Abstract
Introduction
Genetic diversity in the killer immunoglobulin-like receptor (KIR) gene composition and human leukocyte antigen (HLA) class I ligands, such as HLA-C, can affect the activity of natural killer cells and determine anti-cancer immunity. Specific KIR-HLA combinations can enhance cancer predisposition by promoting immune evasion. Studying the relationship between KIR-HLA polymorphisms and thyroid cancer (TC) risk can offer insights into how natural immunity fails, leading to disease development. Therefore, we investigated the association between KIR and HLA-C genotypes and TC risk in Saudi women.
Methods
In this retrospective study, sixteen KIR genotypes and 2 HLA-C allotypes were determined using the polymerase chain reaction-sequence-specific primer (PCR-SSP) method, and the genotypes of 50 Saudi female patients with TC were compared with those of 50 Saudi female healthy controls (HC).
Results
We observed a highly significant decrease in the presence of the KIR2DS2 and KIR2DS4 genes (OR = 0.15, 95% CI = 0.05-0.41, P = 0.0001; OR = 0.06, 95% CI = 0.02-0.2, P = 0.000, respectively) and in the presence of the KIR2DL5A gene (OR = 0.05, 95% CI = 0.02-0.14, P = 0.0000) in the TC group compared to the HC group. The frequency of the HLA-C2C2 allotype was significantly higher in HC compared to patients with TC (P = 0.02). The KIR haplotype group A and AB genotypes revealed a protective effect against TC (P = 0.0003 and P = 0.000, respectively), while the BB genotype showed a risk effect on TC compared to HC. Our results showed significant differences in the KIR gene combinations and KIR-HLA combinations between Saudi female TC patients and HC.
Conclusion
These results suggest that the expression of KIR genes and their HLA-C ligands may influence the risk of TC development in Saudi women.
Plain language summary
Genetic diversity in killer immunoglobulin-like receptors (KIR) gene composition and human leukocyte antigen class I (HLA) ligands such as HLA-C can impact the activity of natural killer cells (NK cells) and determine the results of cancer immunity. Specific KIR-HLA combinations can enhance vulnerability by promoting immune evasion. Studying the relationship between KIR-HLA polymorphisms and thyroid cancer (TC) risk can offer insights into how natural immunity failing leading to disease development. Therefore, we investigated the association between KIR and HLA-C genotypes and TC risk in Saudi women.
Keywords
Introduction
Thyroid cancer (TC) is the most common endocrine tumor worldwide, accounting for 2%-3% of all cancers. 1 However, the incidence of TC is increasing three–four times higher in women than in men, with a mean age of 40 years. 2 In Saudi Arabia, TC is the second most common malignancy, after breast cancer.3,4 To avoid immune monitoring, cancer cells reduce the surface expression of major histocompatibility complex (MHC) molecules, which prevents the detection of cytolytic T lymphocytes (CTLs). In contrast, decreased expression of MHC in tumor cells leads to the activation of natural killer (NK) cell cytotoxicity. 5 NK cells are a type of innate immune cell that develops from lymphoid precursors in the bone marrow. Their function is to eliminate target cells, such as virally infected or tumor cells, regulated by several membrane receptors, including killer immunoglobulin-like receptors (KIRs).5,6
KIRs are cell surface molecules expressed on NK cells. The KIR genes have a variety of genotypic and allelic forms, making them the most polymorphic human family of NK cell receptors.5,7,8 Sixteen KIR genes and pseudogenes have been identified and are clustered on chromosome 19q13.4 within the leukocyte receptor complex. The diversity of the KIR gene complex impacts the presence or absence of individual KIR genes, thereby creating haplotypes. 9 7 of these 16 KIR genes encode inhibitory KIRs (2DL1, 2DL2, 2DL3, 2DL5A/B, 3DL1, 3DL2, and 3DL3), and 7 encode activating KIRs (2DS1, 2DS2, 2DS3, 2DS4, 2DS5, and 3DS1), while the KIR2DL4 receptor has both activation and inhibitory signals. There are also 2 pseudogenes (2DP1 and 3DP1). 9
The KIR family includes members that recognize and interact with human leukocyte antigen class I (HLA). Different KIR-HLA gene combinations, as well as different levels of HLA expression in tumor cells, are associated with variable tumor prognoses and treatment.10,11
The diversity of KIR genes and their HLA ligands play a crucial role in shaping immune responses. 12 These genetic variations impact the immune effector functions of KIR-expressing cells and contribute to their susceptibility to diseases, including cancers. 13
Consequently, an increasing number of genetic studies have demonstrated correlations between particular combinations of KIR receptors with polymorphic MHC-I and the course of numerous diseases across various human cultures.14–22
The relationship between KIR and their HLA ligand genes has been investigated in various types of cancers, including breast cancer, non-small cell lung cancer (NSCLC), uterine cervical disease, and skin cancers. These studies emphasize the importance of understanding the interactions between KIR genes and HLA ligands in the modulation of immune responses against cancer cells.14,23–25
However, KIR gene polymorphisms and their HLA ligands in Saudi patients with TC remain unknown. Hence, we analyzed the frequencies of KIR genes, KIR haplotypes, and Bx subsets to determine their association with TC. In this study, we report the results of our investigation on the association between the frequency of KIR genes and their interactions with HLA-C ligands and the occurrence of TC in Saudi women compared with healthy female controls.
Materials and Methods
Subjects
For this retrospective study, biological samples were obtained from unrelated 50 female patients diagnosed with thyroid cancer (TC) and from randomly selected, unrelated, and 50 healthy unrelated female individuals who were randomly selected. All participants provided informed consent for the use of their samples in accordance with the approval guidelines set by the Medical Ethics Committee at King Khaled Hospital University (KKHU) in Riyadh, Saudi Arabia. The study protocol was reviewed and approved by the Medical Ethics Committee at KKHU (approval number: E−22-7143, dated November 28, 2022) and followed the ethical guidelines of King Saud University. Samples were obtained from formalin-fixed paraffin-embedded (FFPE) tissue blocks TC is diagnosed. Fresh blood samples were collected from healthy Saudi women at King Khalid Hospital, Saudi Arabia, between 2017 and 2020. The mean age was 43 years (range, 14-68 years) in the control group and 32 years (range, 21-45 years) in the TC group.
Genomic DNA Extraction and KIR and HLA-C Typing
Genomic DNA was extracted from FFPE tissue using FFPE (Qiagen, Valencia, CA, USA) according to the manufacturer’s instructions. For control samples, genomic DNA was isolated from peripheral blood using a DNeasy Blood & Tissue Kit (Qiagen, Valencia, CA, USA) according to the manufacturer’s instructions. A KIR polymerase chain reaction (PCR) sequence-specific primer commercial typing kit (Miltenyi Biotec, Inc, Cologne, Germany) was used for KIR genotyping. In accordance with manufacturer’s instructions. A panel of locus-specific oligonucleotide primers included in this kit enabled the amplification of 17 KIR genes (15 genes and two pseudogenes). 2DL5A, B Genotyping of KIRs, HLA-C1, and HLA-C2 was performed using a polymerase chain reaction sequence-specific primer (PCR-SSP) with the same primers as those reported in Tajik et al. 26 For each reaction, PCR was performed as previously described. 27 All PCRs reactions were performed using a Techne thermocycler (Techne TC-Plus Satellite). To determine whether gene-specific amplicons were present or absent, the PCR results were analyzed by electrophoresis on 2% agarose gels stained with ethidium bromide and examined under a UV transilluminator using a gel documentation system (Bio-Rad Gel225 Doc XR+). This study conformed to remark guidelines. 27
Statistical Analysis
Direct counting was used to determine the frequency of each KIR, HLA-C ligand, and KIR-KIR ligand combination in the patient and control groups. Data were analyzed using SigmaPlot version 11. The differences in the distribution of each KIR gene, HLA-C genotype, and KIR/HLA-C combination between the 2 groups were analyzed using a two-tailed Fisher’s exact test. Statistical significance was defined as P < 0.05. The magnitude of the effect was estimated using odds ratios (ORs) and 95% confidence intervals.
Results
Distribution of KIR Genes in TC Patients and HC
Distribution of the KIR Genes in Thyroid Cancer (TC) and Healthy Controls (HC).
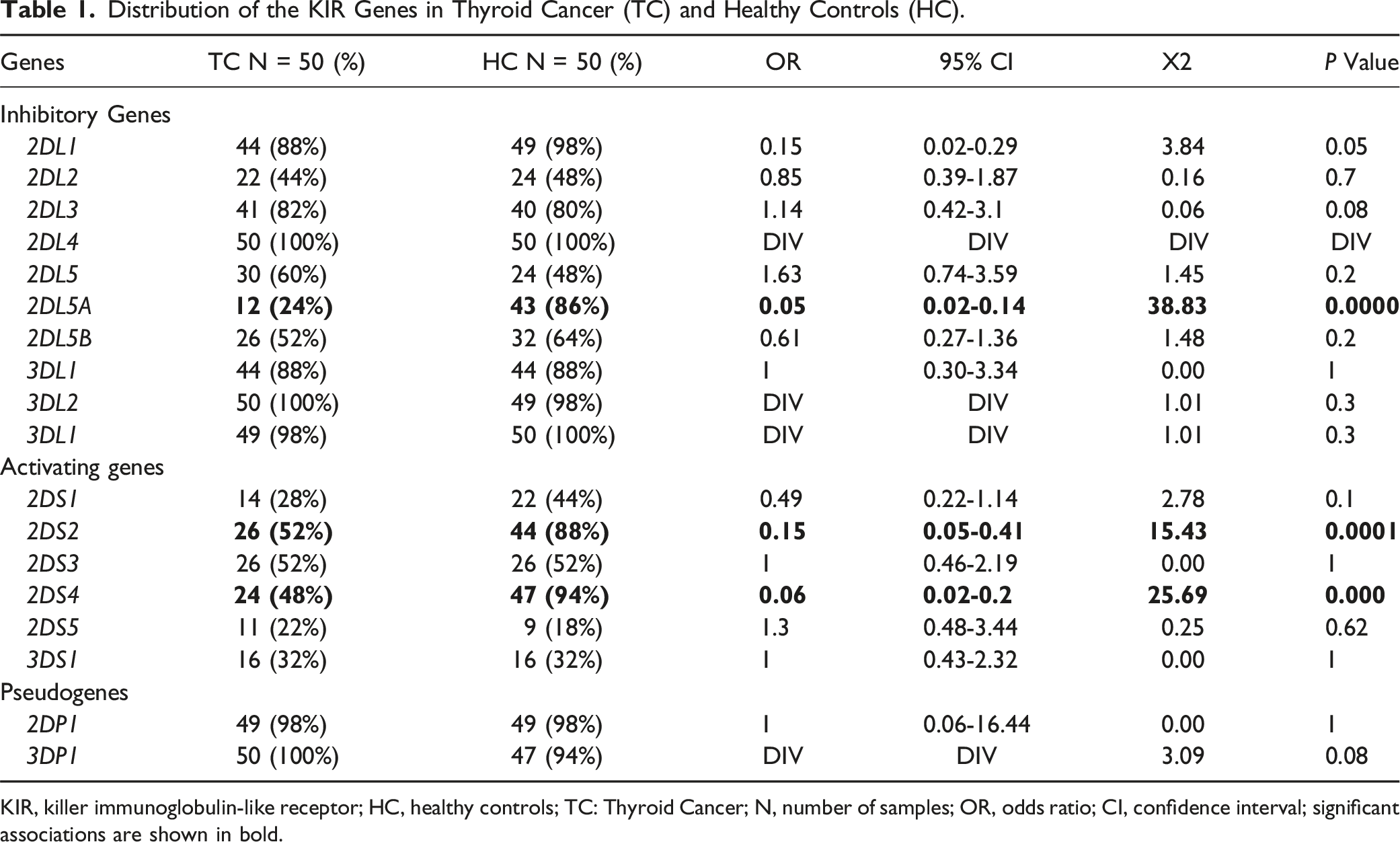
KIR, killer immunoglobulin-like receptor; HC, healthy controls; TC: Thyroid Cancer; N, number of samples; OR, odds ratio; CI, confidence interval; significant associations are shown in bold.
KIR Genotypes in TC Patients and HC
The KIR gene content of a given individual is called ‘‘KIR genotype.” We identified 49 distinct KIR genotypes in 100 study participants from Saudi female (Figure 1). TC patients exhibited only 24 of 49 genotypes, whereas 21 genotypes were found in HC. Four KIR genotypes were common between the 2 groups (genotype ID (GID) 1, 5, 36, and 113) Figure 1. Identification and distribution of KIR genotypes in TC patients compared to HC. KIR: killer immunoglobulin-like receptor; TC: Thyroid Cancer; HC: healthy control; No: number of genotypes; AA: A haplotype; BA and BB: B haplotypes; Shaded box: presence of the gene; White box: absence of the gene; New: New haplotype; C: centromeric genes; T: telomeric genes; 4: number of genes; (%F): percentage frequency.
Distribution of KIR Genotypes in TC Patients and Controls
Distribution of KIR Genotypes in Patients With TC Compared to HC.

Cen + C4+Cx: centromere; Tel + T4+Tx: telomere; 4: number of KIR genes; HC: healthy controls; TC: Thyroid Cancer; N: number samples; OR: odds ratio; CI: confidence interval. Significant associations are shown in bold.
Furthermore, we observed that the AB genotypes were significantly increased in HC (70%) compared to TC (28%), (OR = 0.17, 95% CI = 0.07-0.39; P = 0.000) this suggested that AB genotypes have protective effect against TC. On the other hand, BB genotypes were significantly decreased in HC (28%) compared to TC (64%), (OR = 4.6, 95% CI = 1.96-10.65; P = 0.0003), were found to be associated with the risk of TC and AA genotypes showed no significant statistical difference between TC and HC. In the following analysis, we investigated the 4 Bx linkage genotype subsets (C4T4, C4Tx, CxT4, and CxTx). There were no statistically significant differences between TC patients and healthy controls (HC). Additionally, analysis of centromeric and telomeric gene clusters did not show statistically significant differences between the TC and HC groups.
Distribution of HLA-C Genotypes in Patients with TC Compared to HC
Genotypic Association of HLA-C C1/C2 Alleles Polymorphisms With TC and HC.

HC, healthy control; TC: Thyroid Cancer; OR odds ratio; X2, Pearson’s chi-square test; 95%CI, confidence interval. Significant associations are shown in bold.
Frequencies of KIR/HLA-C Combinations in TC Patients and HC
Association of KIR Genes Combinations With TC and HC.

KIR, killer immunoglobulin-like receptor; HLA, human leukocyte antigen; HC, healthy controls; TC: Thyroid Cancer; N, number of samples; OR, odds ratio; CI, confidence interval. Significant associations are shown in bold.
Association of KIR-HLA Combinations With TC and HC.

KIR, killer immunoglobulin-like receptor; HLA, human leukocyte antigen; C+, allele C; C-, allele C absent; HC, healthy controls; TC: Thyroid Cancer; N, number of samples; OR, odds ratio; CI, confidence interval. Significant associations are shown in bold.
Combination investigations of specific KIR genes with HLA-C1/-C2 ligands revealed that KIR 2DL3+ HLA-C2+, KIR2DS2+ HLA-C1-, KIR2DS2+ HLA-C1C2+, KIR 2DL1+ HLA-C2+, KIR2DL1+ HLA-C2C2+, KIR2DS1+ HLA-C2+, KIR2DS1+ HLA-C1C2+, and KIR 2DS2+ HLA-C2C2+ ((OR = 0.44; 95% CI = 0.19-1; P = 0.045), (OR = 0.34; 95% CI = 0.13-0.88; P = 0.02), (OR = 0.35; 95% CI = 0.12-1; P = 0.045), (OR = 0.26; 95% CI = 0.12-0.61; P = 0.001), (OR = 0.29; 95% CI = 0.1-0.73; P = 0.008), (OR = 0.32; 95% CI = 0.12-0.9; P = 0.03), (OR = 0.22; 95% CI = 0.4-1.09; P = 0.045), (OR = 0.11; 95% CI = 0.3-0.42; P = 0.0002), respectively) were significantly increased in HC compared to TC patients, conferring protection against TC. On the other hand, KIR2DL2-/2DS2-/HLA-C1+ showed a higher frequency in TC patients compared with HC, suggesting an increased risk of TC occurrence (OR = 15.47; 95% CI = 1.93-124.3; P = 0.001) (Table 5). 28
Discussion
KIR genes and their ligands expressed on the surface of NK cells are associated with several disorders, including cancer, autoimmunity, and infections.29,30 To date, only 1 study on TC has demonstrated a role for KIR genes and HLA-C ligands. 5 In this study, we determined for the first time the relationship between the distributions of KIR genes and HLA-C ligands and the development of TC in a Saudi female population. The frequencies of 16 KIR genes were compared between 50 female Saudi patients with TC and 50 female HC.
Our study revealed that activating KIR2DS2 and inhibitory KIR2DL5A gene were negatively associated with TC and could be protective genes against TC. This result is compatible with our previous study 27 ; however, it is similar to our observation of a Chinese Han population, whose findings revealed a lower frequency of KIR2DS2 preeclampsia compared to healthy controls. 31 Furthermore, the decrease in Activating KIR2DS4 genes in TC compared to HC is compatible with the lower KIR2DS4 gene expression in UC compared to the HC Japanese population study. 32
Additionally, our findings showed that the frequency of HLA-C2 haplotype and HLA-C2C2 genotype were lower in patients with TC than in healthy controls, thus conferring protection against TC. This downregulation of the expression of these ligands could help tumor cells to immune escape mechanism by evading NK cell recognition and subsequent elimination. Our results are compatible with those of Goyette et al (2015), Marra et al (2015), and Shimoni et al (2017), who provided strong evidence of a decrease in the HLA-C2C2 genotype in patients compared with the control group.33–35
To evaluate the contribution of KIR gene and KIR_HLA combinations to TC risk, we analyzed the distribution of KIR genes and cognate HLA ligands in patients and HCs.
Combination analyses of selected KIR genes exhibited significant differences between TC patients and controls, indicating that individuals with the presence of the KIR2DS2 gene in the absence of the KIR2DS3/2DL5 genes or the KIR2DL2 gene were significantly less frequently detected in TC than in HC. However, the absence of KIR2DS2 in the absence of KIR2DS3 or KIR2DL2 occurred more frequently in TC subjects than in HC subjects. These results are in agreement with our previous study on the protective effects of KIR2DS2-/2DS3-, KIR2DS2-/2DS3-/2DL5A-, KIR2DL2-/2DS2+, and KIR2DL2-/2DS2- Combinations in BC compared to HC, while differing from KIR 2DS2+/2DS3-/2DL5-gene combinations revealed risks associated with BC and KIR2DS2+/2DS3-and KIR2DS2-/2DS3-/2DL5+; there were no significant differences between BC and HC. 27
Combination analysis of certain KIR genes with HLA-C1/-C2 ligands found that the KIR2DS2+ HLA-C1− combination was decreased in TC patients than in control The presence of KIR2DS2 gene in conjunction with the lack of HLA-C1 ligands may limit the recognition and cytotoxic activity of NK cells. Unlike our study, the study by Mirzazadeh et al, 2023 showed no statistical significance. 36
KIR 2DL1+ HLA-C2+ and KIR2DL1+ HLA-C2C2+ combinations were more frequent in the control group than in the TC group. Consistent with our results, Mirzazadeh et al, 2023 a higher frequency of KIR 2DL1+ HLA-C2+ combinations was observed in the control group than in the AML group. 36 Our results Shahsavar et al, (2010) indicate a higher frequency of KIR2DL1+/HLA-C2+ among patients with ALL. 26 The combination of KIR2DS1+ HLA-C2+ and KIR 2DS2+ HLA-C2C2+ was negatively associated with TC, but was not significantly different. 36 These different results could be explained by the ethnic differences between Saudi and Iranian populations.
In contrast, KIR2DL2-/2DS2-/HLA-C1+ Showed a strong protective effect against TC. These results are compatible with those of our previous study on the protective effects of KIR2DL1+/HLA-C2+, KIR2DL1+/HLA-C2C2+, KIR2DS2+HLA-C1-, and KIR2DL2-/KIR2DS2-/HLA-C1+ in BC and HC. while Our results differed from those of KIR2DS2+HLA-C1C2+, KIR2DS1+/HLA-C2+, and KIR2DS1+/HLA-C1C2 gene combinations, which showed no significant differences between BC and HC. 27 This could be because thyroid and breast cancers have different immune mechanisms linked to the development of cancer. A limitation of this case-control study is its small sample size. To the best of our knowledge, this is the first study to document an association between KIR polymorphisms and thyroid cancer in Saudi women.
Conclusions
In conclusion, the decrease in the activation of KIR2DS2 and KIR2DS4 and the combination of Activating KIR genes with their ligand expression in thyroid cancer may contribute to tumor immune evasion and potentially influence tumor survival. However, our understanding of how these factors influence NK cell function remains limited. Further studies on KIR genes with specific ligands and their functional activities should include a larger number of subjects and more clinical data combinations.
Footnotes
Acknowledgments
The authors would like to thank the Researchers Supporting Project Number (RSP2024R35), King Saud University, Riyadh, Saudi Arabia.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the King Saud university under Grant Number RSP2024R35.
Ethics Statement
The study was approved by the Medical Ethics Committee at King Khaled Hospital University (KKHU) in Riyadh, Saudi Arabia, and followed the King Saud University’s ethical principles (IRB-No. E−22-7143; date of approval: November 28, 2022).
Data Availability Statement
The datasets generated and/or analyzed in the current study are not publicly available due to ethical concerns but can be obtained from the corresponding author upon reasonable request.
