Abstract
Introduction
Cervical cancer (CC) is the third most prevalent malignancy among women worldwide. Candidate gene studies have identified multiple single nucleotide polymorphisms (SNPs) that are associated with an increased risk of CC. The objective of this study was to examine the relationship between 8 specific single-nucleotide polymorphisms (SNPs) and the risk of cervical cancer in the Georgian population.
Methods
The present study employed a prospective case-control design, with 40 patients diagnosed with CC and 45 healthy women. A total of 8 single-nucleotide polymorphisms (SNPs) were genotyped using the TaqMan genotyping assay: rs7579014, rs11263763, rs7726159, rs6897196, rs2853672, rs635634, rs231775, and rs2304204.
Results
Our analysis demonstrated that rs7579014 (BCL11A, G/A), rs7726159 (TERT, C/A), and rs6356634 (ABO, T/A) were associated with an increased risk of cervical cancer in Georgian patients. However, following the implementation of the Benjamini–Hochberg correction, only rs6356634 (ABO T/A) and rs7579014 (BCL11A G/A) remained statistically significant. A lack of statistically significant correlation was identified between the genetic variants rs11263763, rs6897196, rs2853672, rs2304204, and rs231775 and susceptibility to cervical cancer.
Conclusions
This study represents the first attempt to investigate SNP associations in women with cervical cancer in Georgia. The findings indicate that SNP-based analysis may hold promise for the early identification of susceptibility to cervical cancer, and potentially to other cancers. Nevertheless, further research involving larger sample sizes is required to validate and strengthen these preliminary observations.
Keywords
Introduction
Comparison of Cervical Cancer Incidence and Mortality Rates in Different Parts of the World

HPV is recognized as the causative agent in the majority of cervical cancer (CC) cases across most histologic types. 4 Efforts to identify driver mutations in HPV-independent cases have yielded promising data; however, the findings lack reproducibility across studies,5,6 and a consensus on common driver mutations has yet to be established, particularly given the rarity of these tumors.
Current evidence suggests that HPV infection alone is insufficient for the development of cervical cancer (CC), and that genetic predisposition plays a crucial role. This predisposition is regarded as a constellation of gene variants that influence both infection pathogenesis and carcinogenesis. Although each variant individually exerts only a modest effect on cancer risk, their cumulative impact substantially contributes to tumor development and overall disease epidemiology. Meta-analyses have demonstrated that the genetic background influencing susceptibility to cervical cancer (CC) is highly diverse, largely reflecting variations in immune-related genes.7-13 Single nucleotide polymorphisms (SNPs) associated with CC risk are predominantly located in or near immune-related genes, including human leukocyte antigens (HLAs), interleukin-8 (IL-8), interleukin-16 (IL-16), Toll-like receptors (TLR4, TLR9), interferon-gamma (IFNγ), interferon regulatory factors (IRFs), and tumor necrosis factor-alpha (TNFα).14-22 In addition, other SNPs may affect key cellular processes such as DNA repair mechanisms and cell cycle regulation, thereby further modulating the risk of malignant transformation.20,23
To the best of our knowledge, the Georgian population has not previously been investigated for genetic variants associated with cervical cancer (CC) susceptibility, nor has it been included in any genome-wide association (GWA) studies to date. Given the country’s unique geographical, historical, and cultural characteristics, it is reasonable to assume that genetic variants in this population may differ from those reported in populations represented in existing GWAS datasets. Hence, this study represents one of the first small-scale genomic investigations in the Georgian population. In general, GWAS data have demonstrated that a number of tumors share common risk alleles, with gynecologic tumors among them, despite the heterogeneity in the current understanding of their pathogenesis.24,25 Therefore, the aim of our study was to analyze reported cancer risk alleles in relation to their potential impact on CC risk.
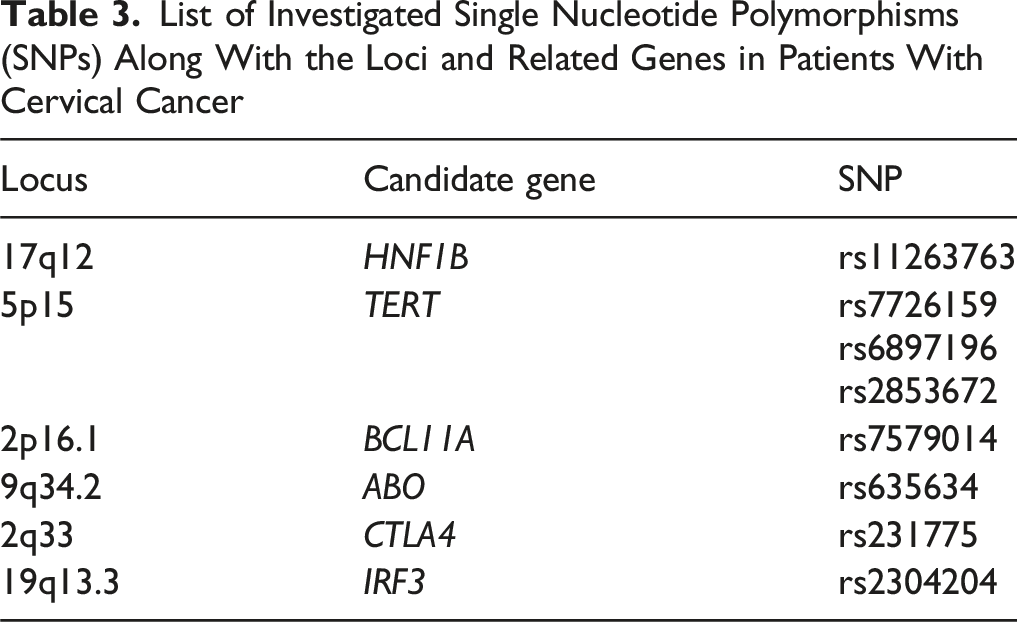
The 8 SNPs investigated in this study, rs7579014, rs11263763, rs7726159, rs6897196, rs2853672, rs635634, rs231775, and rs2304204, were selected based on prior evidence implicating them in cancer susceptibility pathways, immune regulation, or genome stability.26-31
Materials and Methods
Study Subjects
The study was approved by the Ethics Committee of Tbilisi State Medical University (Tbilisi, Georgia) (approval number: N7-2021/91; approval date: 29 November 2021). Written informed consent was obtained from all participants, and patient information was fully de-identified. This prospective case–control study included 40 patients with cervical cancer (CC) (mean age 65.1 ± 10.8 years) and 45 age-matched healthy control subjects (mean age 65.0 ± 10.2 years).
Cervical cancer patients were recruited between December 2021 and November 2024 from the Inova Medical Center, Tbilisi, Georgia.
The inclusion criteria for patients were: (1) histologically confirmed cervical cancer (CC), (2) age ≥20 years, and (3) self-identified Georgian ancestry. Patients with a history of other cancers or significant co-morbidities were excluded.
The selection criteria for controls were: no history of cancer within the past 5 years, no family history of cancer, age ≥20 years, and self-identified Georgian ancestry. Minor illnesses (eg, common cold, headache) were not considered exclusionary.
Demographic and Clinical Characteristics of Cervical Cancer Cases and Controls

Sample Collection and Storage
Peripheral blood samples were collected in vacutainer tubes containing ethylenediaminetetraacetic acid (EDTA). Genomic DNA was extracted from whole blood using a commercially available DNA purification kit (Zymo Research, USA). DNA concentration was quantified using a fluorometer-based method. Extracted DNA samples were initially stored at −20°C and subsequently transferred to −80°C for long-term archiving.
SNP Genotyping Using TaqMan Assay
List of Investigated Single Nucleotide Polymorphisms (SNPs) Along With the Loci and Related Genes in Patients With Cervical Cancer
All TaqMan SNP Genotyping Assays can be optimized with master mixes from different suppliers and require only 3 components for PCR: 1-20 ng of purified genomic DNA per well (final concentration: 0.2 ng/µL), the SNP Genotyping Assay, and a compatible master mix. In this study, the TaqMan Genotyping Master Mix (Thermo Scientific, USA) was used.
Procedures Briefly
PCR reactions were prepared using DNA polymerase, sequence-specific forward and reverse primers (final concentration: 50-500 nM), probes (final concentration: 250 nM), and 10-100 ng of genomic DNA in a final reaction volume of 25 µL. PCR conditions: Polymerase activation was performed at 95°C for 10 min, followed by 40 cycles of denaturation at 95°C for 15 s and annealing/extension at 60°C for 1 min. The post-PCR read and analysis: Allelic discrimination was performed on a QuantStudio 3 Real-Time PCR System (Applied Biosystems, USA). The instrument software generated allelic discrimination (AD) plots, displaying results as scatter plots of Allele 1 (VIC dye) vs Allele 2 (FAM dye). Each reaction well was represented as an individual point on the plot. A typical AD plot showed 3 distinct clusters corresponding to homozygotes for Allele 1, homozygotes for Allele 2, and heterozygotes, while no-template controls clustered separately. The points within each cluster grouped tightly and were clearly distinguishable from one another.
Genotyping quality control included visual inspection of allelic discrimination plots to confirm well-separated genotype clusters and the absence of systematic plate effects. All 85 DNA samples yielded genotype calls for all 8 SNPs, resulting in a 100% genotyping success rate and no missing genotypes in the final dataset. No duplicate samples were included; therefore, we could not directly estimate technical error rates, but the clear cluster separation and internal controls supported high genotyping reliability.
Histology Revision
In all selected cases, tumor histology was re-evaluated to ensure consistent assessment and to obtain a clearer understanding of the histological subtypes.
Statistical Analysis
The study and control groups were analyzed separately. Statistical analyses were performed using GraphPad Prism version 9.3.1 for macOS (GraphPad Software, San Diego, CA, USA) and R (R Foundation for Statistical Computing). Hardy–Weinberg equilibrium (HWE) for each SNP was assessed using the Chi-square goodness-of-fit test. Associations between SNPs and cervical cancer risk were evaluated by calculating odds ratios (ORs) and 95% confidence intervals (CIs) using unconditional logistic regression. A P-value < 0.05 was considered statistically significant. To correct for multiple testing, adjusted P-values were obtained using the Benjamini–Hochberg false discovery rate (FDR). All statistical tests were two-sided. Post hoc power analyses further confirmed that, with 40 cases and 45 controls, the study was well powered to detect the relatively large effects observed for rs7579014 and rs635634 but had limited power (⁓60%) to detect more modest effect sizes (eg, OR ≤2.0), particularly for SNPs with lower minor-allele frequencies. Prospective power calculations were also performed to assess the study’s ability to detect typical genetic effect sizes. With the current sample size (40 cases and 45 controls), the study has <25% power to detect a modest effect size (OR≈1.5) and approximately 58% power to detect a larger effect size (OR≈2.0), assuming a minor allele frequency around 0.30.
Results
Histological assessment revealed that the majority of cases were squamous cell carcinomas, with only 6 cases identified as cervical adenocarcinomas. Given the relatively small sample size, stratification of results by histologic type, tumor grade, FIGO stage, or demographic characteristics was not considered meaningful. Therefore, analyses of polymorphism and genotype frequencies were performed for the cohort as a whole.
Genotype Frequencies
Summary of the Risk of Genotype Frequencies and Distribution in Patients With Cervical Cancer

Abbreviations: SNP = Single Nucleotide Polymorphisms; OR = Odds Ratio; CI = Confidence Interval.
astatistically significant after Benjamini–Hochberg correction.
A significant difference in genotype distribution between cases and controls was observed for rs7726159 (TERT, C/A). Individuals carrying the heterozygous CA genotype had a markedly increased risk of developing cervical cancer (OR = 5.84, P = 0.006). The AA genotype also suggested an elevated risk (OR = 4.96, P = 0.02). However, after correction for multiple testing using the Benjamini–Hochberg method, neither the CA nor the AA genotype associations remained statistically significant. The allele frequency difference (A: 0.625 in cases vs 0.44 in controls) corresponds to a post hoc power of ⁓65%, indicating that the study may have been underpowered to confirm modest effects for this locus.
For rs635634 SNP (ABO T/A) genotype distribution differed significantly between cases and controls, indicating a possible association. In additions, individuals with AA genotype had a significantly higher risk of cervical cancer compared to those with TT (OR = 3.5; CI: 1.33-9.24; P = 0.018). However, this association did not remain significant after correction for multiple testing (adjusted P = 0.88). In the additive model, the rs635634 (ABO, T/A) variant was associated with an increased risk of cervical cancer (OR = 3.38, 95% CI: 1.73-6.61; raw P < 0.001; adjusted P = 0.002). Given the large observed allele frequency difference (A: 0.48 in cases vs 0.21 in controls), post hoc calculations showed ⁓95% power to detect this association at α = 0.05. Although the significant association observed for ABO rs635634, the finding should be interpreted with caution, as this SNP showed a strong deviation from Hardy–Weinberg equilibrium (HWE) in controls. Such deviation may reflect potential genotyping error, population stratification, or, less commonly, a true biological effect.
For rs7579014 (BCL11A, G/A), the AA genotype was strongly associated with increased cervical cancer risk (OR = ⁓ 12.0, raw P = 0.005), and importantly, this association remained significant after FDR correction (adjusted P = 0.024). However, the wide confidence interval (CI: 2.37-60.65) indicates uncertainty due to the small number of AA carriers, suggesting that the effect size is likely overestimated. The GA genotype was associated with a moderate, but not statistically significant, increase in risk (OR = 2.56; raw P = 0.176). In the additive model, each additional A allele conferred an approximately 2.5-fold increase in cervical cancer risk (OR = 2.46, CI: 1.33-4.56), which remained statistically significant after FDR correction (adjusted P = 0.024). Allele frequencies (A: 0.59 in cases vs 0.37 in controls) yielded ⁓82% post hoc power, suggesting adequate ability to detect the observed effect despite the small sample size.
Prospective calculations indicated low power (<25%) to detect modest effect sizes (OR≈1.5), and only moderate power (∼58%) for OR≈2.0.
Discussion
Despite advances in screening programs and prophylactic human papillomavirus (HPV) vaccination strategies, cervical cancer (CC) remains a major global health concern. Its etiology is multifactorial, underscoring the contributory role of genetic variation. Incidence and mortality rates of CC show considerable geographical heterogeneity.1-4 The identification of genetic polymorphisms that predispose individuals to an increased risk of CC holds promise for improving risk stratification and guiding the development of targeted preventive strategies.
In this case–control study, we investigated the potential association between 8 selected SNPs and overall cervical cancer (CC) risk in the Georgian population. To our knowledge, this is the first study in Georgia to evaluate germline genetic variants and their impact on CC susceptibility, thereby contributing to the growing evidence for the role of genetic background in cervical carcinogenesis. As the Georgian population has not been represented in GWAS or other large-scale studies, our preliminary findings provide novel and valuable insights into genetic susceptibility within this understudied Caucasus population.
Our findings revealed significant associations between CC and 3 of the tested variants: rs7726159 (TERT, C/A), rs7579014 (BCL11A, G/A), and rs635634 (ABO, T/A). After applying the Benjamini–Hochberg correction, only rs635634 (ABO, T/A) and rs7579014 (BCL11A, G/A) remained statistically significant. No significant associations were observed for the remaining SNPs, including variants in CTLA4 and IRF, which have previously been reported to increase CC risk in other populations.18,20 For example, Hu et al. investigated CTLA-4 +49 A/G (rs231775) in 2 large independent Chinese case–control cohorts (hepatocellular carcinoma and cervical cancer) and demonstrated that this variant contributes to cancer susceptibility; the cervical cancer cohort confirmed the association of rs231775 in an East Asian population. 20 The discrepancies between our findings and prior studies may reflect population-specific genetic architecture, limited sample size, or potential gene–environment interactions unique to the Georgian cohort.
In our study, only one SNP, rs6356634, showed a significant deviation from Hardy–Weinberg equilibrium in the control group. While this deviation may reflect a true association with CC risk, as suggested by our results, it requires cautious interpretation, as it could also arise from population substructure at this locus. Therefore, the findings for this SNP should be validated in larger, independent cohorts. For BCL11A rs7579014, the Hardy-Weinberg equilibrium test in controls yielded a borderline result (P = 0.05). Although not formally significant at the α = 0.05 level, this suggests that the genotype distribution was only marginally consistent with HWE, likely reflecting the small number of AA carriers and the resulting statistical instability.
The rs7726159 SNP is located in an intronic region of the telomerase reverse transcriptase (TERT) gene. This variant, with a minor allele frequency of approximately 0.3-0.4, has been associated with an increased risk of breast and ovarian tumors.33-35 Although it was long considered that this intronic variant exerted no functional effect on the TERT gene, several hypotheses have been proposed to explain its role in cancer susceptibility. More recently, rs7726159 has been linked to alterations in telomere length, with carriers of the C allele exhibiting shorter telomeres,36,37 suggesting that it may indeed exert a direct effect on TERT protein function. Our observation of an association between rs7726159 and cervical cancer in the Georgian cohort further supports its potential role in carcinogenesis, although larger studies will be required to validate this finding.
The rs635634 SNP is located in the ABO gene (alpha 1-3-N-acetylgalactosaminyltransferase and alpha 1-3-galactosyltransferase), which determines the ABO blood groups. This variant has been associated with susceptibility to ovarian, breast, bladder, and prostate cancers.38,39 In addition, solid evidence supports its role in obesity and in modulating the relationship between obesity and cancer risk among carriers of the risk allele. 39 Our results indicate that rs635634 may also play a role in cervical cancer susceptibility in the Georgian population.
An additional strength of this study is the careful selection of SNPs based on prior oncogenomic literature relevant to gynecologic tumors beyond CC. For example, BCL11A rs7579014, included in our SNP panel, has been implicated in endometrial cancer risk34,40 and may exert its oncogenic effects through transcriptional repression of tumor suppressor pathways. Moreover, the role of BCL11A in immune response regulation may influence the persistence of oncogenic HPV infection, which underlies the majority of CC cases. 41 Taken together, these biological roles provide a plausible explanation for our observed association of BCL11A rs7579014 with cervical cancer risk in the Georgian cohort.
Post hoc power calculations indicated that the study had adequate power to detect the observed effects for rs7579014 (⁓82%) and rs635634 (⁓95%), but only moderate power for rs7726159 (⁓65%). Prospective estimates further showed limited power (<25% for OR≈1.5; ∼58% for OR≈2.0) for SNPs with modest effects or lower minor-allele frequencies, suggesting that the lack of associations for several variants may largely reflect insufficient statistical power rather than true null effects.
Overall, this study provides a foundation for further research into CC polygenic risk modeling and biomarker identification in Georgian population. In further studies will plan to expand the sample size and incorporate additional SNPs to improve statistical power and detect subset genetic association. We also aim to investigate patients with different gynecologic tumors (viz., endometrial and ovarian cancers) and integrate relevant environmental and lifestyle factors such as smoking status, diet, stress level and other relevant exposures.
Overall, this study provides a foundation for further research into polygenic risk modeling and biomarker identification for CC in the Georgian population. In future studies, we plan to expand the sample size and incorporate additional SNPs to enhance statistical power and enable detection of subgroup-specific genetic associations. We also aim to extend our investigations to patients with other gynecologic tumors, including endometrial and ovarian cancers, and to integrate relevant environmental and lifestyle factors such as smoking status, diet, stress level, and other exposures.
In conclusion, our results suggest that rs7579014 (BCL11A, G/A) and rs635634 (ABO, T/A) may be associated with cervical cancer susceptibility in Georgian women. These preliminary findings should be interpreted with caution, as validation in larger, multicenter studies is required to confirm and further evaluate their potential as genetic markers for cervical cancer in this population. Ultimately, such insights may contribute to improved risk stratification and the development of precision prevention strategies tailored to the Georgian and other understudied populations.
Limitations
This study is limited by its small sample size, which reduces statistical power and may inflate effect estimates, particularly for rare genotypes. One SNP (ABO rs635634) also deviated from Hardy–Weinberg equilibrium in controls, raising concerns about potential genotyping error or underlying population structure. Incomplete clinical data further limited the ability to perform stratified analyses. Post-hoc power analyses indicated that the study was adequately powered only for large effect sizes, limiting the detection of modest genetic associations. Therefore, these findings should be interpreted with caution and validated in larger, independent cohorts.
Footnotes
Ethical Considerations
The study was approved by the Ethics Committee of Tbilisi State Medical University (Tbilisi, Georgia) (approval number: N7-2021/91; Approval date: 29/11/2021).
Consent to Participate
A signed informed consent was collected from every study participant.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by Shota Rustaveli National Science Foundation of Georgia (grant #FR-21-17599).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The data supporting the findings of this study are available from the corresponding author upon request.
