Abstract
Purpose
Identifying actionable oncogenic mutations have changed the therapeutic landscape in different types of tumors. This study investigated the utility of comprehensive genomic profiling (CGP), a hybrid capture-based next-generation sequencing (NGS) assay, in clinical practice in a developing country.
Methods
In this retrospective cohort study, CGP was performed on clinical samples from patients with different solid tumors recruited between December 2016 and November 2020, using hybrid capture-based genomic profiling, at the individual treating physicians’ request in the clinical care for therapy decisions. Kaplan–Meier survival curves were estimated to characterize the time-to-event variables.
Results
Patients median age was 61 years (range: 14–87 years), and 64.7% were female. The most common histological diagnosis was lung primary tumors, with 90 patients corresponding to 52.9% of the samples (95% CI 45.4-60.4%). Actionable mutations with FDA-approved medications for specific alterations correspondent to tumoral histology were identified in 58 cases (46.4%), whereas other alterations were detected in 47 different samples (37.6%). The median overall survival was 15.5 months (95% CI 11.7 months-NR). Patients who were subjected to genomic evaluation at diagnosis reached a median overall survival of 18.3 months (95% CI 14.9 months-NR) compared to 14.1 months (95% CI 11.1 months-NR) in patients who obtained genomic evaluation after tumor progression and during standard treatment (P = .7).
Conclusion
CGP of different types of tumors identifies clinically relevant genomic alterations that have benefited from targeted therapy and improve cancer care in a developing country to guide personalized treatment to beneficial outcomes of cancer patients.
Keywords
Introduction
The development of new genomic targeted therapies has revolutionized the treatment of cancer. Advancing technologies, including whole exome, genome, and RNA sequencing combined with bioinformatics, have contributed to the discovery and molecular alterations and their application as molecular therapeutic targets. 1
Currently, 40% of predictive and 63% of prognostic biomarkers performed globally are applied for cancer therapies and have an estimated clinical response of 11 to 18%.2,3 It is predicted that with the correct use of precision medicine, more than 5.5 million patients with cancer per year patients could benefit in terms of survival and quality of life. According to these statistics, precision medicine in oncology could significantly affect Latin-American countries. By 2018, 1,1412,732 cases of cancer were registered in Latin America and the Caribbean. Approximately 340,000 cancer patients could benefit from precision medicine in this region. Furthermore, 101,893 cancer cases were reported in Colombia; hence, 18,340 patients are potential precision oncology candidates within this country.4,5 Genetic alterations are considered actionable if they are susceptible to a Food and Drug Administration (FDA) approved therapy or experimental agents in clinical trials. 6 Targeted therapy against driver genomic alterations has improved outcomes for patients with many different cancers, including lung cancer, melanoma, breast cancer, among others. 7 Comprehensive genomic profiling (CGP) by next-generation sequencing (NGS) enables the identification of all classes of genomic aberrations. It is increasingly being utilized to select patients that could benefit from specific targeted therapies appropriately.8,9 The correct identification of patients with actionable genomic alteration has increasingly become essential for clinical care and improved outcomes.
The National Comprehensive Cancer Network (NCCN) guidelines recommend broad molecular profiling, including BRAF, ERBB2 (HER2), MET, RET, NTRK, and ROS1, in addition to EGFR and ALK for metastatic non-small cell lung cancer (NSCLC).10,11 Indeed, most patients (83%) with advanced breast and head and neck cancers harbor actionable genetic alterations identified by NGS. 12 Furthermore, different clinical trials have demonstrated that targeted therapy results in improved outcomes, including overall response rate and progression-free survival compared to traditional chemotherapy regimens.13,14
Several in-house and commercial testing panels are now available with rapid turnaround times.7,15–17NGS-based platforms are being utilized in the care of cancer patients since FDA approval of 2 assays in November 2017 for patients with advanced-stage. They are subject to national coverage by the Centers for Medicare and Medicaid Services (CMS).18,19 Interestingly, the use of CGP in clinical practice is not uniform; consequently, patient outcomes and cost-effectiveness of these technologies remain unclear.20,21 The most critical challenges to adopting and generalizing the use of CGP in developing countries are the lack of patient affordability and a long turnaround time. Furthermore, the lack of clinical guidelines and recommendations from local or regional clinical oncology organizations on which cancer indications will benefit, the insufficient education of the clinical community, the poor education of patients, and the absence of conclusive cost-benefit studies makes it difficult for policymakers to determine the need for funding support of NGS-based diagnostics. 21
To evaluate the utility and impact of the incorporation of CGP in routine health care in the setting of a developing country, we reviewed a cohort of 170 patients with different types of solid tumors undergoing sequencing by CGP. Our primary aim was to determine the potentially actionable genetic alterations and their subsequent impact on treatment selection.
Materials and Methods
Samples Collection and Clinical Data
This retrospective cohort study was approved by the Institutional Review Board. Informed consent was obtained from all enrolled patients (ONCOLGroup Platform - Registration No. 38792, Cayre, Bogotá, Colombia), approved on the 3rd of February 2021. This study corresponds to a sub analysis of the institutional registry of cancer patients subjected to comprehensive genomic testing and their corresponding clinical outcomes.
A total of 170 patients with a histologically confirmed diagnosis of malignancy in Colombia’s main cities from December 2016 to November 2020 were identified and included in the study. One hundred fifty-three patients had tissue availability, and 99 patients had a complete clinical follow-up. Age, gender, Eastern Cooperative Oncology Group (ECOG) performance status, TNM, and metastases were collected. First and second-line treatment, date of diagnosis, start/end of treatment, response, the rationale for NGS (diagnosis or resistance profiling), and NGS impact on treatment selection (access to on and off-label drugs or inclusion in clinical trials) was also collected. All tumor histologies, disease stages, subsequent or previous treatments, or performance status were included. All tests were provided free of cost to the patients. Patients with incomplete information were excluded.
Genomic Profiling
CGP was performed using an NGS-based (FoundationOne® CDx) clinical trial in a reference laboratory accredited by the College of American Pathologists (Foundation Medicine, Cambridge, MA, http://www.foundationmedicine.com), with validated methods.19,21 Samples were collected in a reference center (Foundation for Clinical and Applied Cancer Research (FICMAC) and were subjected to initial evaluation. Formalin-fixed paraffin-embedded samples were subjected to an assessment by a trained pathologist/oncologist, and tumoral samples were microdissected to guarantee malignant sample representation. In the case of tissue nonavailability, CGP was conducted as a liquid biopsy to evaluate cell-free DNA in whole blood (FoundationOne® Liquid CDx). Patients consented to the realization of CGP and notification of results to treating oncologists.
Statistical Analysis
For descriptive purposes, continuous variables were summarized according to central tendency and dispersion measures. 95% confidence intervals were reported for inferences. Time-to-event analysis was conducted with the Kaplan and Meier methodology with hypothesis testing using the log-rank test. All analyses were conducted using R version 4.02 (the R Foundation, Vienna, Austria).
Results
Patient Characteristics
A total of 170 tumor samples were collected during the time of the study. The median age for the whole cohort was 61, ranging from 14 to 87 years, with 64.7% female representation (95% CI 57.2-71.9%). A total of 17 samples could not be processed due to low specimen quality. With regards to testing choice, Foundation CDx ® accounted for 120 cases (70.6%), followed by Foundation Liquid ® in 43 (25.4%), and finally Foundation Heme ® in 7 (4%). On the other hand, 11 patients had no mutation detected from this panel.
The most common tumor evaluated was lung cancer (57.5%), followed by tumors of unknown primary (10%), tumors of the female reproductive system (6%), gastrointestinal tract tumors (5%), pancreas (6%), and Central Nervous System Tumors (4%). The most common histological diagnosis evaluated were primary tumors of the lung, with 90 patients corresponding to 52.9% of the samples (95% CI 45.4-60.4%). Figure 1 presents information on tumoral type among location and gender. Absolute number of sampled lesions depending on organ and sex of patients.
Representation of Histologies among Sampled Lesions of the Lung.

Comprehensive Genomic Profiles
Percentage of Actionable Mutations and Approved Treatments Based on Histology.

Abbreviations: LDT, lower digestive tract; CNS, central nervous system.
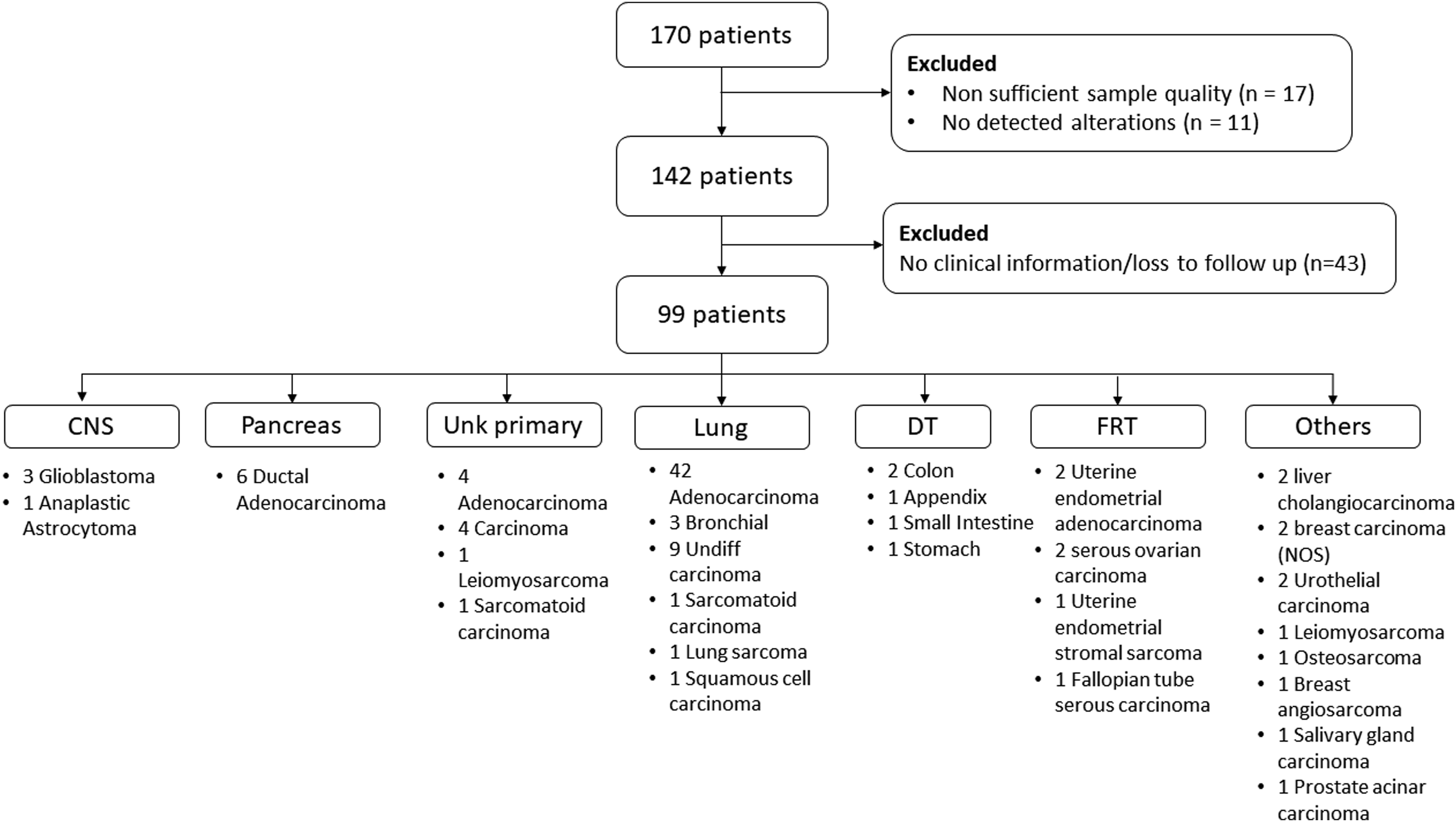
CONSORT flow of included patients, organs of origin, and histological diagnosis.
Median waiting times to genomic evaluation after diagnosis was 8.3 months (95% CI 4.93-14.27 months). Molecular analysis was offered in 2 settings, the first one at diagnosis (37 patients, 35.6%) and after standard disease progression (67 patients, 64.4%). The median time to the ordering of genomic testing for the first scenario was 2.7 months (95% CI 1.52- 6.25 months) compared to 14.73 months (95% CI 8.33-19.79 months) in the latter setting (P = .02). Ninety-five patients (95.9%) had an ECOG performance score <3, and of those, 89 had at least one site of metastatic involvement (89.9%). In this subset of individuals, 45 (45.4%) received genomic-driven treatment, 12 as first-line therapy (26.7%) and 32 (71.1%), 23 (71.9%), 8 (25%), and 2 (6.3%) as second, third-, and fourth-line therapy, respectively. The overall response rate was 36.3%. In the subgroup of patients with lung cancer, a significant relationship was found between the absence of tobacco exposure and the possibility of finding potentially modulable genomic alterations (P = .031).
Tumor Mutation Burden Analysis
With regards to molecular characteristics, different mutational profiles were identified depending on primary histology. Tumor mutational burden (TMB) based on histology as well as microsatellite status is presented in Figure 3. Across the different mutational loads, 2 cases of cholangiocarcinoma were reported with the lowest TMB close to zero. Lung cancer case, which also represented the most extensive sampled lesions, corresponding to the highest dispersion of TMB, ranging from 0 to 81 mut/Mb. Regarding tumors originating in the female reproductive tract, 2 cases represented the only ones with microsatellite instability in the whole, also carrying elevated TMB’s of 35 and 43 mut/Mb and corresponding to endometrial adenocarcinomas. TMB and microsatellite status according to the organ of origin. Points represent individual Tumoral mutational burden (TMB) measurements, whereas the red line depicts the mean (TMB) per individual organ origin. Abbreviations: CNS, central nervous system; IDT, inferior digestive tract; Unk Primary, primary of unknown origin; FRT, female reproductive tract.
Responses Based on Molecular Profiles
Response Types Based on Molecular Pathway Alterations Identified in Genomic Evaluation.

Alteration pathways are presented according to gene ontology. Abbreviations: PR, partial response; CR, complete response; CD, stable disease; PD, progressive disease; ORR, overall response rate (CR rate + PR rate).

Common and concurrent alterations among lung cancer patients.
Seven patients were documented to have a TMB >10, of whom 6 received some form of an immune checkpoint inhibitor (ICI) as monotherapy or combined with chemotherapy (4 pembrolizumab [66%] and 2 atezolizumab [33%]). Overall response rates for ICI-exposed patients in this setting reached 66%. Of the 2 patients who achieved stable disease, one harbored mutations in Rb1, loss of RBXW7, Notch 1, and TP53, and the other had alterations in RBM10, ARID1A, TSC1, ERBB2, KDM61, MCL1, and a TERT promoter mutation.
Survival Analysis
With regards to survival, the whole cohort reached a median overall survival of 15.5 months (95% CI 11.7 months-NR). With regards to patients who were subjected to genomic evaluation after diagnosis, it reached a median of 18.3 months (95% CI 14.9 months-NR) compared to 14.1 months (95% CI 11.1 months-NR) in patients after standard treatment progression (P = .7). Survival curves are presented in Figure 5. OS based on genomic evaluation timing.
Time to progression (TTP) in the subgroup of patients who received a medication based on a recommendation made by genomic results, with only three progression events in this group, a median value failed to be met. Figure 6 presents TTP according to Kaplan–Meier methodology. TTP of patients allocated to approved treatments based on genomic profile.
Discussion
Precision oncology has been revolutionized since the introduction of advanced sequencing technologies. It has already been studied that high-depth sequencing of cancer gene panels is susceptible to new genomic-driven therapies when they correctly identify patients with actionable mutations. However, the clinical utility of CGP in developing countries in Latin America has not been studied. Our study reports retrospective research to evaluate the impact of personalized genomic medicine by using advanced CGP platforms in Colombia and Latin America.
This study reports that 58 cases (46.4%) of patients harbored clinically actionable genomic alterations. Of this group, 45 patients, many with potentially actionable mutations, went on to receive genotype-directed therapy (45/58, 77.5%). Our study demonstrates that most solid tumors (90%) had adequate tissue for CGP. Further, of the 153 patients with sufficient tissue, 84% harbored actionable genomic alterations that could be prosecuted by therapies with clinical benefit in patient´s tumor type and FDA-approved, on-label drug (46.4%) or by treatments with clinical use in another tumor type (37.6%). This finding is similar to observations in prospective studies tumors where 75% of patients have a genomic alteration that was pharmacologically tractable with an approved drug. 23
Many of the somatic mutations identified in cancer patients are oncogenic pathway drivers of pathways that play a role in initiation or tumor progression. Consequently, tumor molecular profiling has become a key component in precision oncology practice as it enables the detection of genomic alterations that are therapeutic targets. 24 In 2017, Gutierrez et al conducted a large retrospective study with a cohort of 814 patients diagnosed with stage IIIB (11%) and IV (89%) NSCLC; in this study they found that only 59% of the patients got the EGFR and ALK biomarker testing according to the national guidelines. They also reported that only 8% of the patients underwent testing for all 8 NCCN recommended genomic alterations. The authors concluded that genomic testing involves a lot of challenges like sample handling, long turnaround times, test reimbursement confusion, limited access to target therapies, and insufficient tissue sample. 10
In contrast, large academic centers have reported successful implementation of CGP in several groups of patients; these studies also reported that many patients have at least one potentially actionable genomic alteration on CGP. However, they also reported that the implementation of CGP did not necessarily have an impact on therapeutic regimens. For example, the study conducted by Hilal et al in 2017 reported that 92% of patients from a cohort of 125 pediatric and adult patients had genomic alterations and were candidates of CGP. Only 15 patients (12%) received molecularly targeted therapy, with a small minority being benefited from it. They also reported that the main reasons for not receiving targeted therapy were stable disease, lack of access to clinical trials, poor performance status, and ongoing standard of care treatment. 25
Another example is the prospective trial by Hirshfield et al, they included 100 patients with rare and refractory cancers and evaluated the clinical actionability of CGP in those patients, as determined by recommendations by a molecular tumor board. Ninety-two patients underwent successful genetic sequencing, and 96% had 1 genetic alteration. In 31% of the patients, a change in treatment was led by CGP. Nevertheless, some subset cases were treated with cytotoxic chemotherapy due to a lack of driver mutations. They reported that the main barriers to access CGP were the clinical status of the patients and demanding access to appropriate clinical trials. 18
Another prospective study by Sholl et al assessed the feasibility of implementing CGP for all cancer patients; they reported results of 3727 patients who were successfully sequenced with their in-house gene panel. They also reported that only 73% of the patients had at least one clinically actionable genetic alteration; still, only 19% of those patients received standard of care therapeutic recommendations. However, this study did not report the impact on patient management. 7
A prospective, single-arm study by Wheler et al enrolled 500 patients with refractory cancers, and 339 patients underwent CGP. Of these patients, 93.5% had at least one potentially actionable molecular alteration. A matching score based on the number of drug matches and genomic alterations per patient was calculated, and 24.4% of patients received matched therapy, and 13.2% received unmatched treatment. Stable disease and partial or complete remission were associated with high matching scores compared to low scores (22% vs 9% respectively, P = .024). High matching scores were also associated with prolonged survival (HR .65; P = .05). 26
In 2018, Chawla et al reported that patients with matched therapy had a more extended treatment (1.5 months), a longer overall survival of 2.4 months, therefore, higher drug treatment costs (by $38K) (P < .01). They also reported that sixty-six percent of increased drug costs were attributable to more extended treatment rather than higher monthly drug costs. They also reported that patients who received matched therapies as an earlier line of therapy derived fewer treatment costs when compared to those who received them as the last line of treatment (4 and above). 8 Our study evaluated the overall response rate of 36.3% and the overall survival of 15.5 months among patients who underwent CGP. These findings highlight the role of CGP in routine clinical and its impact on improved clinical outcomes.
Recently, Mascarenhas presented the first report of the prevalence of driver mutations in Brazilian NSCLC patients using CGP. 27 A total of 513 CGP results were analyzed, 457 (89.1%) from tumors and 56 (10.9%) from plasma ctDNA. The median number of mutated genes by the sample was 3. The most common genomic alterations were single nucleotide variations (SNVs) (81.0% of the samples) followed by copy number variations (CNVs) (49.7% of the samples), frameshift mutations (31.4% of the samples), indels (19.3% of the samples), splice-site mutations (19.1% of the samples), and rearrangements/fusions (12.5% of the samples), considering that samples could have one or more mutations types. Globally, TP53 mutations were identified in 53.6% of tumor samples, KRAS mutations in 24.2%, EGFR activating mutations were detected in 22.5%, STK11 mutations in 11.6%, PIK3CA mutations in 8.8%, ALK rearrangements in 5.4%, BRAF mutations in 5.2%, ERBB2 alterations in 4.9%, Met alterations in 4.7%, Ret alterations in 2.4%, ROS1 rearrangements in 2.2% and NTRK rearrangements in .6%. In addition, the TMB analysis was available for 80.5% of samples tested; low TMB (<5 mutations/Mb), intermediate TMB (5-9 mutations/Mb), and high TMB (≥10 mutations/Mb) were described in 42.7%, 32.4%, and 5.5% of samples, respectively. 28 Unfortunately, this study did not describe the impact of CGP on the main outcomes, and the molecular characteristics of adenocarcinomas were compatible with findings from our study.
Additionally, among our cohort, we reported that 44% of patients had a low TMB, 22.6% had an intermediate TMB, and 2% had high TMB. Higher TMB has been associated with increased response to checkpoint inhibitors in solid tumors. 29 Our findings suggest that the evaluation of TMB is a tool that is useful to detect patients that will benefit from immunotherapy. These data are consistent with another prospective study that described 235 patients who underwent integrated NGS profiling and identified that the median TMB was low. 23
The analysis of the cost-effectiveness of CPG in patients with lung cancer and other selected pathologies is ongoing in Colombia; however, recently, Schluckebier et al evaluated the pharmacoeconomic profile of NGS in patients with lung cancer in Brazil, a country with enormous social and health heterogeneity, but which has similarities with our environment. 30 The study found that NGS added 24% extra positive cases and additional costs attributed to the molecular testing. The ICER (incremental cost-effectiveness ratio) comparing NGS with sequential tests was US$ 3479.11 per correct case detected. In addition, the use of NGS improved a slight gain in life years and QALYs (quality-adjusted life-years). 30
This study has several limitations. Retrospective samples can have limited quality, decreased tumor content, and variable intra-tumoral heterogeneity that can affect the results of the CGP. Additionally, in several cases, CGP results did not impact patient therapy regimens. Prior studies have elucidated CGP implementation issues into routine clinical care, including the lack of drug access and clinical trial availability. However, this study also has important strengths. To the best of our knowledge, our study is the first to evaluate the performance and impact of CGP among cancer patients in Colombia. These findings support the need to develop public health policies to implement genome sequencing techniques to improve precision medicine further and, hence, impact cancer-associated cost burden and improve patient outcomes. Appropriately and cost-effectively select patients that can benefit from targeted therapies. Additionally, this study demonstrates the urgent need for molecular tumor boards that can significantly contribute to the effective implementation of CGP and dramatically improve precision oncology in developing countries. All these considerations were included within the prioritization framework for using biomarkers in Colombia that was recently published. 28
Conclusion
In conclusion, CGP is a technology that dramatically improves the use of personalized therapies in developing countries. The high frequency of clinically relevant genomic alterations in this patient population can potentially result in personalized, targeted therapies for routine care and increase the participation of patients from these countries in ongoing clinical trials. Our data confirm the potential utility of CGP in clinical practices as a form to improve cancer care in a developing country.
Supplemental Material
Supplemental Material - Application of Comprehensive Genomic Profiling-Based Next-Generation Sequencing Assay to Improve Cancer Care in a Developing Country
Supplemental Material for Application of Comprehensive Genomic Profiling-Based Next-Generation Sequencing Assay to Improve Cancer Care in a Developing Country by Claudia Cifuentes, Milton Lombana, Henry Vargas, Paola Laguado, Alejandro Ruiz-Patiño, Leonardo Rojas, Uriel Navarro, Carlos Vargas, Luisa Ricaurte, Oscar Arrieta, Lucia Zatarain-Barron, Leandro Zapata, Guido González, Carlos Ortiz, Laura Bernal, Juan G. Restrepo, Lucia Viola, Fabio Grosso, Ricardo Zapata, William Mantilla, Hernán Carranza, Iván Bustillo, Néstor Llinas, Ricardo Duarte, July Rodríguez, Pilar Archila, Jenny Ávila, Maritza Bermúdez, Tatiana Gámez, Carolina Sotelo, Jorge Otero, Elkin Forero, Mauricio Lema, Catalina Limpias, Camila Ordóñez-Reyes, Sergio Mejía, Christian Rolfo, Rafael Rosell, Andrés F. Cardona; ONCOL Group and CLICaP in Cancer Control.
Footnotes
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Oscar Arrieta reports personal fees from Pfizer, grants and individual fees from Astra Zeneca, grants and individual fees from Boehringer-Ingelheim, personal fees from Lilly, individual fees from Merck, personal fees from Bristol-Myers Squibb, grants and personal fees from Roche, outside the submitted work.
Christian Rolfo is a current employee of the Center for Thoracic Oncology at Tisch Cancer Institute, Mount Sinai Health System and Icahn School of Medicine. He reports receiving supported research funding from Lung Cancer Research Foundation-Pfizer; receiving non-financial research support from Guardant Health; providing advisory services to ARCHER, Inivata, EMD Serono, BMS, Novartis, Boston Pharmaceuticals, Pfizer, Mirati, and Eisai; providing speaker services to MSD, AstraZeneca, Roche, and Guardant Health; and participating in the Safety Monitoring Board for EMD Serono.
Andrés F. Cardona disclose financial research support from Merck Sharp and Dohme, Boehringer Ingelheim, Roche, Bristol-Myers Squibb and The Foundation for Clinical and Applied Cancer Research – FICMAC. Additionally, he was linked and received honoraria as advisor, participate in speakers' bureau and gave expert testimony to Merck Sharp and Dohme, Boehringer Ingelheim, Roche, Bristol-Myers Squibb, Pfizer, Novartis, Celldex Therapeutics, Foundation Medicine, Eli Lilly and Foundation for Clinical and Applied Cancer Research – FICMAC.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was carried out thanks to the unrestricted support provided by the Foundation for Clinical and Applied Cancer Research – FICMAC (Bogotá, Colombia – Grant #072020), and by Foundation Medicine, Roche Diagnostics (Bogotá, Colombia).
Ethical Approval
This study was approved by the Institutional Review Board. Informed consent was obtained from all enrolled patients (ONCOLGroup Platform - Registration No. 38792, Cayre, Bogotá, Colombia), approved the 3rd of February 2021.
Informed Consent
Written informed consent was obtained from the patients for their anonymized information to be published in this article.
Human and Animal Rights Statement
All procedures in this study were conducted in accordance with the Institutional Review Board ONCOLGroup Platform - Registration No. 2018/21903, Cayre, Clínica del Country, Bogotá, Colombia, approved protocols.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
