Abstract
Thirteen American Hereford cattle were reported blind with presumed onset when ~12-mo-old. All blind cattle shared a common ancestor through both the maternal and paternal pedigrees, suggesting a recessive genetic origin. Given the pedigree relationships and novel phenotype, we characterized the ophthalmo-pathologic changes associated with blindness and identified the responsible gene variant. Ophthalmologic examinations of 5 blind cattle revealed retinal degeneration. Histologically, 2 blind cattle had loss of the retinal photoreceptor layer. Whole-genome sequencing (WGS) of 7 blind cattle and 9 unaffected relatives revealed a 1-bp frameshift deletion in ceroid lipofuscinosis neuronal 3 (CLN3; chr25 g.26043843del) for which the blind cattle were homozygous and their parents heterozygous. The identified variant in exon 16 of 17 is predicted to truncate the encoded protein (p. Pro369Argfs*8) battenin, which is involved in lysosomal function necessary for photoreceptor layer maintenance. Of 462 cattle genotyped, only blind cattle were homozygous for the deletion. A query of WGS data of > 5,800 animals further revealed that the variant was only observed in related Hereford cattle. Mutations in CLN3 are associated with human juvenile neuronal ceroid lipofuscinosis (JNCL), or Batten disease, which results in early-onset retinal degeneration and lesions similar to those observed in our cases. Our data support the frameshift variant of CLN3 as causative of blindness in these Hereford cattle, and provide additional evidence of the role of this gene in retinal lesions, possibly as a model for human non-syndromic JNCL.
Blindness has a negative impact on livestock welfare. 34 There are many potential causes of acquired vision loss in livestock, including vitamin A deficiency, lead poisoning, sulfate intoxication,10,13,30 and eye infection.1,34 Cataracts and neuronal ceroid lipofuscinoses (NCLs) are examples of blindness attributed to identified recessive mutations in specific cattle breeds.3,9,11,32 For example, a single base duplication in CLN5 was associated with recessively inherited severe neuronal loss and retinal degeneration in Devon cattle, very similar to human CLN5 disease. 11 In Braunvieh cattle, a missense mutation in CNGB3 was associated with recessively inherited achromatopsia and loss of cone photoreceptor function. 9 In a study designed to identify deleterious mutations, a recessive frameshift variant in the retinitis pigmentosa-1 gene was identified as causative of vision loss in Normande cattle, akin to retinitis pigmentosa in humans and mice. 21
Three blind Hereford cows (19-, 23-, 24-mo-old) were presented to the Nebraska congenital disease investigation program (Lincoln, NE, USA). The cows had no perceived ocular issues as calves and were suspected to be blind at ~12 mo of age. After the initial cases, 10 additional Hereford cattle with suspected blindness were identified.
The 3 initially reported cattle (V2, V10, V11) were born from a single sire (IV3; Fig. 1). Two (V10, V11) were full-siblings through embryo transfer. All blind cattle shared a common ancestor (I1) through both the maternal and paternal pedigree; a recessive variant was therefore the possible cause of blindness in these cattle. In this investigation, we characterized the ophthalmo-pathological changes in the blind cattle and performed genomic analyses to identify the cause of the condition.
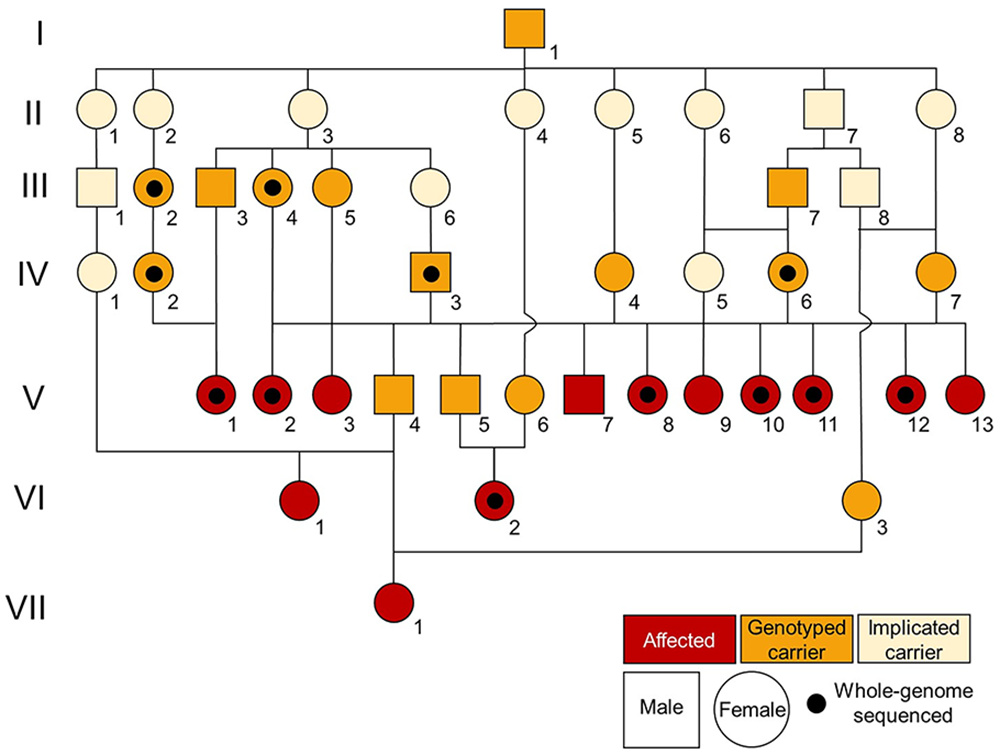
Pedigree of blind cattle.
Materials and methods
Our study was approved by the Institutional Animal Care and Use Committee at the University of Nebraska–Lincoln (protocol 2427), which is accredited by AAALAC International.
Blind animals
The 13 blind cattle were all registered Herefords, born between 2017 and 2022, and included 3 pairs of full-siblings. Ten of the blind animals were from embryo transfer (V3, V7, V8, V9, V10, V11, V12, V13, VI1, VII1). Cows V10 and V11 had previously calved successfully and were pregnant when they were reported to the Nebraska congenital disease program.
Control cattle
Six cows at a commercial abattoir were selected for postmortem tissue collection. Five of the cattle were red with white faces and other white markings, consistent with Hereford breeding. The sixth cow was black with a white face. Tissues from 4 levels of the brain (cerebellum, cerebrum, hippocampus, brainstem), eyes, and masseter muscles were collected 20–30 min postmortem (Table 1).
List of Hereford cattle sampled for DNA isolation, histopathology, and eye examinations.
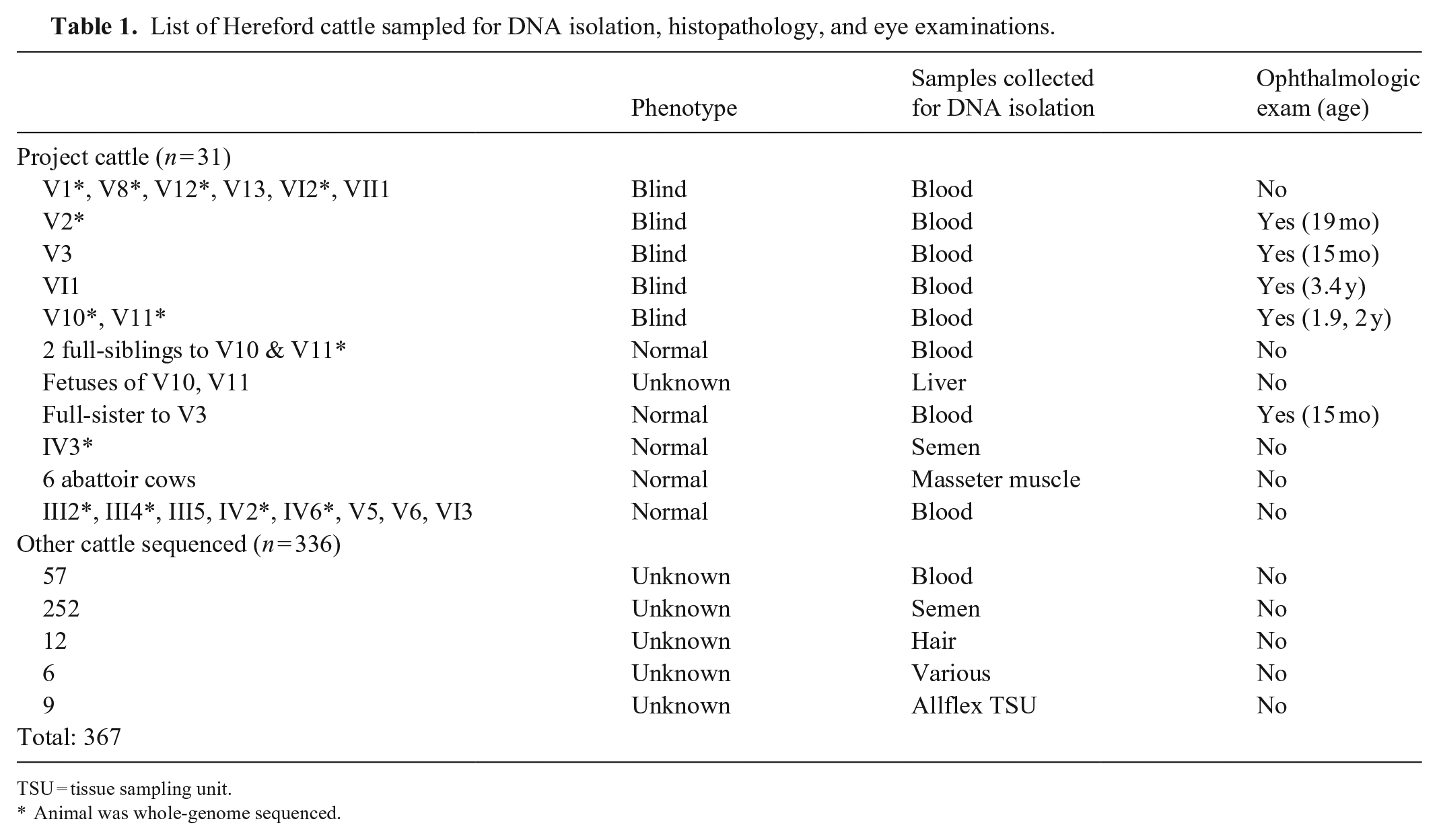
TSU = tissue sampling unit.
Animal was whole-genome sequenced.
Antemortem testing
Ophthalmologic examinations were performed on 4 blind animals and 1 phenotypically normal full-sibling of blind heifer V3. Three blind cows (V2, V10, V11) were examined by a diplomate of the American College of Veterinary Ophthalmologists (ACVO) from the Kansas State University–College of Veterinary Medicine (JM Meekins; Manhattan, KS, USA) to determine the severity of their blindness. Blood was collected from all 3 cattle for serum vitamin A evaluation. A blind heifer (V3) and her phenotypically normal full-sister (both from embryo transfer) underwent an ophthalmic examination by an ACVO diplomate at the Veterinary Teaching Hospital–University of Georgia (KA Diehl; VTH-UGA; Athens, GA, USA). One suspected blind cow (VI1) was examined by an ACVO diplomate from the University of Wisconsin–School of Veterinary Medicine (J Seth Eaton; Madison, WI, USA).
Electroretinogram
The cattle examined at the VTH-UGA underwent electroretinogram (ERG) tests. Tropicamide 1% ophthalmic solution was instilled in both eyes for pharmacologic mydriasis. Specific dark adaptation was not performed, but all artificial barn lighting was turned off and nearby windows were covered with blackout curtains. Retinal stimulation was performed (BPM-300 electrodiagnostic system; RetinoGraphics). Standard placement of ground (~2.5 cm medial to the medial canthus) and reference (~2.5 cm lateral to the lateral canthus) needle electrodes was performed. After the instillation of proparacaine hydrochloride 0.5% ophthalmic solution for topical/corneal anesthesia, a contact lens (ERG-jet; Fabrinal) amply coated with eye lubricant plus ophthalmic lubricant (Optixcare; Aventix) was used for the recording electrode. The eyelids were manually retracted with care to not disrupt the electrodes and allow continuous visualization to ensure continued appropriate placement throughout the test. A manufacturer preprogrammed short ERG protocol was used to provide an average of 4 “yes or no” mixed rod and cone responses in response to a standard flash in a relatively scotopic environment.
Tissue collection
Blind cows V10 and V11 were ~31-mo-old when they were harvested at the USDA-inspected Loeffel Meat Laboratory at the University of Nebraska–Lincoln (Lincoln, NE, USA). After passing federal inspection, tissues from 4 levels of the brain (cerebellum, cerebrum, hippocampus, brainstem), cardiac muscle, semitendinosus muscle, liver, kidney, lung, spleen, and lymph node were collected for pathology examination as well as flash-frozen in liquid nitrogen.
Histopathology
Ocular histologic examination was performed on V10, V11, their fetuses, and the 6 normal abattoir cattle.
Scanning electron microscopy
Retinas from V10 and V11 were processed for scanning electron microscopy (SEM) at the University of Nebraska–Lincoln Microscopy Core Research Facility of the Center for Biotechnology (Lincoln, NE, USA). Eyes from V10 and V11 were bisected and fixed in 2.5% glutaraldehyde in 100 mM cacodylate buffer (pH 7.4) for 2 h at room temperature and post-fixed in 1% osmium tetroxide in distilled water for 1 h. After rinsing 3 times in deionized water, each sample was processed through a series of ethanol dehydration (30%, 50%, 70%, 2 × 95%, 2 × 100%). The eye cups were immediately processed for critical point dry (Samidri-795 system; Tousimis). The samples were mounted onto a SEM stub and coated with a thin layer of chromium (Desk V Sputter; Denton). Side-view SEM images were collected (S4700 field-emission SEM; Hitachi).
DNA isolation
DNA was isolated from the cattle (Table 1) using the following protocols appropriate for the type of sample collected. DNA was isolated from blood and semen (Gentra Puregene blood kit; Qiagen) following a published protocol. 26 DNA was isolated from hair roots (Gentra Puregene blood kit) following a published protocol. 25
DNA was isolated from tissue (masseter muscle, liver; Gentra Puregene blood kit). Three hundred μL of cell lysis buffer and 7 μL of Qiagen proteinase K were added to 30 mg of tissue, and then incubated at 55°C overnight. For tissue sampling units (TSUs; Allflex), 75 μL of the liquid buffer from each sample was added to 225 μL of cell lysis buffer and 7 μL of proteinase K. All samples were then incubated overnight at 55°C, after which they were transferred to ice for 1 min. One hundred μL of protein precipitation solution was added and vortexed (20 s) before incubation on ice for > 5 min and centrifugation (5 min at 15,000 × g at 15°C). The supernatant was poured into a new tube with 650 μL of 100% isopropanol. Tubes were inverted to mix and then centrifuged (1 min at 15,000 × g). The supernatant was discarded, and the tube placed upside down on a paper towel to drain. The pelleted DNA was washed with 300 μL of 70% ethanol and inverted several times. Samples were centrifuged (1 min at 15,000 × g), the supernatant was discarded, and the tube was placed upside down to drain for 5 min; 200 μL of DNA hydration solution was added to the DNA pellet and incubated for 5 min at 55°C before overnight hydration at room temperature. DNA concentration was determined (BioTek Epoch 2 microplate spectrophotometer; Agilent).
Whole-genome sequencing
Isolated DNA from 7 blind cattle (V1, V2, V8, V10, V11, V12, VI2) and 9 unaffected relatives (III2, III4, IV2, IV3, IV6, 1 offspring of IV3, 2 full-siblings to V10 and V11, 1 offspring of V12) was sent to Admera Health (South Plainfield, NJ, USA) for library preparation (KAPA; Roche) and 150-bp paired-end sequencing (NovaSeq; Illumina). Raw data were processed by trimming adapters and removing poor-quality bases using TrimGalore. 16 Reads were mapped to the ARS-UCD1.2 genome with BWA-MEM, 19 and duplicates were marked with Samtools. 8 GATK was implemented to realign indels, and variants were called using GATK Haplotype Caller. 29 A combined variant file was created by merging the data of the 16 related cattle sequenced for our project with 133 control cattle previously sequenced from other projects, including PRJNA513064, PRJNA663547, and PRJNA994471, PRJNA1042650, and PRJNA1042814. A case control script (SnpSift) 5 was run with the 7 blind cattle as cases and the 142 other cattle assumed to be unaffected. Hypothesizing that blindness was attributed to a recessive genotype, the variant file was filtered to identify loci where the p-value assuming a recessive model was < 1 × 10−6.
The variant call format file (VCF) was then filtered to keep loci for which all 7 blind cattle were homozygous for the alternative allele. Loci for which unaffected cattle were homozygous for the alternative allele were removed. The remaining loci were run through a variant effect predictor (VEP). 20
Data pulled from the NCBI Sequence Read Archive database (SRA; https://www.ncbi.nlm.nih.gov/sra) were used to estimate the frequency of the candidate variants. Cattle sequences from the SRA labeled as Hereford (n = 327) were aligned to the ARS-UCD1.2 genome. Variants were called using the VariantFrequency pipeline, available on GitHub (https://github.com/SichongP/VariantFrequency).
In addition, genotypes (VCF) of 95 Herefords from whole-genome sequencing were provided by the American Hereford Association (https://hereford.org/).
Sanger sequencing
Sanger sequencing was performed to determine the ceroid lipofuscinosis neuronal 3 (CLN3) candidate variant genotype in 336 additional cattle (Table 1). These animals included both those targeted for genotyping due to pedigree relationships with known carriers as well as samples on hand from unrelated Hereford cattle used in prior studies. 26 PCR primers (Suppl. Table 1) were designed using Primer3. 28 PCR was performed (FastStart kit; Sigma-Aldrich) in 12-μL reactions containing 4.45 μL of water, 0.25 μL of 25 mM MgCl2, 1.2 μL of 20 mM 10× buffer with MgCl2, 0.5 μL of dNTP (10 mM of each dNTP), 0.1 μL of Taq (5 U/μL), 0.75 μL of 20 μM forward and reverse primer, and 4 μL of 5 ng/μL DNA template. Thermal cycling conditions consisted of 94°C for 4 min, 32 cycles of 94°C for 30 s, annealing temperature (Suppl. Table 1) for 30 s, 72°C for 45 s, a final extension at 72°C for 10 min, then a 10°C hold. PCR product cleanup was performed (0.75 μL ExoSAP-IT; Applied Biosystems) per 4 μL of PCR product and incubation thermal cycling at 37°C for 30 min, 80°C for 15 min, and a 15°C hold.
Relative quantitative real-time PCR
RNA was isolated from the retina of 2 affected cows (V10, V11) and 2 abattoir cows (RNeasy fibrous tissue mini kit; Qiagen) following the manufacturer’s protocol. RNA was washed and eluted on columns with DNase I treatment (Direct-zol RNA MiniPrep Plus; Zymo) according to the manufacturer’s instructions; 2.3 µg of RNA total for each sample was converted into cDNA (Cytiva First-Strand synthesis kit, cat. 27926101; Millipore Sigma) following the manufacturer’s protocol.
The expression of CLN3 was quantified (SYBR Green PCR master mix kit, cat. 4309155, Applied Biosystems; CFX384 real time PCR, Bio-Rad). The quantitative real-time PCR (qPCR) was performed in 10-μL reactions containing 5 μL of 2× SYBR Green master mix, 0.5 μL of 20 µM forward primer, 0.5 μL of 20 µM reverse primer, 3 μL of water, and 2 μL of cDNA template for CLN3 and 1 μL of cDNA template for the reference gene actin beta (ACTB). Samples were run in triplicate with negative controls. The qPCR primers (Suppl. Table 2) were designed using IDT PrimerQuest Tool (www.idtdna.com). The CLN3 forward primer crosses between exons 1 and 2. Thermal cycling conditions consisted of 95°C for 10 min, 40 cycles of 98°C for 15 s, 58°C for 30 s, 72°C for 30 s, then a final extension of 72°C for 60 s. Mean normalized expression (MNE) was calculated using the cycle-crossing thresholds of the replicates, incorporating qPCR efficiencies of each gene. The CLN3 MNE values for each animal were then log10 transformed and used to compare the phenotype groups by t-test.
Results
Animals
Of the 13 blind animals reported, samples from 11 animals were available for genetic testing. In total, DNA was obtained from 367 Herefords (Table 1).
Ophthalmic examinations
At the time of examination, cattle V2, V10, and V11 were unable to navigate the unfamiliar holding pen and bumped into stationary objects. Menace response was bilaterally absent, confirming functional blindness. Neuro-ophthalmic findings were consistent with retinal localization; the dazzle reflex was delayed, resting mydriasis was present, and direct and consensual pupillary light reflexes were slow and incomplete. The ocular adnexa and anterior segment were normal. Patchy, widespread tapetal hyper-reflectivity was present with moderate retinal vessel attenuation and mid-peripheral tapetal pigment clumping (Fig. 2). The optic disc appeared dark. The 3 cattle were ophthalmoscopically diagnosed with severe retinal degeneration. Serum vitamin A concentrations (0.24, 0.25, 0.29 ppm) were just below our laboratory acceptable range of 0.3–0.7 ppm.
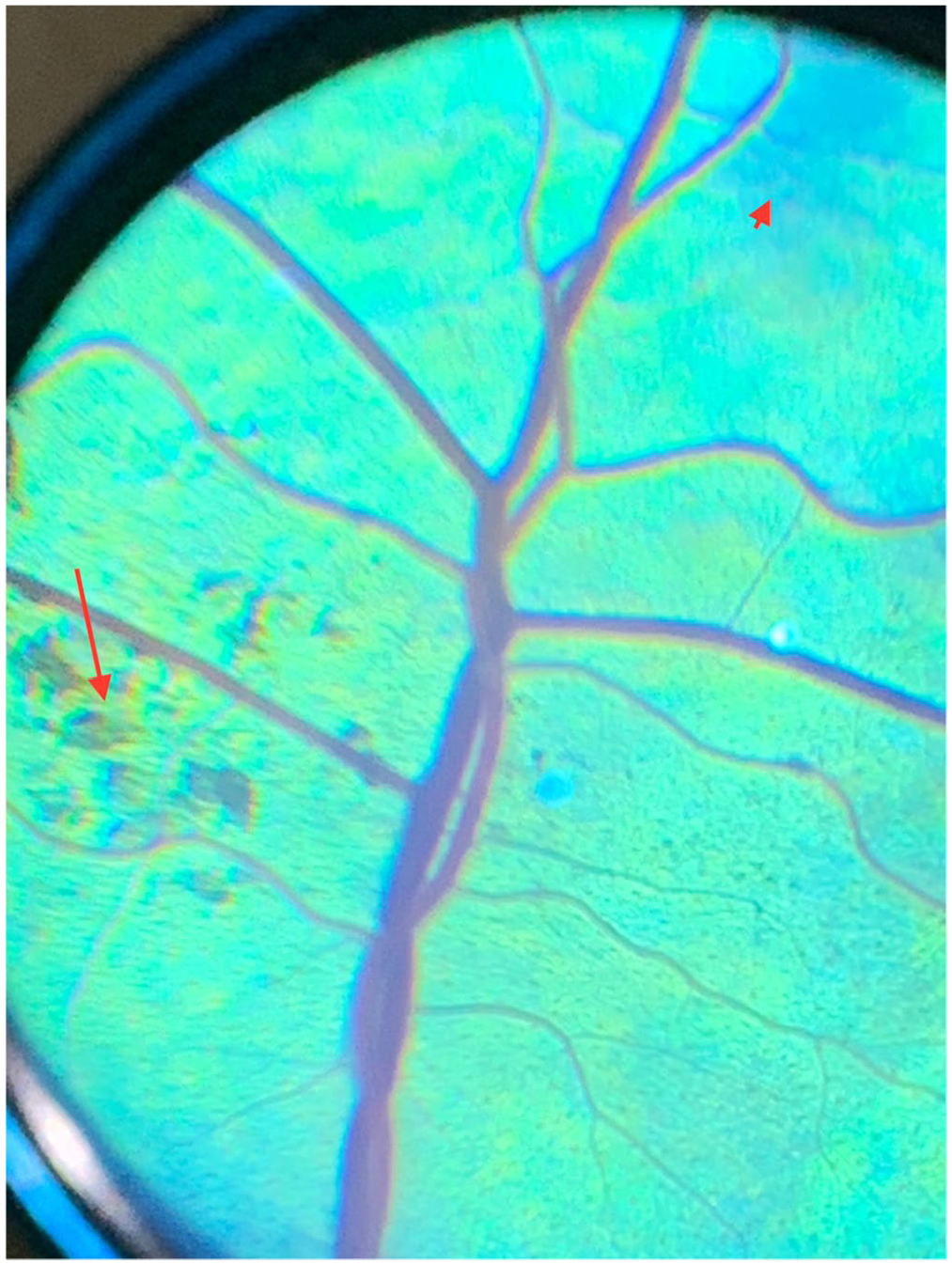
Focal area of the tapetal ocular fundus along the dorsal retinal venule and arteriole of blind Hereford cow V10 with moderate retinal vessel attenuation in addition to diffuse altered tapetal hyper-reflectivity (short arrow) and multifocal pigment clumping (long arrow).
Ophthalmic examination found that V3 was nonvisual with absent menace response. Dazzle reflexes and pupillary light reflexes were delayed and incomplete. On ophthalmoscopic examination, diffuse mottling of the tapetum, multifocal areas of altered tapetal reflectivity, and darkened optic nerve heads were noted (Fig. 3); there was only mild retinal vascular caliber attenuation relative to the clinically normal sibling cow. Both eyes also had multifocal pigment clumping in the tapetal fundus, although this was also seen in her sibling. The remainder of the ophthalmic examination of the sibling was normal. The electroretinogram (ERG) of the clinically affected heifer was extinguished with only regular electrical cycle noise and no discernable or measurable a- or b-wave amplitudes or implicit times (Suppl. Fig. 1). The ERG of the clinically normal sibling heifer was not able to be comprehensively recorded and measured due to excessive environmental electrical noise. However, the waveforms obtained and seen during attempted acquisition appeared normal and with adequate amplitude. The nonvisual status, diffuse tapetal alterations on fundic examination, and absence of waveforms on the electroretinogram were consistent with a diagnosis of retinal degeneration.
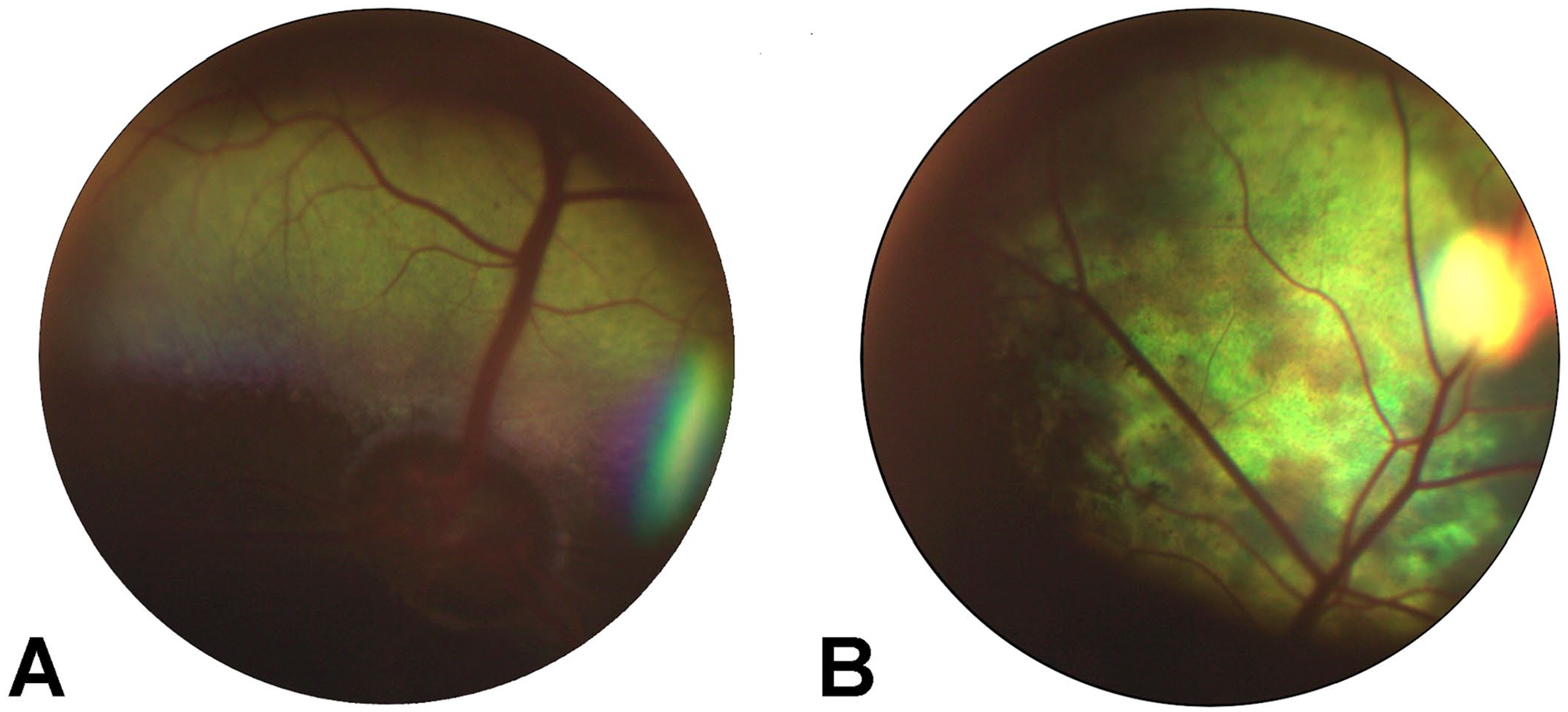
Fundus photographs of clinically normal heifer (A) and clinically affected full-sibling heifer (B).
Examination of blind cow VI1 confirmed her blindness and reported similar fundoscopic changes consistent with retinal degeneration.
Pathology report
The retinas and brain tissues of the 6 control cattle were examined and determined to be normal. These animals served as controls for pathology and RNA expression.
Ocular histology confirmed the clinical diagnosis of blind cows V10 and V11. The anterior segment was normal in both cows. The retina easily detached and retained connection only near the optic disc and ora ciliaris retinae through processing. Marked retinal degeneration was present, evidenced by the absence of rod and cone cells in the outer and inner segments (Fig. 4). The retinal pigment epithelium (RPE) was intact. The outer nuclear layer was absent, and the outer plexiform layer was diminished. There were rare cellular processes extending from the mildly depleted inner nuclear layer into the outer plexiform layer where a few glial cell nuclei remained. The inner plexiform layer, ganglion cell layer, and nerve fiber layers were intact. No vacuolated lymphocytes were observed in the blood. The histology of the nervous system was also normal. Our histologic diagnosis was severe retinal atrophy with loss of the photoreceptor layer, outer nuclear layer, and outer plexiform layer, with a diminished inner nuclear layer. The 2 cows were gravid, and each fetus had normal fetal retinas. In the 2 cows and their fetuses, other tissues (brain, cardiac muscle, skeletal muscle, liver, kidney, lung, spleen, lymph node) were normal.
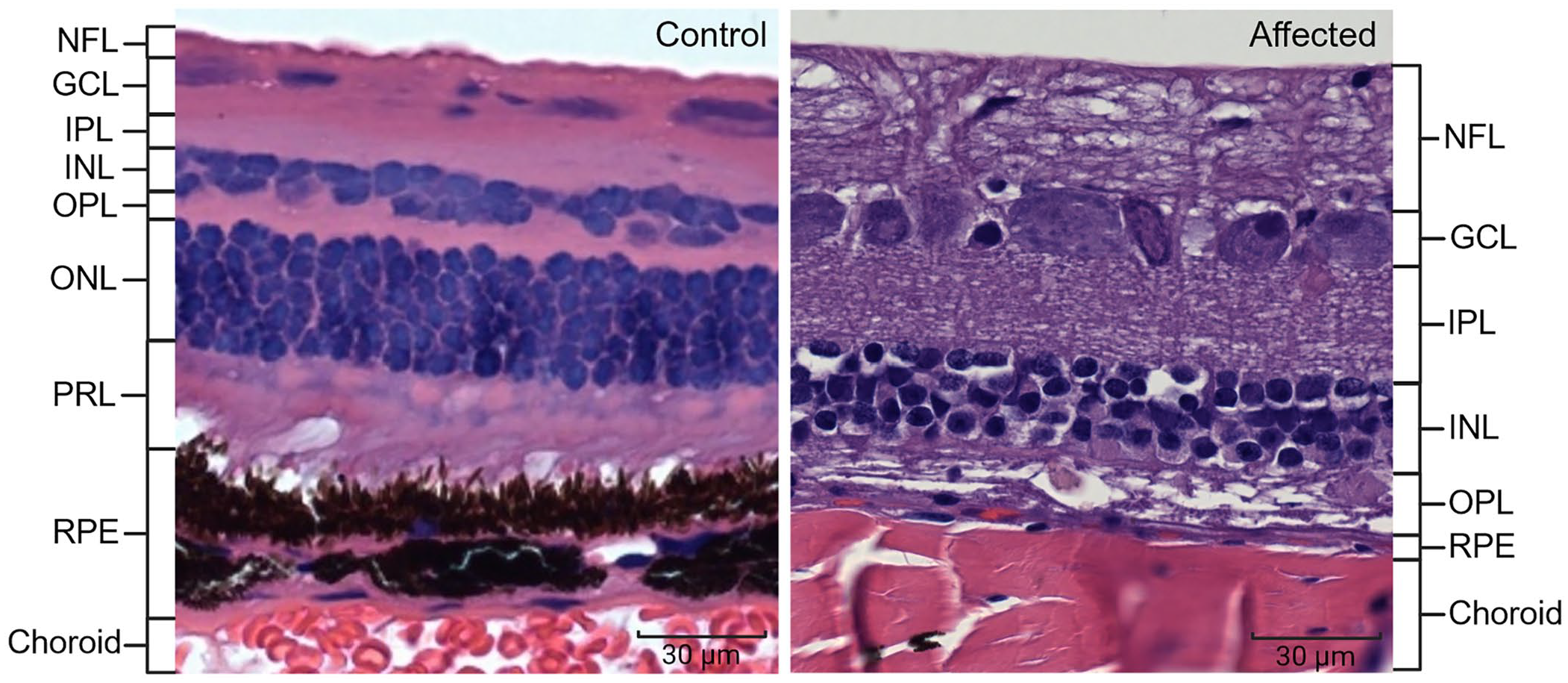
Retinal layers of a clinically normal cow (left) and a clinically affected cow (right). The outer nuclear layer (ONL) and photoreceptor layer (PRL, rods and cones) are absent in the affected retina. GCL = ganglion cell layer; INL = inner nuclear layer; IPL = inner plexiform layer; NFL = nerve fiber layer; OPL = outer plexiform layer; RPE = retinal pigment epithelium. Bars = 30 μm.
A histologic view of the retina from affected cow V10 showed that the outer nuclear, and inner and outer photoreceptor layers, were missing (Suppl. Fig. 2).
Sequencing results
After filtering whole-genome sequence data of 9 blind cattle and 7 unaffected relatives for loci that were homozygous for the alternative allele only in the blind cattle, 280 loci (Suppl. Table 3), all in chromosome 25, matched the criteria. Of those, all but 37 variants had been previously documented as determined by reference single-nucleotide variation (rs) identifiers. Three loci, one novel, were predicted to impact protein function (Table 2).
Variants fitting filter criteria with a predicted impact (variant effect predictor) on gene function. Variant positions are in relationship to reference genome ARS-UCD1.2.
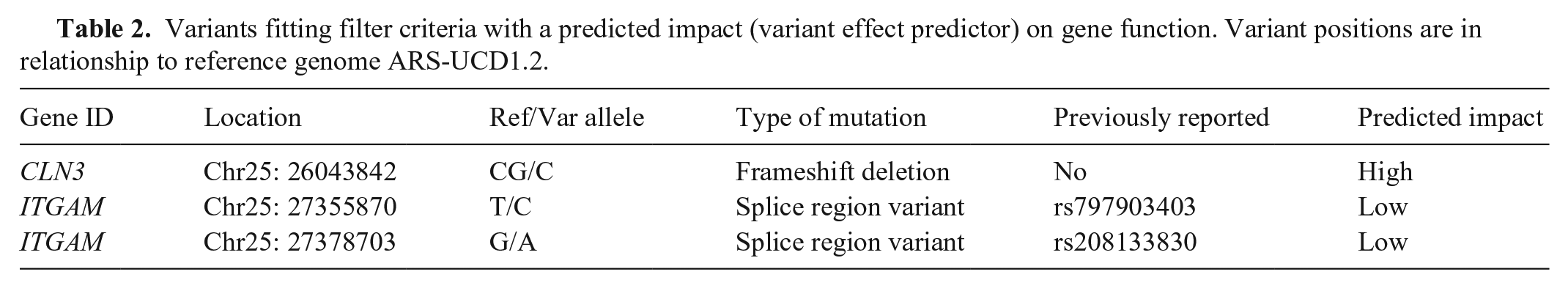
For the 2 candidate loci in Table 2 that were previously documented, the SRA search of 327 samples labeled “Hereford” found 8 heterozygotes for rs797903403 and 20 heterozygotes plus 2 homozygotes for the rs208133830 variant. Whole-genome sequence data from the University of Missouri, which included ~5,500 cattle (Robert Schnabel, pers. comm., 2023), found the rs797903403 variant present in 14 different breeds (allele frequency = 0.004), and the rs208133830 variant in 153 breeds (allele frequency = 0.335). The rare rs797903403 variant was not homozygous in any animals; the more common rs208133830 variant was homozygous in 1,144 animals. Given the distribution of these variants across several breeds, both ITGAM variants were determined to be unlikely as causative of blindness in these Hereford cattle. In searches of both the SRA and University of Missouri databases, the novel frameshift deletion in CLN3 (chr25 g.26043843del) was only observed in Hereford animals. The 18 Herefords in which it was found were each heterozygous. This variant in CLN3 thus became the primary candidate due to being restricted to the Hereford breed and only being observed in the homozygous state in cattle reported to be blind. The CLN3 frameshift deletion (c.1106del) in exon 16 (ENSBTAT00000023952.6) is predicted to result in amino acid changes (p. Pro369Argfs*8) until a premature stop codon truncates the normally 438 amino acid protein at residue 375.
Through either WGS or Sanger sequencing, 462 Hereford cattle were genotyped for the CLN3 mutation (Table 3); 367 were genotyped in our laboratory, and 95 genotypes from WGS were provided by the American Hereford Association. All blind cattle available for genotyping (n = 11 of 13) were homozygous for the variant. All parents of the blind cattle that were available for genotyping (n = 12 of 14) were confirmed carriers of the deletion. The common ancestor, I1, was confirmed to be a carrier as was his maternal grandsire; DNA was not available from other ancestors of I1. The full-sibling of V3, 2 full-siblings of V10 and V11 (through embryo transfer), and the fetuses of V10 and V11 were carriers of the variant. The 6 abattoir cattle genotyped as wild type. A query of the American Hereford Association herd book, which includes ~4.5 million animals, identified 1.2 million descendants (dead and living) of the oldest genotyped carrier of the CLN3 variant. Genotyping performed for research reduced the number to ~700,000 potential carriers by identifying influential descendants that did not inherit the mutation.
Genotypes for 462 Hereford cattle genotyped for the CLN3 mutation (chr25 g.26043843del).
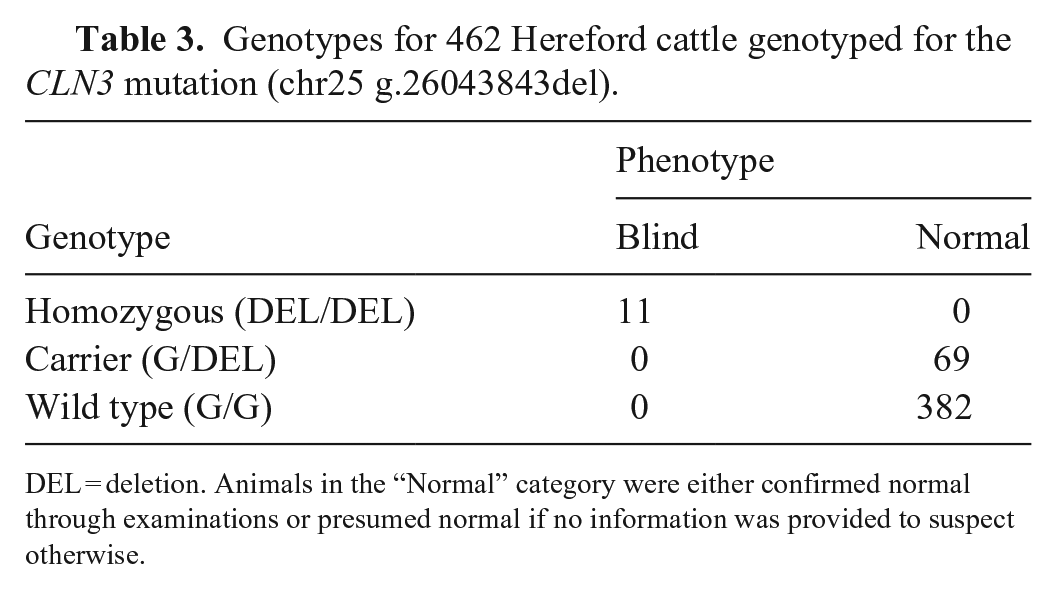

DEL = deletion. Animals in the “Normal” category were either confirmed normal through examinations or presumed normal if no information was provided to suspect otherwise.
Expression of CLN3
Relative qPCR demonstrated that CLN3 was expressed in the retina of the blind cattle with no significant difference in the expression of CLN3 compared to control cattle (Suppl. Table 4).
Discussion
We found that delayed-onset retinal degeneration and blindness in Hereford cattle was associated with a homozygous deletion in CLN3 predicted to truncate the protein. The deletion was only observed in Hereford cattle and could be traced to a common ancestor. Though the gene was expressed in the retina of affected individuals similarly to normal cattle, the observed retinal degeneration provides evidence that the truncated CLN3 protein is dysfunctional, leading to blindness.
The protein encoded by CLN3, battenin, is a membrane transporter expressed in the RPE that plays a role in photoreceptor recycling.6,23,36 Targeting studies found that battenin localizes to lysosomes and endosomes,14,17 but aberrant proteins may instead be retained in the endoplasmic reticulum or may have altered expression.22,23 One explanation for altered localization of the variant protein may be the loss of a lysosomal targeting motif in the cytoplasmic C-terminal tail, which normally guides the protein to lysosomes.7,18 However, human studies suggest that the truncated CLN3 protein may possess partial function if the N-terminal is retained. 7 Therefore, despite the predicted truncation of the cytoplasmic tail, the variant Hereford protein may retain some functionality due to the unaltered N-terminus.7,18,23 Although we did not assess protein function and expression, our real-time PCR results demonstrated that expression of CLN3 in the retinas of the blind cattle was similar to that of the control cattle.
In human medicine, recessive variation in CLN3 has been identified as causative of juvenile-onset retinal and neuronal neurodegeneration, named juvenile neuronal ceroid lipofuscinosis (JNCL), or Batten disease. 24 JNCL is part of a group of neurodegenerative disorders termed neuronal ceroid lipofuscinoses (NCL) that cause blindness due to retinal degeneration and may include seizures and death due to neuronal atrophy; the disorders are classified by the age of onset. 12 The causative genes of JNCL disorders are members of the CLN family: CLN1–3, CLN5, CLN6, and CLN8.12,33 In human JNCL patients, one of the most common disease-causing variants is a 1.02-kb deletion of exons 8 and 9 of CLN3, although missense and nonsense mutations, duplications, and other deletions have also been implicated.22,23,35 Several studies have associated mutations in CLN3 with lysosomal dysfunction.15,35 Normally, the RPE serves to continuously remove waste produced by the outer segments of rods and cones utilizing lysosomes for waste degradation.6,36 Lysosome dysfunction leads to accumulation of degradative byproducts (lipofuscins), particularly in the eye, causing atrophy of the diffuse RPE of the macula.2,24,31
Although we did not observe lipofuscin accumulation in the retina of the blind cattle, the retinal degeneration is very similar to that reported in JNCL patients. The loss of the photoreceptor layer in humans results in undetectable rod and cone responses during ERG recordings,27,35 another result observed in our blind cattle. JNCL patients have sub-retinal dots or sub-foveal deposits, which may translate to the altered tapetal reflectivity observed in blind cattle V3, V10, and V11. 27 Though overt tapetal hyper-reflectivity was observed in blind heifers V10 and V11, the tapetum of blind heifer V3 was diffusely mottled with altered, but not overt, hyper-reflectivity. Retinal vascular attenuation in blind heifer V3 was also only mild. These minor differences are possibly due to her younger age (15 mo) at the time of examination compared to cows V10 and V11 (24 mo). Another common fundic finding in JNCL patients is retinal vessel attenuation. 27 There was minor observer variance due to subjectivity of fundic examination findings in the interpretation of the level of retinal vessel attenuation in the blind cattle; the consensus was that all examined blind cattle had some level of retinal vessel attenuation. The blind cattle lacked the signs of neurodegeneration such as seizures and ataxia reported in human patients with JNCL. 12 However, there is variation in the JNCL phenotype; patients can have mild-to-severe retinal dystrophy but lack neurologic symptoms. 27
There is evidence in the literature that the progression and severity of CLN3 disease are influenced by sex, though it is not yet understood why. In humans, female JNCL patients have a later onset but faster progression of the disease than males. 4 Most blind cattle in our study were female (n = 12 of 13), which may support the hypothesis that sex influences the severity of CLN3 disease. However, this discrepancy may also be due to differences in the management of male and female cattle in beef systems, as males are more often sent to a feedlot to ultimately be harvested whereas a higher proportion of females are retained for breeding. Additional studies are necessary to elucidate whether sex impacts the expression of disease in these cattle as well as if carriers of the mutation may have a subclinical phenotype. Even so, the similarity of these blind cattle to JNCL patients presents the possibility of cattle serving as a model for human non-syndromic JNCL disease.
Our study provides evidence that a 1-bp deletion in CLN3 caused the observed retinal degeneration and blindness in Hereford cattle. Although the cattle were otherwise healthy, blindness can create challenges in navigating unknown terrain, in interactions with other animals, and can cause issues with animal handling, generally impacting the animals’ overall well-being. Given the number of cattle in the herd book that are descendent of carriers, commercial genotyping for the CLN3 variant is ongoing.
Supplemental Material
sj-pdf-1-vdi-10.1177_10406387241239918 – Supplemental material for A recessive CLN3 variant is responsible for delayed-onset retinal degeneration in Hereford cattle
Supplemental material, sj-pdf-1-vdi-10.1177_10406387241239918 for A recessive CLN3 variant is responsible for delayed-onset retinal degeneration in Hereford cattle by Rachel R. Reith, Mackenzie Batt, Anna M. Fuller, Jessica Meekins, Kathryn A. Diehl, You Zhou, Patrick S. Bedwell, Jack A. Ward, Stacy K. Sanders, Jessica L. Petersen and David J. Steffen in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank the Hereford breeders and American Hereford Association staff/personnel for their cooperation and contribution of information and samples. Daniel Garrick, Theta Solutions and The Helical Company, facilitated data transfer. Lianna Walker contributed to the RT-PCR analysis. Harvest was facilitated by the faculty and staff of the University of Nebraska’s Loeffel Meat Lab. Portions of this work were completed utilizing the Holland Computing Center of the University of Nebraska, which receives support from the Nebraska Research Initiative. SEM was carried out at the Microscopy Research Core Facility of the Center for Biotechnology, which is partially funded by the Nebraska Center for Integrated Biomolecular Communication COBRE grant P20 GM113126, and the Nebraska Research Initiative.
Data availability
Whole-genome sequence files are available at the NCBI Sequence Read Archive (accession PRJNA1013498). Variant data for this study have been deposited in the European Variation Archive (EVA) at EMBL-EBI under accession PRJEB71833.
Declaration of conflicting interests
Funding was provided in part by the American Hereford Association. The funders assisted with sample acquisition and pedigree evaluation. The experimental design and analyses of data were independent of the funding agency.
Funding
This project was funded in part by the American Hereford Association.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
