Abstract
Canine schistosomiasis caused by Heterobilharzia americana can lead to severe morbidity and eventual mortality, in part due to the deposition of fluke ova in the liver and gastrointestinal tract, which promotes an influx of peri-ova inflammatory cells. Although fluke eggs can be identified in H&E-stained histologic sections, cases exist in which only fragments of the ova persist, or the egg is obscured by inflammatory infiltrates, which can confound definitive histologic diagnosis. Unfortunately, antibodies specific to Heterobilharzia are not commercially available for immunohistochemical labeling. Therefore, we aimed to use an RNA in situ hybridization strategy to fluorescently label Heterobilharzia ova. Using the H. americana 18S rRNA sequence, we developed an RNA probe and validated its performance on archival formalin-fixed, paraffin-embedded canine tissue. A positive signal was observed for all identifiable ova, fragmented and whole. Use of this methodology could aid understanding of the pathogenesis of H. americana infection in dogs. This technique augments standard diagnostic methodology, enabling spatial colocalization of fluke ova and inflammatory infiltrates when using fluorescent techniques.
Heterobilharzia americana is the causative agent of canine schistosomiasis. This trematode parasite is enzootic in the Gulf Coast and southern Atlantic states of the United States.5,10,13 Manifestation of disease is dependent on parasitic burden and overall host health. 9 Infection occurs when Heterobilharzia cercariae, extruded from the aquatic intermediate host Galba cubensis, transdermally penetrate susceptible hosts.2,4 Once within the mammalian host, cercariae shed their tail and develop into schistosomula.2,4 Schistosomula travel through the vasculature to the lungs and subsequently the liver, where they mature into adults.2,4 Adults migrate to the mesenteric and intrahepatic portal vasculature to form copula.2,4 The intestine, liver, and pancreas are the primary sites of egg deposition.2,4,9,14 Mature ova produce proteolytic enzymes that enhance migration into the bowel lumen.2,4 Aberrant migration and entrapment of ova results in local tissue injury.2,4,16
Current detection methods include fecal saline sedimentation (FSS), molecular techniques (PCR, quantitative real-time PCR [qPCR]) using DNA or RNA isolated from stool or tissue, and identification of ova in H&E-stained, formalin-fixed, paraffin-embedded (FFPE) tissue.3,8,12,15 FSS is the most common method used by general practices to screen for Heterobilharzia ova, but it lacks sensitivity (50%). 12 A conventional fecal PCR and a qPCR test are available through a fee-for-service laboratory at the Texas A&M School of Veterinary Medicine (College Station, TX, USA). Despite conventional PCR being more sensitive (58.3%) than FSS, both methods are subject to inconsistent results due to intermittent shedding of ova in feces.3,12 A sensitive (83.3%) and specific (100%) qPCR assay has been developed, but it does not account for the spatial relationship of ova within the tissue. 12
Histology is the gold standard for the identification of H. americana ova. Histologic lesions associated with infection have been well-documented in case reports and retrospective studies, but the sensitivity and specificity of this method have not been described.2,6,9,10,14 Histologic ova identification is a common inclusion criterion in Heterobilharzia-related investigations, but rarely is it compared to other screening methods.4,9,14 Eggs may be inadvertently removed during sectioning, leaving behind indistinguishable fragments. 2 A commercial H. americana–specific antibody is not available for immunohistochemistry, and although molecular techniques may be used to confirm diagnosis, the spatial relationships to inflammatory infiltrates and tissue architecture are lost with this methodology.
In situ hybridization (ISH) techniques have been developed for the characterization of host–parasite interactions in human-infective schistosomes. 17 RNAscope is an RNA in situ hybridization (RNA-ISH) technology compatible with FFPE tissues.1,11,18 Our objective was to investigate whether RNAscope could serve as a useful tool to identify H. americana ova in histologic samples. RNAscope uses modifiable RNA detection probes capable of recognizing any gene of interest. 1 Use of a double “Z” design allows recognition and hybridization to short sequences and improves the specificity beyond a typical RNA-ISH method. 1 These signals are subsequently amplified in a process that allows up to 8,000× signal amplification, contributing to the high sensitivity of the technique. 1 To enhance spatial investigations, labeled probes can be conjugated to chromogenic or fluorescent detection molecules. 1
The Louisiana Animal Disease Diagnostic Laboratory (LADDL; Baton Rouge, LA, USA) database was searched for canine liver tissue with and without H. americana ova. H&E-stained slides from 2 Heterobilharzia-positive livers and 2 Heterobilharzia-negative livers were reviewed by an ACVP-certified veterinary anatomic pathologist (Emi Sasaki, LADDL) to confirm the presence or absence of H. americana ova. Sections (5 µm) of the corresponding paraffin-embedded liver specimens were mounted onto positively charged slides (Superfrost Plus; VWR) by the LADDL Histology Service. Both livers were confirmed to be Heterobilharzia-positive using a validated qPCR assay and further evaluated using Sanger sequencing (Tables 1, 2).2,12 Eight, 10 µm-thick scrolls were sectioned from the Heterobilharzia-positive case, and FFPE DNA was extracted by the LADDL Histology core laboratory. Briefly, extracted DNA was amplified using custom reverse and forward 250 nM primers (TaqMan universal PCR master mix; Applied Biosystems, Integrated DNA Technologies [IDT]), and a 125 nM probe specific to H. americana (IDT) as described. 12 Sample analyses were run in duplicate (QuantStudio 12K flex real-time PCR system; Applied Biosystems) along with known Heterobilharzia-positive and -negative DNA controls. Canine GAPDH (glyceraldehyde-3-phosphate dehydrogenase) for each sample was amplified according to the manufacturer’s protocols (PerfeCTa SYBR Green FastMix; Quantabio) to test for PCR inhibitors. Sanger sequencing was utilized to further validate the presence of H. americana DNA sequences within the sample. Conventional PCR amplified the target sequence using a mastermix (DreamTaq PCR master mix; Thermo Scientific) and previously validated Heterobilharzia-specific forward and reverse primers. 2 A sequencing primer was designed using Lasergene v.17.3 (DNAstar) and BLAST (https://blast.ncbi.nlm.nih.gov/Blast.cgi) corresponding to GenBank HQ339878.1. 2 PCR cycling conditions were 2 min at 95°C, 35 cycles of 30 s at 95°C, 30 s at 55.4°C, and 90 s at 72°C, followed by 5 min at 72°C, and finally, a hold at 4°C. Samples were then purified (QIAquick PCR purification kit; Qiagen) and submitted to the Louisiana State University–School of Veterinary Medicine GeneLab (Baton Rouge, LA, USA) for Sanger sequencing. Previously confirmed Heterobilharzia-positive and -negative sequencing controls were run in parallel.
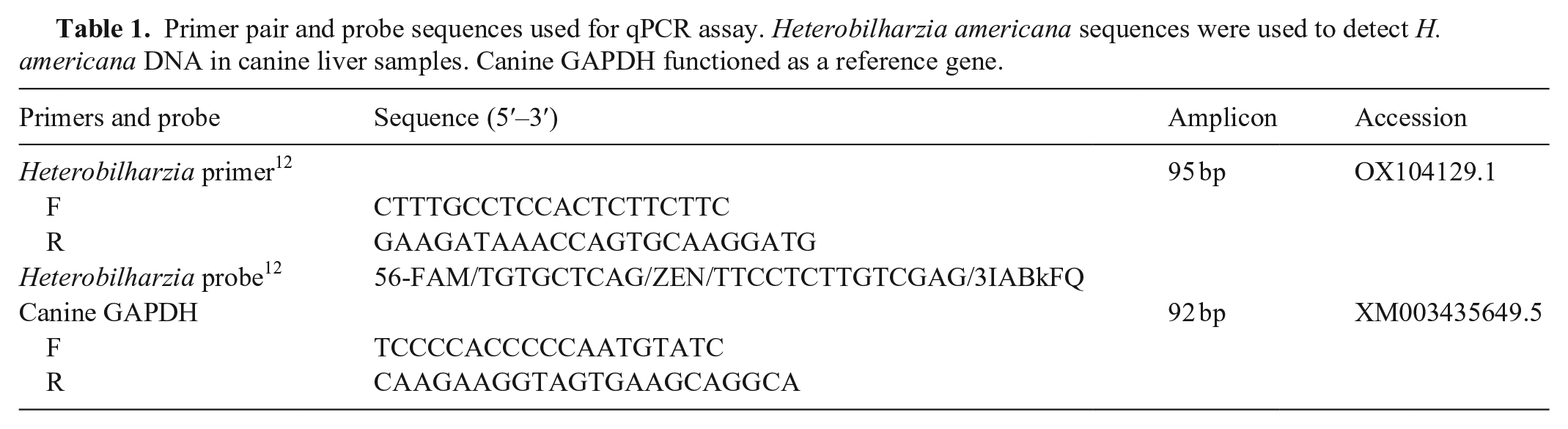
Primer pair and probe sequences used for qPCR assay. Heterobilharzia americana sequences were used to detect H. americana DNA in canine liver samples. Canine GAPDH functioned as a reference gene.
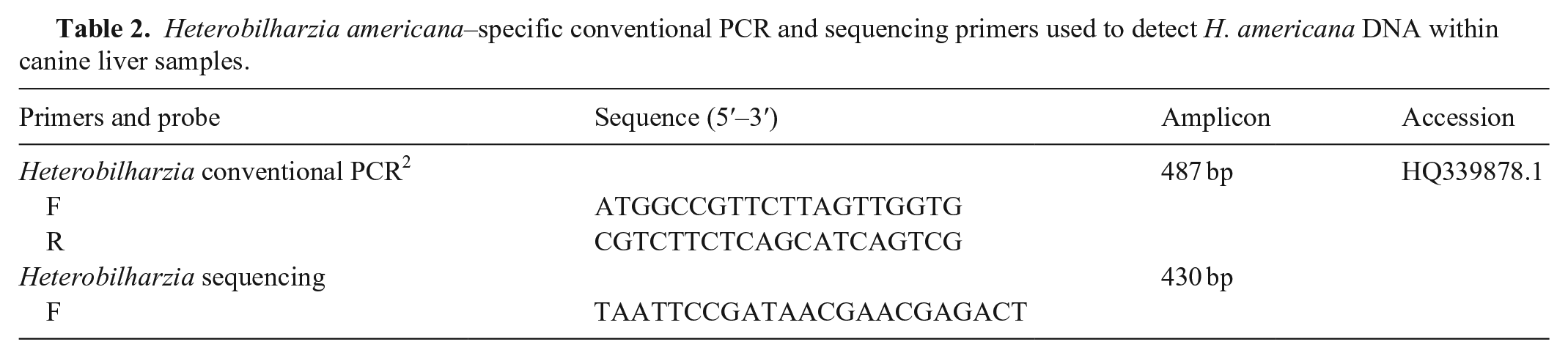
Heterobilharzia americana–specific conventional PCR and sequencing primers used to detect H. americana DNA within canine liver samples.
A probe targeting the 18S small subunit ribosomal RNA gene (GenBank AY157220.1) unique to H. americana was developed by Advanced Cell Diagnostics (ACD; RNAscope Probe-Ham-18SrRNA-No-XCl-C1). The species specificity of the target sequence was confirmed using BLAST. Two histologically confirmed Heterobilharzia-negative liver samples served as negative tissue controls and were assayed simultaneously. These samples were hybridized with the Heterobilharzia RNA probe, a positive control probe specific to canine PPIB (peptidylprolyl isomerase B), and a negative control probe specific to bacterial DapB (dihydrodipicolinate reductase) to validate tissue integrity. The Heterobilharzia probe was conjugated for detection by fluorescence microscopy (Opal 520; Akoya Biosciences). FFPE sections were labeled per manufacturer instructions (RNAscope multiplex fluorescent V2 assay, ACD; Fig. 1), and nuclei were stained with 4′,6′-diamindino-2-phenylindole (DAPI). Fluorescent images were acquired (Biotek Cytation 5; Agilent) at 4× and 20× magnification using DAPI (377 nm excitation, 447 nm emission) and green fluorescent protein (GFP; 469 nm excitation, 525 nm emission) filters or the Axio Observer Z1 (Zeiss) at 20× magnification using reflector cubes (Zeiss 34: 390 nm excitation, 460 nm emission; Zeiss 38: 470 nm excitation, 525 nm emission; Chroma 49004: 545 nm excitation, 605 nm emission). The corresponding H&E-stained slides were imaged at 40× magnification using a whole-slide scanner (NanoZoomer 2.0-HT; Hamamatsu Phototonics).
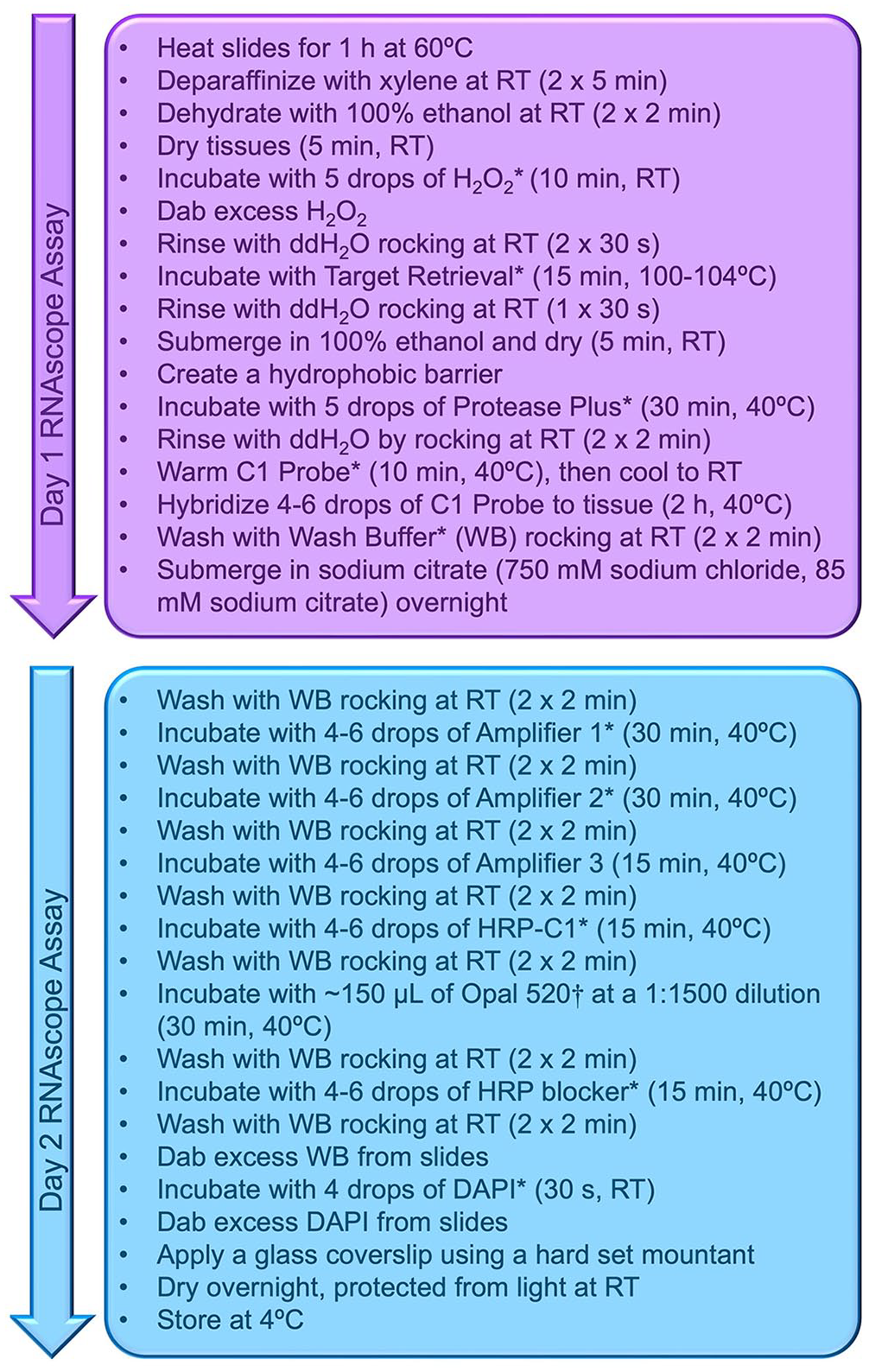
RNAscope protocol workflow divided into 2 d.
H. americana DNA was detected by qPCR in both histologically positive samples. Sample mean Cq values for the housekeeping primers and Heterobilharzia-specific primers were below the cutoff of 35. 7 Sanger sequencing of one Heterobilharzia-positive sample obtained a sequence that was a 100% match to GenBank AY157220.1 and HQ339878.1. The other Heterobilharzia-positive sample did not obtain a sequence, indicating inadequate DNA quality for Sanger sequencing. This finding motivated us to identify an alternative confirmatory test. We investigated the use of RNAscope to identify Heterobilharzia ova within liver tissue.
For fluorescent labeling, true-positive ova were defined as having at least 2 of the following criteria: a thin, fluorescent membrane; a developing or degenerate miracidium; or a round-to-irregular-to-fragmented shape, often with a mildly undulating or folded membrane. Fluorescently labeled ova were compared to the ova on the corresponding H&E-stained slides (Figs. 2A, 2C, 3A). The RNAscope probe was capable of labeling visible ova and remnants of ova that were not readily visible on H&E (Figs. 2B, 2D, 3B). Negative probe control (bacterial DapB) did not bind to Heterobilharzia-positive or -negative tissues (Fig. 3C). Any visible fluorescence was due to crosstalk between channels. Negative control liver tissue lacked signals matching the positive detection criteria.
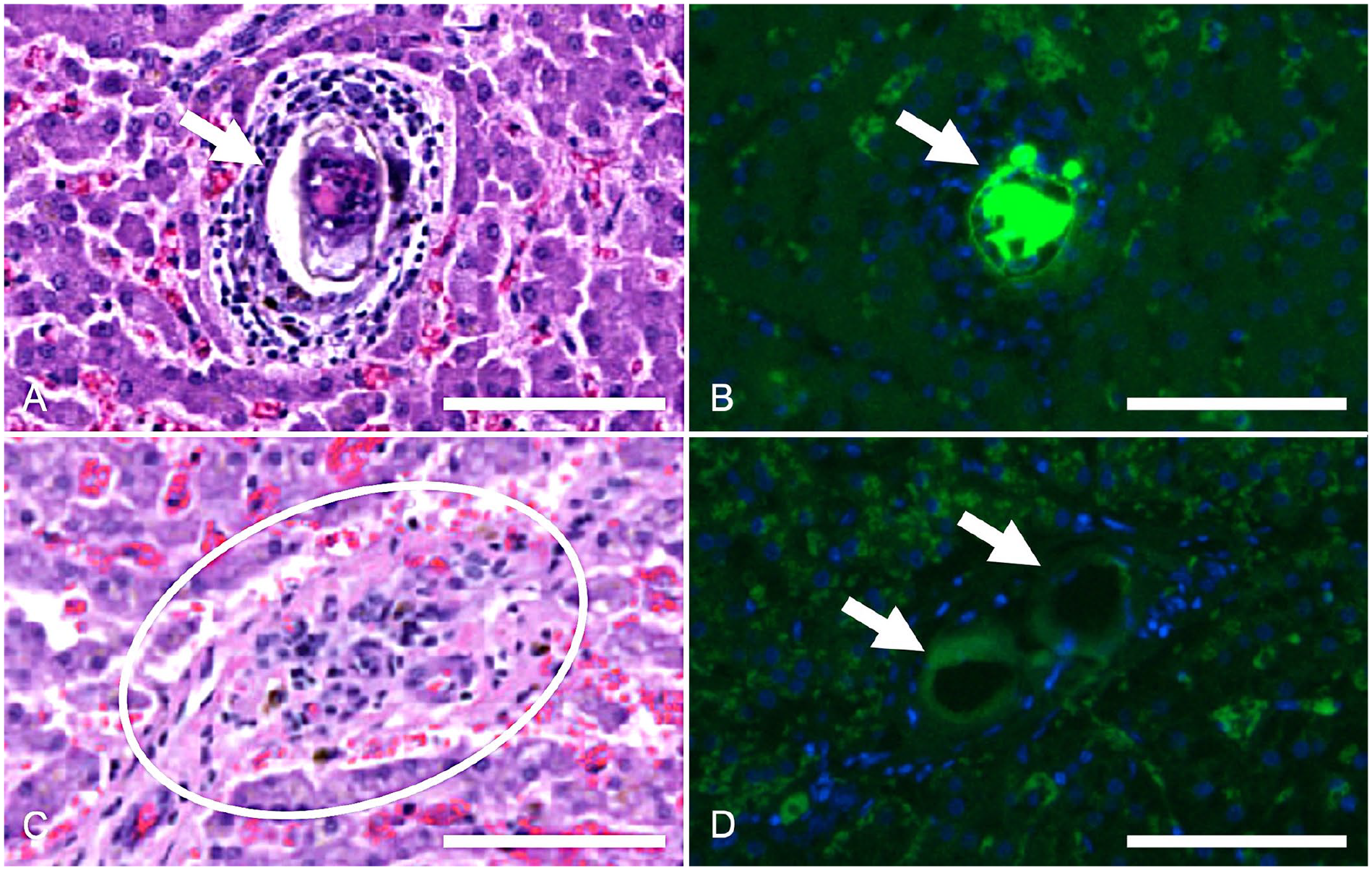
Heterobilharzia americana ova in histologic canine liver sections.
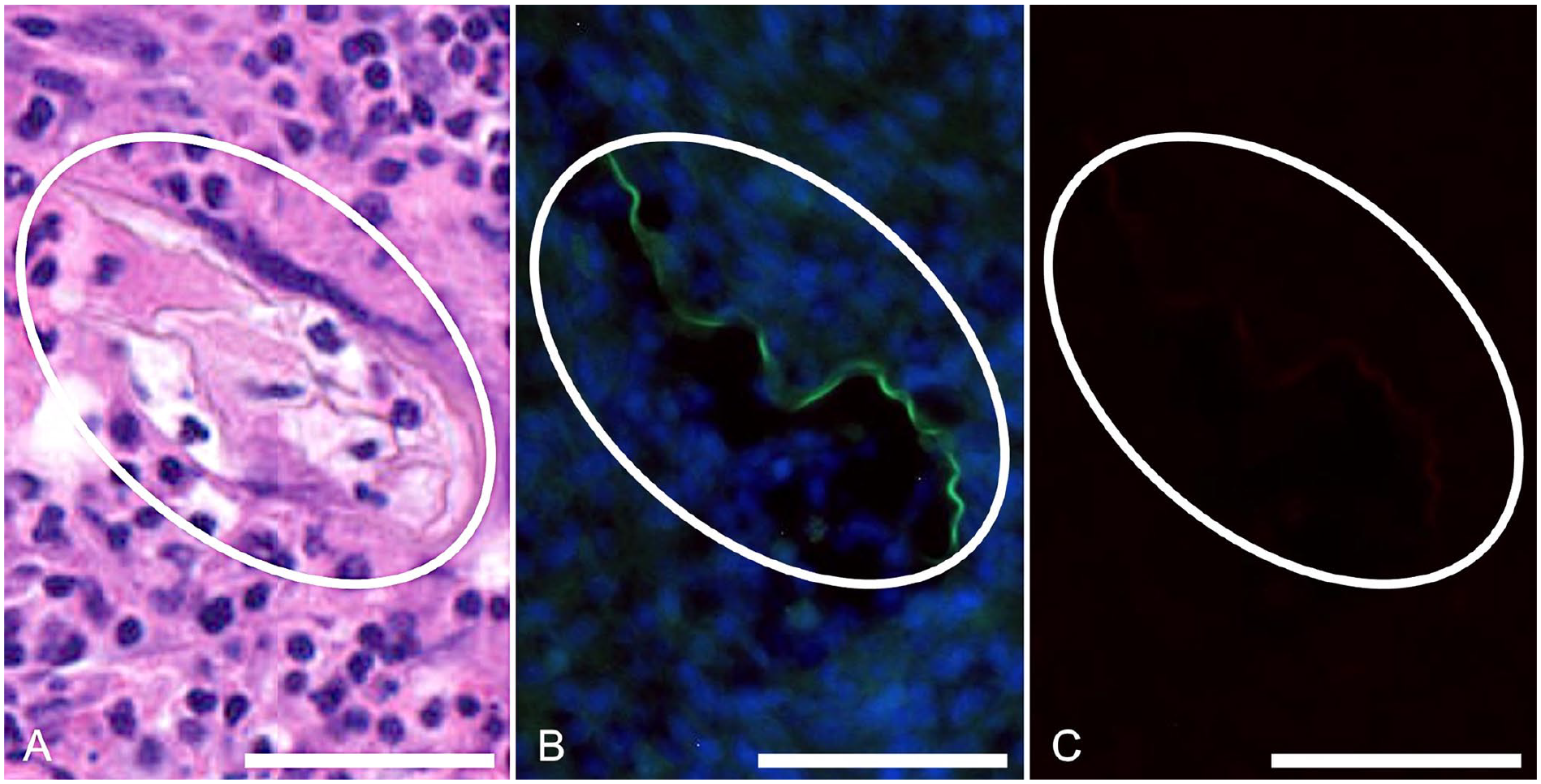
Remnant Heterobilharzia americana ova in histologic canine liver sections.
A limit to the adoption of our method is the need for fluorescence microscopy. Initial costs to purchase the reagent kit and probe will also limit the use of our method; however, once acquired, the assay is simple to run. The strengths of this methodology include its reproducibility and assay duration, which is comparable to other staining techniques such as immunohistochemistry. RNA is sensitive to degradation and time stored. Manufacturers of the RNAscope assay have validated successful staining procedures for archival FFPE tissue specimens prepared within the last 3 y and have reported successful staining in 25-y-old samples. At the time of our experiment, one Heterobilharzia-positive tissue block had been embedded for a little more than 1 y and the other for 5 y. The Heterobilharzia RNA probe designed for our study was assigned to channel 1, a green fluorescence channel. This channel is sensitive to tissue autofluorescence. We used at least 2 inclusion criteria for Heterobilharzia-positive signals to ensure the reproducibility of our experiment. Additional studies will be needed to determine which fluorescence channel may best complement this probe.
Footnotes
Acknowledgements
We thank Dr. Emi Sasaki for histologic review, Louisiana Animal Disease Diagnostic Laboratory histology service for tissue processing, and the Advanced Cell Diagnostics technical support team for assistance with assay optimization.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
We received financial support from the 2022–2023 Louisiana State University–School of Veterinary Medicine–Veterinary Clinical Sciences (VCS) Competitive Organized Research Program (CORP).
