Abstract
Patient safety event (PSE) reports, which document incidents that compromise patient safety, are fundamental for improving healthcare quality. Accurate classification of these reports is crucial for analyzing trends, guiding interventions, and supporting organizational learning. However, this process is labor-intensive due to the high volume and complex taxonomy of reports. Previous work has shown that machine learning (ML) can automate PSE report classification; however, its success depends on large manually-labeled datasets. This study leverages Active Learning (AL) strategies with human expertise to streamline PSE-report labeling. We utilize pool-based AL sampling to selectively query reports for human annotation, developing a robust dataset for training ML classifiers. Our experiments demonstrate that AL significantly outperforms random sampling in accuracy across various text representations, reducing the need for labeled samples by 24% to 69%. Based on these findings, we suggest that incorporating AL strategies into PSE-report labeling can effectively reduce manual workload while maintaining high classification accuracy.
Keywords
Introduction
In healthcare organizations, patient safety events (PSE), referring to incidents that compromise patient safety, such as medical errors, near misses, and adverse events pose a significant threat and can cause avoidable harm, injuries and even death (Wolf & Hughes, 2008). In 2000, the study “To err is human” estimated that clinical errors cause about 44,000 to 98,000 deaths annually in the USA, with later studies raising this estimate to 251,454 deaths, making medical errors the third-leading cause of death in the USA (Donaldson et al., 2000; Makary & Daniel, 2016). The global cost of medication errors alone is estimated at US$ 42 billion annually (World Health Organization, 2021). Following the Global Patient Action Plan by the WHO in 2020 (World Health Organization, 2021), healthcare organizations have widely adopted incident reporting systems to document PSE reports. PSE reporting systems enable healthcare personnel to voluntarily log incidents, including medical errors and unsafe conditions, in the form of structured (event type, harm level, date, and location) and unstructured data (textual event descriptions and outcomes; Albolino et al., 2010; Ong et al., 2010). These reports are then reviewed by hospital staff such as risk managers and patient safety analysts, playing a crucial role in enhancing healthcare quality by providing insights into factors leading to PSEs and facilitating the development of preventive measures (Herzer et al, 2012).
Accurate classification of PSE reports into corresponding event types is crucial for analyzing trends, guiding interventions, and enhancing organizational learning (Brubacher et al., 2011; Fong et al., 2015). However, the high volume of PSEs reported (Koike et al., 2022) and the complexity of the classification taxonomy, which require specialized knowledge, make the classification process labour-intensive, time-consuming, and expensive (Dimitrakakis et al., 2008; Liang et al., 2017). Several studies have demonstrated the efficacy of supervised Machine Learning (ML) models in automating PSE report classification (Chen et al., 2024; Evans et al., 2020; Wang et al., 2017). Improving the reliability of report classification can enhance safety. However, supervised ML models require high-quality and voluminous manually-labeled datasets for optimal performance. Manually labeling data is time-consuming, costly, and often impractical due to the large volume of data generated (Goudjil et al., 2018). Active Learning (AL) is a semi-supervised learning technique that involves selectively querying the most informative data points for manual labeling, thereby optimizing the use of human resources (Settles, 2009). Our study aims to utilize human labeling resources efficiently to develop a robust dataset for training ML models, seeking to balance labeling costs and classification accuracy. This study investigates the efficacy of AL strategies in improving the performance of ML models in event type classification of PSE reports. Thus, our goal is to enhance the PSE labeling process by integrating Artificial Intelligence with human expertise. This study uses only free-text incident descriptions, assessing four AL strategies and a random selection baseline across both static text representations and deep learning-based contextual text representations. We apply these strategies to Support Vector Machine (SVM) and Logistic Regression (LR) models, assessing classification accuracy improvements through iterative data sampling. Our goal is to determine if AL enhances performance with fewer labeled samples than random sampling, comparing improvements across both static and contextual text representations.
Methods
Dataset
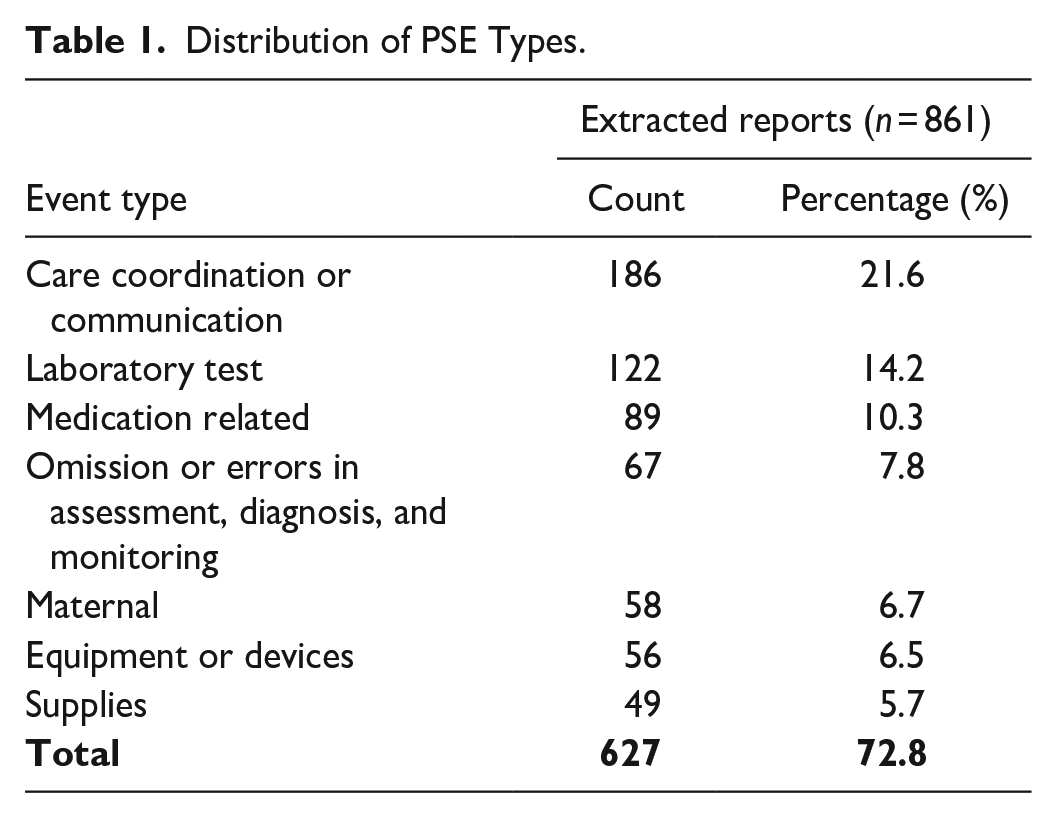
This study utilized 861 PSE reports from a Southeastern U.S. academic hospital’s maternal-care units (Jan 2019–Dec 2020). The study was approved by the hospital’s institutional review board (Pro00105892). Among the 25 distinct event types present in the dataset, the ML classifiers were exclusively trained on PSE-types with at least 40 samples—constituting about 73% of all reports—to ensure sufficient learning data (see Table 1). For event type classification, only the free-text incident descriptions were used as model inputs. To abide by the privacy regulations, all PSE reports were anonymized after data extraction.
Distribution of PSE Types.
Data Preprocessing
To prepare the incident descriptions for classifier input, we apply the following preprocessing steps.
Data Cleaning
PSE texts were anonymized and cleaned for classifier input. For static representations, relying on word occurrence and frequency, this involved lowercasing, stemming, removing stop-words and non-alphabetic characters. However, deep-learning representations, which represent sequences of words, used raw text to preserve context and avoid distortions.
Feature Extraction
We consider two types of static text representations namely binary bag-of-words (CB) and TF-IDF (Manning et al., 2008), and deep-learning contextual representations like RoBERTa (Liu et al., 2019), DeBERTa (P. He et al., 2020). Contextual representations are known to better capture the meaning of words by incorporating their position and context within sentences (Chen et al., 2024; Santiago Gonzalez-Carvajal, 2020; Subakti et al., 2022; Wang, 2020). Additionally, we utilize domain-specific representation, BioMedBERT (Gu et al., 2021) and GatorTron (Yang et al., 2022), trained on biomedical and clinical data, to better capture linguistic patterns.
Data Augmentation
To address the imbalance in our dataset across different event types (see Table 1), which can degrade classifier performance (Hasanin et al., 2019; H. He & Garcia, 2009), we employed the synthetic minority oversampling technique (SMOTE; Chawla et al, 2002). SMOTE was applied to the training dataset after feature extraction at each AL iteration before retraining the classifier, while the validation and test set remained unchanged.
Data Splitting
The dataset was divided into 70% training, 15% validation, and 15% testing sets, using a stratified split. The train set was used for classifier learning, while the validation set facilitated hyperparameter tuning and was used to select the best model across iterations. This approach prevented overfitting and enhanced the stability of AL experiments.
Experimental Setup
In this study, we particularly focus on simulating the pool-based AL sampling approach (Goudjil et al., 2018), by initially introducing a small portion of the labeled dataset to the ML model for training. Then, additional informative samples are selected iteratively by the AL strategy and the model is retrained from scratch to evaluate if the additional samples help improve classification accuracy (see Figure 1).
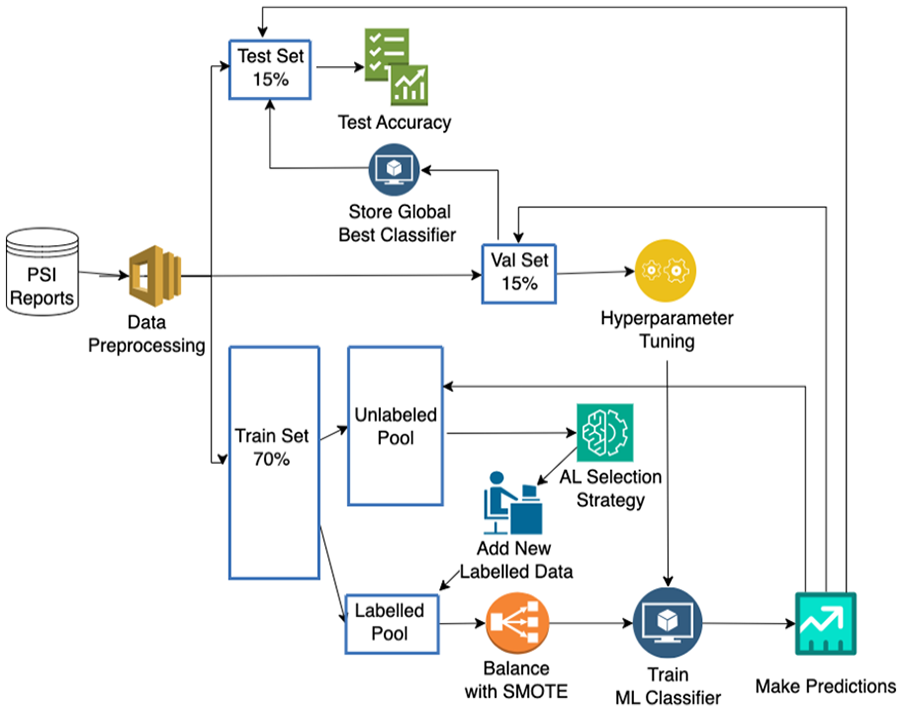
Active learning cycle.
AL Strategies
Several AL selection strategies have been proposed in literature, including uncertainty-based, Query By Committee (QBC), and Information Density (ID) based approaches (Settles & Craven, 2008). Our study explores three uncertainty-based methods: Least Confidence (LC), Margin (M), and Entropy (E) and a cosine similarity-based ID sampling combined with LC (IDCos × LC). Additionally, we implement a random selection (R) as a baseline to compare against AL strategies. This method randomly selects instances from the unlabeled set for labeling, in contrast to AL strategies that select instances in a more informed manner. We provide a brief explanation for each AL strategy: (1) LC queries the instance with the least confident prediction. (2) M measures uncertainty as the difference between the probabilities of the top two class predictions. (3) E measures uncertainty using the probability distribution of an event’s outcome, with high values indicating multiple equally likely outcomes (Settles, 2011). (4) IDCos × LC weighs the informativeness of an instance against its similarity to the entire dataset, using measures such as cosine similarity. It is used in conjunction with a base sampling strategy like LC, querying instances that are both informative and representative of the input space distribution (Settles, 2011).
All experiments start with a balanced set of 35 samples, with 5 samples per event type, drawn from the training set, allocating the rest to the unlabeled pool. In each AL iteration, 10 samples from the unlabeled pool are selected using the specified AL strategy and, consequently, these samples are added to the labeled set to simulate manual labeling of the selected samples. The ML model was then re-trained on this expanded set, after applying SMOTE oversampling to address potential class-imbalance. This process was repeated until all samples from the unlabeled set were exhausted. We tested the models after each AL iteration with an unseen test set, using them to predict the corresponding PSE type. These predictions were evaluated against the ground truth labels using classification accuracy as the metric. To mitigate bias due to initial selection of labeled samples, each experiment was repeated with 10 random balanced initial labeled sets, and the average test accuracy was reported for each iteration.
AL experiments were conducted using two popular models in text classification tasks: Logistic Regression (LR; Kleinbaum et al., 2002) and Support Vector Machines (SVM; Cristianini & Shawe-Taylor, 2000). Four AL strategies, along with a random sampling baseline, were evaluated across six text representations using both SVM and LR classifiers, totaling 60 experiments.
Results
The performance of ML classifiers, using various AL strategies, was evaluated over 41 iterations by analyzing average test set accuracy until all unlabeled samples were used (see Figures 2 and 3).
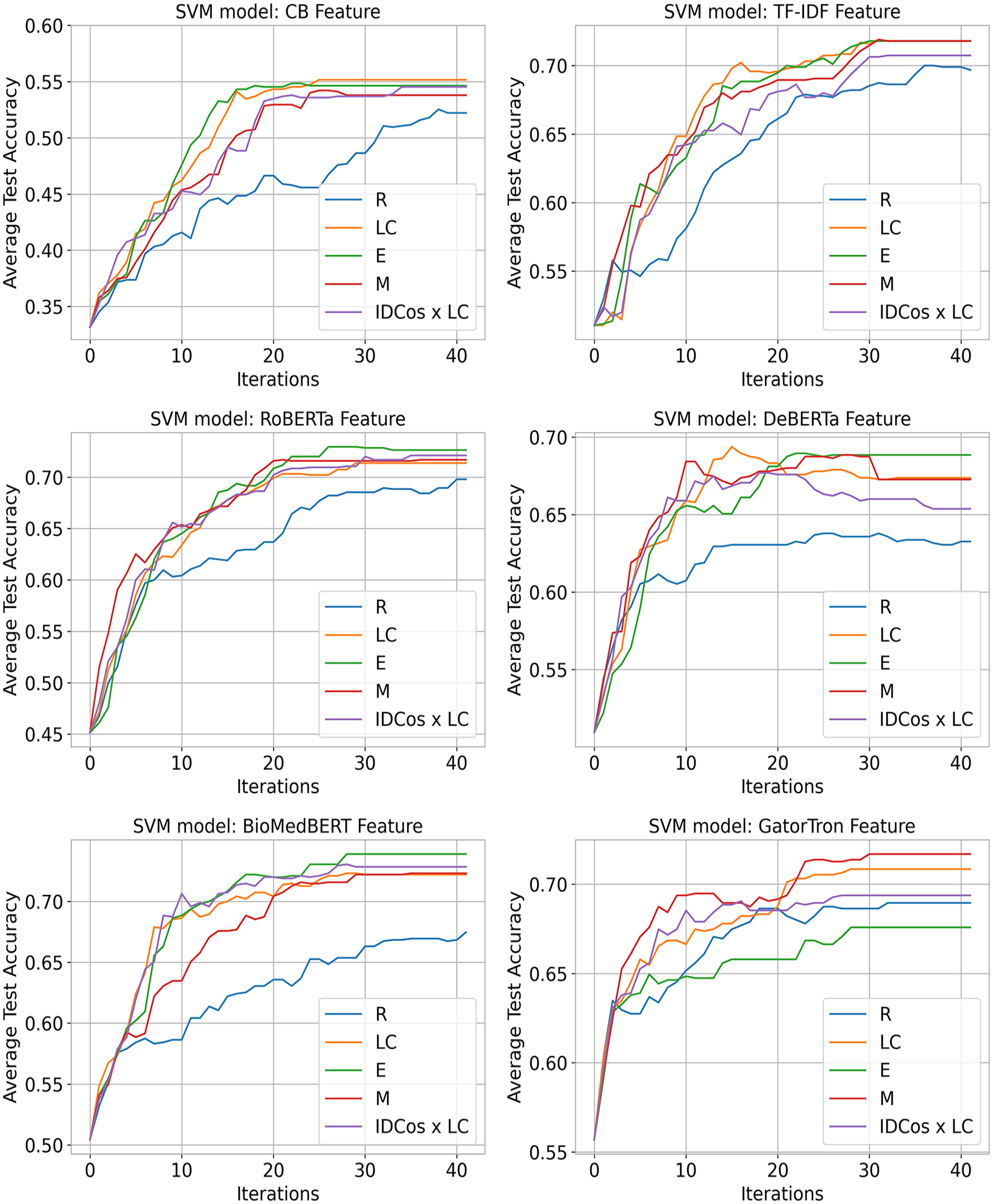
SVM average test accuracy improvements across text representations.
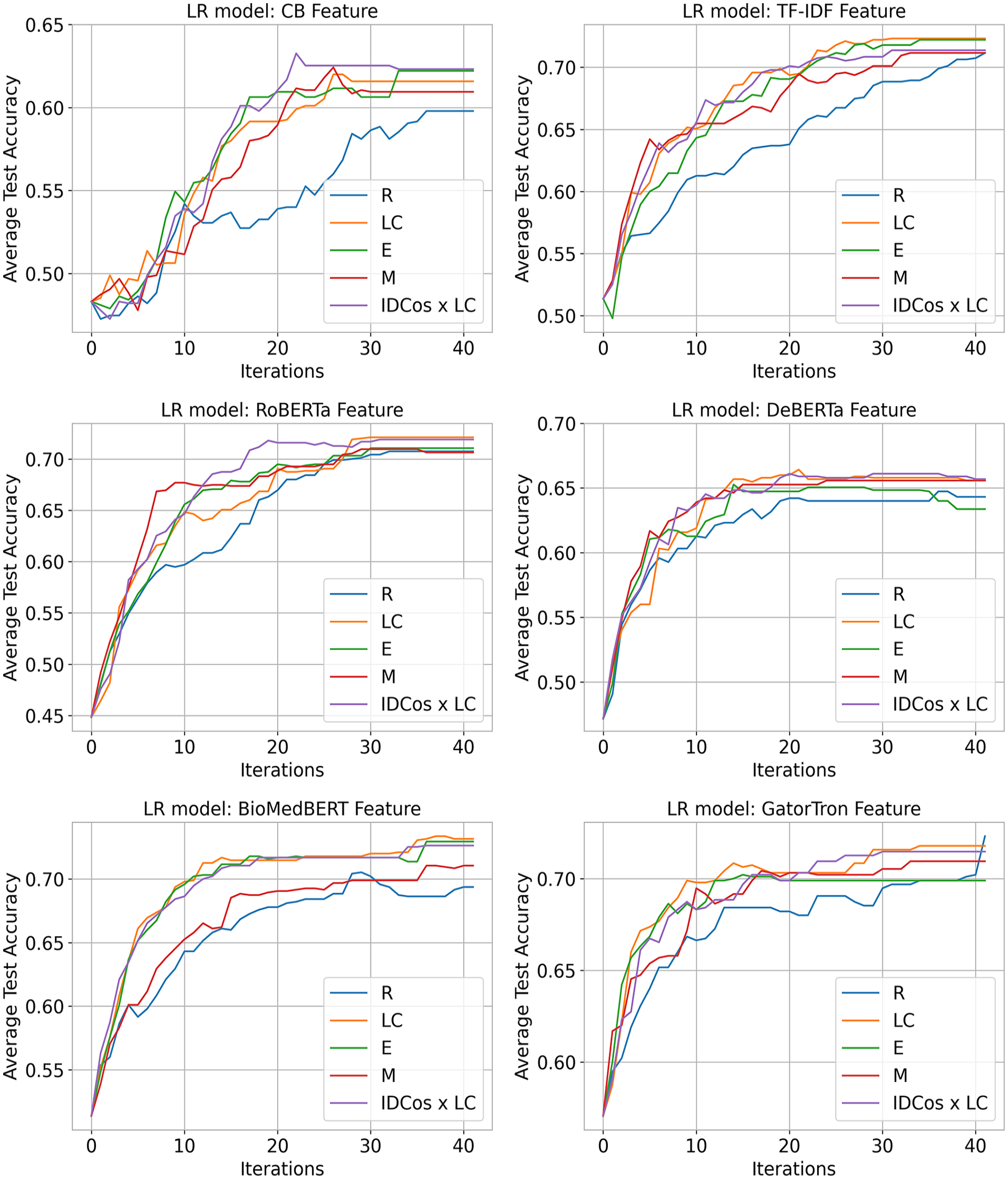
LR average test accuracy improvements across all text representations.
To compare the effectiveness of AL strategies against the random baseline R, we report the test set accuracy achieved at the 20th iteration, as well as average improvements over R at this iteration and between 10 and 30 iterations (see Table 2).
Comparison of Average Accuracy for SVM and LR Using Active Learning.
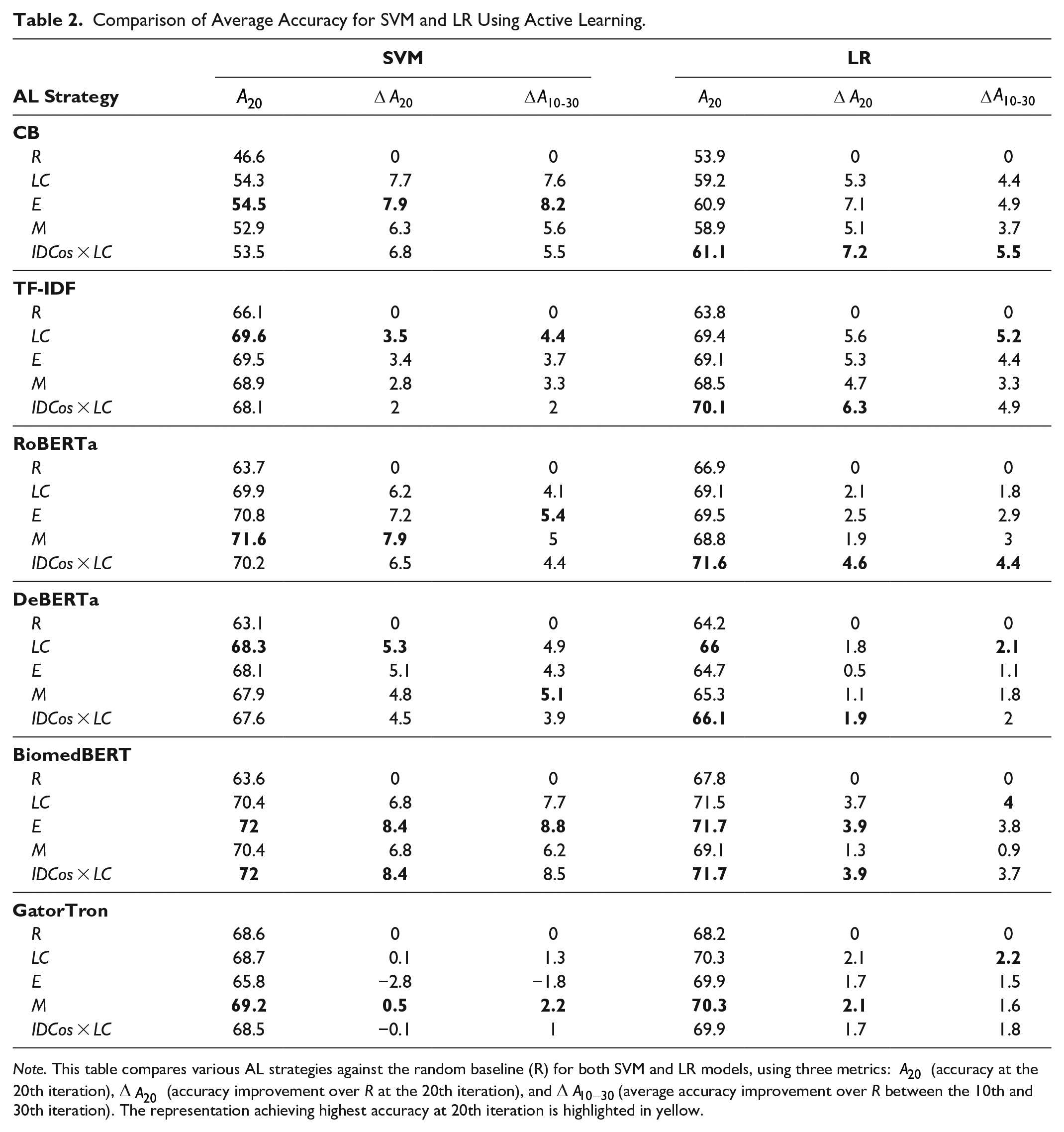
Note. This table compares various AL strategies against the random baseline (R) for both SVM and LR models, using three metrics: A20 (accuracy at the 20th iteration), ∆A20 (accuracy improvement over R at the 20th iteration), and ∆A10−30(average accuracy improvement over R between the 10th and 30th iteration). The representation achieving highest accuracy at 20th iteration is highlighted in yellow.
BioMedBERT features yielded the highest accuracy at 20 iterations, with SVM and LR achieving 72% and 71.7%, respectively, surpassing the R baseline by 8.4% and 3.9% using both E and IDCos × LC strategies. Following BioMedBERT, RoBERTa, TF-IDF, and GatorTron features reported high accuracies, ranging between 69.2% and 71.6% for SVM and 70.1% to 71.6% for LR classifier. The static TF-IDF representation performed comparably to the clinical domain-specific GatorTron embeddings, indicating that static representations can sometimes match the performance of deep learning models. At the 20th iteration, for SVM, LC strategy improves accuracy by 4.93% on average; while for LR, IDCos × LC strategy increases accuracy by 4.26% on average, over the R baseline across all features.
Static CB features reported the lowest accuracy at the 20th iteration, reaching 54.5% with SVM and 61.1% with LR, using E and IDCos × LC strategies, respectively. However, the AL strategies still significantly outperformed the R baseline, with improvements of 7.9% and 7.2% by 20 queries. This demonstrates the effectiveness of AL strategies across both static and deep-learning text representations, with highest accuracy gains seen with domain specific embeddings.
Discussion
Many hospitals across the world use incident reporting systems and the data collected on PSEs play a significant role in quality improvement efforts (Hasegawa & Fujita, 2018). Reliable classification of PSE types is crucial for patient safety, and while ML classifiers can accurately classify PSE-reports (Chen et al., 2024; Wang et al., 2017), they typically require large amount of labeled data to perform effectively. Our study investigated how AL techniques can minimize labeling efforts by utilizing a smaller subset of informative data points. We evaluated the effectiveness of four AL strategies in improving ML model performance for classifying event types in PSE reports. Additionally, we compared the impact of static versus contextual text representations and generic versus domain-specific embeddings.
AL Strategies Versus Random Sampling (R)
Our analysis revealed that most AL strategies significantly outperformed random selection (R) in terms of accuracy. AL strategies enhance labeling efficiency across all text representations, achieving near peak performance within 10 to 30 iterations (see Figures 2 and 3), reducing the required number of labeled samples by 24% to 69%, considering the total number of samples at the last iteration is 438. These strategies lead to highest accuracy gains particularly for domain-specific embeddings, notably BioMedBERT, improving SVM and LR accuracy to 72% and 71.7% respectively within 20 iterations, using only 54% of PSE-reports. Hence, integrating AL into PSE-report classification can streamline labeling, reduce workload for PSE analysts, and optimize both classifier accuracy and resource allocation. It will enable analysts to focus on annotating fewer incidents for model development, fostering human-AI collaboration.
Best AL Strategy
Among the AL strategies, Entropy (E) and the combination of Information Density (ID) Cosine with Least Confidence (IDCos × LC) were notably effective for SVM and LR, respectively. This demonstrates the efficacy of AL strategies, particularly when integrated with representativeness measures such as ID.
Static Versus Contextual Representations
For both ML models, static CB representations consistently underperformed other features in all experiments. However, static TF-IDF features sometimes matched or even surpassed the performance of contextual DeBERTa and GatorTron embeddings. Therefore, we cannot definitively state that contextual text representations always outperform static ones. Nonetheless, contextual features can represent text more efficiently in lower-dimensional spaces than TF-IDF enabling faster training, often with comparable or superior accuracy.
Generic Versus Domain-Specific Representations
BioMedBERT domain-specific contextual embedding slightly outperforms generic English RoBERTa embeddings by 20 iterations, which may be attributed to their ability to better capture medical terminology present in the text. With SVM, BioMedBERT achieves the largest overall improvement from the R baseline between 10 and 30 iterations, leading to a 7.8% increase in accuracy averaged across all AL strategies. However, with LR, static TF-IDF features yield the highest overall improvement from R, increasing accuracy by 4.5%. Thus, we conclude, while domain specific contextual embeddings generally lead to better performance compared to static text representations, they do not necessarily result in larger improvements using AL strategies across all models. Interestingly, when SVM is used with GatorTron features, the random baseline R performs relatively well, even outperforming the Entropy strategy and performing comparably with other AL strategies.
Regardless of the text features used, AL strategies prove to be effective tools in helping improve labeling efficiency over random sampling. By 20 iterations of AL sampling, we observe average improvements over the random baseline of 5.03% in accuracy for SVM, and 3.48% for LR, respectively, across all features.
Limitations
The ML models developed in our study were trained using PSE reports from maternal care units of one hospital in the United States, potentially limiting the generalizability of the results to different settings. Additionally, the scope of the analysis was limited by the small quantity of PSE reports available. Only the seven most frequent classes were used for this study. Thus, by not accounting for rarer event types, the performance of our models might be overestimated. We recommended that future research assess the efficacy of ML models integrated with AL strategies over more extensive and disparate datasets.
Conclusion
PSE labeling is particularly challenging as it requires specialized knowledge and human intervention, making full automation difficult. AL aims to support rather than replace manual labeling by filtering instances that are most likely to improve classification accuracy. AL also helps address the imbalanced-data problem typical of PSEs, by prioritizing rare or difficult-to-classify incidents, thus learning from a diverse incident set (Attenberg et al., 2013). PSE reports can change over time due to evolving medical practices and emerging safety issues. AL systems can adapt to these changes more effectively than static models, as they can incrementally learn and re-train from the newly labeled data (Perkonigg et al., 2021). Therefore, AL emerges as an essential tool in the classification of PSE-reports, offering more insightful, efficient, and effective use of this critical data in improving patient safety.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by the Agency for Healthcare Research and Quality (AHQR) [Grant no. 1R03HS027680].
