Abstract
Purpose
This study enhances the efficiency of predicting complications in lung cancer patients receiving proton therapy by utilizing large language models (LLMs) and meta-analytical techniques for literature quality assessment.
Materials and Methods
We integrated systematic reviews with LLM evaluations, sourcing studies from Web of Science, PubMed, and Scopus, managed via EndNote X20. Inclusion and exclusion criteria ensured literature relevance. Techniques included meta-analysis, heterogeneity assessment using Cochran’s Q test and I2 statistics, and subgroup analyses for different complications. Quality and bias risk were assessed using the PROBAST tool and further analyzed with models such as ChatGPT-4, Llama2-13b, and Llama3-8b. Evaluation metrics included AUC, accuracy, precision, recall, F1 score, and time efficiency (WPM).
Results
The meta-analysis revealed an overall effect size of 0.78 for model predictions, with high heterogeneity observed (I2 = 72.88%, P < 0.001). Subgroup analysis for radiation-induced esophagitis and pneumonitis revealed predictive effect sizes of 0.79 and 0.77, respectively, with a heterogeneity index (I2) of 0%, indicating that there were no significant differences among the models in predicting these specific complications. A literature assessment using LLMs demonstrated that ChatGPT-4 achieved the highest accuracy at 90%, significantly outperforming the Llama3 and Llama2 models, which had accuracies ranging from 44% to 62%. Additionally, LLM evaluations were conducted 3229 times faster than manual assessments were, markedly enhancing both efficiency and accuracy. The risk assessment results identified nine studies as high risk, three as low risk, and one as unknown, confirming the robustness of the ChatGPT-4 across various evaluation metrics.
Conclusion
This study demonstrated that the integration of large language models with meta-analysis techniques can significantly increase the efficiency of literature evaluations and reduce the time required for assessments, confirming that there are no significant differences among models in predicting post proton therapy complications in lung cancer patients.
Plain Language Summary
Using Advanced AI to Improve Predictions of Treatment Side Effects in Lung Cancer: This research uses cutting-edge artificial intelligence (AI) techniques, including large language models like ChatGPT-4, to better predict potential side effects in lung cancer patients undergoing proton therapy. By analyzing extensive scientific literature quickly and accurately, this approach has proven to enhance the evaluation process, making it faster and more reliable in foreseeing complications from treatments.
Keywords
Introduction
This study is dedicated to developing and evaluating advanced predictive models to increase the accuracy of predicting complications after proton therapy for patients with lung cancer. Proton therapy is a sophisticated radiation therapy technique that focuses the radiation dose precisely on the tumor target, significantly reducing damage to surrounding healthy tissues. 1 It has been clinically proven to be highly effective for treating lung cancer patients. However, despite these advantages, posttreatment complications such as radiation esophagitis and radiation pneumonitis can still severely impact the quality of life of patients.2,3
In recent years, numerous studies have utilized genetic markers, pharmacological interventions, imaging diagnostics, and invasive techniques to improve the diagnostic accuracy of these complications.4-7 By accurately predicting such complications, it is possible to reduce the occurrence of adverse effects and enhance treatment outcomes, effectively decreasing mortality rates. This growing body of research underscores the need for innovative approaches that combine the precision of modern medical technologies with predictive modeling to better manage and mitigate the risks associated with proton therapy.
Owing to the relatively small sample size of lung cancer patients receiving proton therapy, statistical significance is often not apparent. Therefore, the use of meta-analysis methods to quantitatively assess the overall effects of all studies can increase the credibility of the research. With the increasing volume of medical literature, as demonstrated by a search using the keywords “machine learning” and “cancer” in the Web of Science database, which yielded 51,398 articles, there is a clear upward trend in research related to predictive models. This necessitates meticulous evaluation of each study’s results and bias risks to ensure its clinical applicability.
Assessing bias risk is crucial in the meta-analysis process, as biased results directly affect the credibility and applicability of the research. 8 At least two independent reviewers are required to minimize the influence of individual bias in bias risk assessment. 9 However, previous studies have shown that interrater reliability in bias risk assessment ranges from 48% to 90%.10,11 As the number of studies increases, the time required for assessment can range from several days to weeks . 9 To ensure the quality of the literature and accurate assessment of bias risk, we employed the prediction model risk of bias (PROBAST) Assessment tool to evaluate the bias risk and applicability of the predictive model studies. This tool covers 20 signaling questions that help researchers systematically identify and quantify biases in the literature, thereby improving the accuracy and reliability of predictive models.
In recent years, natural language processing (NLP) has become a powerful tool,12-14 with generative AI and large language models (LLMs) such as OpenAI’s ChatGPT and Meta AI’s Llama series taking center stage.15,16 These technologies have demonstrated remarkable capabilities in processing and generating natural language, garnering widespread attention. These LLMs can now perform a variety of complex NLP tasks, and increasing research is emerging on the application of ChatGPT in the health and medical fields. 17 However, these models face numerous challenges in specialized medical literature assessment, such as generalizability, accuracy in understanding specialized terminology, and objectivity and consistency in evaluating literature quality. 18 While NLP technology has been widely applied in many medical fields, such as in our previous research, to improve the efficiency and accuracy of literature assessment, 19 its application in predicting complications after proton therapy remains relatively limited. Current studies rely mostly on traditional statistical methods, which may not fully utilize the hidden potential information in complex datasets.
To address these challenges, we combined meta-analysis methods with LLMs to investigate the performance of different predictive models in terms of complications after proton therapy for lung cancer. We evaluated the effectiveness of various models through meta-analysis and heterogeneity analysis and conducted in-depth literature quality assessments via LLMs, comparing the consistency of bias risk assessment between humans and the ChatGPT-4 and Llama models.
Materials and methods
Research Framework
This study comprises two main components. The first component focuses on exploring and comparing predictive models for identifying complications following proton therapy in lung cancer patients. We conducted detailed analyses of various complications, such as radiation esophagitis and radiation pneumonitis, and compared different predictive models to determine the most effective models for specific complications. Subgroup analyses further help in identifying optimal model choices for each complication type. The second component uses LLMs to increase the accuracy and efficiency of literature quality assessment. We employ three models, ChatGPT-4, Llama2-13B, and Llama3-8B, to perform preliminary evaluations of literature quality. These assessments are further deepened with the use of tools such as PROBAST, aiming to refine the accuracy and reduce the time needed for literature assessments. The overall research framework is visually represented in Figure 1, which illustrates our comprehensive approach to integrating technological advancements with medical research to improve treatment planning and literature evaluation efficiency. This article was conducted in accordance with the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines,
20
and was registered with INPLASY. The registration number for this study is INPLASY202480103. Overall research flowchart.
Data Sources and Search Strategy
We utilized PubMed’s Medical Subject Headings (MeSH) as a basis for our search keywords and extended our searches across databases, including Web of Science, PubMed, and Scopus. The search cutoff date was April 1, 2024. Our strategy, which is structured around the PICOS framework, 21 aims to encompass a broad and relevant spectrum of research. The search terms included “lung cancer” AND “proton therapy” AND “machine learning” AND “complications” OR “radiation pneumonitis” OR “radiation esophagitis,” with the use of Boolean operators “AND” and “OR” to maximize the retrieval of pertinent literature (detailed keywords can be found in Supplemental Table S1). The collected data were managed and analyzed via EndNote X20 (Clarivate Analytics, X20, Philadelphia, USA) software.
Inclusion and Exclusion Criteria
In this study, we established strict inclusion and exclusion criteria to ensure the scientific validity and relevance of the literature. The inclusion criteria mandated that studies must detail machine learning models predicting complications of proton therapy for lung cancer, encompassing any patient undergoing such treatment. We considered all proton therapy techniques and comparative analyses with other treatment modalities, with the requirement that the studies report posttreatment complications and embrace various research designs. The exclusion criteria were meta-analyses, systematic reviews, studies lacking complete model construction or model outcome evaluation, studies with unclear methodologies, studies with incomplete data reporting, studies in languages other than English, and studies with inaccessible full texts.
Data Extraction
In this study, the data extraction phase was conducted after strictly adhering to the inclusion and exclusion criteria, with full-text evaluations carried out independently by two authors. Any disagreements between the two authors were resolved through consultation with a third author, ensuring fairness and consistency in the data extraction process.
The extracted data items included the first author’s name, which records the primary researcher responsible for the study to facilitate subsequent analysis and citation; the year of publication to identify the timeliness and relevance of the study; the country indicating the geographic location where the study was conducted; aiding in analyzing regional differences’ impact on the results; the type of complications, detailing various complications caused by lung cancer proton therapy such as radiation esophagitis and radiation pneumonitis; the radiation therapy method, noting the specific radiation therapy techniques used, including details of the proton therapy method; the sample size, extracting the number of patients involved in the study, a key factor in evaluating the statistical significance of the results; overfitting prevention methods, recording the techniques or methods used to prevent model overfitting, ensuring the generalizability of the model; feature selection methods, extracting the methods used to increase model predictive capability and reduce variables; and the type of models used, describing the types of machine learning models employed, such as support vector machines, decision trees, etc.
Through this data extraction process, the study ensures that the collected information is comprehensive and accurate. This process also helps standardize data extraction methods, increasing the transparency and reproducibility of the research.
Quality Assessment and Risk of Bias
We employed PROBAST to evaluate the quality of the studies and the risk of bias. 22 The assessment covers four main domains—participants, predictors, outcomes, and analysis—which together comprise 20 signaling questions to facilitate a structured judgment of risk of bias (ROB).
Each signaling question is answered with “Yes”, “ Probably Yes”, “ Probably No”, “No” or “No Information.” If any signaling question is answered with “No” or “ Probably No,” there may be a risk of bias. If all four domains are rated as having low risk, the overall risk of bias can be considered low. If at least one domain has a high risk of bias, the overall risk of bias is assumed to be high. If at least one domain has an unclear risk of bias and the other domains have low risk, the overall risk of bias is deemed unclear.
Through this structured assessment process, we can systematically identify and quantify potential biases in the literature, further enhancing the transparency and scientific rigor of the research. This approach not only helps improve research quality but also ensures the reliability and applicability of the study results.
Statistical Analysis
This study utilized the following methods for statistical analysis, all of which were performed via Python (version 3.11.5). The effect size was measured via the area under the curve (AUC), which was used to assess the predictive ability of the models. We employed the inverse-variance-weighted average method to calculate the weights for the combined effect size, 23 assigning weights to each study’s effect size on the basis of its variance, with smaller variances receiving larger weights.
To evaluate the heterogeneity among studies, we applied degrees of freedom (DF), Cochran’s Q test, and the I2 statistic. An I2 value greater than 50% indicates significant heterogeneity. When significant heterogeneity was present, we used a random effects model because, regardless of the degree of statistical heterogeneity, there are differences in treatment methods and types of complications among studies.
To identify the sources of heterogeneity, we conducted subgroup analyses by evaluating different complications to investigate potential sources of heterogeneity and compare the performance of different subgroups. The overall effect size was expressed with a 95% confidence interval (CI), 24 and a P value of 0.05 was used as the significance level. We also plotted forest plots to display the results, and Publication bias was evaluated using a funnel plot
Large Language Model Design
In this study, we utilized transformer-based LLMs to increase the efficiency and accuracy of risk assessment. 25 These models include OpenAI’s ChatGPT-4 and META’s Llama3-8b and Llama2-13b, which are known for their superior capabilities in understanding and generating language, making them well suited for learning language-related features from large volumes of text and applying them to various language processing tasks.
The design phase of the LLMs is described as follows: 1. Model Selection and Application: • The chosen models include ChatGPT-4, Llama3-8b, and Llama2-13b, which are all based on the transformer architecture and are capable of handling complex language understanding and generation tasks. • These models were used to analyze and answer questions regarding literature quality and risk assessment, specifically addressing the 20 questions in the PROBAST risk assessment. • To assess model performance, use AUC, accuracy, precision, recall, F1 score, and time efficiency as evaluation metrics, with accuracy being the primary evaluation metric. 2. Data Preprocessing and Model Fine-Tuning • Initially, the 20 questions in the PROBAST risk assessment were organized (detailed in Supplemental Table S2), providing structured inputs for model evaluation. • These questions were integrated into the prompts for the LLMs, and the models were fine-tuned according to specific assessment needs to enhance their adaptability and accuracy in evaluating professional literature. 3. Interactive Interface and Ensuring Independence: • ChatGPT and AnythingLLM were used as user interaction interfaces, allowing researchers to interact with the models through natural language, pose questions, and receive answers. • To ensure the independence and accuracy of the assessments, a new conversation window was restarted after each article evaluation to prevent the memory of the previous interaction from influencing subsequent assessments.
Through this design, we not only improved the efficiency of risk assessment but also ensured the reliability and accuracy of the evaluation results. Additionally, the high degree of automation in this system allows for the processing of large volumes of literature in a short time, significantly enhancing research efficiency.
Results
Literature Screening and Selection
A total of 1569 articles were identified across the search databases, including Web of Science, Scopus, and PubMed. After 791 duplicate articles were removed, 634 articles were further excluded on the basis of title and abstract screening. For the full-text evaluation, 81 articles were initially included. At this stage, articles that did not align with the research topic or lacked model prediction outcomes for complications were excluded, resulting in 11 articles being retained. Additionally, 2 new studies that met the inclusion criteria were identified from the references of these 11 articles. Ultimately, 13 articles met the inclusion criteria for the systematic review. The flowchart of the literature screening process is illustrated in Figure 2. Literature screening flowchart.
Characteristics of the included studies
Studies-Included Characteristics.

Abbreviation: PSPT, Passive-Scattering Proton Therapy; IMRT, Intensity-Modulated Radiation Therapy; PBT, Proton Beam Therapy; PBS, Pencil Beam Scanning; DS, Double Scattering; 3D-CRT, Three-Dimensional Conformal Radiation Therapy; VMAT, Volumetric Arc Therapy; PCIRT, Proton and Carbon Ion Radiation Therapy.
Studies-Included Methodological Characteristics.

Abbreviation: sLKB, standard Lyman–Kutcher–Burman; gLKB, generalized Lyman–Kutcher–Burman; MLR, Multivariable Logistic Regression; LR, Logistic Regression, SVM, Support Vector Machine; LASSO, Least Absolute Shrinkage and Selection Operator; NTCP, Normal Tissue Complication Probability; RF, Random Forest; K-NN, K-Nearest Neighbors; EN, Elastic Net; LDA, Linear Discriminant Analysis; QDA, Quadratic Discriminant Analysis; NR, No Report; GB, Gradient Boost.
Performance Analysis of Models Predicting Complications after Proton Therapy for Lung Cancer
According to the results depicted in Figure 3, both fixed effects and random effects models were applied to estimate the effect sizes in the analysis of the prediction of complications after proton therapy for patients with lung cancer. The fixed-effects model (Figure 3(a)) showed an effect size of 0.77 with a 95% confidence interval (CI) ranging from 0.75--0.78, suggesting a relatively stable predictive effect when considering the consistency across proton therapy methods and types of complications. Forest plot of the model used to predict complications after proton therapy for lung cancer (a) Fixed-effect model (b) Random-effect model. Abbreviation: sLKB, standard Lyman–Kutcher–Burman; gLKB, generalized Lyman–Kutcher–Burman; LR, Logistic Regression; SVM, support vector machine; LASSO, least absolute shrinkage and selection operator; RF, random forest; K-NN, K-Nearest Neighbors; GB, gradient boost; AUC, area under the ROC Curve; ROC, receiver operating characteristic.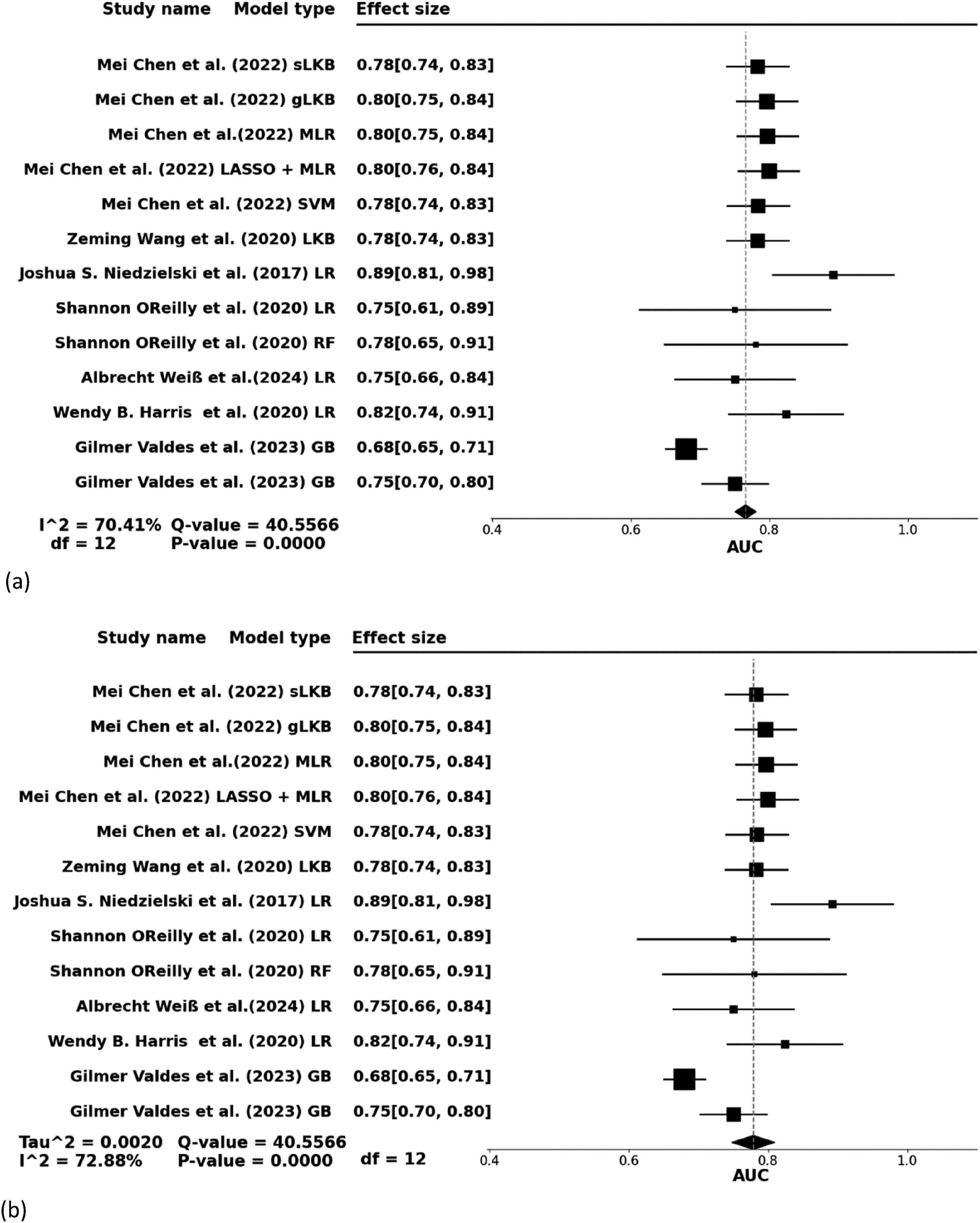
Nevertheless, despite the similarity in the types of treatments and complications studied, significant differences were observed in the study outcomes. The random-effects model (Figure 3(b)) demonstrated an effect size of 0.78 with considerable heterogeneity (I2 = 72.88%, P < 0.001). This indicates substantial variability among the studies, likely due to differences in study design, patient demographics, treatment methodologies, or methods of complication assessment (The funnel plot is included in Supplemental Figure S1).
Subgroup analyses were conducted to delve deeper into the sources of this heterogeneity, identifying specific conditions or variables that influence the predictive performance of the models. These analyses not only assess the practical application performance of the predictive models but also underscore the significant discrepancies that exist even under similar treatment and complication scenarios, highlighting potential areas for future research.
Subgroup Analysis of Radiation Esophagitis and Radiation Pneumonitis Patients
The subgroup analysis detailed in Figure 4(a)–(c) illustrates the stability and variability of the predictive models for radiation esophagitis and radiation pneumonitis. In the analysis of radiation esophagitis (Figure 4(a)), the model predicted an effect size of 0.79, with a confidence interval ranging from 0.78--0.81 and a heterogeneity index of I2 = 0%, P = 0.45. This finding indicates a high level of consistency among the studies in predicting radiation esophagitis, with no significant heterogeneity observed, suggesting that the models used can consistently reflect the risk of this complication after treatment. Forest plot of subgroup analysis (a) Analysis of radiation esophagitis (b) Analysis of radiation pneumonitis (c) Analysis of radiation pneumonitis after excluding heterogeneous studies.
In contrast, the initial analysis for radiation pneumonitis (Figure 4(b)) presented more challenges, with an effect size of 0.74, a confidence interval from 0.70 to 0.80, and significant heterogeneity (I2 = 68.65%, P = 0.0125), indicating considerable differences among the studies. After the exclusion of studies with high heterogeneity, the effect size for the prediction of radiation pneumonitis was adjusted to 0.77, with a confidence interval ranging from 0.73--0.80, and the heterogeneity was significantly reduced (I2 = 0%, P = 0.56). This adjustment demonstrated that addressing sources of heterogeneity significantly enhances the consistency and reliability of model predictions for radiation pneumonitis.
These analyses not only highlight the effectiveness of proton therapy predictive models for different complications but also emphasize the importance of further exploration of sources of heterogeneity to optimize predictive accuracy.
Comparison of LLMs and manual evaluations in quality and risk assessment
Figure 5(a)–(d) and Table 3 elucidate the disparities between LLMs such as ChatGPT-4, Llama2-13b, and Llama3-8b and traditional manual evaluation methods in terms of quality and risk of bias assessment. The data underscore notable differences in risk grading outcomes (high risk, low risk, unknown risk) across the various evaluation techniques. Notably, ChatGPT-4 achieved an impressive 90% accuracy rate in risk assessments, significantly outstripping the performance of Llama2-13b and Llama3-8b, which registered accuracies of 62% and 44%, respectively. This difference is also mirrored in the AUC, precision, recall, and F1 score metrics: ChatGPT-4 posted AUC is 0.88 and precision, recall, and F1 scores all at 0.86, markedly superior to those of Llama2-13b and Llama3-8b. Furthermore, while manual evaluations typically require 3423 seconds, LLM assessments are completed in approximately 1.06 seconds, as detailed in Supplemental Table S3. This substantial disparity highlights LLMs’ tremendous advantage in time efficiency, particularly demonstrated by the Words Per Minute (WPM) metric, which highlights LLMs' ability to swiftly process extensive data volumes and execute rapid assessments. Risk of bias in predictive models included in the study (a) manual assessment (b) ChatGPT-4 assessment (c) Llama2-13b assessment (d) Llama3-8b assessment. Note: High risk “-“; Low risk “ + “; Unclear or ambiguous ‘?”. Abbreviation: ChatGPT, chat generative pretrained transformer; Llama, large language model Meta AI. Performance and efficiency comparison of LLMs versus manual bias assessment. Abbreviation: b, billion; LLM, Large Langue Model; ChatGPT, Chat Generative Pretrained Transformer; Llama, Large Language Model Meta AI; AUC, Area Under the Roc Curve.
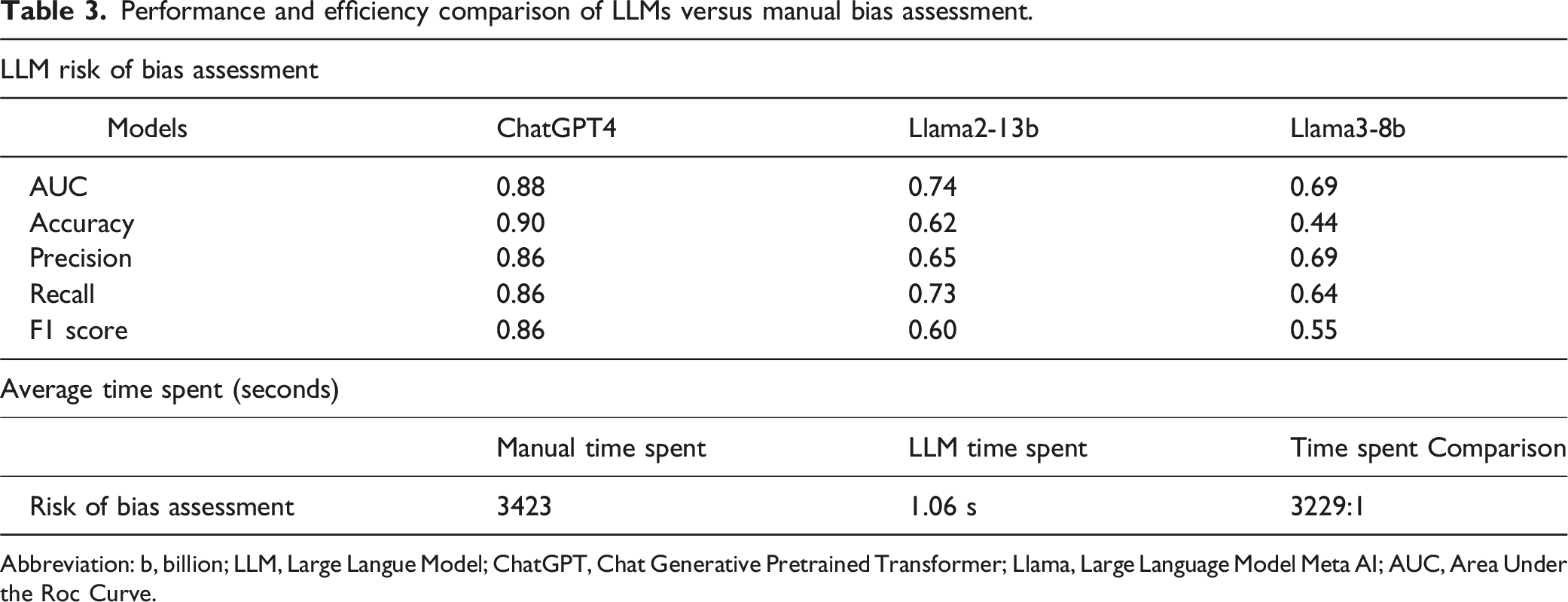
Independent evaluations via the PROBAST tool revealed that 9 studies were classified as high risk, 3 as low risk, and 1 as unknown risk. The results presented in Figure 5 indicate that LLMs more frequently marked studies as high risk and had a higher frequency of marking studies as unknown risk. This may reflect differences in the strictness of the evaluation standards and the sensitivity of the models.
These findings underscore the feasibility of using LLMs for risk assessment, demonstrating that they can provide high-accuracy evaluations and significantly improve efficiency. LLMs have been shown to be effective tools in the research quality assessment process, offering considerable advantages in both accuracy and speed.
Discussion
Results of the Meta-Analysis
Our study investigated the efficacy of predictive models for complications following proton therapy for lung cancer through meta-analysis combined with an evaluation of LLMs. The meta-analysis revealed that these models exhibit moderate to strong predictive capabilities, as evidenced by an overall effect size of 0.78. Despite this, a significant level of heterogeneity (I2 = 72.88%, P < 0.001) was noted, indicating considerable variability in the performance of these models across different studies. Subgroup analysis further suggested that no single model consistently outperformed others across all scenarios, underscoring the complexity of predicting complications in proton therapy.
Additionally, the use of LLMs, particularly ChatGPT-4, has significantly shortened the time required for risk assessment evaluations, drastically enhancing the efficiency of the literature review process. ChatGPT-4 demonstrated superior performance across all the evaluation metrics, indicating notable advancements in the application of artificial intelligence in medical research. This integration of LLMs not only streamlines the evaluation process but also highlights the potential for these technologies to improve the accuracy and speed of clinical research assessments.
When evaluating the complex dynamics of predictive models, particularly the efficacy of LLMs compared with manual evaluations, it is essential to consider a comprehensive set of evaluation metrics. Beyond just accuracy and time efficiency, our analysis now includes precision, recall, the F1 score, and the AUC. These metrics are critical for a thorough assessment of model performance in clinical environments. Precision is vital for minimizing the clinical risk associated with false positives, whereas recall is crucial for ensuring that all relevant cases are identified, preventing missed diagnoses. The F1 score, which balances precision and recall, is essential in clinical settings where the consequences of both false positives and false negatives are significant. Moreover, the AUC provides insights into the model’s ability to differentiate between outcomes across various thresholds, which is crucial for customizing clinical interventions on the basis of risk stratification. This expanded evaluation framework not only deepens our understanding of the models’ capabilities but also identifies areas for future improvement, enhancing their utility in clinical practice. By detailing these metrics, we aim to clarify how effectively these models can support clinical decision-making, guiding further enhancements in their development and application.
For the detailed analysis, we conducted subgroup analyses for radiation esophagitis and radiation pneumonitis. The results showed that the logistic regression model used in Joshua S. Niedzielski’s study 28 performed better in predicting radiation esophagitis than other models that relied on traditional clinical and dosimetric factors. The inclusion of new biomarkers, such as CT imaging, significantly improved the predictive accuracy. However, for radiation pneumonitis, Shannon O'Reilly’s study 30 revealed that even with the use of biomarker imaging, the results did not significantly differ from those of other studies. This could be due to insufficient model training and sample size. In Gilmer Valdes’s study, 33 the use of data imputation methods to handle missing values, although helpful in addressing data incompleteness, may have introduced biases that affected the accuracy of the model results.
From the PROBAST risk assessment, we found that most studies indicated high or unclear risk results, which could significantly affect the reliability of our study outcomes. The prevalence of high-risk biases suggests that the results might be unreliable, particularly since the majority of studies in our analysis did not employ methods to validate overfitting or were deficient in predictive factors. This scenario potentially leads to overly optimistic outcomes, aligning with other research findings.39-41 In fact, most studies reported only partial indicators of their models, which could contribute to the overestimation of performance metrics and excessive reliance on model predictions. The lack of comprehensive reporting underscores the need for more rigorous evaluation methods to ensure accuracy and reliability.
Moreover, to enhance the generalizability of models, it is crucial to validate them across different contexts and populations. 42 The ongoing development of PROBAST-AI, a more rigorous tool specifically designed for evaluating artificial intelligence and machine learning models, 43 represents a significant step toward addressing these issues, although it is still under development and not yet available.
Results of Large Language Model Analysis
In this study, we employed LLMs such as ChatGPT-4 to assess the quality of the literature, achieving an accuracy rate of up to 90%, which is significantly higher than the 44% to 62% accuracy rates of smaller models such as Llama2 and Llama3. This significant difference in performance is evident not only in accuracy but also in AUC, precision, recall, and F1 score: ChatGPT-4 achieves 0.86 in all three metrics and an AUC of 0.88, far outperforming Llama2-13b and Llama3-8b, which display lower metrics in these areas.44,45 These models demonstrated unique advantages in handling large datasets and extracting complex features, 46 particularly in terms of time efficiency, being 3229 times faster than manual methods in the literature.
ChatGPT-4’s superior performance is attributed to its larger model parameters, which enable it to more effectively understand and integrate complex contextual information within professional medical literature. 47 Conversely, smaller parameter models such as Llama2-13b and Llama3-8b, even after adjustments to their PROMPTs to enhance prediction accuracy, still have limitations. Their lower capacity for learning and fewer parameters render them less suitable for complex professional assessment tasks. These findings highlight the critical importance of selecting and applying appropriate LLMs for medical literature evaluation, especially in scenarios requiring high accuracy and efficiency.
Impact of Model Heterogeneity on Predicting Complications after Proton Therapy for Lung Cancer
Through detailed meta-analysis, our study revealed significant heterogeneity in the performance of different predictive models for complications following proton therapy for lung cancer. This finding highlights substantial differences in the effectiveness and applicability of models across various clinical and research settings. This heterogeneity primarily stems from variations in study design, sample size, treatment methods, and patient populations. For example, traditional statistical models such as logistic regression tend to perform better with smaller datasets or when there is a strong correlation between variables, whereas machine learning models, particularly deep learning models, may demonstrate greater advantages in handling large datasets and identifying complex, nonlinear relationships. 48
By employing subgroup or sensitivity analysis, researchers can better understand and interpret heterogeneity, thereby enhancing the reliability and applicability of model predictions. 49 These findings improve the assessment of therapeutic outcomes and the management of complications for patients undergoing proton therapy for lung cancer. Analyzing these heterogeneity factors in depth not only reveals the limitations of various models but also provides practical guidelines for selecting more effective models in specific clinical contexts. For clinicians, understanding the performance of different models in predicting complications for particular patient groups or conditions can guide more precise treatment decisions and the design of personalized treatment plans.
Bridging the Gap: From Theoretical Models to Clinical Implementation
While our study highlights the theoretical advantages of using LLMs for predicting complications from proton therapy, we recognize the importance of discussing the practical implications of these findings for clinical practice. The integration of LLMs, such as ChatGPT-4, has shown promising results in reducing the time required for literature assessment and increasing the accuracy of predictive models. However, the transition from theoretical models to clinical application involves several crucial steps.
First, the clinical implementation of these models necessitates rigorous validation and calibration processes to ensure that they are tailored to the specific needs and contexts of clinical environments. This includes extensive testing against existing standards of care to confirm that model predictions align with real-world outcomes.
Second, to facilitate the adoption of these models in clinical settings, it is essential to develop user-friendly interfaces that allow clinicians to interact with the models efficiently. These interfaces must be designed to integrate seamlessly into existing medical workflows, ensuring that they augment rather than disrupt clinical practice.
Moreover, there is a need for ongoing training and support for medical professionals to use these models effectively. This training should focus on understanding model outputs and integrating these insights into clinical decision-making processes.
Finally, ethical considerations, particularly concerning patient data privacy and the interpretability of model decisions, must be addressed to ensure that the use of LLMs in clinical settings upholds the highest standards of patient care and transparency.
By focusing on these aspects, we can bridge the gap between the theoretical improvements offered by LLMs and their practical implementation, ultimately enhancing patient outcomes and the efficiency of medical practices.
Limitations and Challenges for Future Research
This study revealed several limitations and challenges in evaluating the performance of predictive models for complications following proton therapy for lung cancer. First, differences in sample size, study design, or predictive models may affect the generalizability of the results, leading to high heterogeneity in our meta-analysis. Second, the subgroup analysis did not reveal significant differences in model superiority, indicating that no single model consistently performs best in all the scenarios. Additionally, our intervention measures were confined to proton therapy. However, many studies do not distinguish between patients treated with proton therapy and those treated with photon therapy, resulting in research data that may not fully meet the needs of this study. Finally, the issue of bias in data imputation remains a challenge, as imputed values may not fully reflect the real situation, 50 thereby affecting the accuracy of model predictions. Furthermore, the application effectiveness of smaller parameter models in specific domains is limited by their scale and complexity.
These limitations and challenges highlight the need for further improvements and refinements in research methodologies in future work. This includes selecting more suitable models, improving data collection and processing methods, and enhancing the generalizability of models. Future research should also consider a more granular approach to distinguish between different types of treatments and carefully address potential biases in data handling to ensure the reliability and accuracy of predictive models.
LLMs excel at generating coherent and contextually relevant responses, but their performance heavily depends on the quality, diversity, and representativeness of the training data. Accuracy remains a major concern, as these models can produce outdated or entirely fictitious responses, a phenomenon known as “hallucination”.51-53 The models may inadvertently learn and perpetuate biases and inaccuracies present in the data, leading to biased outputs and diagnostic discrepancies. 54 Additionally, these language models lack a true understanding of fundamental medical concepts, as their responses are based solely on statistical patterns, making it difficult for them to address more complex evaluation questions. 55 Relying solely on responses from LLMs in scenarios requiring deep background understanding can result in inaccurate or incomplete information.
Future research should focus on several areas of improvement. First, the sample size of the literature should be increased, and the study design should be optimized to enhance the representativeness and reliability of the results. Second, the performance of different predictive models should be further explored and compared, and integrated models that combine their strengths should be developed. Addressing the limitations of LLMs involves targeted improvements, including enhancing the quality and diversity of training data and fine-tuning the models to improve their understanding and response accuracy to specialized medical questions, thus reducing hallucinations and increasing reliability. Finally, interdisciplinary collaboration and the implementation of clinical trials should be promoted to ensure that research findings can be effectively translated into clinical applications. Through these concerted efforts, future research can more effectively address the prediction of complications following proton therapy for patients with lung cancer, thereby having a profound impact on treatment decisions and patient management.
Conclusion
This study highlights the transformative potential of integrating LLMs, such as ChatGPT-4, with meta-analytic techniques for predicting posttreatment complications in lung cancer patients receiving proton therapy. ChatGPT-4 demonstrated a high degree of accuracy, achieving an agreement rate of 90% with manual evaluations, and significantly accelerated the literature review process, which enhances the overall efficiency of the research.
The benefits to clinical practice are clear: these advanced predictive models can potentially refine treatment planning, contribute to personalized patient care, and reduce the incidence of adverse effects. However, despite these advances, this study identified key areas for further research. Given the high heterogeneity observed across studies, future efforts should aim to standardize research methodologies to reduce variability and improve outcome predictability.
Moreover, refining the ability of LLMs to accurately understand specialized medical terminologies and assess risk in an unbiased manner remains imperative. Future research should continue to focus on enhancing the training processes of these models, particularly in terms of their ability to analyze complex medical literature effectively. This approach will not only optimize the practical application of LLMs in clinical settings but also pave the way for their broader adoption in precision medicine and clinical decision support systems.
In summary, while this study confirms the feasibility and benefits of using LLMs in the medical field, it also lays the groundwork for substantial future advancements. The ongoing refinement and application of these models are essential for realizing their full potential in enhancing medical research and patient care.
Supplemental Material
Supplemental Material - Improving Prediction of Complications Post-Proton Therapy in Lung Cancer Using Large Language Models and Meta-Analysis
Supplemental Material for Improving Prediction of Complications Post-Proton Therapy in Lung Cancer Using Large Language Models and Meta-Analysis by Pei-Ju Chao, Chu-Ho Chang, Jyun-Jie Wu, Yen-Hsien Liu, Junping Shiau, Hsin-Hung Shih, Guang-Zhi Lin, Shen-Hao Lee, and Tsair-Fwu Lee in Cancer Control.
Footnotes
Acknowledgments
This study was partially supported by a grant from the National Science and Technology Council (NSTC) of the Executive Yuan, Republic of China (Grant No. 113-2221-E-992-011-MY2). A portion of these results has been submitted in abstract form to the ICMLSC 2025 Conference.
Author Contributions
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Science and Technology Council (NSTC) of the Executive Yuan of the Republic of China; 113-2221-E-992-011-MY2.
Ethical Statement
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
