Abstract
Rabies virus (RABV; Lyssavirus rabies) is a neurotropic virus that can be transmitted to mammals by the hematophagous bat Desmodus rotundus. An accurate, accessible method for the detection of RABV in cattle is necessary in Paraguay; thus, we evaluated the detection of RABV using 4 techniques: fluorescent antibody test (FAT), immunochromatography rapid detection test (RDT; Anigen Rapid Rabies Ag test kit; Bionote), a reverse-transcription PCR (RT-PCR) assay, and histologic lesions in different portions of the CNS of 49 Paraguayan cattle to determine the most sensitive and specific technique. By FAT and RDT, 15 of 49 (31%) samples were positive. By RT-PCR amplification of N and G genes, 13 of 49 (27%) and 12 of 49 (25%) were positive, respectively. RDT had high agreement with FAT (kappa = 1); sensitivity was 100% (95% CI: 97–100%) and specificity was 100% (95% CI: 99–100%). The amplification of the N and G genes resulted in substantial agreement (kappa of 0.9 and 0.8, respectively) compared with FAT, and the sensitivity and specificity of the N gene were 87% (95% CI: 66–100%) and 100% (95% CI: 98–100%), respectively, and those of the G gene were 80% (95% CI: 56–100%) and 100% (95% CI: 98–100%), respectively. Histologic lesions observed were lymphoplasmacytic meningoencephalitis, gliosis, and neuronophagia. The agreement observed between the FAT and RDT tests suggests that RDT is an accurate tool for the detection of RABV. Histopathology can be used to confirm lesions caused by RABV and to rule out other conditions; the RT-PCR assay is useful for molecular epidemiology studies.
Rabies virus (RABV; Rhabdoviridae, Lyssavirus rabies), the causative agent of classic rabies in all mammals, including humans, is a neurotropic virus that causes fatal encephalomyelitis once the infection is established and reaches the CNS. RABV is distributed world-wide among various species of mammals, including carnivores and bats, which are the main reservoirs. 27 Hematophagous bats are widely distributed in Latin America and transmit RABV to different species of herbivores, mainly cattle and equids. 9 Hematophagous bats, Desmodus rotundus, feed preferentially on cattle. Infected cattle may develop signs of restlessness or excitement with sudden onset of paraplegia, which progresses to the forelimbs. Drooling is commonly observed but is thought to be due to swallowing difficulties rather than excessive production of saliva. Wasting is observed in animals that survive longer, but the disease is invariably fatal. 8
Transmission of RABV by bats to animals of economic interest is one of the most serious issues concerning rabies in Paraguay. 17 A national campaign against bovine rabies has been available since 1978 in Paraguay, and in 2015, the National Rabies Control Program in susceptible animals was approved, establishing that the control of rabies in the country is carried out through the vaccination of susceptible species, which is mandatory in high-risk areas. 21 However, data obtained from the annual statistics of the National Service of Quality and Animal Health (SENACSA) in 2021 reported 43 of 342 (12.5%) positive animals in Paraguay, of which 31 of 43 (72%) were bovids, indicating that the prevalence was high in cattle in that year. 20
The development of a RABV detection method that is fast, accurate, and accessible is highly desirable, considering the limited access to specialized veterinary services in remote parts of Paraguay. Currently, the most widely used test for the detection of RABV is the fluorescent antibody test (FAT), which is the reference test recommended by the World Organisation for Animal Health and the World Health Organization. 26 The FAT is based on the detection of viral antigen in impressions of different portions of the CNS stained with a fluorophore bound to conjugated antibodies and observed through a fluorescence microscope. 4
However, other viral antigen detection techniques, such as rapid detection testing by immunochromatography (RDT), are alternatives to conventional detection methods and have the potential to strengthen prevention and control efforts in low-resource situations or cases of inaccessibility. 14 RDT offers qualitative detection of viral antigens through the use of CNS tissue, 1 and has been shown to have high concordance with the FAT.5,12,22,24 A commercial RDT (Anigen rapid rabies Ag test kit; Bionote) has exhibited very good performance in the detection of different RABV strains, achieving almost perfect agreement with the FAT and good reproducibility in interlaboratory studies. 19
Molecular techniques such as reverse-transcription PCR (RT-PCR) have become an important tool in the diagnosis of rabies given that they can detect RNA in samples with not only a low viral load but samples in advanced states of decomposition.2,16 RT-PCR is considered an auxiliary method of detection given its high sensitivity.3,6,22 Also, real-time RT-PCR is a highly accurate method to detect RABV RNA, and is useful to quantify the viral load even with low viral concentrations.4,18,23 N gene–based RT-PCR has been used widely for detection purposes, because the N gene is one of the most conserved regions in RABV, and it can be used for the identification of different genetic lineages. 23 Detection of the gene encoding the G protein has been used for the investigation of viral pathogenicity, and the gene is useful in molecular epidemiology given its potential for adaptation to the host. 18
Histologic examination is a widely used technique to detect lesions caused by RABV in H&E-stained histologic sections of the CNS and is used in cases negative for rabies by other techniques. 11 The CNS lesions associated most frequently with RABV are monocytic and lymphocytic infiltration in the perivascular spaces and meninges, gliosis, neuronal degeneration, necrosis, and the characteristic intracytoplasmic inclusion bodies (ICIBs) in the neuronal cytoplasm, which occur in 30% of bovine rabies cases.7,11,25
Our objective was to compare the detection of RABV by various techniques (FAT, RDT, RT-PCR, and histology) under routine conditions, and to determine the most sensitive and specific technique for use in Paraguayan cattle.
Materials and methods
Study population and origin of the samples
Forty-nine pooled samples of different portions of the CNS of cattle (Table 1) that died with nervous signs compatible with rabies from different livestock farms in 13 departments of Paraguay (Fig. 1) were sent to the CEDIVEP laboratory (San Lorenzo, Paraguay) for the detection of RABV by FAT and histology from January 2021 to April 2022. Once received, the samples were stored at −20°C until their subsequent use for RDT and RT-PCR testing, also conducted in the CEDIVEP laboratory.
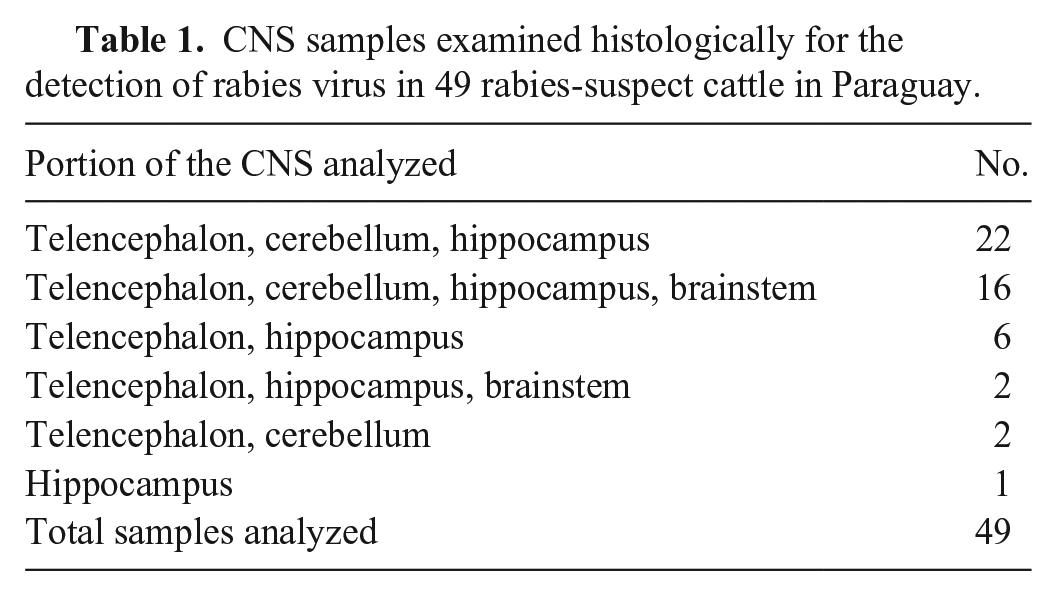
CNS samples examined histologically for the detection of rabies virus in 49 rabies-suspect cattle in Paraguay.

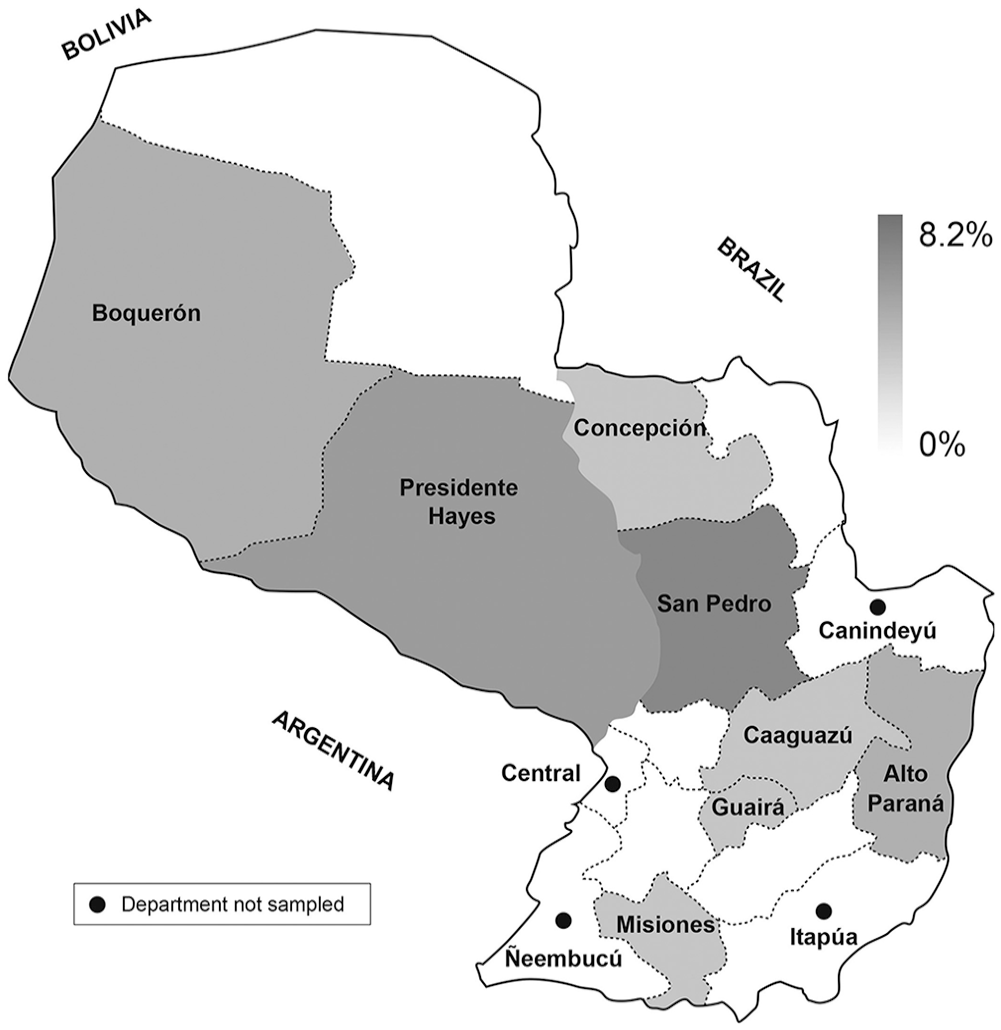
Geographic distribution of samples that were positive for the detection of rabies virus in cattle in Paraguay, by at least one of the techniques analyzed.
For FAT, RDT, and RT-PCR tests, one sample of 1 g from each portion of the CNS (Table 1) was homogenized, obtaining a single pooled sample from the CNS of each animal.
Fluorescent antibody test
The FAT was performed with a polyclonal anti-nucleocapsid antibody to RABV conjugated with fluorescein isothiocyanate (Bio-Rad), which detects all Lyssavirus genotypes; this technique has been validated in our laboratory and is accredited by our local sanitary authorities (SENACSA). Pooled portions of CNS of each animal were taken with a swab, placed on a slide, and fixed in acetone at −20°C for 30 min. Next, 20 μL of the conjugate was placed on the slide and incubated for 30 min in a wet chamber. Two successive 5-min PBS washes were then performed, and a final wash was carried out with distilled water. Subsequently, the slides were coverslipped with buffered glycerin. A fluorescence microscope (BH; Olympus) was used for the readings by a trained technician.
Histology
CNS samples (portions of the telencephalon, cerebellum, hippocampus, and/or brainstem) were fixed in 10% neutral-buffered formalin, processed routinely, and 3-μm sections were stained with H&E.
Rapid detection tests
The frozen CNS samples were left at 25°C until thawed. The commercial RDT (Anigen Rapid Rabies Ag test kit; Bionote), which is an immunochromatographic assay for the qualitative detection of RABV, was performed according to the manufacturer’s instructions. One g of pooled portions of CNS of each animal were taken with a swab, placed in a tube containing 9 mL of PBS, and mixed until the sample was homogenized. The suspension was left to stand for 10 s, and then 4 drops of the suspension were deposited on the reading cassette. After 10 min, the reading was performed. Two bands indicated a positive result; one band on line C (control) indicated a negative result.
RT-PCR assay
For RNA extraction, 1 g was taken from the tissue sample pool, placed in 1 mL of TRIzol (Invitrogen), homogenized in a vortex mixer for 5 min, and centrifuged at 12,000 × g for 5 min at 4°C. The supernatant was transferred to a new tube, incubated for 5 min, and then 0.2 mL of chloroform was added and mixed by vortexing for 3 min. The sample was incubated for 5 min until dissociation of the layers was observed, and the sample was then centrifuged for 15 min at 12,000 × g at 4°C. After obtaining complete dissociation of the layers, the aqueous phase was placed in a new tube. To isolate RNA, 500 μL of isopropanol was added, incubated for 10 min, and then centrifuged for 10 min at 12,000 × g at 4°C; the supernatant was discarded. Then, the RNA was washed in 1 mL of 75% ethanol, mixed by vortexing for 1 min, and then centrifuged at 7,500 × g for 5 min at 4°C. The supernatant was discarded, and the pellet was dried for 5–10 min. The RNA pellet was resuspended in 20 μL of RNase-free water and incubated in a water bath at 60°C for 10 min before being stored at −20°C.
For RT-PCR, genes N and G were amplified, using the following pairs of primers: N gene, forward IPF-1 (5′-ACGCTTAACAACAARATCARAG-3′), reverse P784 (5′-CCTCAAAGTTCTTGTGGAAGA-3′ 23 ); G gene, forward Ga-3222-40 (5′-CGCTGCATTTTRTCARAGT-3′), reverse Gb-4119-39 (5′-GAGGGCACCATTTGGTMTC-3′ 18 ). For the synthesis of cDNA, we used an RT kit (AccuPower RT PreMix; Bioneer). Seven microliters of extracted RNA were mixed with 1 μL (10 μM) from each pair of primers for each of the N gene and the G gene; 2 μL (10 μM) of the primer was used. The mixture was incubated at 70°C for 5 min and then placed on ice. The mixture was then transferred to a tube with premix (AccuPower RT PreMix) and mixed with 11 μL of RNase-free, diethylpyrocarbonate (DEPC)-treated (Thistle Scientific) water several times by pipetting. Then, the mixture was incubated (GTC 96S thermal cycler; Thistle Scientific) at 42°C for 60 min, and reverse transcriptase inactivation was performed at 94°C for 5 min.
For the amplification of N and G genes, 2 μL of cDNA (concentrations ≤ 100 ng/μL) was taken, quantified (NanoPro; MaestroGen), and mixed in a tube containing premix (AccuPower Taq PCR PreMix), previously diluted with 16 μL of DEPC-DW and 1 μL (10 μM) of each pair of primers, to amplify each gene, reaching a final volume of 20 μL. Cycling conditions were 94°C for 5 min, 35 cycles of 94°C for 45 s, 55°C for 45 s, 72°C for 2 min, and a final extension at 72°C for 10 min. The expected products were 900 bp for the G gene and 869 bp for the N gene. The products were resolved in 2% agarose gel (6x RunSafe; Thistle Scientific) and run in an electrophoresis device (MultiSub horizontal midi; Thistle Scientific) for 45 min at 100 V. The gels were then visualized through gelLITE (Thistle Scientific).
Sequencing and bioinformatic analysis
Two amplicons of samples of gene N and one from gene G were purified and sequenced. The bands observed in the agarose gels were more intense and, when quantified, had values > 10 ng/μL. The amplicons obtained were quantified (NanoPro quantifier; MaestroGen). Products with the correct band size and required concentration were sent to Macrogen (Korea) for purification and Sanger sequencing. The chromatograms obtained were cleaned by removing the regions at the ends that were not legible. The sequences were analyzed with BLAST (https://blast.ncbi.nlm.nih.gov/Blast.cgi) to confirm the target gene.
The genetic sequences obtained were edited, and for homology analysis, 650 bp of the RABV nucleoprotein were used. Sequences retrieved from GenBank were from different Latin American countries (KX090640, KX090637, KX090634, KX090628, KX090593, KX090591, EF363745, EF363751, GQ160918, GQ160921, GQ160923, GQ160954, EF428579, EF428576, EF152275, EF152264 from various regions of Brazil; EF363727 and EF363728 from Ecuador; EU981923 and EU981928 from Uruguay; KF864386 and JX648434 from Argentina; OL451916, OL451908 and OL451907 from Paraguay), and 695 bp of the RABV glycoprotein were used. Sequences retrieved from GenBank from Brazil were included (AB518495, AB17428, AB247412, AB247414, AB110666, KY824722, KY824721, KY824715, KY824710, KY824661, FJ649042, FJ649039, FJ649035, FJ649033, FJ649029, FJ649020, FJ649017, FJ649012, FJ648997). These sequences were aligned with ClustalW (v.7.1.3.0; BioEdit), and the phylogenetic tree was constructed using the maximum-likelihood T92 G+I method with MEGA 7 (https://www.megasoftware.net/) and FIGTREE (https://github.com/rambaut/figtree) software.
Statistical analysis
The test sensitivity and specificity with CI values for RDT relative to FAT, and for RT-PCR (N-G) relative to FAT, were calculated using EpiData v.3.1 (https://epidata-analysis.software.informer.com/). The overall test agreement between RDT and FAT, and RDT and RT-PCR (N-G), was calculated using the kappa value, where kappa measured the agreement between the tests. The kappa value was interpreted as follows: 0.0 = poor; 0.00–0.20 = slight; 0.21–0.40 = fair; 0.41–0.60 = moderate; 0.61–0.80 = substantial; 0.81–1.00 = almost perfect agreement.
Results
We analyzed 49 CNS samples from cattle. The samples originated from 13 of the 17 departments of Paraguay. Samples from 8 departments were positive by at least one of the techniques. Positive cases were more frequent in the departments of San Pedro (4 of 49; 8.2%), Pte. Hayes (3 of 49; 6.1%), and Boquerón and Alto Paraná (2 of 49; 4.1%). One of 49 (2%) positive cases was determined in each of Concepción, Caaguazú, Guairá, and Misiones departments (Fig. 1).
For both the FAT and RDT, 15 of 49 (31%) samples were positive, and 34 of 49 (69%) were negative; the same 15 positive samples were detected by both assays. For RT-PCR amplification of N and G genes, 13 of 49 (27%) and 12 of 49 (25%) positive samples were obtained, respectively (Table 2). These positive samples belonged to the group of samples that were positive by FAT and RDT (Table 3).
Determination of the kappa value comparing the fluorescent antibody test (FAT) and other techniques for the detection of rabies virus in cattle in Paraguay.
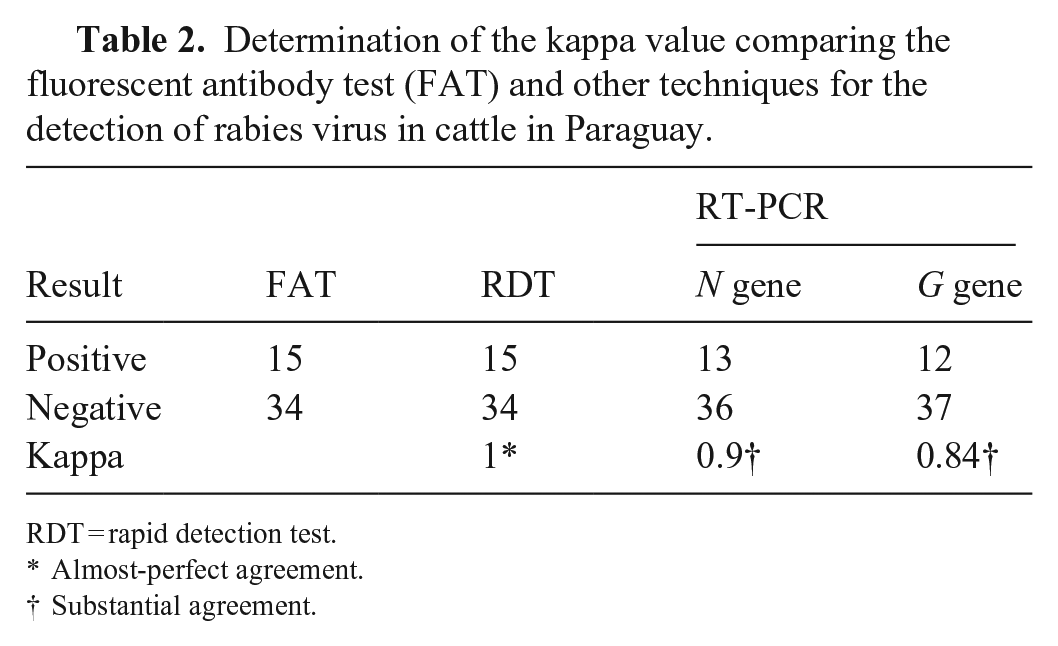
RDT = rapid detection test.
Almost-perfect agreement.
Substantial agreement.
Histologic lesions and test results in 49 rabies-suspect cattle in Paraguay.
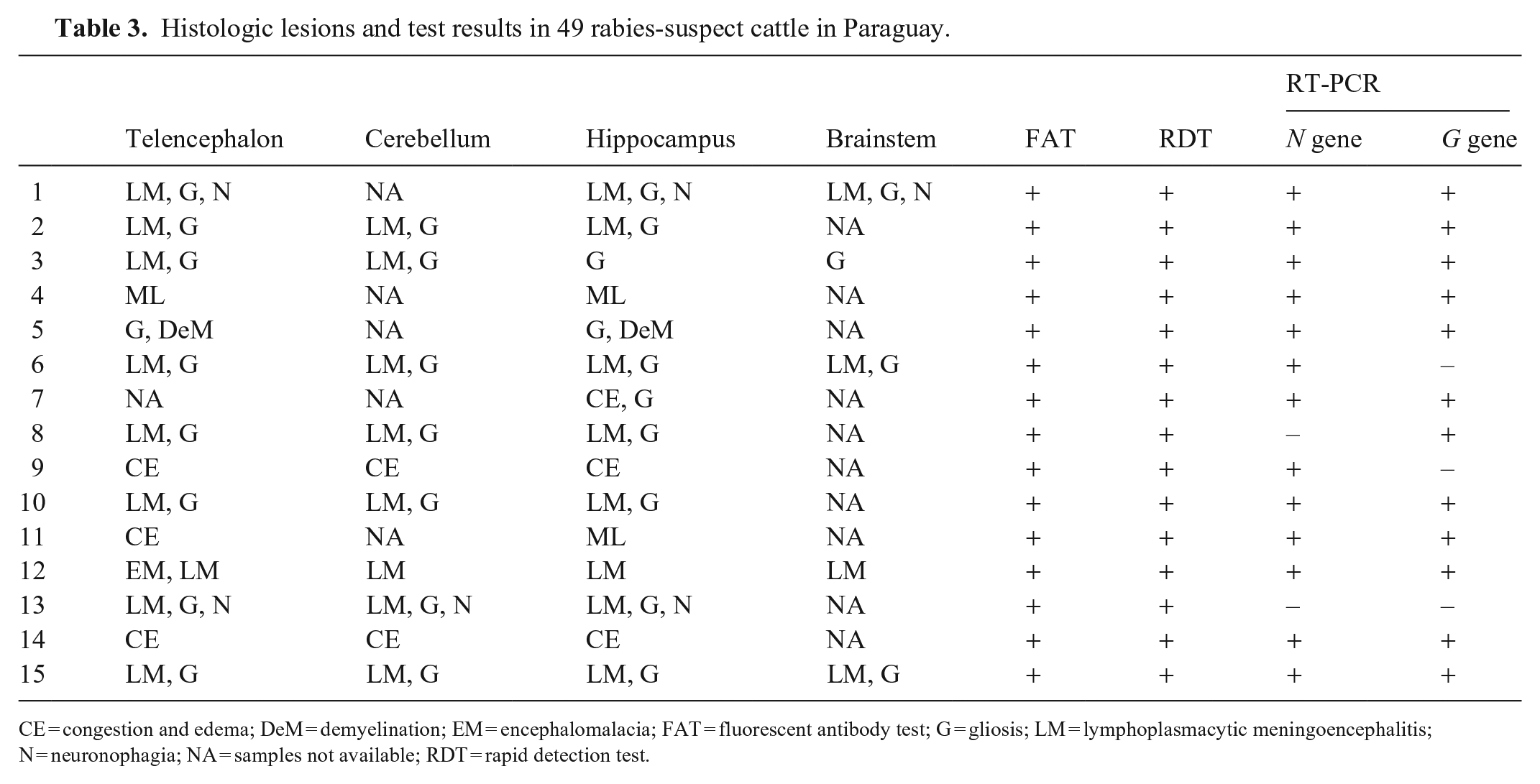
CE = congestion and edema; DeM = demyelination; EM = encephalomalacia; FAT = fluorescent antibody test; G = gliosis; LM = lymphoplasmacytic meningoencephalitis; N = neuronophagia; NA = samples not available; RDT = rapid detection test.
RDT results were in high agreement with FAT results (kappa = 1); the RDT sensitivity was 100% (95% CI: 97–100%) and specificity was 100% (95% CI: 99–100%). For RT-PCR of N and G genes, the concordances showed substantial agreement (kappa values of 0.9 and 0.8, respectively; Table 2). Compared with FAT, the sensitivity and specificity for gene N were 86.7% (95% CI: 66–100%) and 100% (95% CI: 98–100%); and for the G gene were 80% (95% CI: 56–100%) and 100% (95% CI: 98–100%), respectively.
In the samples analyzed histologically, the lesions most frequently observed in positive cases of rabies (by at least one of the techniques analyzed) were lymphoplasmacytic meningoencephalitis, gliosis, and neuronophagia (Table 3). No ICIBs were found in the histologic lesions.
To confirm the RT-PCR assay results, the amplicons obtained from the amplified target genes N and G were analyzed, obtaining sequences of ≥ 900 bp for both genes. We observed 2 clusters: cluster 1-RABV, with a genetic lineage characteristic of the hematophagous bat D. rotundus; cluster 2-RABV, with a genetic lineage characteristic of dogs. Samples CDV97 and CDV98 were grouped in cluster 1, with the other RABV sequences having genetic lineage characteristics of D. rotundus hematophagous bats (Fig. 2). Sample CDV89 was grouped in cluster 1, with the other RABV sequences having genetic lineage characteristics of D. rotundus hematophagous bats (Fig. 3).
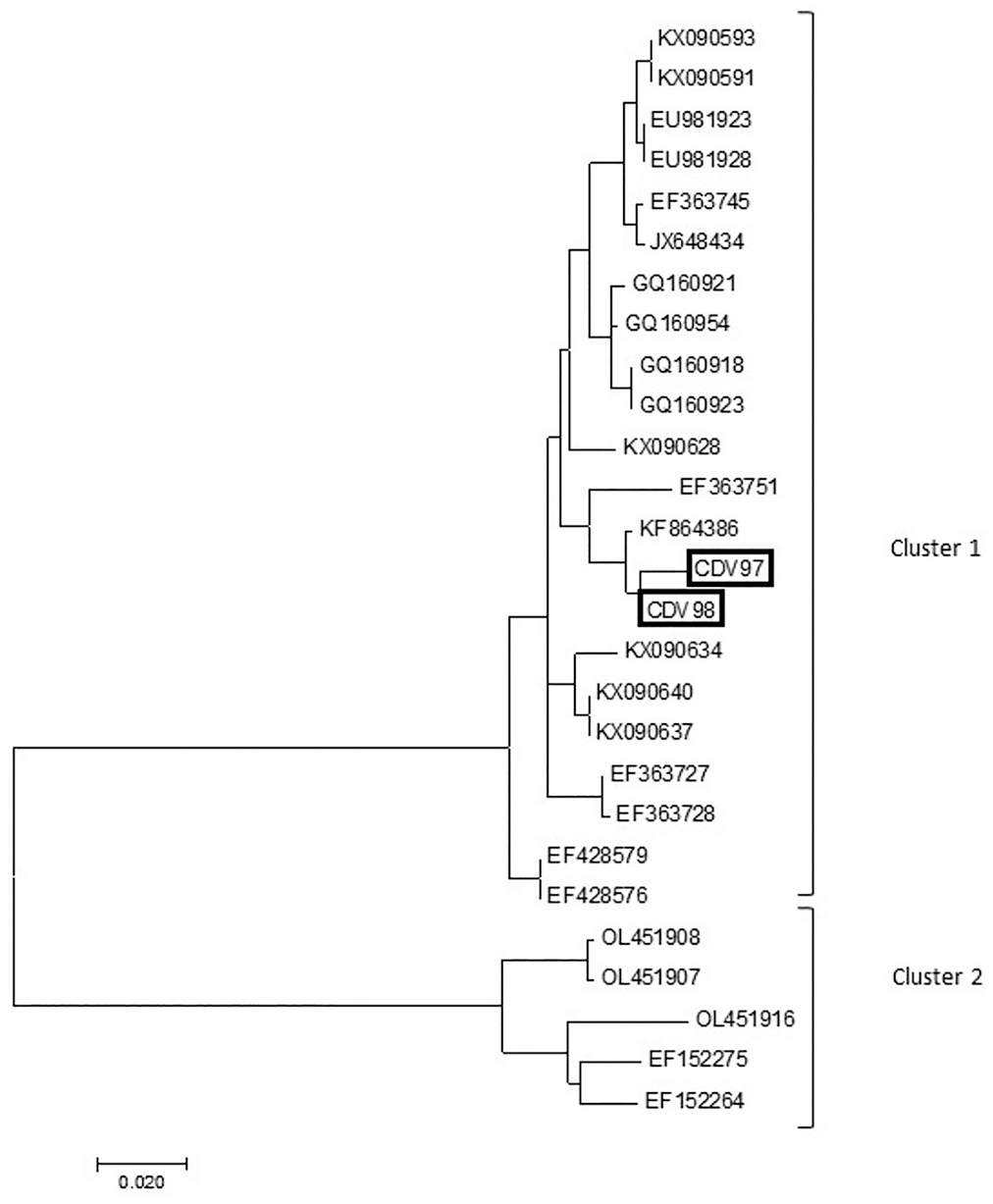
Phylogenetic tree based on the partial 650-nt region within the N gene of rabies virus (RABV). Tree was generated using the maximum-likelihood method based on the Tamura–Nei model, with MEGA 7 software. Cluster 1 (RABVs from hematophagous bats) and cluster 2 (RABVs from dogs); sample sequenced in our study is marked in black.
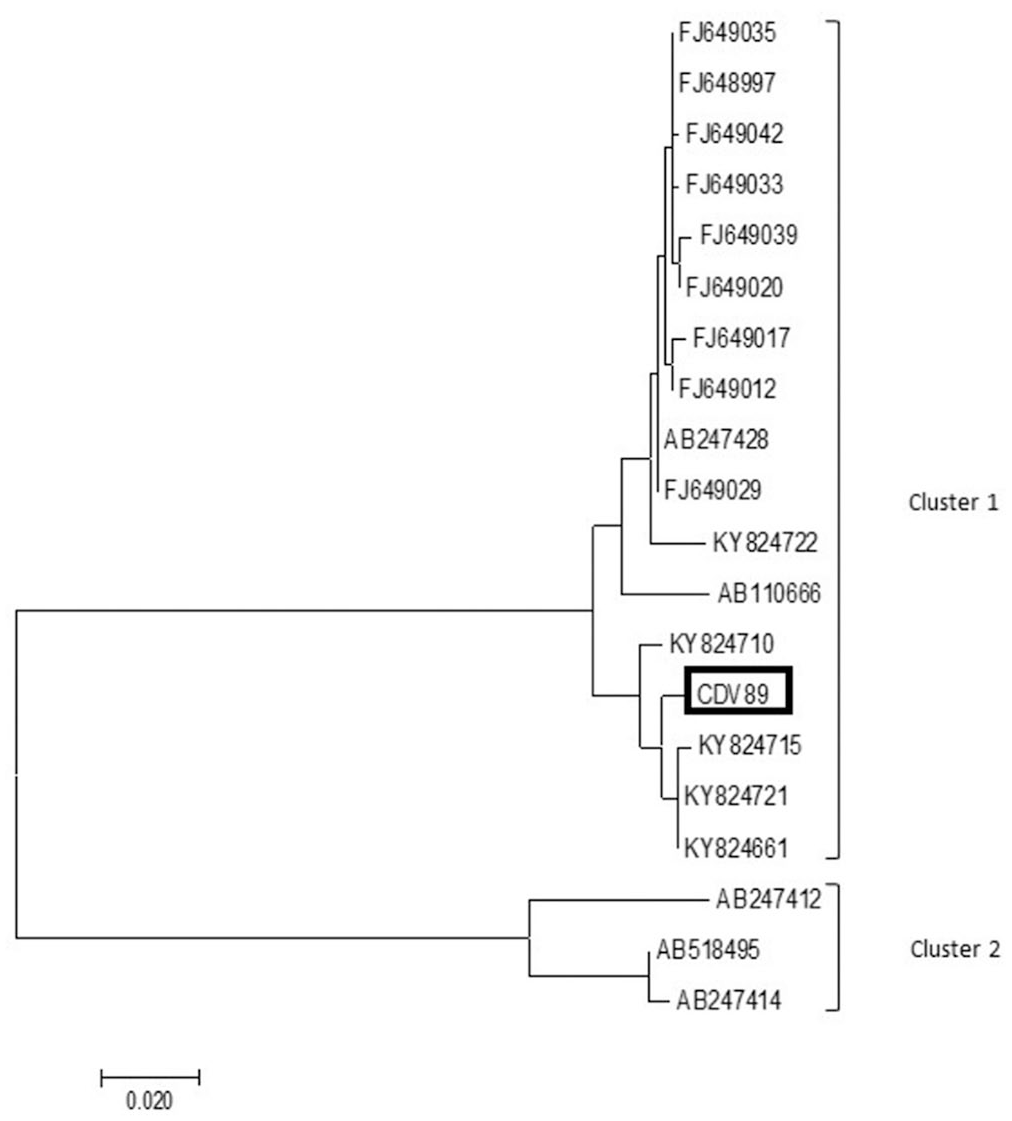
Phylogenetic tree based on the partial 695-nt region within the G gene of rabies virus (RABV). Tree was generated using the maximum-likelihood method based on the Tamura–Nei model, with MEGA 7 software. Sample CDV89 was grouped in cluster 1, with the other RABV sequences with genetic lineage characteristics of D. rotundus hematophagous bats. Sample sequenced in our study marked in black.
Discussion
Only FAT is used as a confirmatory test for the detection of RABV in Paraguay. The almost-perfect agreement that we observed between the FAT and RDT suggests that the use of rapid tests could be a valid tool for the detection of RABV in bovine samples. The RDT has had the same performance as FAT, obtaining a specificity and sensitivity of 93.3–100% and 95.3–100%, respectively, in laboratory conditions and field studies conducted by others. 12 We did not conduct field studies; however, we determined that the RDT could be useful in field conditions, without the need for a laboratory structure; this represents an advantage over the FAT, which must be conducted in optimal building conditions, especially considering that the conjugate must be kept frozen at −20°C and the test requires the use of a fluorescence microscope by a trained operator. Unlike FAT, the Anigen Rapid Rabies Ag test kit can be stored at room temperature and is simple and practical to use. We recommend the use of the RDT for surveillance and to confirm suspected cases of rabies for rapid decision-making, particularly in outbreaks or in conditions of scarce resources. 24
The agreement obtained between the standardized conventional RT-PCR assay and the FAT was lower than that obtained with the RDT and variable when both N and G genes were amplified (27% and 25%, respectively); this discrepancy may be because the N gene is the most conserved of the RABV genes, and because the use of primers specific to that region increases the possibility of detecting RABV. 23 One of the possible causes of the disparity in the results between the 2 genes could be that the gene that encodes the G protein is related to the ligation of cell receptors and is targeted for the neutralization of the virus; therefore, it has a higher rate of mutation, and divergences in the sites of annealing of the primers could be a probable cause of nonamplification of the samples that amplify the G gene.15,27 Although molecular techniques have great advantages, their routine use presents some difficulties, such as the loss of integrity of viral RNA due to freezing and thawing, as well as the need for storage at −80°C. 1 The samples that we analyzed were frozen and thawed for processing; therefore, this could be a factor in RNA degradation. The presence of inhibitors such as RNAses and the size of the target gene segment could cause a difference in viral RNA detection. 10 However, the RT-PCR assay is useful for genetic characterization and epidemiologic monitoring of the strains detected in Paraguay.
ICIBs were not observed. However, histologic lesions typical of RABV infection (lymphoplasmacytic meningoencephalitis, gliosis, and neuronophagia) 11 could be useful in resolving results that are discordant with other methods (FAT, RDT, RT-PCR). Studies in Brazil found ICIBs in 68% of cattle that survived > 4 d after the development of clinical signs, specifically in Purkinje neurons, brainstem, and spinal cord neurons. 11 There is no relationship regarding the duration of the course of the disease and the intensity of inflammatory lesions in the CNS tissues of rabies-positive animals; however, ICIBs are observed in a higher percentage in cattle whose death is natural than in animals that were slaughtered. 25 Other factors, including strain, viral load, and distribution, could influence inflammatory lesions and the development of ICIBs. 13
Of the techniques that we studied, RDT is the screening technique most suited for cases in which there are neither laboratories suitably equipped with trained personnel nor resources for the implementation of more advanced techniques, or in remote places with low accessibility. The detection of RABV by FAT could be used to confirm RDT results, or in negative cases in which clinical signs are compatible with rabies. Histologic examination is a useful support, especially to confirm other diagnoses in rabies-negative cattle with nervous signs. A RT-PCR assay is needed for molecular epidemiology studies.
Footnotes
Acknowledgements
We thank the laboratory technicians of CEDIVEP.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
