Abstract
Since ~1980, the Animal Health Laboratory (AHL) in Ontario, Canada, has isolated animal mollicute species by culture. Data for the most recent 17 y (2007–2024) captures over 90,000 test results. Advancements in PCR, qPCR, and DNA sequencing have shifted the percentage of testing by PCR from 18.7% in 2007 to 91.1% in 2024. The bulk of this shift is due to the uptake of molecular testing as a screening tool for clinically normal animals, but this shift has not been universal, particularly for ureaplasma testing. Culture remains the gold standard for the detection and identification of rare pathogens and plays a key role in research through our mycoplasma cryobank, which includes 40+ y of isolates. Synergizing the microbiologic and molecular techniques developed over the AHL’s multi-decade history has presented novel opportunities for detection, characterization, and local eradication of animal mollicutes, including the development of new assays, tracking of historical trends for antimicrobial resistance (AMR), and identifying AMR-associated mutations in Mycoplasmopsis (Mycoplasma) bovis.
Bacterial species of the order Mollicutes—mycoplasmas, ureaplasmas, and acholeplasmas—are small and lack a cell wall, contributing to their classic “fried egg” colony appearance under light microscopy and resistance to beta-lactam antibiotics.12,17 Animal mollicutes can cause primary or secondary infections in mammals, birds, and reptiles, occurring as pneumonia, joint and soft tissue infection, abortion, infertility, non-regenerative anemia, and chronic unthriftiness. 12 Their veterinary recognition dates to 1898 with the isolation of the causative agent of contagious bovine pleuropneumonia, later identified in the 1960s as Mycoplasma mycoides subsp. mycoides SC. 17 Several mycoplasma species are World Organisation for Animal Health (WOAH)-listed pathogens,16,17 and even non-listed species such as Mycoplasmopsis (Mycoplasma) bovis are subject to stringent import/export testing requirements. 9
Isolation by culture and identification by fluorescent antibody testing (FAT) remains the gold standard for the characterization of mycoplasma strains. 19 This technique is technically challenging and specialized work with a low throughput and a lengthy turnaround time (TAT); the development of mycoplasma-specific PCRs has offered significant cost and time savings if the diagnostic goals do not require isolation of live mycoplasma.3,19
The Animal Health Laboratory (AHL; University of Guelph, Guelph, Ontario, Canada) has provided veterinary mycoplasma detection services since the 1970s. In the early years, the laboratory detected bovine abortion and neonatal death associated with Ureaplasma diversum, 15 and developed protocols for the detection of animal mollicutes. 16 AHL’s current laboratory information management system (LabVantage; LabVantage Solutions) captures case information from 2007 to the present, in a searchable format that preserves temporal and geographic information for the cases submitted as well as test results. From May 2007 to December 2024, 16,138 samples were cultured for animal mollicutes (Fig. 1), with 2,945 of 16,138 (18.2%) yielding positive results. We identified 4,958 isolates to the species level and 1,535 to the genus level; 52 species were identified by culture (Table 1), with M. bovis the most common, isolated 1,205 times from milk, semen, and tissue samples. The discrepancy between the number of positive submissions and the number of positive isolates is due to coinfections of individual samples by multiple mollicute species.
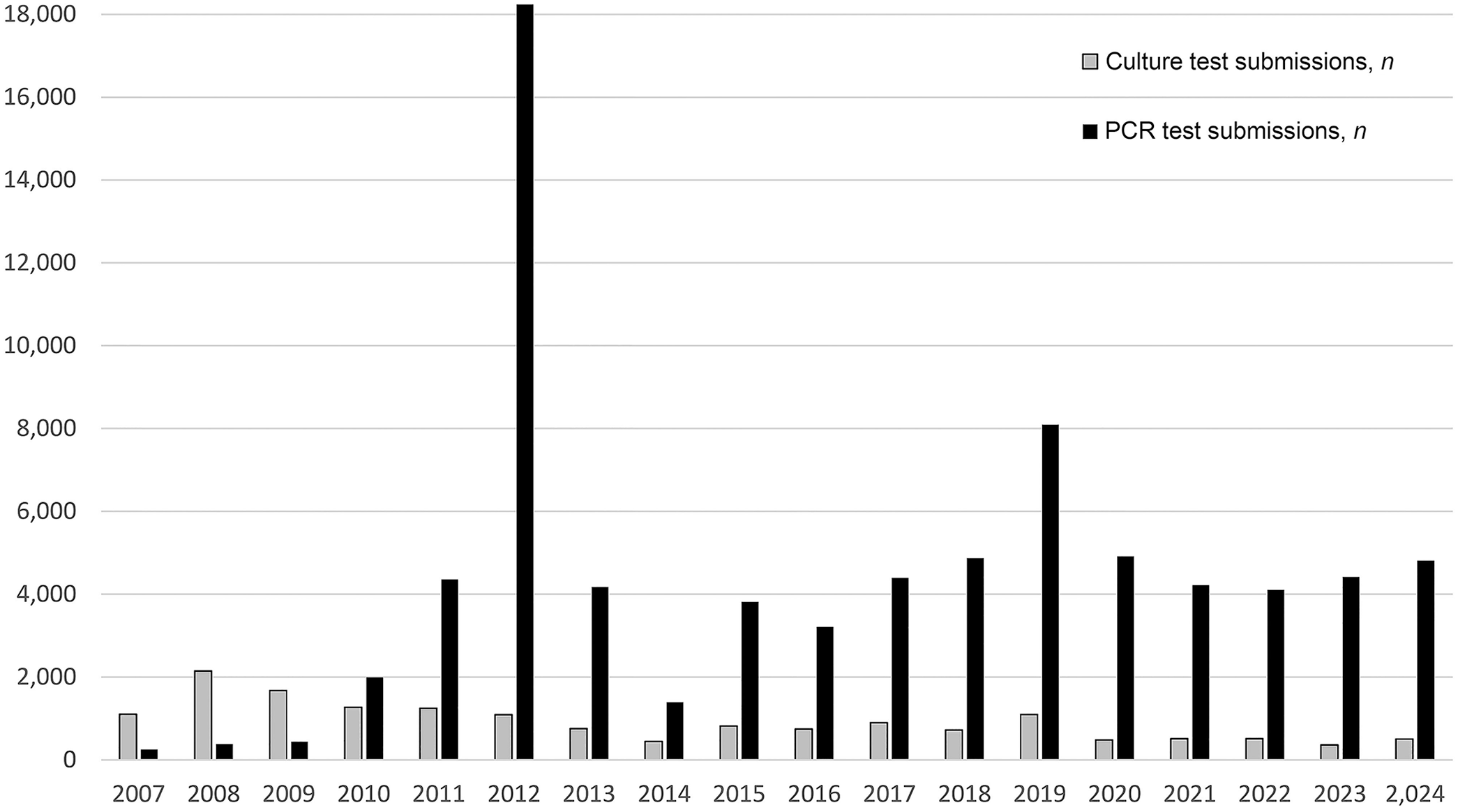
Annual procedure count (2007–2024) of animal mollicute culture and PCR.
Count of animal mollicutes identified at AHL and method of detection, 2007–2024.
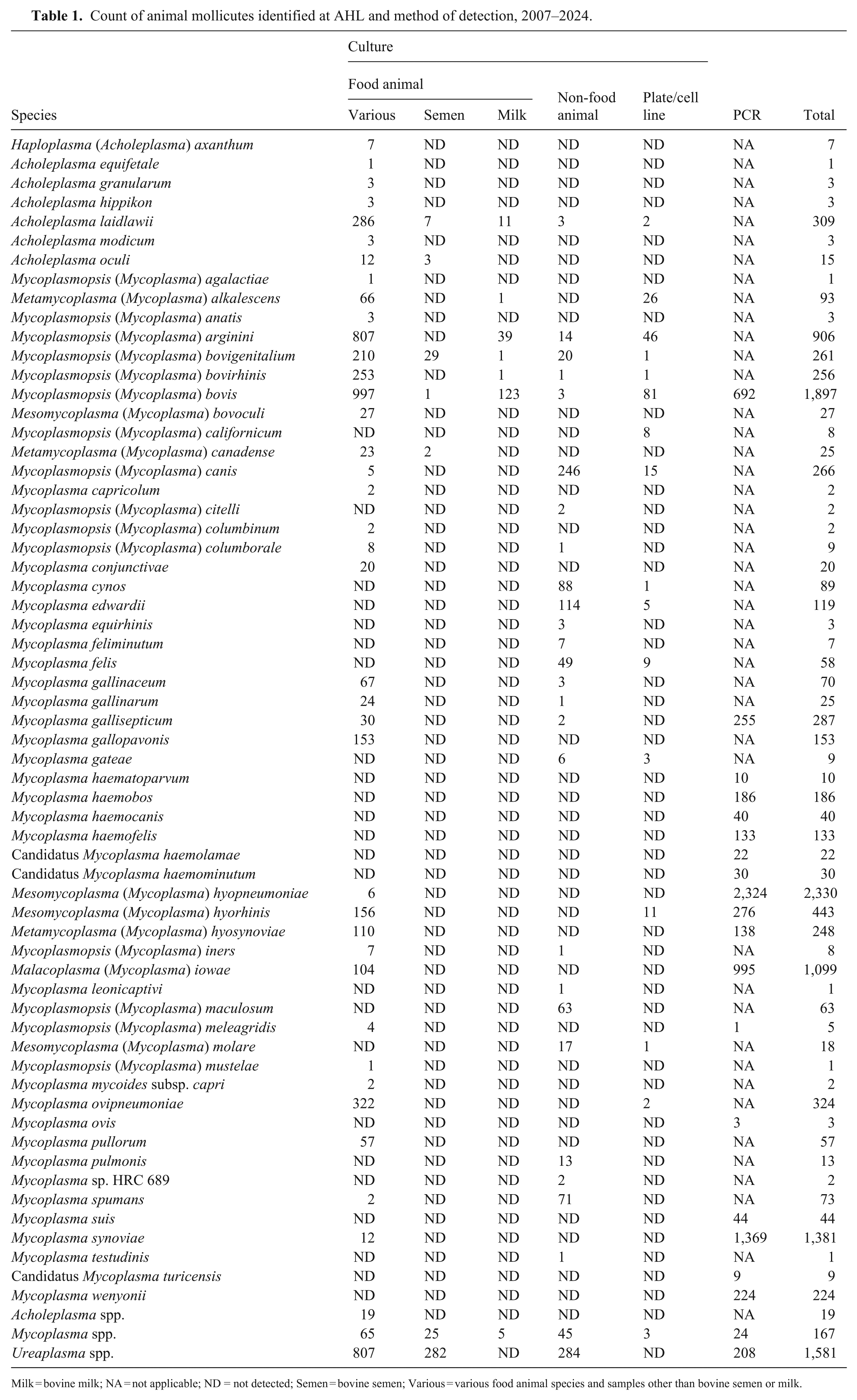
Milk = bovine milk; NA = not applicable; ND = not detected; Semen = bovine semen; Various = various food animal species and samples other than bovine semen or milk.
Submissions for culture are grouped into 5 categories (Table 2) based on the test code assigned to them in our LIMS: semen, milk, cultures/cell lines, food animal general (food animal thereafter), and non-food animal samples. Within the 5 categories, culture methods and selective media are determined by host species and/or sample type, per the foundational texts used in the development of our in-house protocols; for non-milk mammalian samples, ureaplasma-selective medium is included in addition to various mycoplasma-selective media.16,19 MALDI-TOF mass spectrometry (MS) is used as an optional intermediate step in identification, as mollicutes are not well characterized by this method, in addition to requiring greater volumes of pure isolate and a more involved preparation phase compared to other bacteria (unpublished data). However, MALDI-TOF MS remains useful as a guide for selecting appropriate controls for identification by FAT (unpublished data).
Annual procedure counts for animal mollicute culture and sample category, 2007–2024.
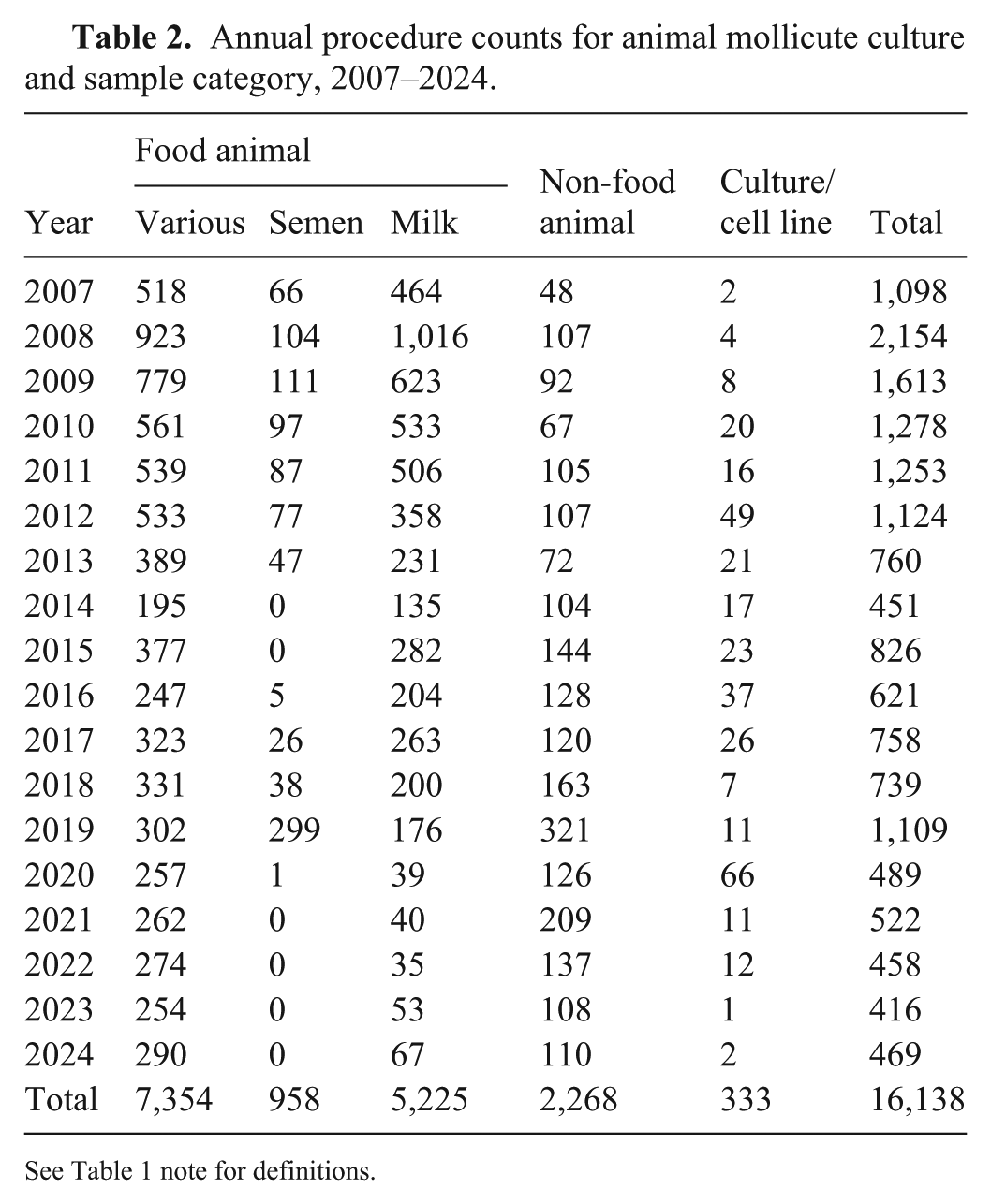
See Table 1 note for definitions.
In total, over the 17-y period (Table 2), food animal samples were the most common submission for culture (7,354; 45.6%) followed by milk (5,225; 32.4%), non-food animals (2,268; 14.0%), semen (958; 5.9%), and cultures/cell lines (333; 2.1%). When considered year-over-year (Fig. 2), we see a shift from predominantly food animal and milk submissions in 2007, towards a testing landscape in 2024 in which food animal submissions are even more predominant but the percentage of non-food animal submissions has increased 5-fold and the percentages of both milk and semen submissions have declined significantly. In the case of semen, no samples have been submitted for culture since 2020 (Table 2). The changing landscape may reflect an increased awareness of mollicutes as disease agents in non-agricultural settings, but it is also likely shaped by the shift toward PCR-based testing, as discussed below.
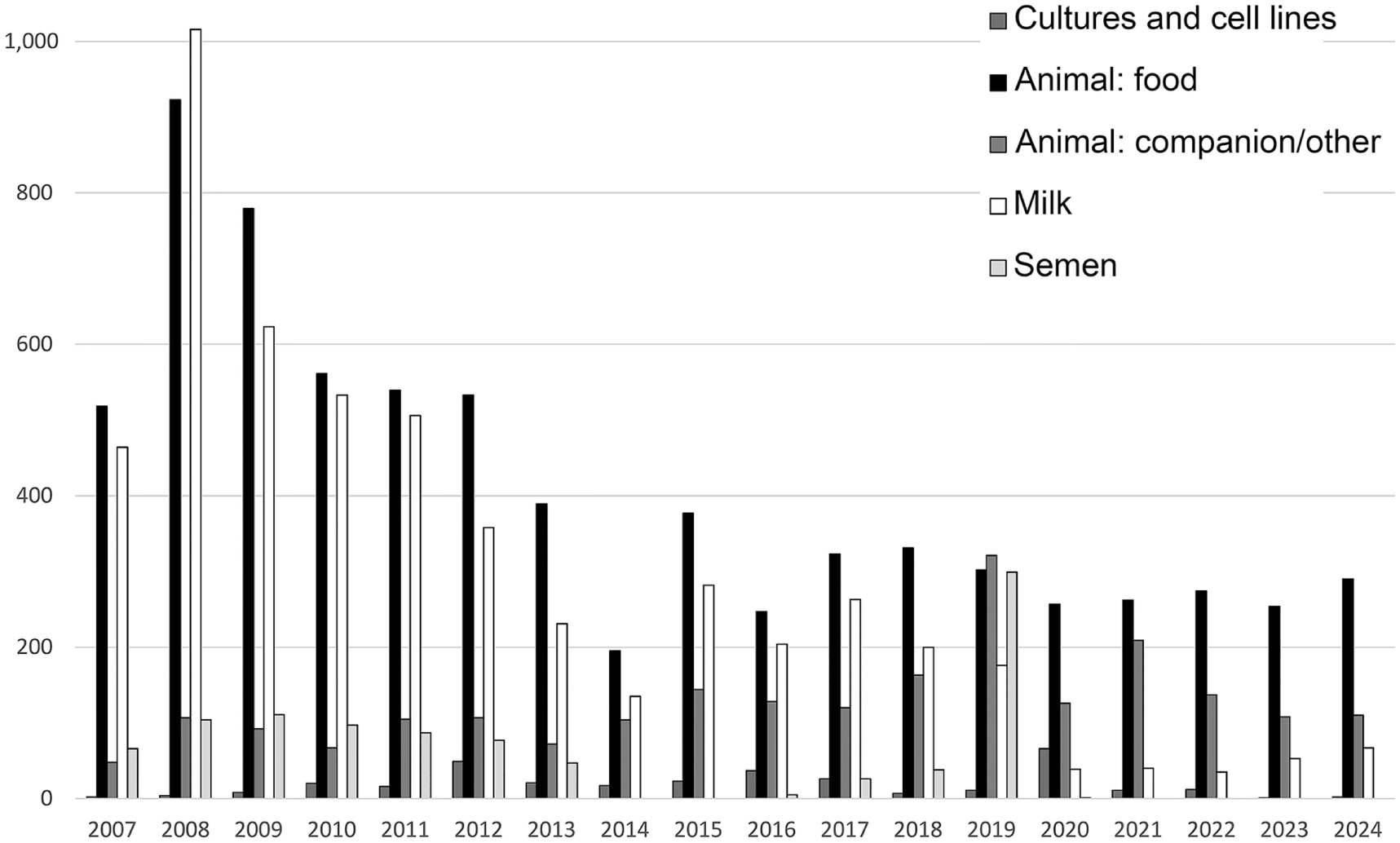
Annual procedure count of animal mollicute cultures at the AHL, by sample source.
The variability in animal mollicute growth rates results in culture TATs ranging from 24 h (Ureaplasma spp.) to 21 d [Mycoplasmoides (Mycoplasma) gallisepticum]. 19 PCR, in contrast, often provides same-day or next-day results, offering significant time savings. 3 This change is captured in our LIMS data: in 2024, the internal metrics for TAT (the time between sample receipt and result reporting) averaged 14.9 d for culture and 0.74 d for the 5 most commonly ordered PCR assays—M. bovis, 9 M. iowae, 3 M. gallisepticum, 5 M. synoviae,2,13 and M. hyopneumoniae. 18 The detection by PCR of intact DNA rather than live organisms also allows testing of samples unsuitable for culture. PCR has seen a rapid uptake relative to culture as a detection tool, representing 18.7% of mycoplasma testing at the AHL in 2007 versus 91.1% in 2024 (Fig. 1).
Between 2007 and 2024, we have used 13 individual PCR assays at the AHL for the detection of mollicutes, either adapted from published literature or developed in-house (Table 3). We conducted 78,169 PCR tests for mollicutes over this time; 6,983 of 78,169 (8.9%) tests were positive, which is lower than our culture-positive rate, primarily because we use PCR as a screening tool for clinically normal animals. We detected 18 animal mollicute species using species-specific PCR assays (Table 1), including 9 hemotropic mycoplasmas unable to be isolated by routine culture.12,16 We determined 232 samples to be positive at the class (Mollicutes spp.: 24) or genus level (Ureaplasma spp.: 208) using 2 pan-species PCRs (Table 3).
Categorized list of animal mollicute PCRs in use at the AHL, 2007–2024, with procedure counts, percent positivity, and associated method development references.
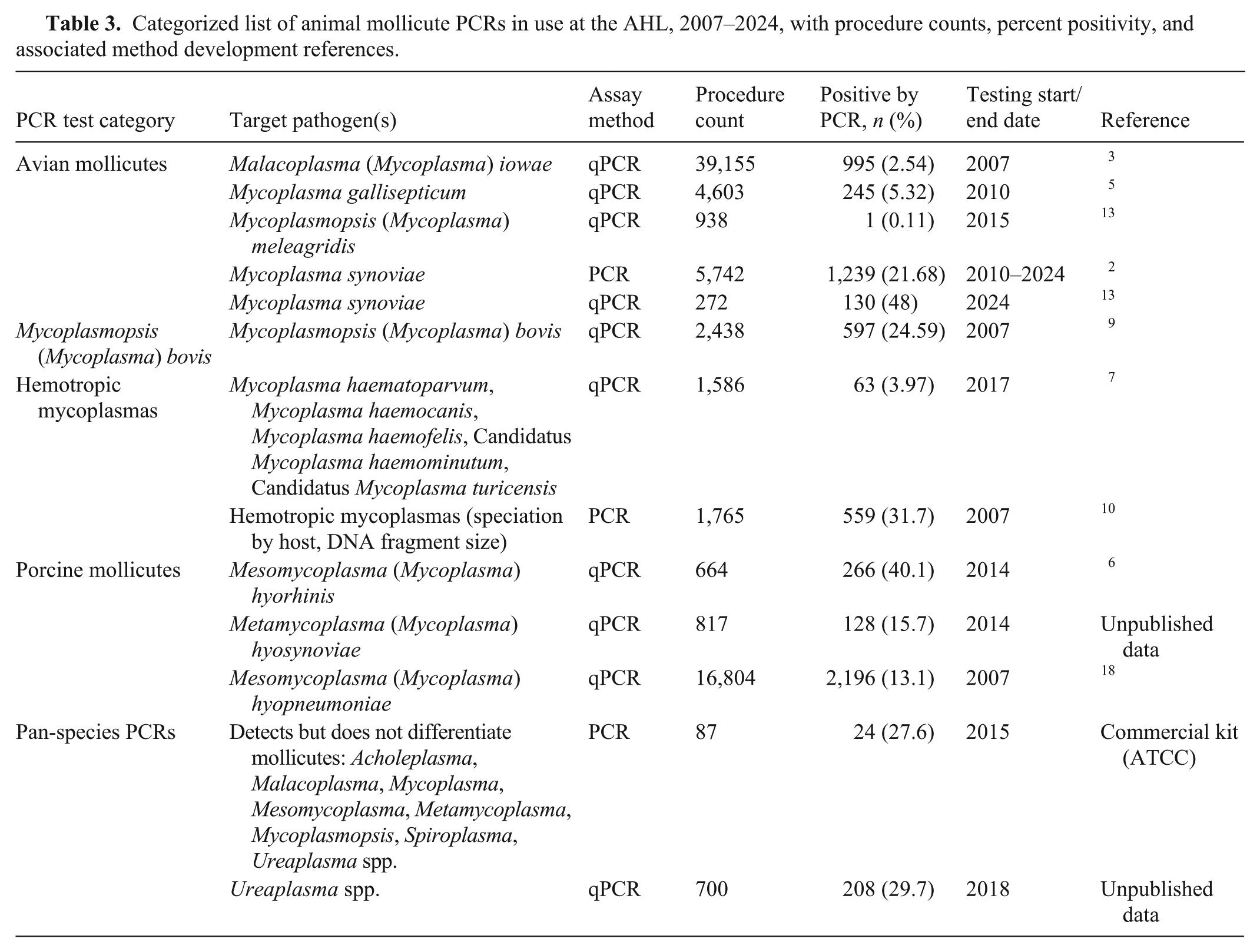
Of interest, although the Ureaplasma spp. PCR (introduced in 2018) lags in procedure count compared to culture, and the total percent positivity is higher for PCR [208 of 700 (29.8%)] than for culture [1,373 of 8,786 (15.6%)], culture-based Ureaplasma detections have been increasing in recent years. The percentage of positive reactions by culture and PCR-based detection showed similar trends for 5 of the 7 y in which both detection methods were available (Fig. 3).
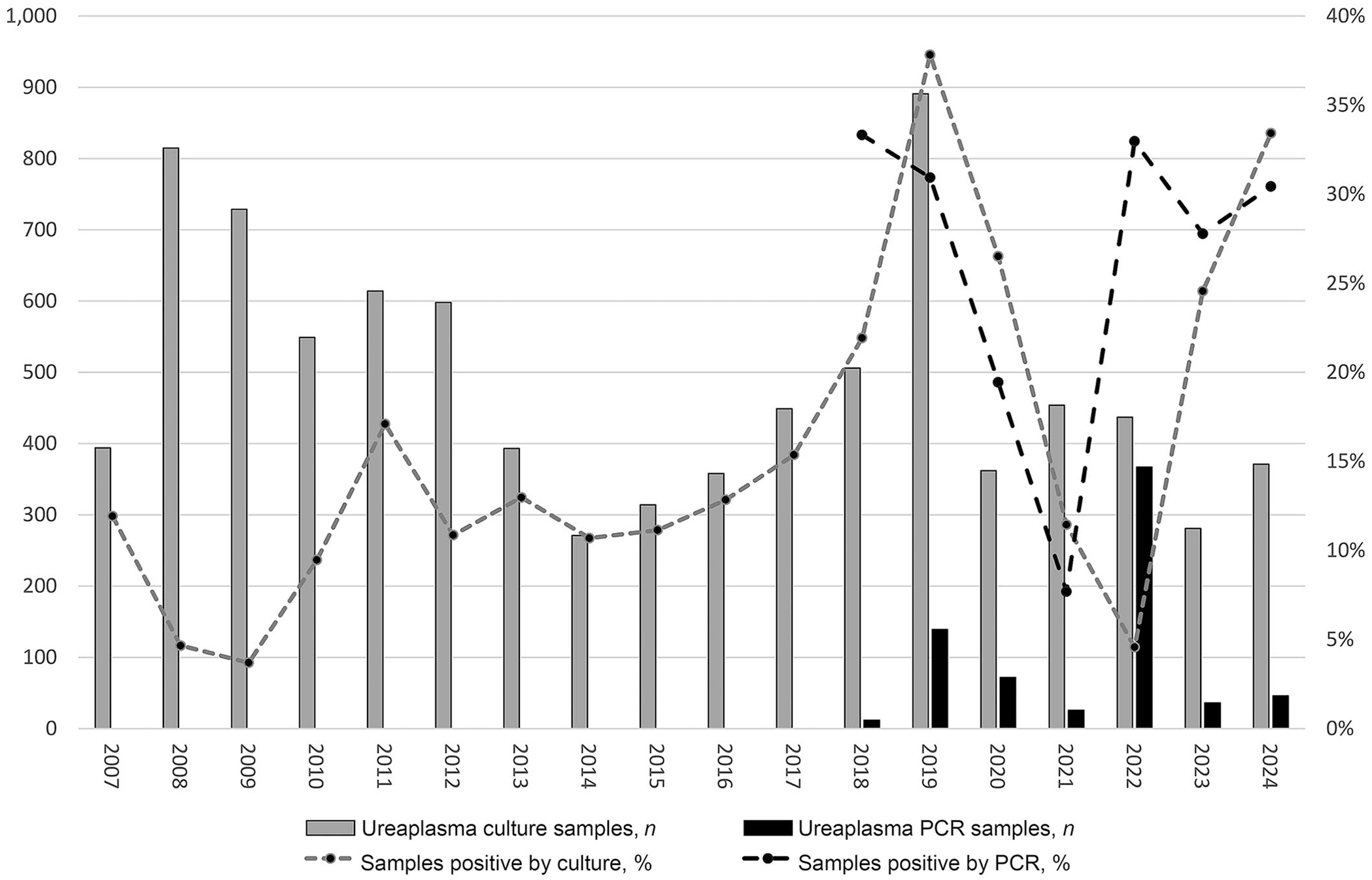
Annual procedure counts (left y-axis) and sample percent positivity (right y-axis) for culture versus PCR-based detection of Ureaplasma spp. The 2022 spike in both PCR procedure count and percent positivity derives from a single large research project submission.
When PCR tests are used primarily for screening clinically normal animals, the expected percentage of positive samples is low [i.e., canine and feline hemoplasmas (3.97%), or M. iowae (2.54%)] (Table 3). The 29.8% average percentage positivity for Ureaplasma spp. PCR suggests that, unlike these PCR assays, the Ureaplasma spp. PCR assay is used primarily for confirmation when Ureaplasma is already suspected as a causative agent of disease. As well, our data suggest that PCR-based screening of clinically normal animals for Ureaplasma infection has yet to see significant uptake among clinicians.
The 10 most common species detected (Table 4) accounted for 4,438 of 6,493 (68.4%) culture isolates, and 6,119 of 6,983 (87.6%) positive PCR results. Identified solely through culture (Table 1), Acholeplasma equifetale, Mycoplasmopsis (Mycoplasma) agalactiae, Mycoplasma leonicaptivi, Mycoplasmopsis (Mycoplasma) mustelae, and Mycoplasmopsis testudinea (Mycoplasma testudineum) were the rarest species detected, isolated a single time each.
Count of positive detections of the 10 most common animal mollicutes identified at the AHL and method of identification, May 2007–December 2024.
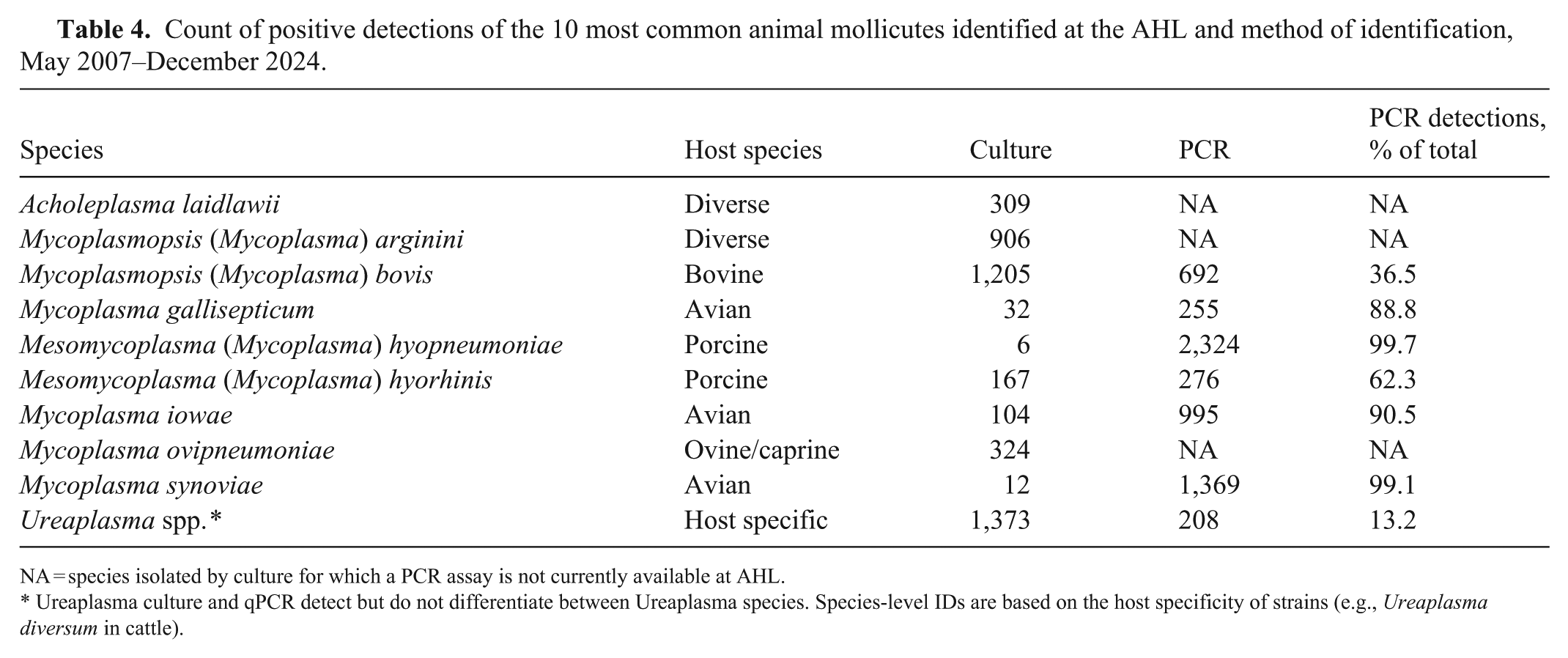
NA = species isolated by culture for which a PCR assay is not currently available at AHL.
Ureaplasma culture and qPCR detect but do not differentiate between Ureaplasma species. Species-level IDs are based on the host specificity of strains (e.g., Ureaplasma diversum in cattle).
Although submissions for culture and isolation have decreased from 2007–2024 (Fig. 2), culture remains valuable in assay development. In 2007, we used positive cultures to develop a Malacoplasma (Mycoplasma) iowae qPCR assay, which ultimately demonstrated cost savings for PCR over culture and FAT. 3 The scalability of PCR enables high-throughput screening, resulting in the use of the M. iowae PCR to test 24,000 samples between 2011–2013 (Fig. 4), and contributing to the eradication of M. iowae from a high-value turkey-breeding facility. 3 Rigorous PCR monitoring for M. iowae at the facility has consistently yielded negative results for over a decade (unpublished data). PCR assays are also used for pre-shipment certification (M. iowae, M. hyopneumoniae, M. bovis) and for rapid screening of animal blood donors (feline and canine hemotropic mycoplasmas). 7 In a 2019 M. synoviae study, we used a vlhA gene PCR assay and sequencing directly from clinical samples to determine the prevalence and diversity in Quebec poultry flocks, leading to the discovery of a novel strain (AHL strain QC-1). 2 Similarly, M. hyopneumoniae strain typing via p146 sequencing revealed the diversity of the pathogen in Ontario. 20
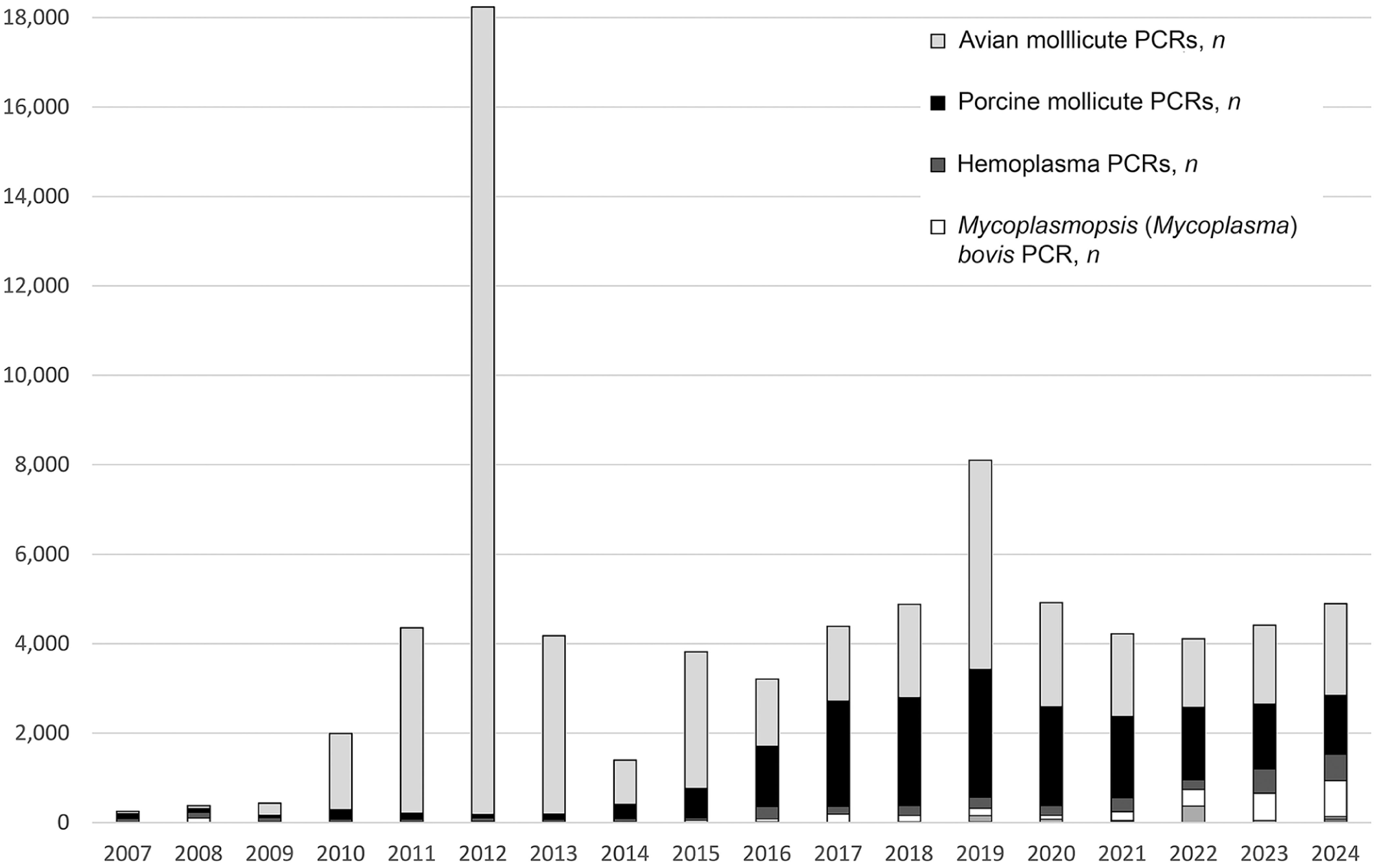
Annual procedure counts of animal mollicute PCR testing, grouped by category (Table 3). The 2011–2013 dominance of avian mollicute PCRs resulted from qPCR screening for Malacoplasma (Mycoplasma) iowae as part of an eradication project.
Pure isolates are preferred for DNA sequencing and are frequently used for PCR method development, demonstrating complementary roles for culture and PCR. The AHL maintains a cryobank of positive isolates, which has facilitated the development of a multi-locus sequence typing (MLST) scheme for M. bovis, 1 a high-resolution melting curve analysis to identify commonly isolated mycoplasma species from ruminant, avian, and canine samples, 14 the study of antimicrobial resistance (AMR) trends in M. bovis over 3 decades, 4 and the identification of AMR genes in M. bovis using whole-genome sequencing (WGS). 11 For the latter, we revived cryobanked isolates with high, low, and typical AMR levels identified previously, 4 and confirmed with qPCR and minimum inhibitory concentration testing before sequencing, identifying 55 potential AMR mutation sites, including 3 linked to M. bovis AMR and 16 associated with AMR in other bacterial species. 11 WGS of cryobanked isolates has also produced the first complete genomes of M. cynos and M. felis. 8
In addition to molecular and microbiologic testing, we perform immunologic testing, including ELISA, to assess prior mycoplasma exposure. From 2007–2024, >230,000 animal mollicute ELISAs were conducted, demonstrating even greater uptake than PCR-based detection. Our integrated microbiology, molecular testing, and immunology has provided comprehensive mollicute testing tailored to diverse veterinary diagnostic and research needs.
Footnotes
Acknowledgements
We are grateful for the technical support from former and current technicians, Tanya Brock, Colin Cameron, Hamid Haghighi, Jeff Hornby, Afreeda Mahesaniya, Leah Read, and Ana Rita Rebelo. We acknowledge the leadership of former laboratory supervisors Louise Ruhnke, Gaylan Josephson, and Tony van Dreumel.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our study was supported in part by the Ontario Animal Health Network (OAHN), Ontario Ministry of Agriculture, Food and Agribusiness (OMAFA), and the University of Guelph Animal Health Laboratory (AHL).
