Abstract
We performed a diagnostic disease investigation on a cohort of coho salmon (Oncorhynchus kisutch) fingerlings in Alaska exhibiting anorexia, gaping mouths, anemia, and increased mortality. Histologic examination revealed mild-to-severe myocardial degeneration and lymphohistiocytic and neutrophilic myocarditis, moderate splenic histiocytosis, and mild renal histiocytosis. Piscine orthoreoviruses 1 and 3 were not detected by molecular methods, and no other viruses could be cultured on 3 common diagnostic fish cell lines. De novo assembly produced a viral genome of 10 linear segments with >80% homology to piscine orthoreovirus 2 (PRV2) encoding all 11 PRV2 proteins. An in situ hybridization probe using RNAscope was developed against 697 viral nucleotides identified by sequencing, which revealed viral genome in heart, spleen, gill, kidney, liver, blood, and the lamina propria of the intestines. Our findings are supportive of a novel piscine orthoreovirus most closely related to PRV2 associated with morbidity and mortality of coho salmon in the northeastern Pacific.
Keywords
Piscine orthoreovirus (PRV; Spinareoviridae, Piscine orthoreovirus) is a segmented, double-stranded RNA virus associated with cardiovascular and systemic diseases in salmonid fishes. Genomic phylogeny supports 3 genotypes of PRV: PRV1 (found in the northern Atlantic, northeastern Pacific, and Chile), PRV2 (in Japan), and PRV3 (in the northeastern Atlantic and Chile).7,22 Although many full-length and partial genomes have been published for PRV1 and PRV3, only one genome has been published for PRV2 despite more than a decade of PRV sequencing. 26 In the Pacific Northwest of North America, where PRV1 is endemic, PRV has been detected in wild and farmed salmonids absent of apparent disease or mild myocarditis. 22 An association has been suggested for PRV1 with heart and skeletal muscle inflammation (HSMI) in Atlantic salmon (Salmo salar) and HSMI-like disease in coho salmon (Oncorhynchus kisutch) in Norway and Chile. 8 HSMI has also been described in association with naturally PRV3-infected rainbow trout in Norway, and challenge studies have demonstrated variably reproducible disease in rainbow trout and Atlantic salmon.19,29,30 In Japan, PRV2 RNA was detected in peripheral blood and tissues of fish sampled during an epizootic event of erythrocytic inclusion body syndrome (EIBS) in coho salmon, and development of erythrocyte cytoplasmic inclusion bodies was observed alongside increasing PRV2 RNA in peripheral blood of EIBS-naïve fish after experimental inoculation with infectious material purified from EIBS-affected fish. 26 It is important to note that, although both HSMI and EIBS have been associated with the detection of PRV materials, controlled laboratory infections have failed to recapitulate diseases, suggesting that additional factors may be implicated in causing these diseases in salmonids. 26 EIBS conditions have been identified in Pacific Northwest salmonids, although different causes have been implicated.15,17,24
PRV is not currently reportable to the World Organisation of Animal Health (WOAH), and evidence suggests that each genogroup may be endemic in its respective niche. Due to the possibility that PRV contributes to diseases with significant mortality, fish health professionals in affected regions are continuously vigilant. Our objectives here were to describe the viral genomics, tissue distribution, and pathology associated with an orthoreovirus detected by sequencing of diseased coho salmon during a disease investigation in which increased morbidity and mortality were of concern.
Whole bodies of 12 coho salmon fingerlings 10–12.5 cm long from the southeastern region of Alaska were submitted on ice to the Washington Animal Disease Diagnostic Laboratory (WADDL; Washington State University, Pullman, WA, USA) for autopsy and disease investigation because of an increase in morbidity and mortality in January 2021. The submitter reported that animals had anorexia of 4-mo duration, pale gills, gaping mouths, and hematocrits of <0.10 L/L (<0.20 L/L = anemia). No parasites were observed on skin scrapes, and bacteria were not identified cytologically in systemic organs. Fish were randomly segregated into 3 equal groups, and fresh tissue pools of heart, spleen, and kidney were collected. Complete sets of systemic organs were also collected in identically pooled groups and preserved in 10% neutral-buffered formalin. Fixed tissues were processed and embedded in paraffin wax within 24 h. H&E-stained slides were prepared from 9 fish and included gills, heart, kidney, intestines, pancreas, liver, and spleen.
Histologic examination revealed mild-to-severe myocardial degeneration and lymphohistiocytic and neutrophilic myocarditis in both the compact and trabecular myocardium (Fig. 1) in 4 of 9 fish, moderate splenic histiocytosis in 4 of 7 fish (2 spleens were lost in processing), and mild renal histiocytosis in 5 of 9 fish. Given the histopathologic findings, tissue pools were tested for PRV1 and PRV3 by a reverse-transcription PCR (RT-PCR) assay.
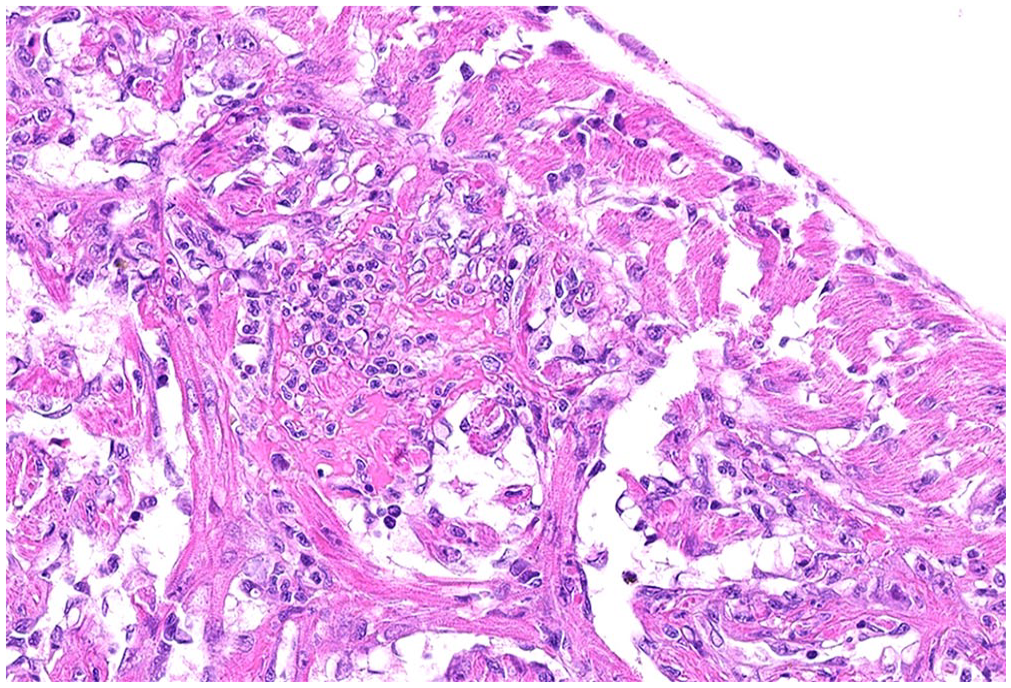
Myocardial degeneration and lymphohistiocytic and neutrophilic myocarditis in both the compact and trabecular myocardium of a coho salmon (Oncorhynchus kisutch) fingerling. H&E.
For molecular testing, a 10% tissue homogenate was prepared using 0.1 g sampled from each tissue pool homogenized in 900 µL of Dulbecco modified eagle medium (DMEM; Invitrogen) with silica beads for 2 min (Mini-Beadbeater-96; BioSpec). To clear cell debris, homogenates were centrifuged for 5 min at 4,000 × g at 4°C. Total nucleic acids (TNAs) were extracted from 200 µL of the homogenate supernatant (MagMAX-96 viral RNA isolation kit; ThermoFisher) per the manufacturer’s instructions, with 65°C warmed elution buffer. TNAs were subjected to PRV1 and PRV3 assays, as described previously.19,20 PRV1 and PRV3 were not detected in any of the pooled tissue samples. With no identified cause of disease, and histopathology suggestive of infection, this case was recommended for investigation by virus isolation (VI) and next-generation sequencing.
VI was attempted from each tissue pool homogenized following methods in the Manual of Diagnostic Tests for Aquatic Animals. 32 Homogenates (10%) were gently vortexed to re-suspend, and then cleared again by centrifugation for 15 min at 2,800 × g at 4°C to pellet cell debris. Supernatants were used to prepare two 10-fold dilutions so that 10%, 1.0%, and 0.1% homogenate supernatants were used as inoculums for VI. Cleared supernatants were used to inoculate epithelioma papulosum cyprini (EPC), Chinook salmon embryo-214 (CHSE), or fathead minnow (FHM) cells previously seeded onto tissue culture–treated, clear, 24-well cell culture plates (Costar; Corning) with the following modifications. 5 EPC cells were pretreated with 50 µL of 7% polyethylene glycol (in MEM-HEPES [Corning] + 1% Pen Strep [MilliporeSigma] + 10% Fungizone [MilliporeSigma]) for 1 min before inoculation with 100 µL of the 10%, 1.0%, and 0.1% diluted inoculum; 100 µL of the 10% and 1.0% inoculum was added to CHSE and FHM plates, 1 well per dilution of each sample. Each plate included 2 negative control wells inoculated with 100 µL of processing medium only. CHSE-214 cells were fed with RPMI-5 + 1% Pen Strep, 0.1% gentamicin + 0.5% Fungizone. FHM cells were fed MEM-Hank (Corning) + 1% Pen Strep, 0.1% gentamicin + 0.5% Fungizone. EPC cells were treated with MEM containing 0.6% methylcellulose, 0.3% tricine, and 0.01% sodium bicarbonate, 0.5% gentamicin, and 1% Fungizone + 1% Pen Strep + 1% L-glutamine (Gibco GlutaMAX; ThermoFisher). Inoculated plates were incubated at 15°C for 14 d, with each sample well observed 3 times per week for cytotoxic effect (CTE), microbial contamination, and cytopathic effect (CPE). At the incubation period of the second passage, no CPE was observed in any well, and the VI attempt was concluded. CTE and microbial contamination were not observed in either primary or subculture throughout the incubation periods.
For whole-genome sequencing (WGS), homogenates prepared for single-agent PCR and VI assays above were thawed on ice and vortexed to resuspend. After clearing centrifugation (15 min at 32,800 × g at 4°C), the supernatant was mechanically filtered through a 0.45-µm syringe filter. TNAs were extracted from 500 µL of filtered supernatant (QIAmp UltraSens Virus kit; Qiagen). A fraction of the purified TNA (11 µL or up to 500 ng of viral genomic RNA and total mRNA) was taken for double-stranded complementary DNA (ds-cDNA) synthesis by sequence-independent single-primer amplification (SISPA) to determine if the SISPA procedure affected detected viral reads. 2 We used 48 µL of synthesized ds-cDNA as templates for library prep (Ligation sequencing kit SQK-LSK109; Oxford Nanopore Technologies [ONT]) using the included short fragment buffer with a ratio of 1:1 sample volume-to-beads (AMPure XP; Beckman Coulter) to clean the end-prepped template in the first step and 1:0.6 sample-to-beads to clean up the adapter ligation reaction. Based on the mean fragment length of previous runs (500 bp), libraries quantified by Qubit (ThermoFisher) were normalized to 50 fmol in 12 μL for loading (16.2 ng/μL). Prepared libraries from each tissue pool were sequenced individually (new MinION flow cells; R9.4.1; ONT) having at least 800 health pores, run with no reloading, on a sequencer (GridION; ONT) for up to 72 h. The mode read length was ~350 bp, which contributed to above-normal pore attrition rates.
For viral discovery, raw reads were classified by translated amino acid sequence similarity to virus-specific proteins using Diamond BLASTx as described 5 with the Reference Viral Database (RVDB v.21.0) 9 and by map-filtering against all NCBI viral RefSeq (“Viruses”[Organism] NOT “cellular organisms”[Organism] NOT wgs[PROP] NOT gbdiv syn[prop] AND (srcdb_refseq[PROP] OR nuccore genome samespecies[Filter]) AND refseq[filter]; retrieved 2021 Feb 18). Diamond translated nucleotide search against the RVDB protein database with - - max-target-seqs = 1 and default scoring parameters returned 152,483 high-scoring query-subject pairs in BLAST tabular format (- - outfmt 6). Subject IDs (column 2, sseqid in Diamond output) were sorted and deduplicated, then used as a namelist to pull the full sequence header from the database fasta file (seqtk subseq U-RVDBv21.0-prot.fasta name.lst | grep ‘>’ > uniqViralProteins_fullnames.list; https://github.com/lh3/seqtk). The viral protein list was further deduplicated, resulting in a list of 474 viruses. This list of viruses was manually sorted to rank each virus by the probability of infection, where “likely” included viruses infecting fish and aquatic vertebrates, “somewhat likely” included viruses infecting terrestrial vertebrates, and “least likely” was everything else including phage, phytoviruses, and viruses infecting insects (Suppl. Table 1). Starting with viruses infecting fish and aquatic vertebrates, raw reads were re-mapped to individual candidate genomes, one at a time, prioritized by their hit frequency from the Diamond output. 1 Diamond BLASTx identified 3,355 reads with significant identity to all 11 proteins of PRV2. 26 All raw reads mapped against PRV2 produced nearly full coverage of each segment (Fig. 2); raw reads mapped to segments of PRV1a/b (Figs. 3, 4) and PRV3 (Fig. 5) produced low-coverage, low-quality alignments. Viral reads covering candidate viral genomes were de novo assembled using Canu v.2.1.1 12 during the discovery stage to avoid reference bias, and these assembled contigs were blasted against the NCBI nr/nt database to ensure they only matched PRV2. A reference-guided draft assembly was built by Pomoxis v.0.3.6 (https://nanoporetech.github.io/pomoxis/) using the published PRV2 genome as a guide, for comparison of ORFs and segment lengths. 10
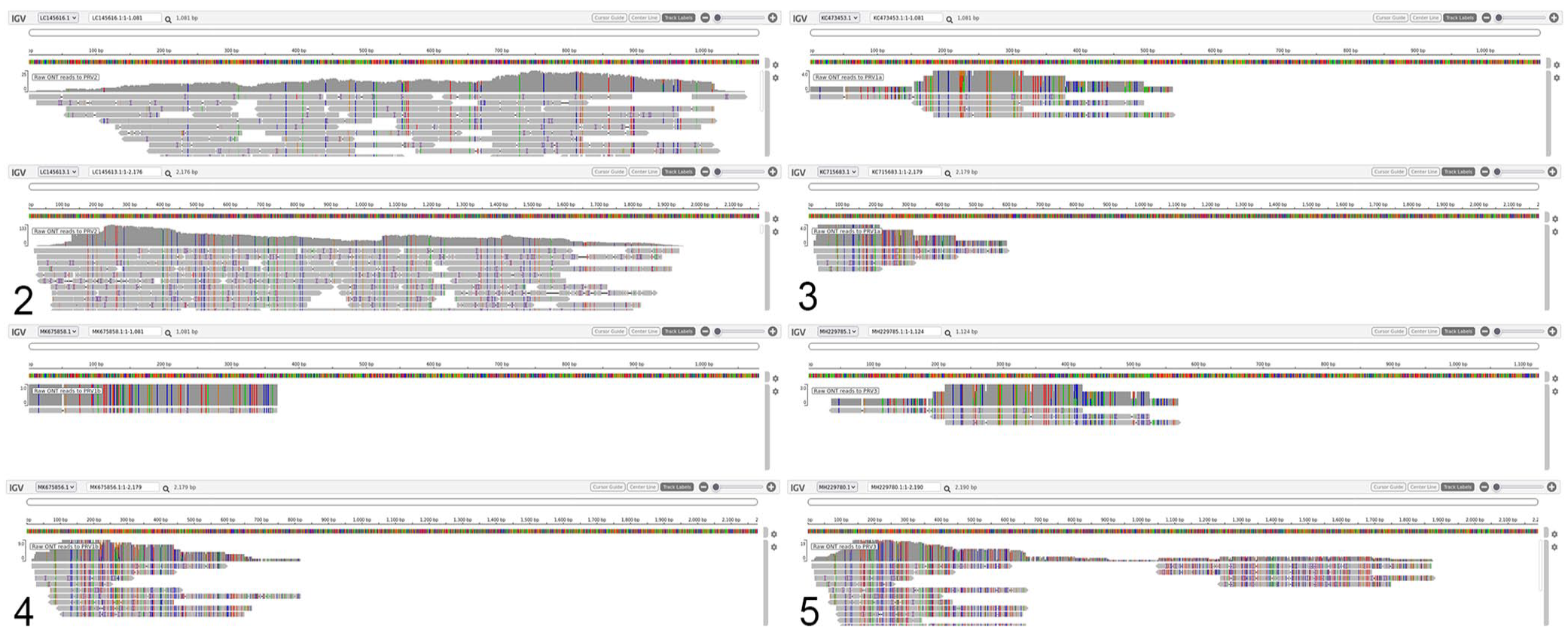

Raw Oxford Nanopore Technologies (ONT) reads mapped to piscine orthoreovirus (PRV) segment S1 (upper panel) and M2 (lower) using minimap2, visualized on IGV (https://igv.org/app/), shows high concordance with PRV2 (Fig. 2) and low concordance with PRV1a (Fig. 3), PRV1b (Fig. 4), and PRV3 (Fig. 5). Colored positions highlight discordance of basecalls against reference, and gray indicates concordance to reference.
Of the 3 tissue pools, only tissue pool 1 produced significant coverage of the PRV2-like virus by WGS (Table 1). Pool 2 produced 6 reads mapping to PRV2, a total of 3.3 kbp mapped across segments LC145613.1 (segment M2) and LC145609.1 (segment L3); pool 3 produced a single read that covered 236 bp of LC145610.1 (segment L1). For genome finishing, 2 additional rounds of ONT sequencing on new MinION flow cells as above, and an Illumina library, were generated from tissue pool 1 using the same ds-cDNA that was used to make the first ONT library, following methods as described previously. 6 Briefly, sequencing adapters were added to SISPA-cDNA (DNA Prep library kit; Illumina) and sequenced on a full lane of the PE300 cycle kit (MiSeq v.3; Illumina) producing 27,064,820 short reads; 6,264 reads mapped to PRV2 with 90–99% coverage of each reference segment and mean depth of 7.6–88.6 reads per base, with segment length loosely correlating with depth at non-zero coverage positions (R2 = 0.91). Hybrid assembly was performed in a short-read first approach with Unicycler v.0.4.8 (https://github.com/rrwick/Unicycler) and polished by Racon v.1.5.0 28 and Medaka v.1.2.1.
Pairwise nucleotide and amino acid similarities between a novel piscine orthoreovirus 2–like virus and representative PRV strains.
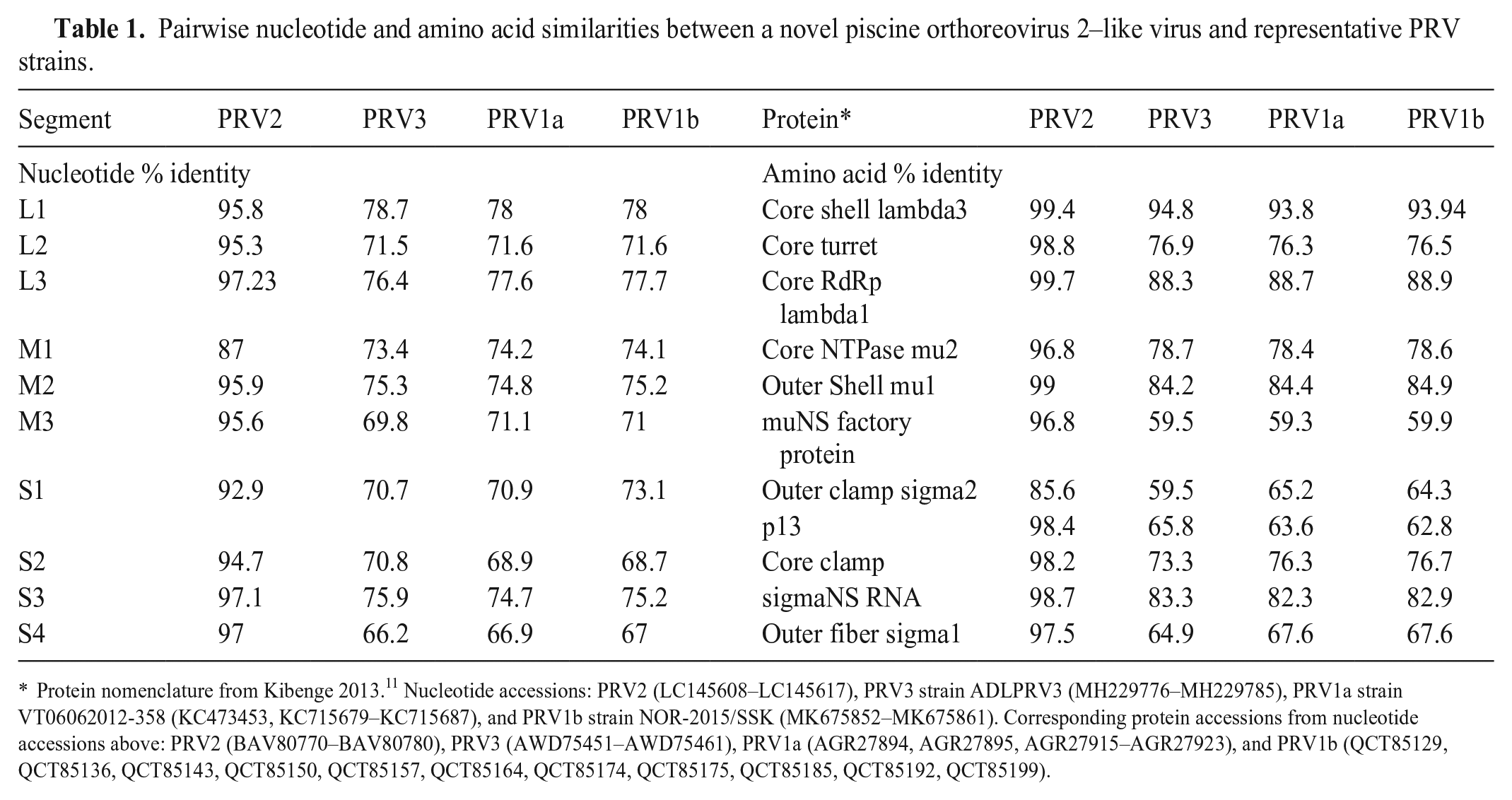
Protein nomenclature from Kibenge 2013. 11 Nucleotide accessions: PRV2 (LC145608–LC145617), PRV3 strain ADLPRV3 (MH229776–MH229785), PRV1a strain VT06062012-358 (KC473453, KC715679–KC715687), and PRV1b strain NOR-2015/SSK (MK675852–MK675861). Corresponding protein accessions from nucleotide accessions above: PRV2 (BAV80770–BAV80780), PRV3 (AWD75451–AWD75461), PRV1a (AGR27894, AGR27895, AGR27915–AGR27923), and PRV1b (QCT85129, QCT85136, QCT85143, QCT85150, QCT85157, QCT85164, QCT85174, QCT85175, QCT85185, QCT85192, QCT85199).
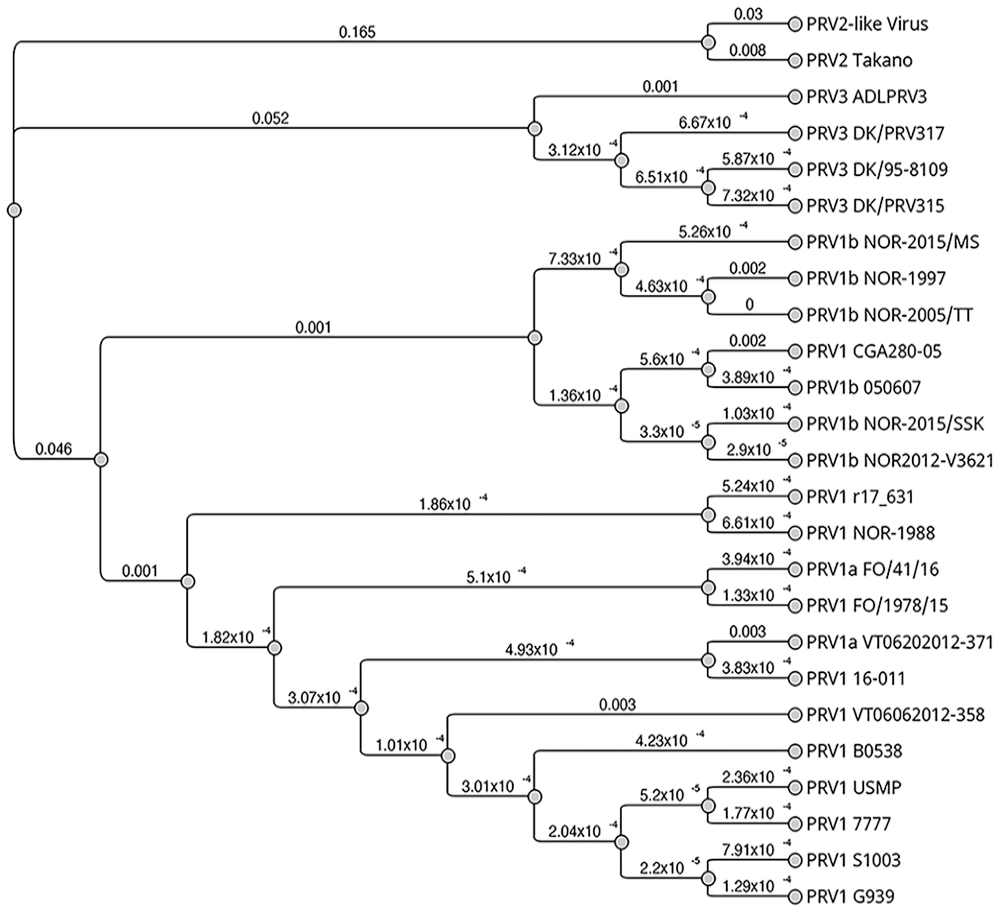
The polished, de novo assembled draft genome for this new PRV2-like virus contained 10 segments, each with ORFs encoding protein homologous to those of other PRVs (Table 1). Assembled contigs from the draft genome were deposited in GenBank (OR872657–OR872666). Segments were compared first based on coding sequence similarity, then by nucleotide identity between segments encoding the same (or similar) proteins. PRV isolates (n = 24) submitted to GenBank complete with 11 proteins were concatenated and aligned using Clustal Omega v.1.2.3 (http://www.clustal.org/omega/) using full-distance matrix calculation for guide tree construction and alignment iteration. A Jukes–Cantor neighbor-joining phylogenetic tree was constructed using Geneious Tree Builder on Geneious Prime 2023.1.1 (Dotmatics; Fig. 6). The concatenated coding sequences of our PRV2-like virus were 88.9% identical to that of a published PRV2, 26 and on average 72.3% identical to PRV1 and 72.2% identical to PRV3 viruses used for the phylogeny (Suppl. Table 2).
Once the PRV2-like virus was identified by WGS, previously extracted TNAs were subjected to PRV2 conventional RT-PCR as described previously, followed by Sanger sequencing if a band was visualized in PCR products run on 1.5% agarose gel with 0.3% ethidium bromide. 26 To establish a deployable and existing PCR machine–compatible real-time assay for PRV2 at WADDL, we developed a new hydrolysis probe real-time RT-PCR. Working in the same locus as described previously, 26 which had high coverage in our untargeted sequence, we developed real-time primers and probe targeting 164 bp of the L1 segment of PRV2 (GenBank LC145610), using Primer3.13,14,27 The forward primer starts at nucleotide 1088 (5′-ATCGTCATGGCGGTGAAGTT-3′) of the L1 segment, the reverse primer ending at nucleotide 1252 (5′-TTAGCGTTGAGTGTGCGTCT-3′), and an internal probe covering bases 1167:1189 (5′FAM-CGGTGTCTGGTCGTCTCTTGCGT-BHQ 3′). Real-time RT-PCR was performed (ABI 7500 Fast, v.1.5.1; Thermo Fisher) with a final volume of 25 µL, consisting of 5 µL of denatured dsRNA template, 12.5 µL of RT-PCR buffer (AgPath ID one-step kit; ThermoFisher), 1 µL of RT-PCR enzyme mix, 1.67 µL of detection enhancer, 12.5 µmol of primers, and 7.5 µmol of probe using cycling parameters as described previously. 6 Real-time RT-PCR was performed on extracted TNA prior to SISPA treatment to demonstrate that viral cDNA were used as a template for sequencing library prep. PRV2-like genome was detected in all 3 of the coho salmon tissue pool samples before and after SISPA treatment, although the Ct values were higher in all SISPA-treated extracts. Interestingly, a lower Ct value did not appear to increase viral reads by WGS. Samples from this case also had detectable PRV RNA when assayed with the Pan-PRV assay performed as described previously (Table 2). 33
Ct values of a novel piscine orthoreovirus 2–like (PRV2-like) genome in extracted nucleic acids by real-time RT-PCR, before and after SISPA treatment, and detection by conventional PCR, compared to all reads and bases by untargeted long-read sequencing.
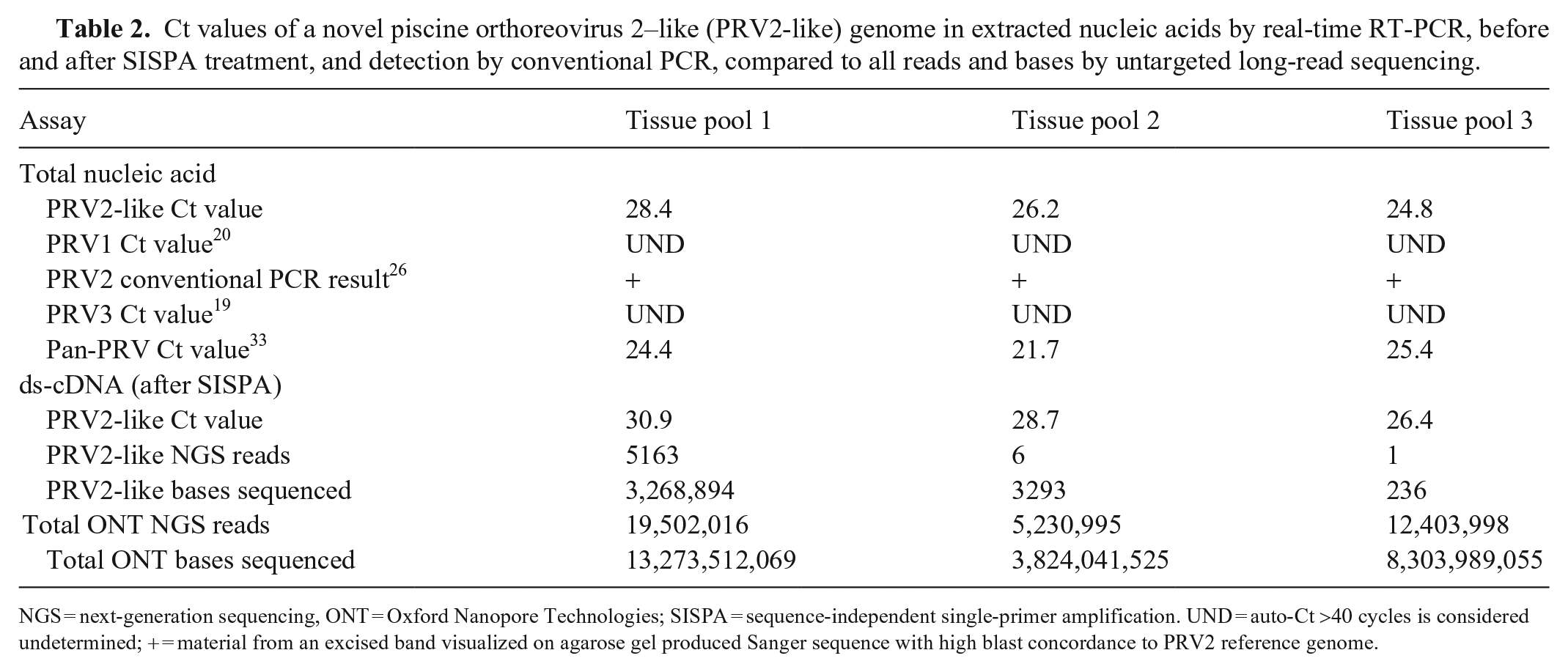
NGS = next-generation sequencing, ONT = Oxford Nanopore Technologies; SISPA = sequence-independent single-primer amplification. UND = auto-Ct >40 cycles is considered undetermined; + = material from an excised band visualized on agarose gel produced Sanger sequence with high blast concordance to PRV2 reference genome.
We used in situ hybridization (ISH; RNAscope, Advanced Cell Diagnostics [ACD]) to visualize viral RNA in tissues harvested from the 9 fish embedded in paraffin wax. A set of anti-sense specific RNA probes comprised of 20 Z pairs targeting a 697 nucleotide-long segment of the L1 region of the viral genome characterized by metagenomic WGS was developed by ACD and performed as described previously for 2.5 HD Red Detection with the following specific conditions. 5 To test the specificity of the PRV2-like probe, the probe was applied to various control fish including 2 Atlantic salmon of unknown age and size submitted to WADDL for HSMI investigations and diagnosed previously with PRV1 infection, a 1-y-old Atlantic salmon experimentally inoculated with infectious salmon anemia virus (ISAV) from a previous research investigation, and a 1-y-old Atlantic salmon that was negative for ISAV, PRV1, and PRV2-like virus from the same investigation. 5 Paraffin-embedded tissues from diseased and control animals were sectioned at 4 µm on charged slides. Samples were submerged in target retrieval solution for 15 min, followed by incubation with protease plus at 40°C for 20 min. A positive control probe for salmon gene peptidylprolyl isomerase B (PPIB, GenBank NM_001140870.2) and a negative control probe for the DapB gene of Bacillus subtilis (GenBank EF_191515) were applied to test and control tissues.
ISH proved to be a very useful tool to detect PRV2-like genomic material in histologically abnormal fish; PRV2-like genome was not detected in fish without histologic lesions except for one fish without obvious myocarditis that had a positive ISH result. ISH detected PRV2-like genomic material in 5 of 9 fish (Figs. 7–12). Four of the ISH-positive hearts had obvious myocarditis on H&E, and interestingly, one ISH-positive heart did not. Positive reactivity in hearts was widespread in the trabecular myocardium, not present in the outer compact myocardium, greater in the ventricle compared to atrium, and, in hearts that contained blood, there was abundant reactivity in blood cells (Fig. 8). PRV2-like genomic material was detected in 5 of 9 kidneys tested, which corresponded with the kidneys with renal histiocytosis (Figs. 9, 10). The hematopoietic tissue of the kidney interstitium had moderate multifocal reactivity among hematopoietic cells, and renal tubules did not have reactivity (Fig. 10). Four of the 7 tested spleens had positive detection of PRV2-like genome. The spleens exhibited histiocytosis and had widespread multifocal ISH positive reactivity, although specific cells could not be identified (Figs. 11, 12). In the gills, reactivity was in the primary and secondary lamellae predominantly in regions of blood vessels, and rarely more superficially. Livers that had positive reactivity had localization to hepatocytes, presumed Kupffer cells, and intravascular blood cells. Positive reactivity was mild when observed in the gastrointestinal tracts, and localized to the lamina propria of the stomach and pyloric ceca. It was not possible to match the positive or negative ISH tissues with any particular fish because tissues from multiple fish were present on the stained slides.
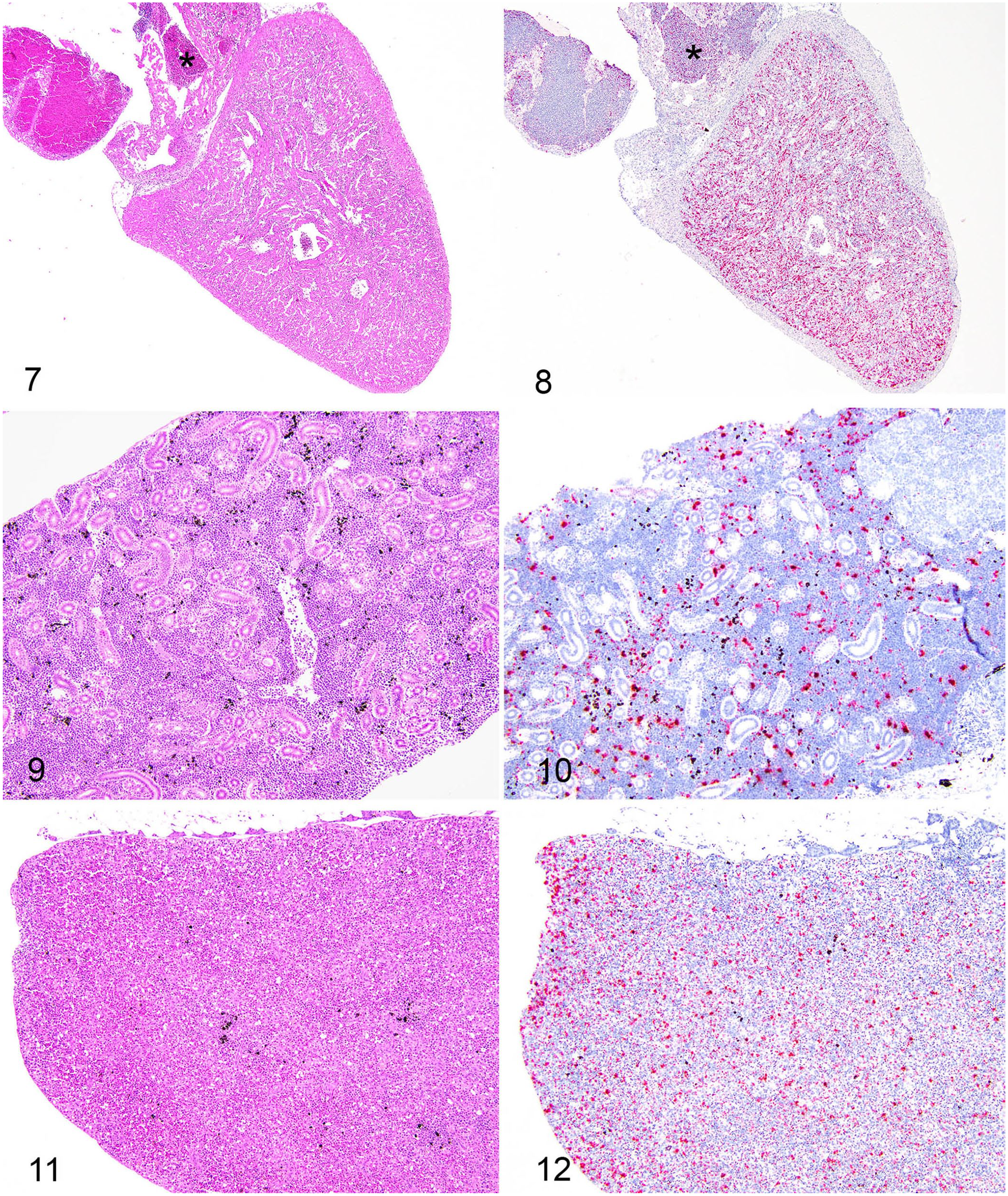
Matching H&E histopathology and tissue distribution of piscine orthoreovirus 2–like (PRV2-like) virus by in situ hybridization (ISH) from multiple infected fish.
Test tissues had positive reactivity with the positive control probe, confirming intact RNA in tissues, and did not detect the DapB gene (negative control). Neither the PRV2-like genome or DapB were detected in the PRV1-infected, ISAV-infected, and uninfected control fish, in each of which intact RNA was detected with the salmon PPIB positive control probe.
Although detection of this novel PRV2-like virus in coho salmon does not prove causation, localization of viral RNA by ISH within the histologic lesions identifies a strong association worthy of further investigation. 18 We used various testing methods in this investigation, including histopathology, VI, real-time RT-PCR, WGS, and ISH. Of these methods, we found that real-time RT-PCR and ISH were of the greatest utility when attempting to detect the PRV2-like virus. Detection of this variant with our new PRV2-like RT-PCR assay is correlated with positivity by a pan-PRV assay, 33 although we did not evaluate sensitivity due to limited samples. Histopathology was also helpful for detecting lesions; however, myocarditis is not specific for this virus. In addition, ISH revealed viral genomic material in one heart without evidence of myocarditis. WGS proved to be important in the discovery and characterization of the virus; however, there were samples with very few viral reads to use this method as an affordable or technically simple test. While it was not determined if the Alaskan coho salmon fingerlings in our case had evident erythrocytic inclusion bodies, the genomic similarity and presence of cardiac inflammation and anemia are similar to that described for Japanese coho salmon that tested positive for PRV2. 26 There is also strong similarity in cardiac lesions, tissue distribution of virus by ISH, and anemia seen in PRV1-infected Chinook salmon in British Columbia, but different from PRV1-infected Chinook salmon in that they have degenerative-necrotic lesions in the kidney. 4 Contrarily, it is important to note that other observational studies have demonstrated weak associations between PRV1 infection and anemia, erythrocytic inclusions, and myocardial lesions in coho salmon. 25
In a 2018 survey of salmon from commercial and enhancement hatcheries in the northeastern Pacific, PRV RNA was detected in only 4 of 757 coho salmon tested with the pan-PRV assay. 23 PRV1 was confirmed in these samples using a 2010 RT-PCR assay 20 followed by amplicon sequencing, although possible coinfections were not investigated once PRV1 was detected. 23 More sampling is needed to determine the geographic distribution, affected populations, and evolutionary timeline of our isolated virus. Our case further highlights the complexity of natural PRV infections, as different strains of PRV have been associated with various disease complexes in different species, often with overlapping lesions, and the prevalence of the virus in fish without apparent disease also confounds its contribution to those diseases.3,16,21,31 As well, factors such as management, coinfections, environmental stressors, or breed-specific host genetics may contribute to the overall virulence of PRV. 21 This highlights gaps in knowledge of both viral kinetics and disease progression, and further studies will be necessary to elucidate the epidemiologic trajectory of this agent in the northeastern Pacific.
Supplemental Material
sj-pdf-1-vdi-10.1177_10406387241250119 – Supplemental material for Detection, sequencing, and tissue distribution of piscine orthoreovirus 2–like virus in diseased coho salmon in Alaska
Supplemental material, sj-pdf-1-vdi-10.1177_10406387241250119 for Detection, sequencing, and tissue distribution of piscine orthoreovirus 2–like virus in diseased coho salmon in Alaska by Chrissy D. Eckstrand, Brandi K. Torrevillas, Rebecca M. Wolking, Daniel S. Bradway, Joetta Lynn Reno, Kathleen M. McMenamin-Snekvik and Kevin R. Snekvik in Journal of Veterinary Diagnostic Investigation
Supplemental Material
sj-pdf-2-vdi-10.1177_10406387241250119 – Supplemental material for Detection, sequencing, and tissue distribution of piscine orthoreovirus 2–like virus in diseased coho salmon in Alaska
Supplemental material, sj-pdf-2-vdi-10.1177_10406387241250119 for Detection, sequencing, and tissue distribution of piscine orthoreovirus 2–like virus in diseased coho salmon in Alaska by Chrissy D. Eckstrand, Brandi K. Torrevillas, Rebecca M. Wolking, Daniel S. Bradway, Joetta Lynn Reno, Kathleen M. McMenamin-Snekvik and Kevin R. Snekvik in Journal of Veterinary Diagnostic Investigation
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to research, authorship, and/or publication of this article.
Funding
Funding for this investigation was from the U.S. Department of Agriculture–National Animal Health Laboratory Network (NAHLN; grant AP20VSD&B000C015), awarded to Chrissy D. Eckstrand.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
