Abstract
African rhinoceros undergo chemical immobilization and prolonged transport during translocations for conservation purposes and, hence, experience several pathophysiologic changes, including skeletal muscle injury. Potential concurrent myocardial injury has not been investigated due to a lack of validated immunoassays. We aimed to use inferred cardiac troponin I (cTnI) amino acid sequences of southern white (Ceratotherium simum simum) and southern-central black (Diceros bicornis minor) rhinoceros to assess the potential usefulness of several commercial cTnI immunoassays for detecting cTnI in African rhinoceros. We extracted RNA from the myocardium of deceased rhinoceros (2 white, 1 black rhinoceros) followed by primer design, cDNA synthesis via RT-PCR, and Sanger sequencing. The inferred cTnI amino acid sequences were obtained from the mRNA transcript sequences. The homology of epitope binding sites recognized by capture and detection antibodies in 6 human immunoassays was visually evaluated using aligned inferred rhinoceros cTnI amino acid sequences. Percentage identity between white and black rhinoceros cDNA nucleotide sequences was 99%; inferred amino acid sequences were identical. There were 5 amino acid differences between humans and rhinoceros in the epitope binding sites of immunoassay antibodies; 5 assays contained antibodies against epitopes that were not conserved. For one assay, the single capture antibody targeted a short heterologous epitope (residue 87–91), and cross-reactivity with rhinoceros cTnI was deemed unlikely. For the other 5 assays, complete antibody-epitope homology, or the inclusion of multiple detection or capture antibodies, or targeting of long epitopes, indicated that these assays could be suitable for further investigation of cTnI measurement in African rhinoceros.
The southern-central black (Diceros bicornis minor) and southern white (Ceratotherium simum simum) rhinoceros are, respectively, listed as critically endangered and near-threatened by the International Union for Conservation of Nature Red List of Threatened Species. 37 Poaching, habitat fragmentation and loss, international trade in illegal rhino products, global financial recessions, and pandemics resulting in revenue shortfall negatively affecting conservation programs are all threats to African rhinoceros conservation.12,36 Translocation is an important conservation strategy and involves capture using chemical immobilization, temporary captivity, transport, and release into a new environment.10,36,37 Several physiologic challenges occur during capture and prolonged transport in particular, including severe hypoxemia, acidemia, tachycardia, hypertension, dehydration, electrolyte imbalances, a negative energy balance, muscle injury, protein catabolism, and stress-induced immunomodulation.37,38 Rhabdomyolysis also occurs due to muscle over-exertion from being chased, the effects of stress and potent opioids used during immobilization, such as etorphine, causing muscle tremors and rigidity resulting in muscle hypoperfusion and hypoxia.31,38 An increased state of muscle contraction (i.e., standing in a transport crate during long journeys) with resultant tissue hypoxia and repeated IM injections of tranquilizers throughout transport are additional factors that contribute to muscle damage. 38 The activities of biomarkers of muscle damage (muscle-specific creatine kinase and nonspecific aspartate aminotransferase) that increase during transport are not specific to skeletal muscle or cardiac muscle damage; hepatocellular injury and hemolysis will also result in increases in activity of these enzymes in the blood.
Furthermore, reported cardiorespiratory effects of chemical immobilization in rhinoceros include severe hypoxemia, hypercapnia, tachycardia, hypertension, and respiratory and metabolic acidosis.9–11,21,53 Cardiac myocytes are very sensitive to hypoxia, with tissue ischemia and concurrent acidosis significantly impairing cardiomyocyte metabolism, resulting in cell necrosis and apoptosis.23,28 To compensate for prolonged increased peripheral resistance secondary to drug-induced vasoconstriction and hypertension, the myocardium enlarges further to cardiomyocyte hypertrophy and resultant fibrosis due to necrosis and/or apoptosis. 42 Thus, some degree of reversible or irreversible myocardial injury will likely result from chemical immobilization in rhinoceros, but this has not been investigated.
Troponins are proteins that regulate muscle contraction in skeletal and cardiac muscle tissue. 52 The troponin complex consists of 3 proteins, namely troponins I, T, and C. 52 There are 3 troponin I isoforms, 2 are present in skeletal muscle; 1 isoform is present only in cardiac tissue. 52 The cardiac isoform is the largest troponin isoform (24 kDa) due to an additional 32 amino acid N-terminal peptide. 52 Cardiac troponin I (cTnI) is highly conserved among mammalian species, with a reported cTnI nucleotide homology in cats, dogs, horses, humans, and rats of 85–95% and amino acid homology of 93–96%.32,39,40 Cardiac troponins are highly sensitive and specific biomarkers for myocardial damage and are used in human and veterinary medicine for the detection of cardiomyocyte injury. 52
Blood troponin concentrations are determined with immunoassays that utilize specific cTnI antibodies that target various epitopes on the cTnI molecule.26,27,52 A variety of cTnI assays, produced by different manufacturers and with different levels of analytical sensitivity, are available.26,27 Cardiac troponin I human assay antibody cross-reactivity for blood cardiac troponin levels in various species has been demonstrated.27,32,39,40 The epitopes on the cTnI molecule that are commonly detected by second- and third-generation immunoassays are highly conserved among cats, dogs, horses, and humans. 40 Even so, validation of a candidate assay is recommended before using it for diagnostic purposes in a novel species. 2 Various human cTnI assays have been validated in domestic and non-domestic animals; however, not in rhinoceros. Information about the epitopes targeted by antibodies contained in cTnI assays is available, 14 and prediction of the degree of antibody binding to these epitopes on the rhinoceros cTnI protein would assist in the screening and selection of assays to be included in full method validation studies.
We aimed to assess the potential use of several commercial candidate human cTnI immunoassays for detecting African rhinoceros cTnI. We did so by determining the mRNA transcript sequence of white and black rhinoceros cTnI and assessing sequence homology. The inferred amino acid sequences were used to evaluate the degree of conservation of the different cTnI epitopes recognized by the capture and detection antibodies used in these immunoassays.
Materials and methods
Animals and sample preparation
Samples of myocardium from the left ventricle were obtained from 2 horses, 2 white rhinoceros, and 1 black rhinoceros immediately after death or euthanasia due to causes unrelated to our study. Black rhinoceros skeletal muscle was also obtained, to serve as a negative control. The samples collected from the horses were used for optimization of the analytical procedures. For each tissue, duplicate samples of ≤5 mm in any dimension were prepared: one was transferred to a tube containing RNA stabilizer solution (NucleoProtect; Macherey-Nagel), stored at room temperature for 12–24 h, and thereafter at −80°C.
cTnI mRNA transcript analysis
Total RNA was extracted (NucleoSpin RNA midi kit; Macherey-Nagel) according to the manufacturer’s instructions. The RNA quantity and quality were assessed spectrophotometrically (Xpose; Trinean). Primers targeting the coding sequence of cTnI genes were designed based on the predicted mRNA transcript sequence of the southern white rhinoceros (NCBI reference sequence XM_004439072.2) and horses (AY819020.1) using Primer-BLAST (https://www.ncbi.nlm.nih.gov/tools/primer-blast/). 55 The forward primer was 5′-TCTTGCCCCTTCTCTGCCTC-3′; the reverse primer was 5′-TTCCTCAGGGCCATYCTCAG-3′, yielding an amplicon of 708 bp. Reverse-transcription PCR (RT-PCR) was performed (PrimeScript One Step RT-PCR kit v.2, dye plus, Takara Bio; Applied Biosystems Veriti 96-well thermal cycler, Thermo Fisher). RT-PCR conditions for rhinoceros RNA consisted of initial reverse transcription for 30 min at 50°C, followed by PCR activation for 2 min at 94°C and 30 cycles of denaturation (94°C for 30 s), annealing (55°C for 30 s), and extension (72°C for 1 min). Thereafter, the samples were kept at 4°C. The PCR conditions for the kit positive control were performed according to the manufacturer’s recommendations.
The integrity of the PCR amplicons was visualized by electrophoresis in 1.5% Tris acetate–EDTA–agarose gel stained with ethidium bromide. Amplified PCR products were subsequently submitted to Inqaba Biotec (South Africa) for purification and sequencing. Briefly, PCR products were purified with exonuclease I–shrimp alkaline phosphatase (ExoSAP; New England Biolabs) according to the manufacturer’s instructions. The purified products were sequenced (BrilliantDye terminator cycle sequencing kit v.3.1, BRD3-100/1,000 cycle sequencing kit, NimaGen; Applied Biosystems 3730xl genetic analyzer, Thermo Fisher). Generated overlapping sequences were aligned and contigs constructed using MEGA (v.11, https://www.megasoftware.net/) followed by BLAST analysis.
Commercial human cTnI immunoassay evaluation
The candidate commercial human cTnI immunoassays were evaluated based on the epitopes recognized by the antibodies, which were previously validated in other veterinary species. These immunoassays were chosen for assessing the degree of identity between different species (Table 1). Additionally, a novel point-of-care (POC) cTnI assay validated in 2022 and released for human medical use was evaluated (Atellica VTLi hs-cTnI; Siemens). 13 The degree of amino acid conservation of cTnI antibody epitopes was assessed in African rhinoceros, the cat, dog, horse, and rat as follows: the predicted amino acid sequences of the African rhinoceros cDNA nucleotide sequences from the BLAST analysis were obtained and aligned with the amino acid sequences of the cat (UniProt identifier Q863B6), dog (Q8MKD5), horse (Q5PYI0), rat (P23693), and human (P19429) using the UniProt platform (https://www.uniprot.org/align). 48 Amino acid sequence identity at the epitopes recognized by the capture and detection antibodies utilized in each immunoassay was visually assessed for amino acid differences between the species listed above.
Summary of candidate human cardiac troponin I (cTnI) immunoassays evaluated for amino acid sequence homology.
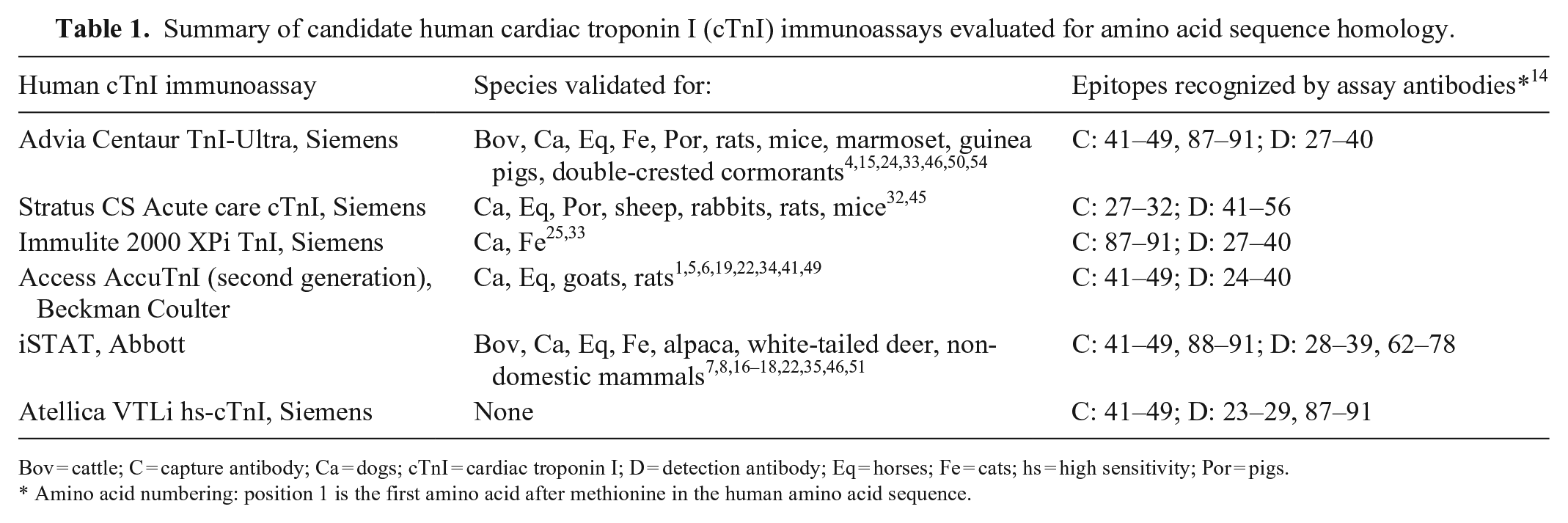

Bov = cattle; C = capture antibody; Ca = dogs; cTnI = cardiac troponin I; D = detection antibody; Eq = horses; Fe = cats; hs = high sensitivity; Por = pigs.
Amino acid numbering: position 1 is the first amino acid after methionine in the human amino acid sequence.
Results
cTnI mRNA transcript analysis
Extracted RNA yields were 21.2–47.0 ng/µL in an elution volume of 500 µL, with optical density (OD260/280) values of 2.04–2.15. Agarose gel electrophoresis of the cDNA had amplicons in base pair size consistent with cTnI (708 bp; Fig. 1)
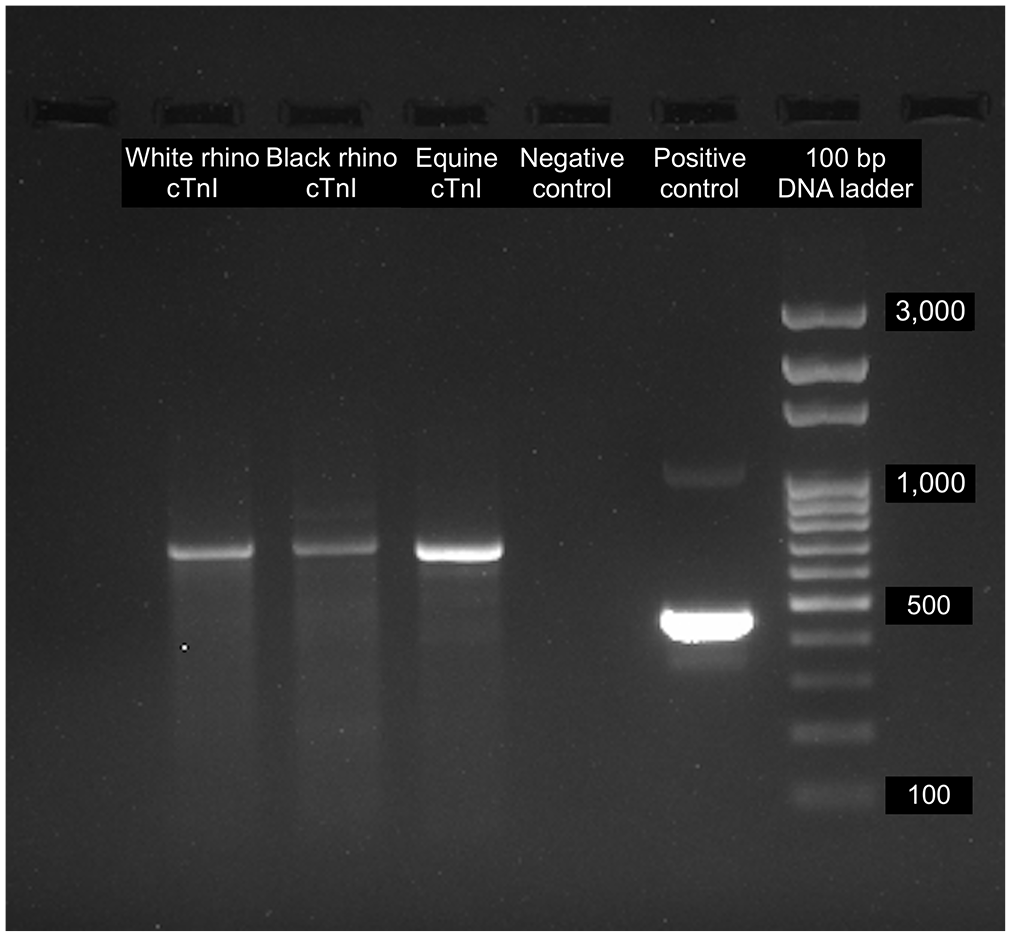
1.5% agarose gel electrophoresis of cDNA PCR products from myocardium of white and black rhinoceros and horse, amplified using cardiac troponin I (cTnI)-specific primers. A 708-bp amplicon product is present. Skeletal muscle RNA from a southern-central black rhinoceros served as a negative control. Positive control RNA (amplicon, 462 bp) included in the kit (PrimeScript one step RT-PCR kit v.2, Dye Plus; Takara Bio). rhino = rhinoceros.
Feature annotation of the rhinoceros nucleotide sequences was performed by the NCBI GenBank User Services (https://www.ncbi.nlm.nih.gov/genbank/) and identified regions confirmed by reviewing well-annotated cTnI transcripts of other domestic species. 43 The southern white rhinoceros cTnI cDNA revealed a coding sequence (CDS) of 618 nucleotides (nt). The partial nucleotide sequence of the southern-central black rhinoceros cTnI revealed a partial CDS of 612 nt. These sequences were submitted to GenBank (OR374027, OR374028).
When comparing the CDS of the cDNA sequence alignment, the percentage of identity between white and black rhinoceros was 99%. Five single nucleotide variants were identified at the following locations, with the start of numbering from the white rhinoceros CDS: C189G; C192T; C207G; G306T; T444C (Fig. 2). Comparing the cDNA sequence alignment between white and black rhinoceros versus horses, the percentage identity was 94% and 95%, respectively, and between white and black rhinoceros versus humans, 91%.
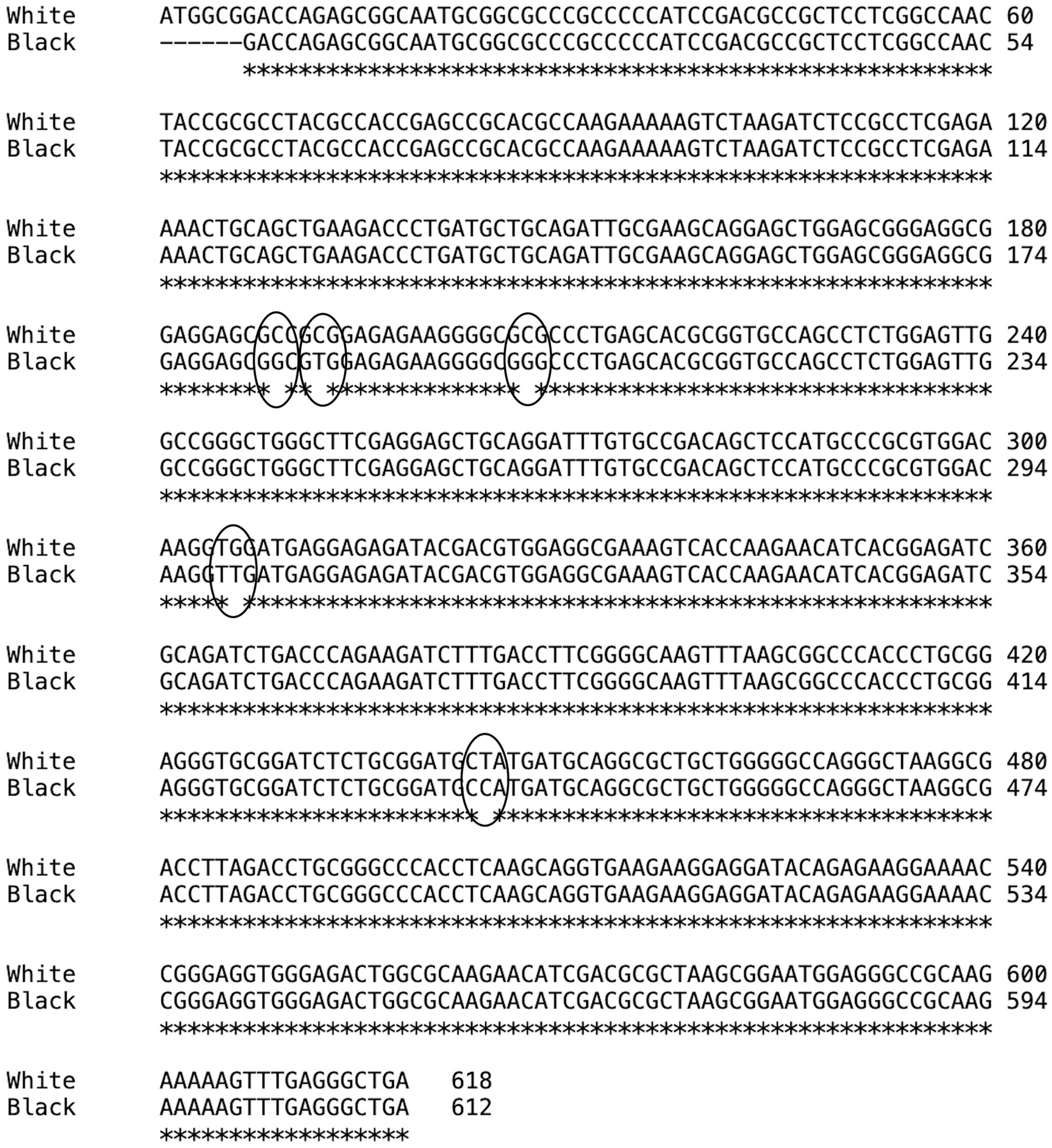
Coding nucleotide sequence alignment of cardiac troponin I mRNA from southern white (White) and southern-central black (Black) rhinoceros. Single nucleotide variants are circled.
cTnI protein analysis
Despite the 5 nucleotide variants, the inferred amino acid sequences of the 2 rhinoceros species were identical (100% identity), with 94% identity between white and black rhinoceros compared to humans (Fig. 3). The percentage identity of the rhinoceros predicted cTnI amino acid sequence compared to the cat, dog, horse, and rat was 94%, 95%, 97%, and 92%, respectively (Fig. 3).

The predicted amino acid sequence of cardiac troponin I (cTnI) from southern white and southern-central black rhinoceros aligned with the cTnI amino acid sequence of rat, human, dog, cat, and horse. The black boxes highlight gaps in the sequences, and the blue boxes highlight regions commonly targeted by cTnI immunoassay capture and detection antibodies. Amino acid numbering is based on the human sequence. The cardiac-specific N-terminal region lies from 1–31. Dash (-) = gap; asterisk (*) = sequence conserved; no dots = no sequence conservation; period (.) and colon (:) = some conservation, with colon indicating all-pyrimidines or all-purines amino acids at position.
Commercial human cTnI immunoassay evaluation
Amino acid sequence regions described here (Fig. 3) represent the epitopes targeted by the selected immunoassay antibodies (Table 1). Specific epitopes within the first amino acid sequence region (22–56) were targeted by most antibodies utilized in the immunoassays, which included 6 detection and 5 capture antibodies, and there was complete amino acid sequence homology between humans and rhinoceros at the epitopes targeted by 4 of 6 detection antibodies and 1 of 5 capture antibodies. The entire second amino acid sequence region (62–78) was targeted by a single detection antibody, with complete amino acid sequence homology between humans and rhinoceros. The short third amino acid sequence region (87–91) contained epitopes targeted by 3 capture antibodies and 1 detection antibody, and sequence homology was not present between humans and rhinoceros.
The same, single amino acid difference was noted in the epitope recognized by the single or one of the pairs of capture antibodies used in the Immulite 2000 XPi TnI, Advia Centaur TnI-Ultra, and iSTAT cTnI immunoassays in rhinoceros, horses, and rats compared to humans, cats, and dogs. For the iSTAT assay, a single amino acid difference, at different locations, was also noted in the epitope targeted by one of the pairs of detection antibodies, in the horse and rat. For the Stratus CS cTnI detection antibody, a single amino acid difference was noted in the epitope target in all species compared to humans. Regarding the Atellica VTLI, a single amino acid difference was present at the epitope for one detection antibody in all species compared to humans, and a single amino acid difference was present at the epitope bound by the second detection antibody for the rhinoceros, horse, and rat versus humans, cats, and dogs. The epitopes recognized by the capture and detection antibodies in the Beckman Coulter Access AccuTnI immunoassay were conserved across all species evaluated.
Discussion
The complete nucleotide CDS of white rhinoceros cTnI cDNA revealed a coding region of 618 nt, with a partial CDS of 612 nt found in the black rhinoceros. This is similar to the equine cTnI cDNA coding region of 618 nt but is slightly smaller than the feline and canine coding region of 636 and 633 nt, respectively.39,40 The southern white rhinoceros and southern-central black rhinoceros are closely related sister-taxa with divergence noted ~5.2 million years ago (Ma) and cessation of ancestral gene flow reported between 3.3 and 4.1 Ma.30,47 The 5 nucleotide variants that were identified when comparing the white rhinoceros cDNA sequence to that of the black rhinoceros represent synonymous codon substitution (i.e., the mutations do not alter the encoded amino acid). This is indicative of purifying selection, with synonymous mutations frequently assumed to be neutral concerning fitness. 3
The inferred amino acid percentage identity among cats, dogs, horses, humans, and rats was 92–97%. Our amino acid sequence alignment using the Clustal Omega program in UniProt showed that white and black rhinoceros cTnI have alanine in the N-terminal region, which is also present in canine, feline, equine, and rat cTnI but missing from human cTnI. 44 Interestingly, cTnI of both rhinoceros species lacked the same 6 amino acids in the N-terminal region as cTnI of horses, a finding not present in all other species studied. 40 Considering the evolution of perissodactyls, with the order including the family Equidae, it is not surprising that identical amino acid deletion was seen in all 3 of these species. 47
Measurement of blood cTnI concentration is facilitated by immunoassays utilizing cTnI-specific antibodies, and monoclonal and recombinant antibodies are considered the most suitable options for both capture and detection antibodies (Abs), offering higher specificity. 29 The use of multiple Abs targeting different epitopes (i.e., ≥2 capture Abs and ≥1 detection Ab) results in higher assay sensitivity and accuracy. 29 Antibodies specific for epitopes on cTnI located within the central region of the protein should be used given that the most stable region of cTnI is between amino acid residues 30 and 110. 20
The epitopes on the cTnI molecule that are commonly detected by second and third-generation cTnI immunoassays have been reported to be highly conserved, with only 8 amino acid differences noted among cats, dogs, horses, and humans in the sequence after Ala19,25,40 Our results confirm these findings, with only 5 amino acid differences noted in the epitope binding sites of the Abs used in the 6 candidate immunoassays among both rhinoceros species, cats, dogs, horses, humans, and rats.
Despite this high homology, 5 of the 6 immunoassays that we evaluated contain antibodies targeted against the cTnI epitopes that are not conserved between humans and several other species, including rhinoceros. Considering that the Immulite 2000 XPi cTnI immunoassay only utilizes a single capture Ab, and the short amino acid sequence of the epitope recognized by this capture Ab (epitope region 87–91; 5 amino acid length), cross-reactivity with cTnI from species other than the dog and cat (for which the assay has been validated) is highly unlikely.25,33 This supposition is supported by a reported lack of detection of equine cTnI (same amino acid sequence as rhinoceros at epitope region 87–91) by 2 immunoassays whose capture antibodies target this epitope. 41 One of these 2 immunoassays was the Immulite 1000 cTnI assay, which uses the same capture Ab as the Immulite 2000 XPi cTnI assay.
A capture Ab used in the Advia Centaur cTnI-Ultra also targets the non-conserved 87–91 epitope, but this assay has a second capture Ab (epitope 41–49) and one detection Ab (epitope 27–40) that target epitopes with complete homology. This explains why this assay has been successfully validated in many animal species, including cats, dogs, horses, and rats; the Abs in this assay could be expected to cross-react with African rhinoceros cTnI.4,24,33,54
The iSTAT cTnI assay has been used with success in many species including the cat, dog, and many non-domestic mammals (but not rhinoceros), likely because the assay utilizes 2 capture Abs8,16,18 and the epitopes recognized by one of these Abs are conserved across all the species that we evaluated. This POC assay therefore shows promise for use in African rhinoceros under field conditions.
The same single amino acid difference (leucine in position 52 replaced by methionine) was present in the epitope recognized by the detection Ab utilized in the Stratus CS Acute care cTnI immunoassay, in all evaluated species; the epitope recognized by the capture Ab was conserved. Despite the single amino acid difference in the detection Ab epitope, this assay has been validated for use in dogs, horses, mice, pigs, rabbits, rats, and sheep, and reference intervals established for cats.32,45 The epitope recognized by the Stratus detection Ab consists of 16 amino acids, with the amino acid difference located in the last third of the sequence. The degree of homology is thus sufficiently high in this relatively long epitope to allow adequate binding of the detection Ab to the captured cTnI antigen. The Stratus CS Acute care cTnI assay is therefore also potentially useful in the evaluation of serum cTnI concentration in rhinoceros.
The epitopes recognized by the capture and detection Abs in the Access AccuTnI immunoassay are conserved across all evaluated species, and the assay has been validated in dogs, goats, horses, and rats.1,5,6,19,34,41,49 The Access AccuTnI immunoassay will also potentially be useful in the evaluation of serum cTnI concentration in rhinoceros.
The Atellica VTLi hs-cTNI is a relatively new POC assay that has capture antibodies bound to superparamagnetic nanoparticles. 13 The capture antibodies target only one epitope, which was conserved across all species. Three detection antibodies potentially bind the nanoparticle–capture Ab–cTnI complex; for one of these detection antibodies, there was one amino acid difference in all species versus humans at the targeted epitope (23–29), and for the second detection Ab, there was one amino acid difference (epitope 87–91) in horses, rhinoceros, and rats compared to humans, dogs, and cats. The third detection Ab binds to binary cTnI–cTnC complexes. This new assay has not been investigated in species other than humans, but also holds promise for cTnI measurement in African rhinoceros.
A limitation of our study is that the cTnI mRNA transcript nucleotide CDS of the black rhinoceros was only determined from a single animal. Furthermore, the 5′-region of the black rhinoceros CDS was incomplete, with 6 nucleotides missing because of poor signal strength of the sequencing data. Obtaining fresh tissue samples from this critically endangered species is challenging. However, a comparison of our cTnI mRNA transcript sequences to the recently added predicted Diceros bicornis minor cTnI mRNA sequence (NCBI reference sequence XM_058530778.1) revealed 100% identity.
By evaluating the homology of rhinoceros cTnI amino acid sequences compared to that of humans and other animal species and using information about the epitopes targeted by cTnI assay Abs, we were able to screen available assays before embarking on further investigations. We identified that 5 of the 6 candidate cTnI immunoassays evaluated may be useful for the measurement of blood cTnI concentration in African rhinoceros and could be selected for full analytical method validation studies in the future. The eventual establishment of reference intervals in healthy populations and assessment of cTnI concentrations using a validated assay in animals at risk for cardiomyocyte injury will enhance the welfare of these animals during conservation-associated activities.
Footnotes
Acknowledgements
We thank Dr. Charles Byaruhanga for his assistance with the RNA extraction and RT-PCR method optimization.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by the Health and Welfare Sector Education and Training Authority (HWSETA) of South Africa. The HWSETA was not involved in the study design, collection, analysis, and interpretation of data; the writing of the manuscript; or the decision to submit the manuscript for publication.
