Abstract
Sarcocystis pinnipedi is an apicomplexan protozoal parasite that was first recognized during a mass mortality event in juvenile grey seals (Halichoerus grypus) in the northwest Atlantic Ocean. Since its identification, this parasite has been reported in various pinniped species and has been associated with fatal necrotizing hepatitis. Little is known of the host range of S. pinnipedi. Here we report 2 cases of California sea lions (Zalophus californianus) in managed care that died following an 8-d history of inappetence, vomiting, diarrhea, and progressive lethargy with elevated hepatic enzyme activities. Postmortem examination identified hepatitis and icterus. Sarcocystis schizonts and zoites were identified in regions of necrosis. Molecular and ultrastructural findings demonstrated the close relatedness of this Sarcocystis to S. canis, which produces a similar lesion in bears.
Two adult, male California sea lions (Zalophus californianus; hereafter, CSL 1 and 2) were kept in managed care and housed at separate, geographically distant aquaria. Both declined rapidly following approximately one week of inappetence, vomiting, diarrhea, and progressive lethargy. Notably, CSL 1 lived in the open ocean for up to 4 y until it was moved indoors where it spent the rest of its life (~15 y); CSL 2 never lived in the open ocean. Treatments for both animals included broad-spectrum antimicrobials, gastroprotectants, analgesic and anti-inflammatory drugs, and fluid therapy. Hematologic and serum biochemical results in both cases indicated hemoconcentration, marked hyperbilirubinemia, and elevated aspartate aminotransferase, alanine aminotransferase, and gamma-glutamyl transferase activities. Both sea lions died within 8 d after initial presentation.
Postmortem findings for both sea lions included icterus (Fig. 1A) and a friable liver with multiple pale or dark-red foci scattered throughout the parenchyma (Fig. 1B). A complete set of tissues was collected at autopsy, fixed in 10% neutral-buffered formalin, and processed by routine methods. The primary histologic lesion identified in both sea lions was severe necrotizing hepatitis. Throughout the liver, there were numerous randomly distributed and coalescing necrotic foci with disruption of hepatic cords, accumulations of cellular debris and fibrin, and low numbers of neutrophils (Fig. 2). In regions of necrosis, numerous 10–15-µm protozoa were present within hepatocytes or Kupffer cells or were free within extracellular spaces. Protozoa ranged from oval, deeply basophilic immature schizonts with lobulated nuclei to faintly basophilic mature schizonts containing abundant <2-µm long, deeply basophilic merozoites that were occasionally arranged in a radiating pattern surrounding central residual bodies (rosette schizonts; Fig. 2). Rare mature schizonts were ruptured and surrounded by loosely arranged free merozoites. Tissue cysts (sarcocysts) were not visualized in any tissue evaluated histologically, and protozoal-related lesions were confined to the liver in both cases.
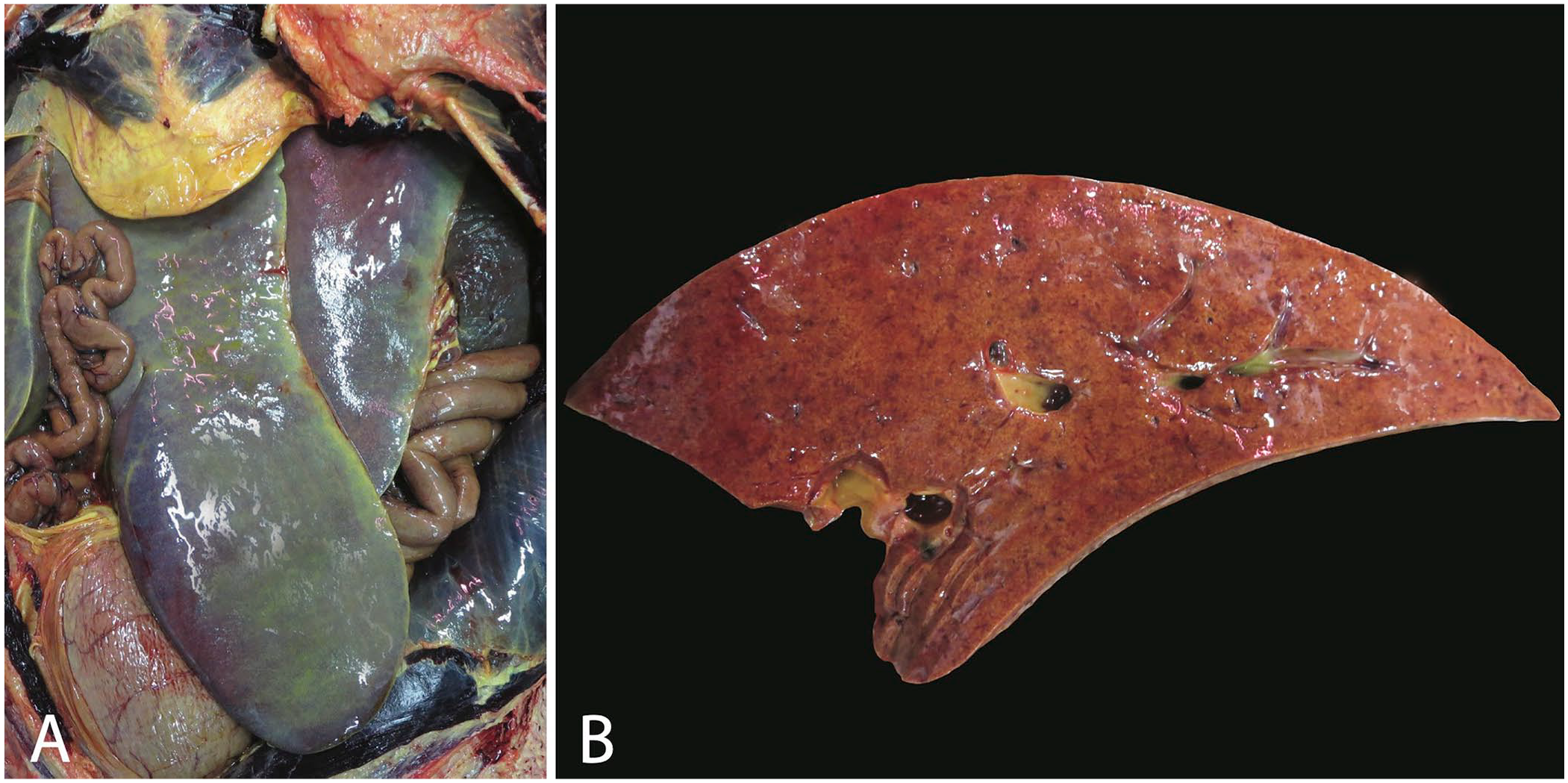
Gross lesions from California sea lion 1.
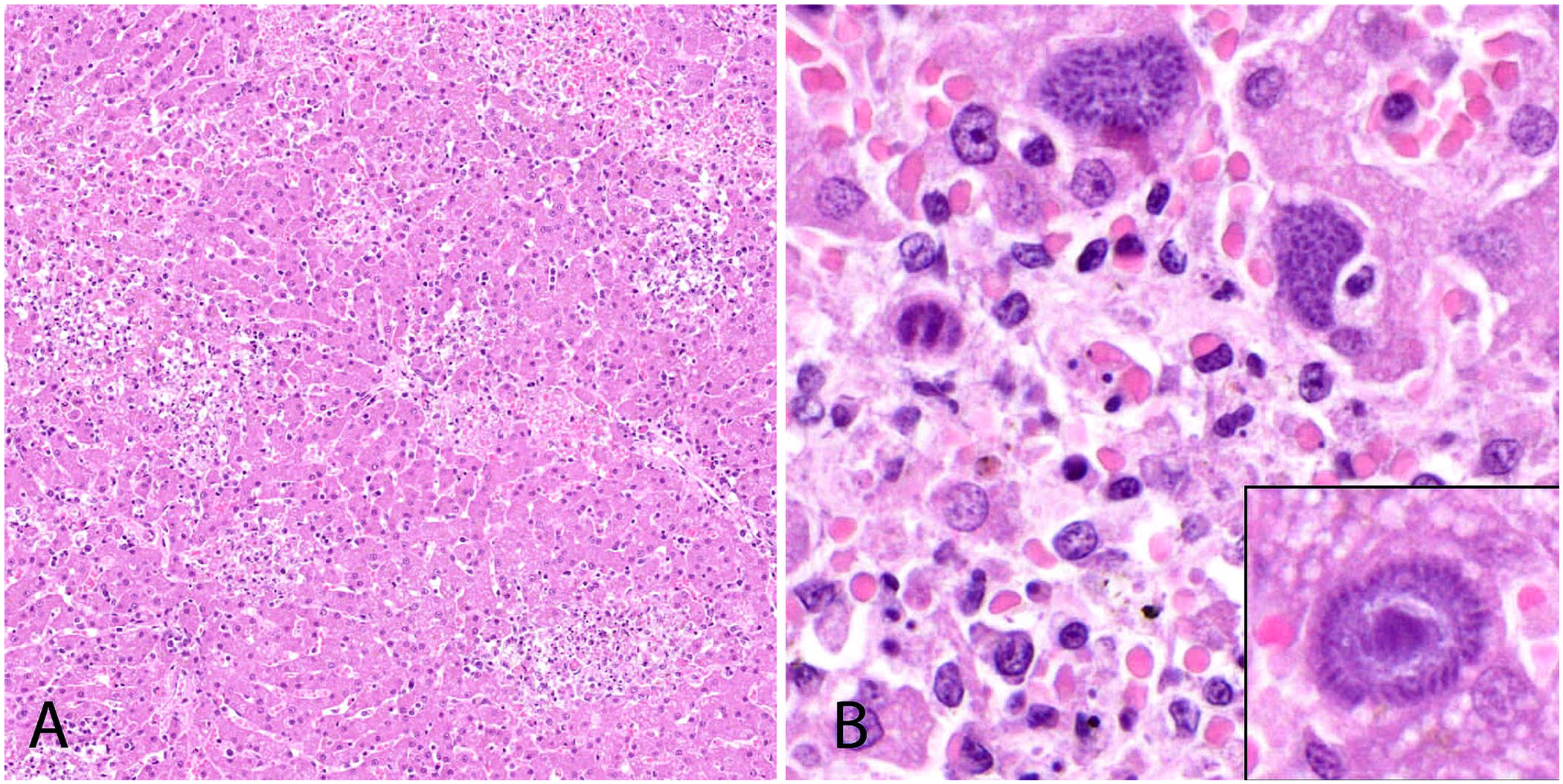
Histology of the liver from California sea lion 1. H&E.
Immunohistochemistry using available antibodies for several protozoan pathogens was performed on sections of liver according to published methods.3,16 Protozoa within the liver were not immunoreactive with antibodies for Sarcocystis neurona (polyclonal; Ohio State University, Columbus, OH, USA), Sarcocystis falcatula (polyclonal; Ohio State University), or Toxoplasma gondii (AR125-SR, polyclonal; BioGenex Laboratories). Antemortem serologic titers assessed using an immunofluorescent antibody test (IFAT; University of California–Davis, Davis, CA, USA [UC Davis]), were also negative for S. neurona and T. gondii.
Sections of liver from both sea lions were submitted in 10% neutral-buffered formalin for ultrastructural analysis. Briefly, liver was trimmed to 1–2 mm3, and multiple sections were processed and embedded in resin as described previously. 2 Thick sections were stained with toluidine blue to detect protozoa within the lesions. Seventy-nm thin, contrast-enhanced sections were transferred to a 200-mesh copper grid (Electron Microscopy Sciences). Organisms were visualized with a transmission electron microscope (1400; Jeol), and images were acquired (AMT Capture Engine v.7.00 camera; Advanced Microscopy Techniques). Only asexual stages were identified in the liver. Schizonts and merozoites were present within hepatocytes. Parasitophorous vacuoles were not identified. Merozoites (n = 9; 3.2 × 1.6 µm) had electron-dense micronemes surrounding the conoid and dense granules around the mid-body and nucleus (Fig. 3A); rhoptries, which are not present in the merozoite stage of Sarcocystis spp., were not identified. 6 Anlagen developed radially by endopolygeny through elongation of the posterior extension around a central lobulated nucleus above a spindle apparatus (Fig. 3B). Collectively, these findings were consistent with apicomplexan members of the genus Sarcocystis.
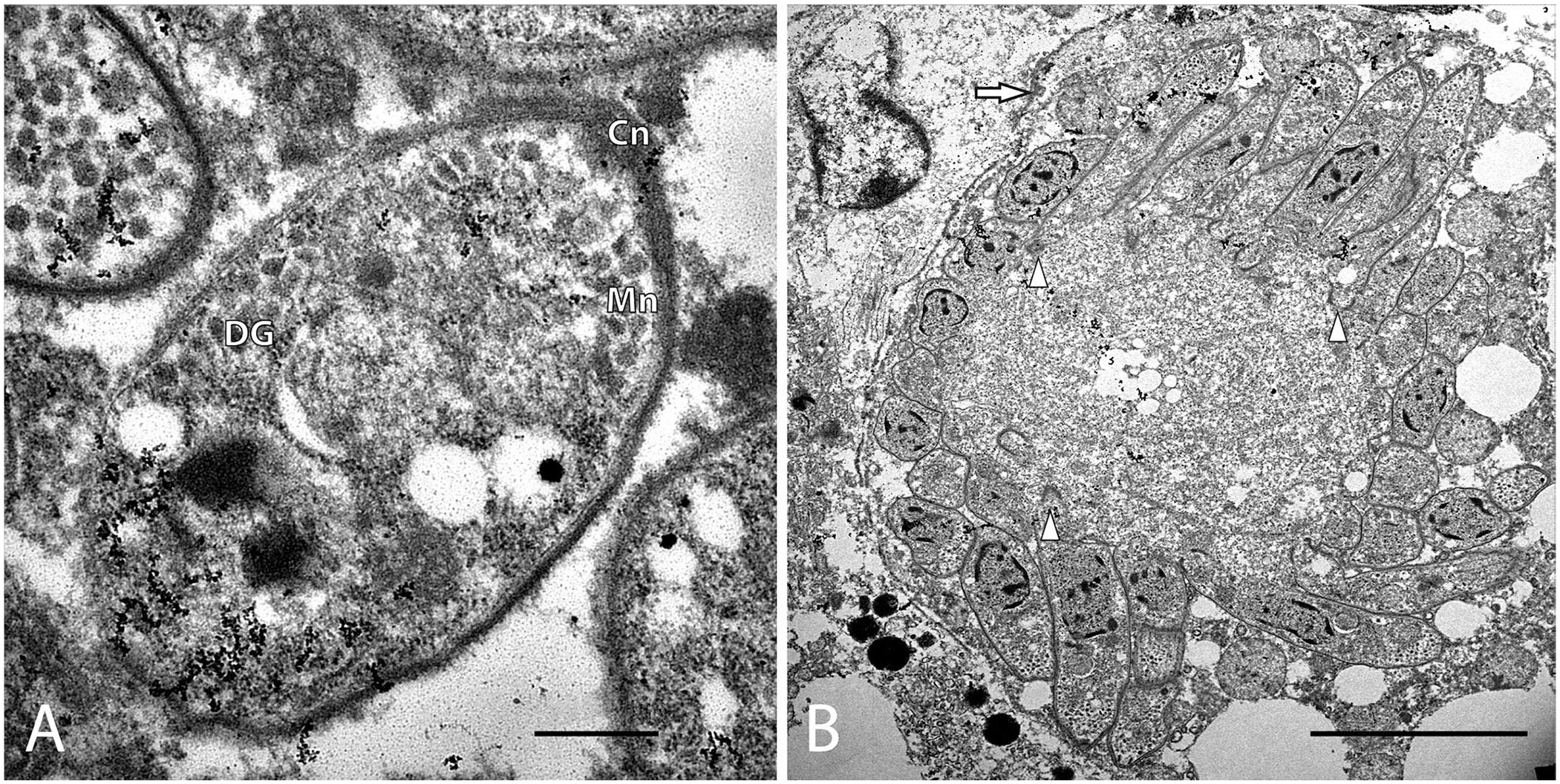
Electron photomicrograph of Sarcocystis pinnipedi.
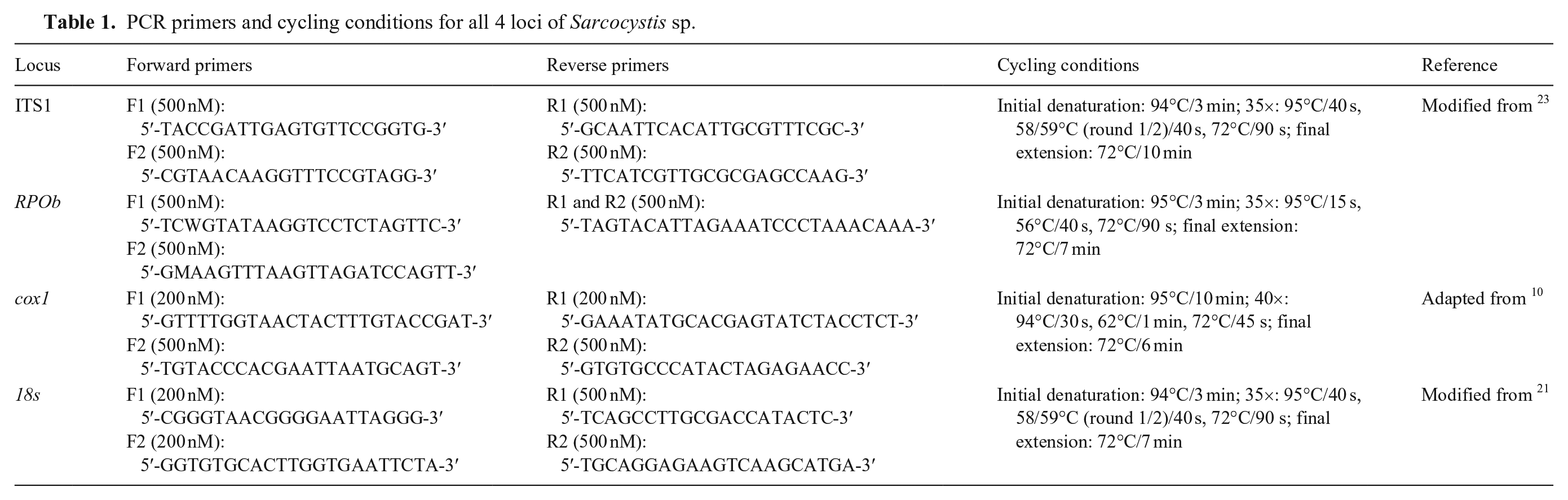
Molecular characterization of the protozoa was performed using conventional PCR that targeted 4 loci (Table 1). DNA was extracted from the liver of each sea lion (DNeasy blood and tissue kit; Qiagen) per the manufacturer’s instructions. Nested PCR assays that targeted the internal transcribed spacer 1 (ITS1) and 18S rRNA loci were performed on each biologic replicate to screen for apicomplexan DNA. Sarcocystis-specific assays, including nested PCR that targeted cytochrome oxidase 1 (cox1) and hemi-nested PCR that targeted the RNA polymerase β subunit (RPOb), were used for further characterization. Each PCR reaction consisted of a 50-µL volume (AmpliTaq Gold 360; Applied Biosystems). PCR products were extracted after electrophoresis from 2% agarose gel, purified (QIAquick gel extraction kit; Qiagen), optimized according to the manufacturer’s direction for sequencing applications, and submitted for Sanger sequencing (UC Davis DNA Sequencing Facility, and GeneWiz). Sequences were deposited in GenBank (OR866194–866196, OR880534–OR880537).
PCR primers and cycling conditions for all 4 loci of Sarcocystis sp.
Sequences from all 4 loci were examined visually, and consensus sequences were constructed from paired forward and reverse sequences in Geneious v.11.1.5 (Dotmatics). Consensus sequences obtained for the ITS1 locus of both sea lions had 100% homology to Sarcocystis pinnipedi recovered from a harbor seal (MT460246) by nucleotide BLAST (https://blast.ncbi.nlm.nih.gov/Blast.cgi), which was distinct from the closest related species S. canis, with 98% nucleotide similarity. Four additional previously termed S. canis–like ITS1 sequences (OR339983–OR339986) shared 99.7% nucleotide identity with the CSL-derived ITS1 sequences. Nucleotide similarities to S. canis at the cox1, 18S, and RPOb loci were 99.8%, 99.7%, and 99.7%, respectively. Reference sequences for related Sarcocystis spp. were downloaded from GenBank and imported into Geneious for comparison. Locus-specific sequences were aligned using MUSCLE 8 in Geneious, and open-reading frames were manually checked for cox1, 18S, and RPOb genes. Best-fit substitution models were determined for all 4 loci by using jModelTest 18 in MEGA v.11.0.13 (https://www.megasoftware.net). Maximum-likelihood nucleotide trees rooted against S. neurona (not always depicted) for all 4 loci were constructed using 1,000 bootstrap replicates. Single-locus phylogenetic trees were constructed with the model calculated to have the lowest Bayesian information criterion available in PhyML 11 implemented in Geneious. The ITS1 tree (Fig. 4) was best modeled with HKY + I. 13 The 18S tree (Suppl. Fig. 1) best-fit model test resulted in a JC model 14 as the best-performing model; these results were compared to trees constructed with the most parameterized model (GTR+G+I) that has been proposed as the de facto model for phylogenetic reconstruction, which resulted in similar phylogenetic relationships.1,19 The RPOb sequence could only be obtained from CSL 1, and the resultant RPOb tree (Suppl. Fig. 2) was best modeled with HKY+G+I, supporting phylogenetic relationships identified by ITS1. 13 The cox1 tree was best modeled with HKY (Suppl. Fig. 3). 13
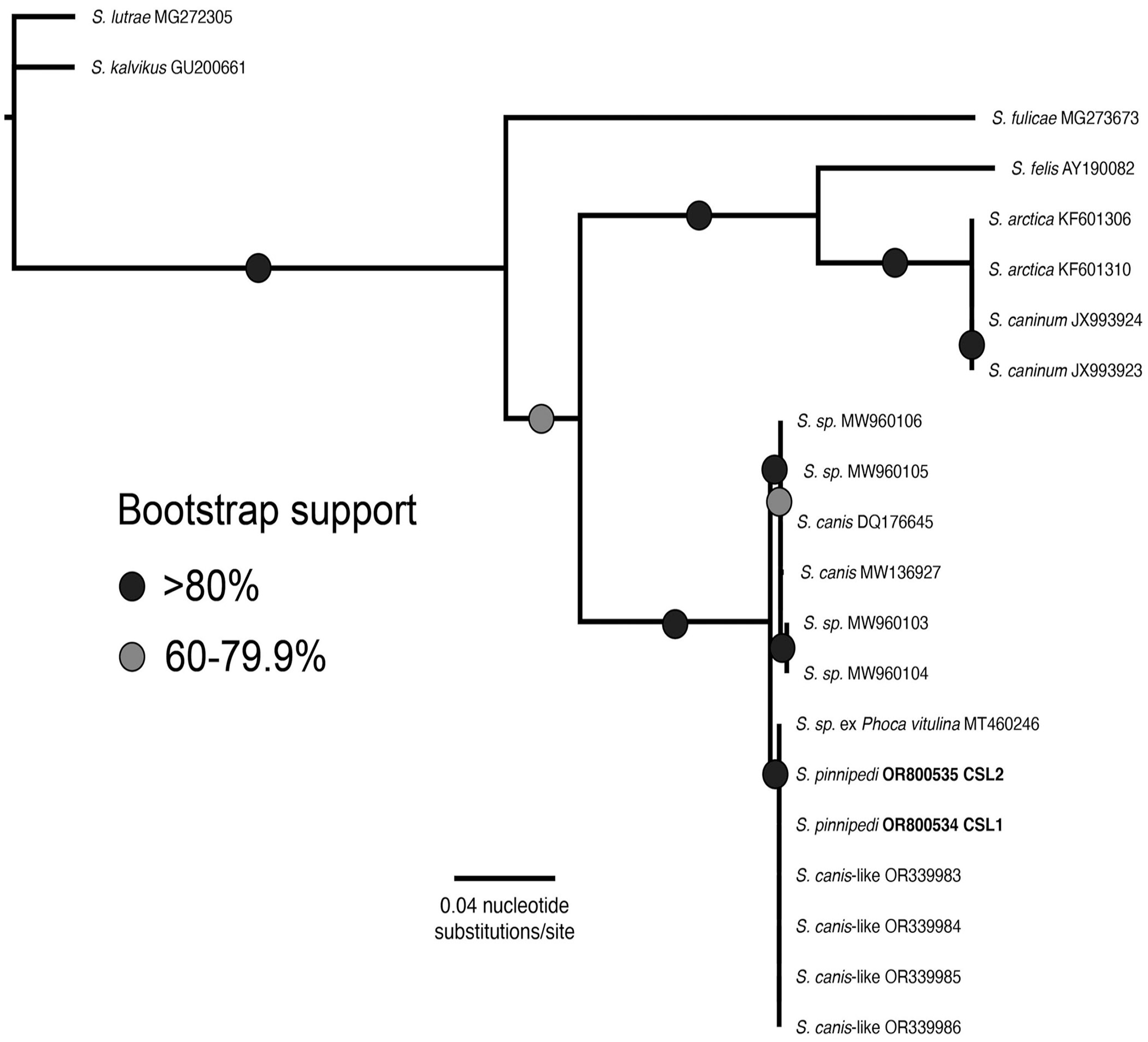
Maximum-likelihood phylogenetic tree for ITS1 (HKY+I) shows that Sarcocystis pinnipedi has been documented previously and is distinct from the closely related S. canis. Samples from our cases (CSL 1 and CSL 2) are in boldface.
Phylogenetic analysis of the Sarcocystis sp. identified in both CSLs classified them as S. pinnipedi based on the ITS1 locus (Fig. 4), forming a monophyletic clade with S. pinnipedi sequenced from a harbor seal and presumptive S. pinnipedi (identified as S. canis–like) described in other CSLs.15,22 It was also considered distinct from the closely related species S. canis based on ITS1, RPOb, and 18S sequences. Relationships between S. canis, our S. pinnipedi sequences from CSL 1 and 2, and another closely related species, S. arctica, could not be resolved at the cox1 gene (Suppl. Fig. 3). S. pinnipedi differs slightly from S. canis morphologically as well as genetically; merozoites of S. pinnipedi are smaller than S. canis merozoites. 7 Unfortunately, sarcocysts were not identified in our cases for ultrastructural comparison between S. pinnipedi and S. canis at this asexual stage.
S. pinnipedi is an apicomplexan parasite that was first documented while investigating a mass mortality event of juvenile grey seals (Halichoerus grypus) in Nova Scotia, Canada in 2012. 12 Affected seals had necrotizing hepatitis with intralesional protozoa similar to those described in our study. Sarcocystis sp.–associated hepatitis has been reported in several pinnipeds and cetaceans.4,17 A lack of definitive molecular, immunohistochemical, and/or ultrastructural characterization led many of these previous reports to classify the protozoa as “Sarcocystis sp.” or “S. canis–like sp.” due to the similar hepatic lesions caused by S. canis in bears.9,15 However, this approach has led to confusion regarding which parasites circulate in various host species. This appears to be the case with S. canis–like organisms identified in an Indo-Pacific bottlenose dolphin and parasites in CSLs (Suppl. Fig. 3). 4 It is possible that some of these cases of Sarcocystis sp.–associated hepatitis in marine mammals were S. pinnipedi infections.
Members of the genus Sarcocystis have an obligate 2-host life cycle. Infection of Sarcocystis sp. in naturally adapted hosts is generally considered an incidental finding. However, infection in aberrant or incidental hosts may develop into severe disease. For example, horses and southern sea otters infected with S. neurona can develop severe neurologic disease when infected with this pathogen. 5 Additionally, S. neurona infection in CSLs has been implicated in the development of immune-mediated polyphasic rhabdomyositis. 20 The definitive and intermediate host range of S. pinnipedi is currently unknown. However, the fatal necrotizing hepatitis observed in CSLs suggests that sea lions (and potentially other pinniped species) may not be the natural intermediate host for S. pinnipedi but rather may serve as aberrant hosts.
The source of infection for our 2 cases remains unknown, and it is interesting that CSL 1 was housed exclusively indoors for over a decade. The acute nature of the hepatitis and lack of tissues cyst stages suggest a recent exposure to S. pinnipedi rather than a chronic infection. Fish purchased for feed were considered a possible source, and limited samples from herring, capelin, and squid, along with water and feces from another CSL, a whale, a dolphin, and a mouse from the facility housing CSL 1 were processed for DNA extraction and PCR at the ITS1 locus. Toxoplasma gondii DNA was detected in the capelin and mouse feces. No other protozoa were detected.
Antemortem tests to determine infection with S. pinnipedi are challenging. To date, an S. pinnipedi–specific serologic assay is not available. Both sea lions were serologically and immunohistochemically negative for S. neurona and T. gondii, suggesting a lack of cross-reactivity in these commonly used assays for marine mammals. Molecular testing remains the most efficient procedure for definitive identification of this parasite but requires sampling of infected tissue, which can typically only be acquired postmortem or via surgical biopsy. Given the apparent impact that this pathogen has on wild and captive marine mammals, further research is needed to better understand the parasite’s life cycle and susceptible intermediate host range, and to develop antemortem tests, surveillance, and preventive or treatment strategies. Based on our findings, S. pinnipedi should be included as a differential diagnosis for rapid-onset hepatic disease in pinnipeds in managed care regardless of past or present housing or perceived exposure risks.
Supplemental Material
sj-pdf-1-vdi-10.1177_10406387251321832 – Supplemental material for Clinicopathologic, molecular, and ultrastructural features of Sarcocystis pinnipedi infection in 2 California sea lions with fatal necrotizing hepatitis
Supplemental material, sj-pdf-1-vdi-10.1177_10406387251321832 for Clinicopathologic, molecular, and ultrastructural features of Sarcocystis pinnipedi infection in 2 California sea lions with fatal necrotizing hepatitis by Elliott S. Chiu, Devinn M. Sinnott, Martha A. Delaney, Michael M. Garner, Lance Adams, Bill Van Bonn, Kathleen M. Colegrove, Katie Haman, Anibal G. Armién and Karen Shapiro in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank the animal husbandry, veterinary, and technical staff from the contributing institutions. Thanks to the University of Illinois Zoological Pathology Program faculty, residents, and staff, the Veterinary Diagnostic Laboratory Histology Lab personnel, Northwest ZooPath staff, and LeRoy Brown of Histology Consultation Services for technical expertise and assistance. Thanks to Leslie Rueda (UC Davis) for laboratory support of molecular testing, as well as Christina Heard (CAHFS) for electron microscopy sample preparation and sections.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
