Abstract
Background and Aims
So far, long noncoding RNAs (lncRNAs) signatures in acute myeloid leukemia (AML) are poorly understood. The present study aims to explore the prognostic significance of eleven cancer-related lncRNAs in bone marrow (BM) samples from adult Egyptian AML patients.
Materials and Methods
In this study, we analyzed eleven lncRNAs using the qRT-PCR assay in the bone marrow (BM) of 79 de novo AML adult patients before receiving any therapy.
Results
Five lncRNAs out of 11 were aberrantly expressed, and two lncRNAs influenced significantly the patient’s overall survival (OS). LncRNA-XIST was favorable when overexpressed (in univariate and multivariate analysis, P-value = .001). LncRNA-GAS5 adversely affected the OS (only in multivariate analysis P-value = .02). Two other lncRNAs (UCA1 and HIFAL) impacted complete remission induction (CR) significantly in univariate analysis (P-value = .046 for both). Furthermore, lncRNA-UCA1 affected CR significantly in multivariate COX regression analysis (P-value = .004). The 4 previously mentioned lncRNAs were among the 9 downregulated lncRNAs. Instead, the only 2 upregulated lncRNAs (SNHG15, MALAT1) did not significantly influence neither CR induction nor OS. LncRNA-PTCSC3, a fifth lncRNA, emerged as the only one that could predict relapse occurrence in an upfront original BM sample.
Conclusion
Two lncRNAs out of eleven (lncRNA-XIST and GAS5) impacted OS, and two other lncRNAs (UCA1 and HIFAL) affected CR in adult de novo AML patients. LncRNA-PTCSC3 predict relapse, however, further validation is still required.
Introduction
AML is an aggressive and molecularly heterogeneous disease with various genetic mutations and molecular characteristics. 1 AML is the most common form of acute leukemia in adults, with a worldwide incidence of approximately 5.4 per 100,000 per year. The incidence of AML increases with age, and the disease is more common in men than women. 2 Minimal residual disease (MRD) on day 14 is currently used to monitor disease. Looking for an upfront marker would help in improving clinical decision-making. Recent research identified various lncRNAs implicated in leukemia diagnosis and treatment. 3
lncRNAs are a class of endogenous, non-protein-coding RNAs with a length longer than 200 nucleotides that are highly linked to various cellular functions and pathological processes.4,5
Recent advances in sequencing technologies have enabled the discovery of an increasing number of lncRNAs. These newly discovered lncRNAs are providing new insights into the mechanisms underlying cancer development and progression. 6 However, the precise mechanisms by which lncRNAs modulate AML development and progression are not fully understood. 7 Expression of many oncogenic lncRNAs can promote cancer.8-10 Potentials as biomarkers, and their functions as oncogenes and tumor suppressors. Several dysregulated lncRNAs in cancer were previously reported like lncRNAs-SNHG15, HIFAL, NKILA, and PTCSC3. 11 So far, these lncRNAs have not been analyzed before in Egyptian AML patients.
LncRNA-urothelial carcinoma-associated 1 (UCA1) is a long non-coding RNA that modulates cell proliferation and apoptosis by regulating the miR-296-3p/Myc axis. 7 Further studies have shown that LncRNA-UCA1 is overexpressed in a wide range of tumors, including breast cancer and acute myeloid leukemia (AML).12,13
The lncRNA reported to influence clinical outcomes in AML included lncRNA-X-inactive-specific transcript (XIST) and lncRNA antisense for X-inactive-specific transcript (TSIX). LncRNA-XIST is a lncRNA that regulates X chromosome inactivation in females, and its overexpression has been associated with dismal outcomes in AML patients. Lower expression of lncRNA-XIST could upregulate the expression of miR-196b and inhibit the proliferation and invasion of KG1a cells.14,15 Meanwhile, lncRNA-TSIX is lncRNA with a substantial role in X chromosome inactivation 16 as being a cancer immune biomarker. 17
Another crucial player in carcinogenesis is lncRNA-HIFAL; HIF1A is a transcription factor that controls oxygen delivery and utilization as a critical modulator of oxygen homeostasis.18,19 HIFAL is crucial for maintaining and promoting HIF-1α-mediated transactivation and glycolysis. 20 Cancer cells adapt to hypoxia by altering their metabolism through gene expression reprogramming and proteomic changes that impact various cellular and physiological functions. 21
LncRNA-Metastasis-associated lung adenocarcinoma transcript1 (MALAT-1) was reported to be overexpressed in AML cells, and its overexpression is associated with poor prognosis in AML patients. Where, knocking down of LncRNA-MALAT-1 inhibits AML cell proliferation and migration, suggesting that LncRNA-MALAT-1 plays a critical role in AML progression. 22
Antisense non-coding RNA in the INK4 locus (ANRIL) has been found to stimulate cell proliferation, migration, and invasion while inhibiting cell apoptosis in AML cells. 23
LncRNA-H19 is encoded by the H19 gene and confirmed as a critical player in cancer progression in many tumors. 24 It is upregulated in AML cells and its overexpression is associated with poor prognosis in AML patients. Previous studies demonstrate that the knockdown of lncRNA-H19 promotes AML cell apoptosis, suggesting that lncRNA-H19 plays a critical role in inhibiting apoptosis in AML cells. 25
Another study concluded that the lncRNA- growth arrest specific 5 (GAS5) polymorphism predicts a poor prognosis of AML in Chinese patients via affecting hematopoietic reconstitution and that the GAS5/BCL2L11 pathway could be a potential therapeutic target for AML. 26
Small nucleolar RNA host gene 15 (SNHG15) is specified as a pivotal regulator in carcinogenesis and various human cancer progression, its downregulation has been reported as a critical biomarker and therapeutic target for cancer patients. 27
Furthermore, LncRNA- Papillary thyroid carcinoma susceptibility candidate 3 (PTCSC3) and LncRNA-NKILA (NF-KappaB Interacting LncRNA) were previously reported as tumor suppressors in some human malignancies.28,29
The present study was conducted to assess the potential prognostic value of these lncRNAs as biomarkers in bone marrow specimens of adult AML patients for clinical management.
Patients and Methods
Patients and Samples Collection
Clinical/Laboratory Data of the Studied Group.

TLC: total leucocytic count, HB: hemoglobin, PB: peripheral blood, BM: bone marrow, AML: acute myeloid leukemia, FAB: French-American-British classification, FLT3-ITD: internal tandem duplication, FLT3-TKD: tyrosine kinase domain, NPM: nucleophosmin, INV 16: inversion chromosome 16, NEG: negative, PML/RARa: promyelocytic leukemia/retinoic acid receptor alpha.
Total RNA Isolation/Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
Total RNA was isolated from BM samples using QIAzol lysis reagent (Qiagen, Hilden, Germany) in line with the manufacturer’s instructions. Extracted RNA was reverse transcribed into cDNA using the miScript II RT kit according to the manufacturer’s instructions. Relative expression of 11 different lncRNAs (lncRNAs-TSIX, XIST, SNHG15, HIFAL, ANRIL, NKILA, UCA1, H19, PTCSC3, MALAT1, GAS5) was performed by qRT-PCR using the miScript SYBR Green kit (Qiagen), where 1 μL of diluted RT product was used as a template in a 10 μL PCR reaction containing 1X SYBR Green master mix, 200 nM lncRNA specific forward primer, and 200 nM universal primer. The qRT-PCR conditions were as follows: 95°C for 10 min, followed by 40 cycles of 95°C for 15 s and 60°C for 30 s. All qRT-PCR reactions were carried out on a ViiA 7 real-time PCR system (Applied Biosystems, Foster City, CA, USA). The expression of glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was used for normalization. The real-time PCR experiments were performed in duplicate. The primers were designed using the Primer-BLAST tool available on the NCBI website (https://www.ncbi.nlm.nih.gov/tools/primer-blast/), and were obtained from (eurofins Genomics, Hamburg, Germany).
Fold Change Calculation
The lncRNA expression levels were evaluated by the (2−ΔΔCT) method. For each lncRNA, first, ΔCT was calculated by subtracting the CT values of the GAPDH from the CT values of the target lncRNA. Second, ΔΔCT was then calculated by subtracting the average ΔCT of the healthy control samples from the ΔCT of the patient’s samples. Then to determine the relative quantitative levels of target lncRNA, the fold change (FC) in the lncRNA expression level was calculated (fold change = 2−ΔΔCT). Lastly, lncRNA relative expression levels are presented by Log2 FC.
Statistical Analysis
All data were statistically analyzed using a statistical package for social science ‘IBM SPSS Statistics for Windows, version 28.’ Categorical data were presented as numbers and percentages (%), while continuous data were presented as mean ± standard deviation (SD) with a 95% confidence interval. Non-normal variables were summarized as the median and interquartile range (IQR) and used the frequencies and percentages for categorical variables. Data normality was assessed statistically using the Shapiro test and visually by examining histograms and Q-Q plots. Comparisons between continuous data were carried out using the Mann-Whitney U-test. Univariate analysis was conducted to determine the prognostic performance of each studied biomarker then multivariate analysis using backward stepwise Cox regression was conducted. Categorical variables were treated as continuous variables in the multivariate Cox regression analysis for the sake of not losing information and to allow for higher statistical power. Survival analysis was carried out using the Kaplan-Meier test and Cox regression analysis. A P value <.05 was considered statistically significant. Adjustment of multiple testing was done using the Benjamini-Hochberg procedure. Overall survival (OS) was calculated from the date of diagnosis till the date of death or last follow-up. Disease-free survival (DFS) was calculated from the date of complete remission till the date of relapse, death, or last follow-up.
Results
Relative Expression of lncRNAs-TSIX, XIST, SNHG15, HIFAL, ANRIL, NKILA, UCA1, H19, PTCSC3, MALAT1 & GAS5 in AML Patients
Relative Expression of Different the Studied lncRNAs in AML Patients Compared to Control Group.
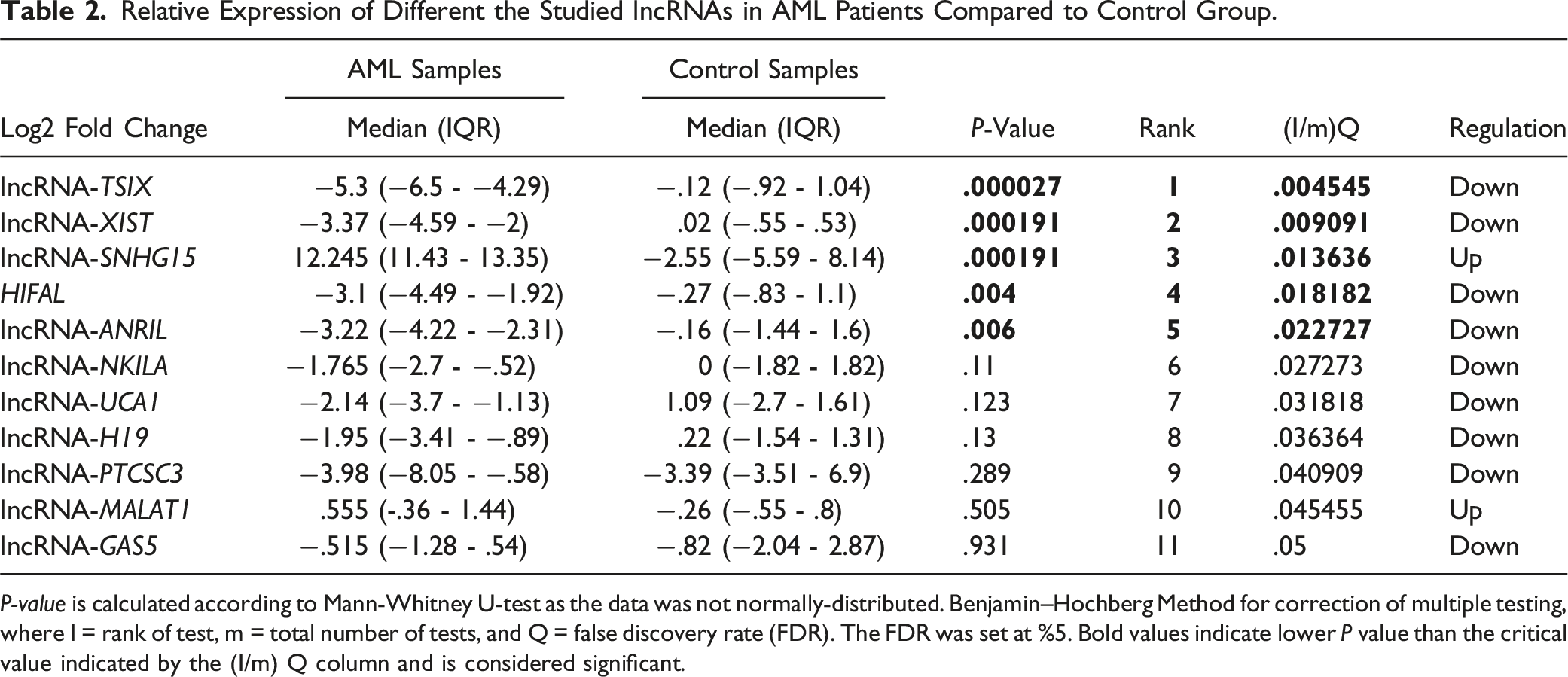
P-value is calculated according to Mann-Whitney U-test as the data was not normally-distributed. Benjamin–Hochberg Method for correction of multiple testing, where I = rank of test, m = total number of tests, and Q = false discovery rate (FDR). The FDR was set at %5. Bold values indicate lower P value than the critical value indicated by the (I/m) Q column and is considered significant.

Heatmap showing the expression of the evaluated lncRNAs in AML patients presented in log2 fold change (FC).
Effect of Different lncRNA Expressions on Response to Chemotherapy in the Studied Group
Among 79 who received the conventional induction chemotherapy, CR was achieved in 50 (63.3%) patients only, relapse occurred in 9 (4.5%) out of 50 patients. The median survival time was 3.68 months for overall survival, 0.95 months for complete remission, and 10.82 months for relapse after complete remission (disease-free survival). At the end of the 55 (69.6%) out of 79 studied patients died. The survival status of the studied patients is shown in Figure 2. OS, CR, and time from complete remission to relapse are shown in Figure 3(A)-(C). Flowchart detailing the survival status of the studied AML patients. Kaplan Meier survival curves showing (A): overall survival of the studied patients, (B): time till complete remission in months, (C): time from complete remission till relapse (disease free survival).

Univariate Cox Regression Showing the Different Measured lncRNAs as Prognostic Factors for Overall Survival and Complete Remission.

HR: hazard ratio; CI: confidence interval.
Bold text: significant values P-value <.05. Higher HR indicates higher risk for the outcome. For overall survival, higher HR means higher risk for death in patients with elevated expression of the lncRNA, which is unfavorable. For complete remission, higher HR indicates higher “risk” for complete remission with elevated expression, which is favorable.

Effect of relative expression of lncRNA-XIST on overall survival. Low and high expression were categorized according to the median Log FC value.
Survival Analysis of the Patient’s Clinical/Laboratory Data as Prognostic Factors for Overall Survival.

TLC: total leucocytic count; HB: hemoglobin; PB: peripheral blood; BM: bone marrow; AML: acute myeloid leukemia; FAB: French-American-British classification; IPT: immunophenotyping; FLT3-ITD: internal tandem duplication; FLT3-TKD: tyrosine kinase domain; NPM: nucleophosmin; INV 16: inversion chromosome 16; NEG: negative; PML/RARa: promyelocytic leukemia/retinoic acid receptor alpha; MST: median survival time; HR: hazard ratio; CI: confidence interval.
aVariable was treated as continuous; no categorization was done and therefore Log-rank test was not performed.
bMST not calculated as one or more categories contained 0 events.
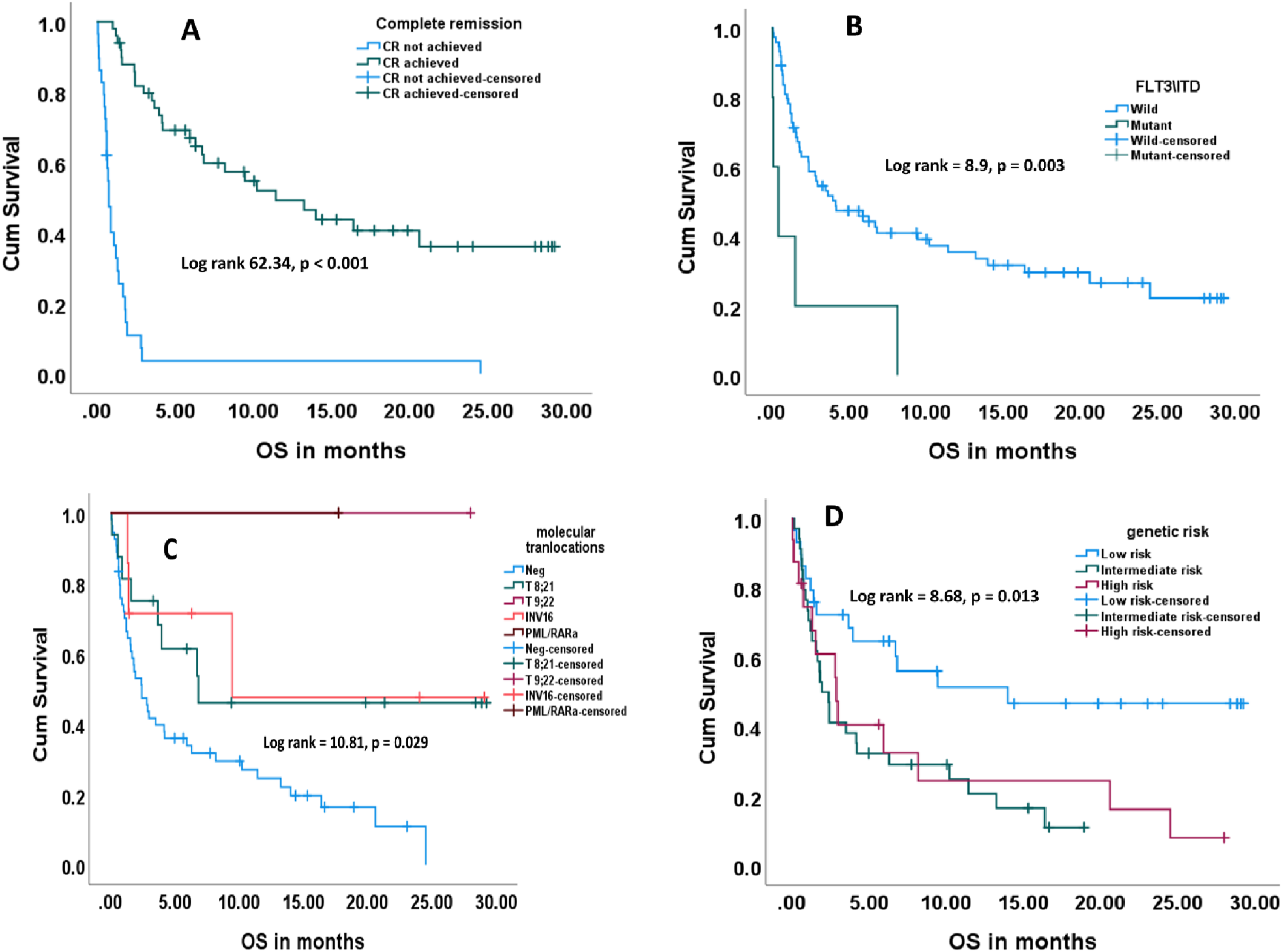
Kaplan meier survival cuves showing the effect of some clinical parameters on patient’s overall survival. (A): Achieving complete remission, (B): FLT3/ITD gene, (C): molecular translocations, (D): genetic risk.
Kaplan Meier Survival Analysis Paired With Log-Rank Test to Examine the Effect of the Clinical Parameters on Patient’s OS
Univariate analysis revealed that FLT3/ITD of the mutant type (HR 3.37, 95% CI 1.47 - 9.47, P-value = 0.006), moderate or high genetic risk (HR = 2.51, 95% CI = 1.3:4.83, P-value = 0.006), higher initial PB blasts (HR = 1.01, 95% CI = 1.0:1.02, P-value = 0.047) were adverse risk factors for OS. On the other hand, t (8; 21) translocation (HR = 0.42, 95% CI = 0.2: 0.9, P-value = 0.026), and achieving CR (HR = 0.12, 95% CI = 0.07:0.22, P-value <0.001) carried favorable prognosis regarding the OS (Table 4, Figure 6(A)-(C)). Moreover, none of the clinical parameters had any effect on the time to CR (Data not shown). Kaplan Meier survival curves showing the effect of expression of some lncRNAs on complete remission ((A): lncRNA-UCA1, (B): lncRNA-XIST, (C): lncRNA-HIFAL).
Multivariate Survival Analysis Including Both Clinical Parameters and Measured lncRNAs Expression
Backwards Stepwise Multivariate Cox Regression Showing Prognostic Factors for Overall Survival.

B: beta coefficient; SE: standard error; aHR: adjusted hazard ratio; CI: 95% confidence interval.
aInitial variables used in the multivariate model: age, sex, TLC at presentation, HB at presentation, PB blasts at presentation, FLT3/ITD, molecular translocations, genetic risk, complete remission, Log2 FC of the following lncRNAs: XIST, and GAS5.
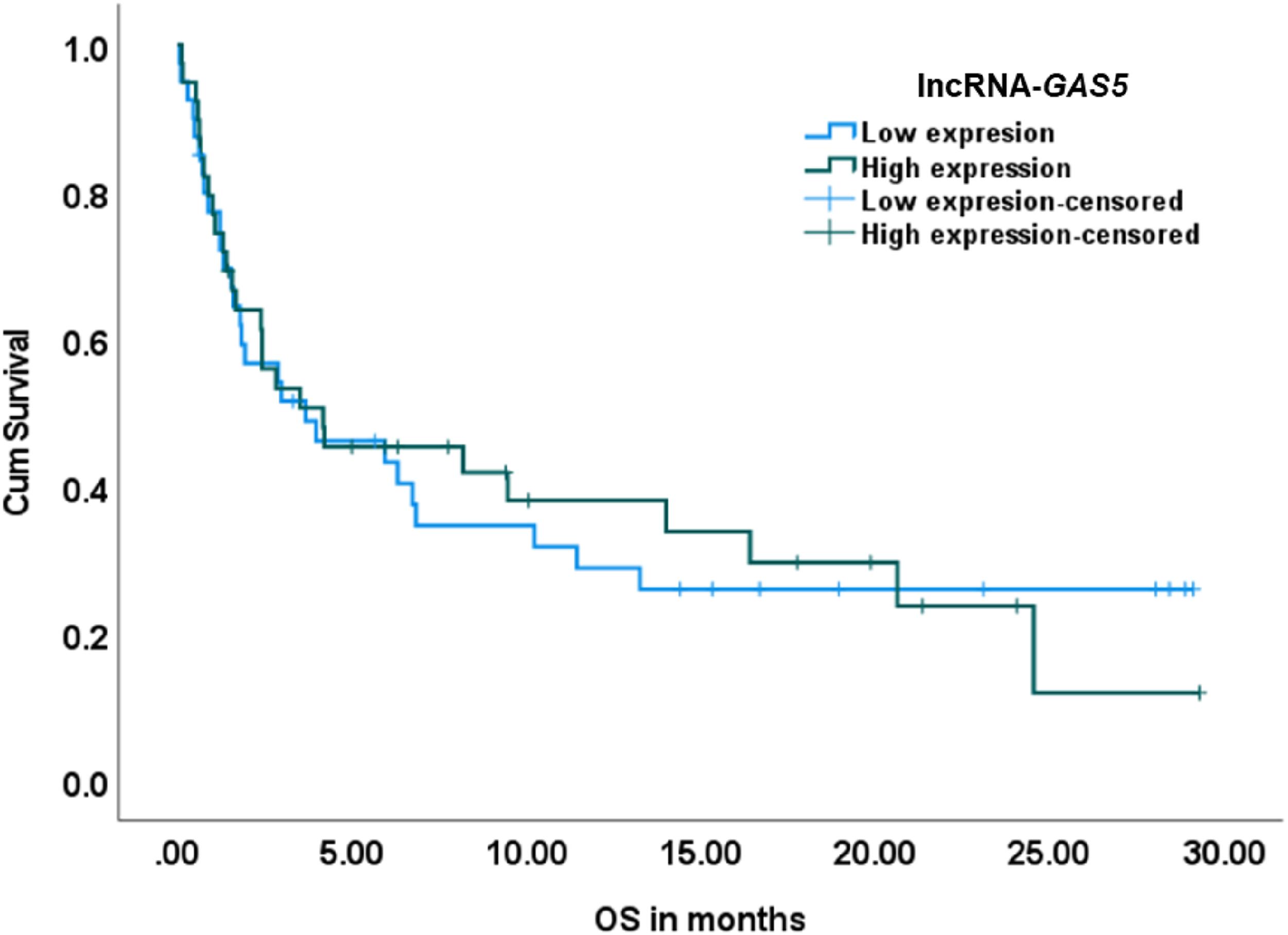
Effect of relative expression of lncRNA-GAS5 on overall survival. Low and high expression were categorized according to the median Log2 FC value. No significant effect was found in the univariate model (Log rank = .009, P = .922).
Backwards Stepwise Multivariate Cox Regression Showing Prognostic Factors for Complete Remission.
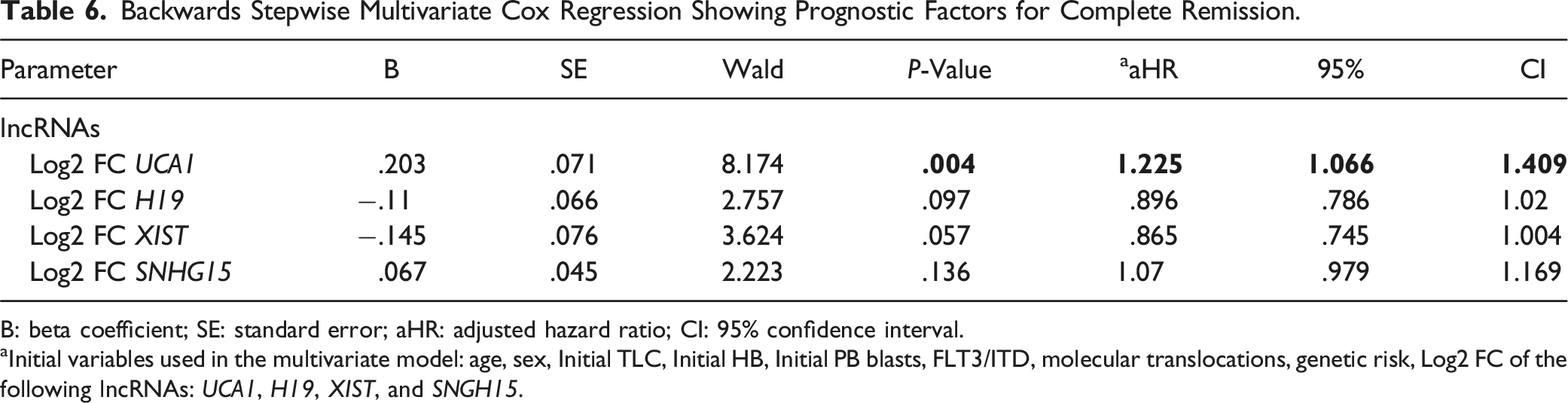
B: beta coefficient; SE: standard error; aHR: adjusted hazard ratio; CI: 95% confidence interval.
aInitial variables used in the multivariate model: age, sex, Initial TLC, Initial HB, Initial PB blasts, FLT3/ITD, molecular translocations, genetic risk, Log2 FC of the following lncRNAs: UCA1, H19, XIST, and SNGH15.
Relative Expression of lncRNAs in Patients Who Developed Relapse
Relative Expression of lncRNAs in Patients Who Had Relapse Occurred and Those Who Did Not.
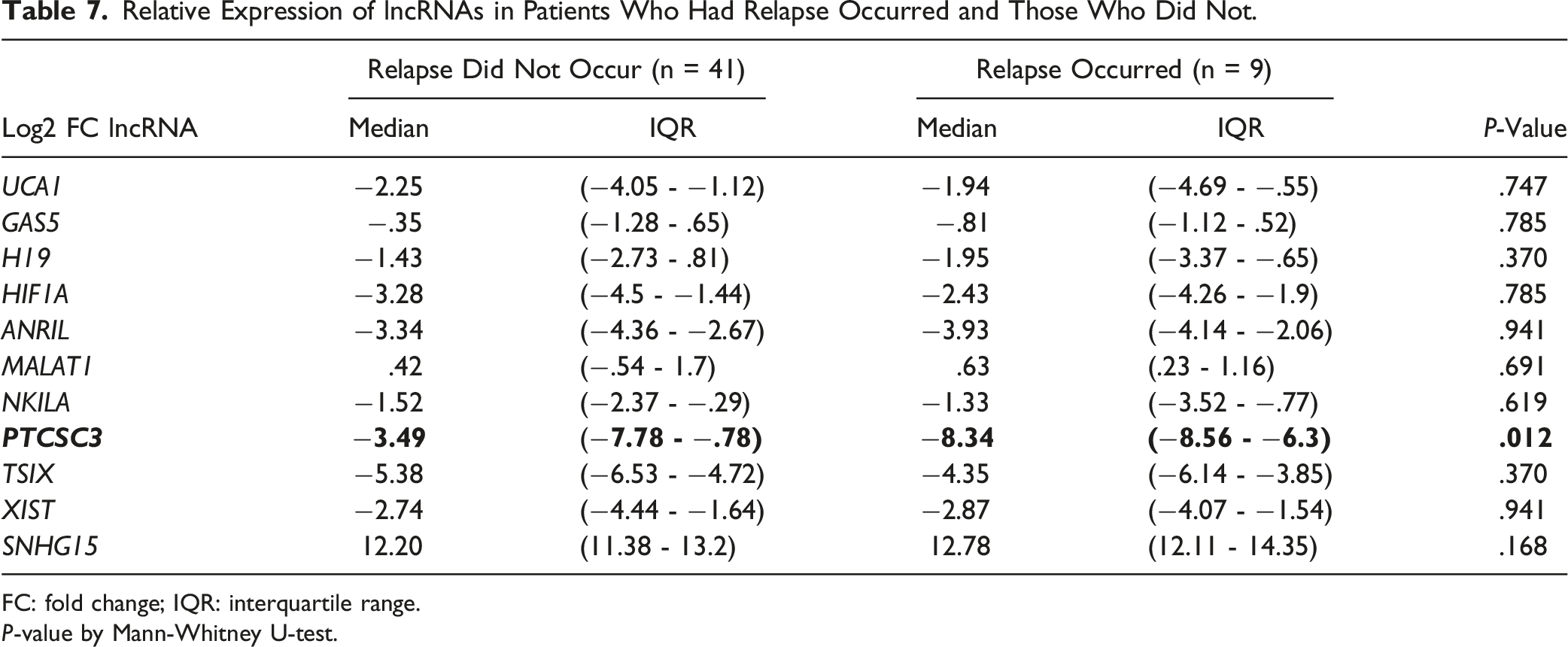
FC: fold change; IQR: interquartile range.
P-value by Mann-Whitney U-test.
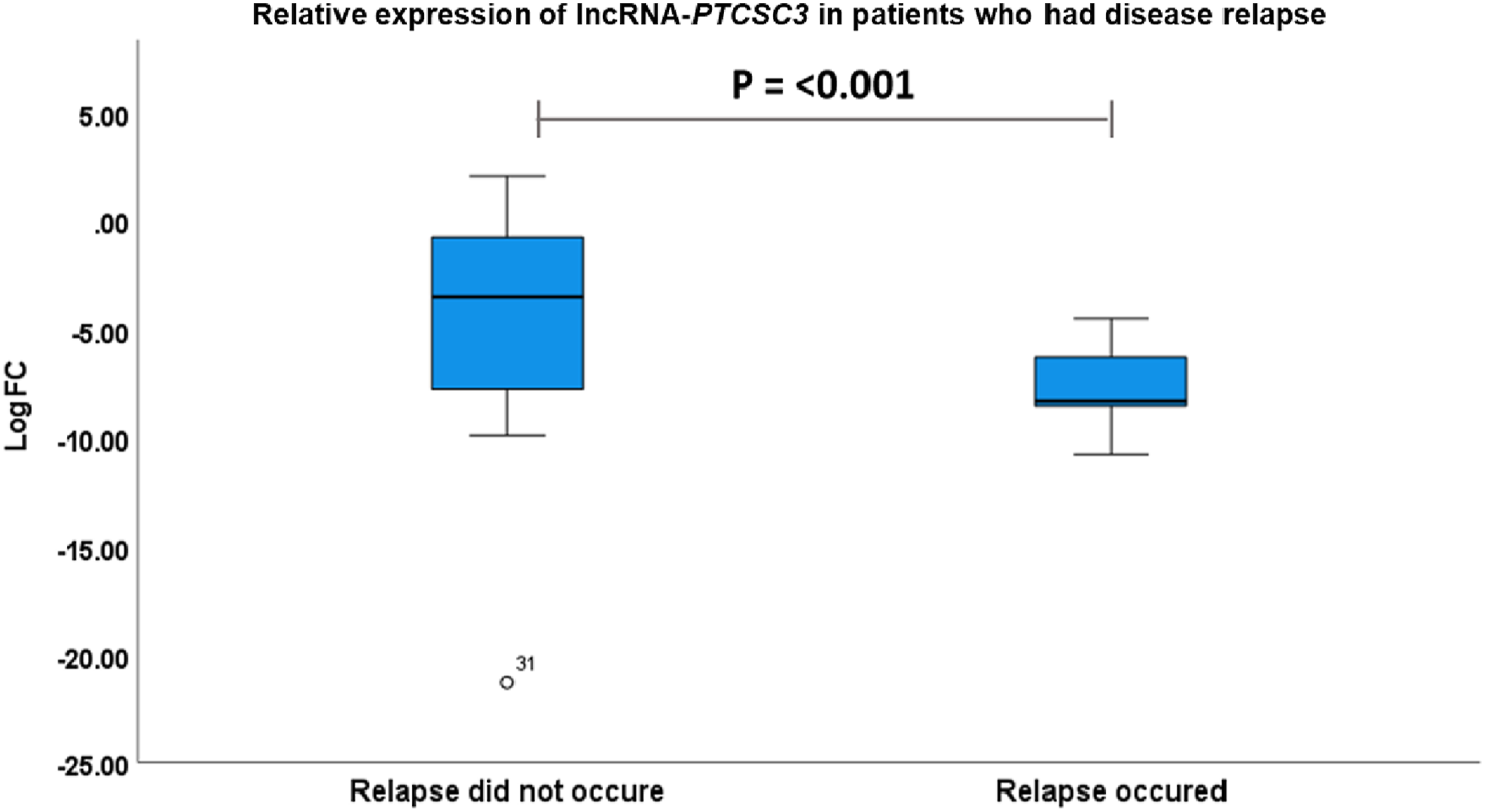
Relative expression of lncRNA-PTCSC3 in patients who had disease relapse. P-value by Mann-Whitney u-test.
Discussion
The differential expression of long non-coding RNAs (lncRNAs) in acute myeloid leukemia (AML) patients compared to control individuals has been a topic of interest in recent years. The dysregulation of lncRNAs has been implicated in the pathogenesis of various cancers, including AML. 31 In AML, lncRNAs have been shown to regulate gene expression and play a role in disease progression. 32
The current study investigated the relative expression of eleven lncRNAs in AML patients compared to controls. Our results identified a significant differential expression difference between AML and normal control samples in five of the eleven studied lncRNA expressions (lncRNA-TSIX, XIST, SNHG15, HIFAL, and ANRIL). Interestingly, all these lncRNAs were downregulated except lncRNA-SNHG15. LncRNA-SNHG15 has been reported to have a role in cell proliferation, differentiation, and apoptosis. Its upregulation in AML may contribute to leukemogenesis by promoting cell proliferation and inhibiting apoptosis. 33 Its dysregulation may contribute to tumorigenesis or suppression depending on the type of cancer. 27 Our findings provide further insight into the dysregulation of lncRNAs in AML and highlight the potential role of lncRNA-SNHG15 in the pathogenesis of the disease.
LncRNA-PTCSC3 expression was previously found to be significantly downregulated in both gastric and papillary thyroid cancers compared to the surrounding paired paracancerous tissue.34,35 In the present study, this was not the case, since the reduced LncRNA-PTCSC3 differential expression in AML was statistically not significant when compared to normal control. However the expression level of lncRNA-PTCSC3 was significantly downregulated in patients who later developed relapse when compared to those who did not. lncRNA-PTCSC3 has been reported to act as a tumor suppressor by inhibiting cell proliferation via induction of apoptosis and autophagy, inducing apoptosis, and suppressing invasion and metastasis in human oral cancer. 36 Moreover, previous studies have shown that LncRNA-PTCSC3 expression is downregulated in thyroid cancer tissues compared to normal tissues. 37 However, applying the Mann-Whitney U-test for differential expression of the eleven lncRNAs clarified that LncRNA-PTCSC3 in relapsed samples versus those who did not develop relapse stands out as a powerful predictor of relapse occurrence, whereby the relatively lower levels of its expression in an upfront BM samples carries out a significant risk of relapse development (P <0.012). To our knowledge, this is the first time to report such a finding. If this biomarker is validated, it could provide high implications for treatment decision-making or even in the development of targeted therapy. Down-regulating its expression could help in relapse prevention. Our findings are consistent with previous studies that have reported downregulation of LncRNA-PTCSC3 in oral and/or thyroid cancer tissues.36,37 It was reported by Wang et al., 2017 PTCSC3 inhibits the proliferation and invasion of glioma cells by suppressing the WNT/β-catenin signaling pathway. 38 WNT gene expression in the same 79 samples was done. WNT expression was downregulated in cases who developed relapse versus those who did not develop relapse (data not shown) which implies that a similar mechanism of downregulation is involved in AML similar to glioma. A key finding in this study is the relapse prediction by low levels of lncRNA-PTCSC3. If such a novel finding is reproducible, this might have enormous implications in AML treatment decision-making or even in the development of targeted therapy.
Consistent with previous reports overall survival was significantly influenced by FLT3/ITD of the mutant type, moderate or high genetic risk, and higher peripheral blood blasts.37-41 Here, lncRNA-XIST overexpression was the only lncRNA that affected OS favorably in both univariate and multivariate analysis. This is in contrast with previous findings that have reported that high expression of lncRNA-XIST is associated with poor prognosis in several other cancers, including glioma, breast cancer, and gastric cancer. 42 Our results are consistent with Zhou et al., 2022 who found that high lncRNA-XIST expression is associated with a better prognosis in AML patients. 43 The mechanism underlying the favorable prognostic effect of lncRNA-XIST in AML is not fully understood. However, one possible explanation is that lncRNA-XIST may act as a tumor suppressor by inhibiting cell proliferation and promoting cell apoptosis. 43 Also, Multivariate Cox regression analysis including both clinical parameters and measured lncRNA expression showed that FLT3/ITD of the mutant type, genetic risk, and Log2 FC lncRNA-GAS5 adversely affect overall survival.
Complete remission is a critical milestone in AML treatment, and patients who achieve complete remission have significantly better outcomes than those who do not. LncRNA-XIST expression however has no significant influence on complete remission induction, while the low expression of both lncRNA-UCA1 and -HIF1A carried a slightly favorable prognosis for achieving complete remission. lncRNA-UCA1 is a long non-coding RNA that has been shown to be involved in the regulation of cell proliferation, invasion, and apoptosis in several cancers, including AML. 7 In support of our study in a cell line-based model study higher expression of lncRNA-UCA1 is associated with resistance to the drug Daunorubicin in AML, via a mechanism involving the MAP kinase pathway response to chemotherapy. 44 Another study showed that lncRNA-UCA1 knockdown inhibited proliferation and induced apoptosis in AML cells. 7 They demonstrated that lncRNA-UCA1 regulated AML progression by miR-296-3p/Myc axis. However, the mechanisms underlying the association between lncRNA-UCA1 and -HIF1A expression and the achievement of complete remission in AML patients are not fully understood.
In the present study, lncRNA-MALAT-1 is one of the two upregulated lncRNAs examined, unfortunately, the upregulation is not statistically significant. In accordance with Do and Kim, 2018 who proved that lncRNA-MALAT-1 was upregulated in AML patients (using serum samples from AML patients and healthy volunteers) and cell lines (HL60 and THP1) 9 . In their study, lncRNA-MALAT-1 knockdown was found to inhibit the proliferation, migration, and invasion of AML cells and induce cell apoptosis. In the same study, however, lncRNA-MALAT-1 was not upregulated in the HS-5 cell line which can explain why our upregulation was not statistically significant, since AML is a very heterogeneous disease. However, increasing the sample size may bring it to a significant level.
The study does have some limitations: further validation in larger, independent cohorts is needed to establish the prognostic value of lncRNAs like PTCSC3. This brings in the issue of generalizability due to the small sample size of 79 AML patients. Also, since lncRNAs act through complex gene regulatory networks, future studies should investigate these interactions for enhanced therapeutic potential. Finally, stratification of patients based on hematopoietic lineage markers could clarify the relevance of lncRNA across AML subtypes, guiding more tailored therapeutic strategies.
Conclusion
CR induction in AML can be affected by lncRNAs (UCA1 and HIFAL. OS was affected adversely by lncRNA-GAS5. While it was positively impacted upon by LncRNA-XIST. The key novel finding of the present study is that lncRNA-PCTSC3 can predict relapse in AML. Taking into consideration the marked heterogeneity of AML, further studies are needed to verify and validate the clinical value of this marker to be applied in personalized medicine.
Supplemental Material
Supplemental Material - PTCSC3, XIST, GAS5, UCA1, and HIFAL: Five lncRNAs Emerging as Potential Prognostic Players in Egyptian Adult Acute Myeloid Leukemia (AML) Patients
Supplemental Material for PTCSC3, XIST, GAS5, UCA1, and HIFAL: Five lncRNAs Emerging as Potential Prognostic Players in Egyptian Adult Acute Myeloid Leukemia (AML) Patients by Asmaa A. El Leithy, Yasser Mabrouk Bakr, Naglaa M. Hassan, Khaled T. Dardeer, Magda Assem, and Abdel Hady A. Abdel Wahab in Journal of Cancer Control.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical Statement
Supplemental Material
Supplemental material for this article is available online.
Appendix
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
