (V) Data analysis and interpretation: Zhang Zhiyu, Xu Jianhao and Song Zhen; (VI) Manuscript writing: A
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The authors disclose receipt of the following financial support for the research, authorship, and/or publication of this article. This work was supported by the Suzhou Science and Technology Project (grant numbers SLJ201906 and SYS2019053) and National Natural Science Foundation of China (grant number 32200533).
Ethical Statement
Data Availability Statement
All original raw data of this study can be accessed from https://figshare.com/s/1cb57c7305879cd80c48. Publicly available data were also obtained from TCGA database (KIRC, https://portal.gdc.cancer.gov/) and GEO database (GSE40435, ![]() ).
).
