Abstract
Highly pathogenic avian influenza (HPAI) is an acute viral disease associated with high mortality and great economic losses. Immunohistochemistry (IHC) is a common diagnostic and research tool for the demonstration of avian influenza A virus (AIAV) antigens within affected tissues, supporting etiologic diagnosis and assessing viral distribution in both naturally and experimentally infected birds. RNAscope in situ hybridization (ISH) has been used successfully for the identification of a variety of viral nucleic acids within histologic samples. We validated RNAscope ISH for the detection of AIAV in formalin-fixed, paraffin-embedded (FFPE) tissues. RNAscope ISH targeting the AIAV matrix gene and anti-IAV nucleoprotein IHC were performed on 61 FFPE tissue sections obtained from 3 AIAV-negative, 16 H5 HPAIAV, and 1 low pathogenicity AIAV naturally infected birds, including 7 species sampled between 2009 and 2022. All AIAV-negative birds were confirmed negative by both techniques. All AIAVs were detected successfully by both techniques in all selected tissues and species. Subsequently, H-score comparison was assessed through computer-assisted quantitative analysis on a tissue microarray comprised of 132 tissue cores from 9 HPAIAV-infected domestic ducks. Pearson correlation of r = 0.95 (0.94–0.97), Lin concordance coefficient of ρc = 0.91 (0.88–0.93), and Bland–Altman analysis indicated high correlation and moderate concordance between the 2 techniques. H-score values were significantly higher with RNAscope ISH compared to IHC for brain, lung, and pancreatic tissues (p ≤ 0.05). Overall, our results indicate that RNAscope ISH is a suitable and sensitive tool for in situ detection of AIAV in FFPE tissues.
Avian influenza is a highly contagious disease caused by influenza A viruses (IAVs; Orthomyxoviridae, Alphainfluenzavirus).16,24 Wild birds, particularly aquatic and migratory species belonging to the orders Anseriformes and Charadriiformes, represent the natural reservoir of avian IAVs (AIAVs), contributing to viral circulation and transmission to commercial poultry.8,10,21 Since 2016, clade 2.3.4.4b highly pathogenic avian influenza A(H5) viruses (H5 HPAIAVs) have been associated with several panzootics, resulting in high morbidity and mortality in both wild birds and poultry.10,17 Between October 2021 and January 2023, 7,173 H5 HPAIA outbreaks were reported worldwide and were responsible for massive die-offs; European and French outbreaks represented 69% and 22% of outbreaks, respectively. 11
Prompt identification of infected birds, leading to the institution of effective control measures, is crucial to limit the spread of HPAIAVs within a susceptible population. Rapid diagnosis typically relies on molecular detection of AIAVs within clinical specimens and laboratory samples by reverse-transcription real-time PCR (RT-rtPCR) and/or sequencing analysis.1,28 In situ detection of viral antigens or nucleic acids within infected tissues, although more time-consuming and technically demanding, is important to understand the pathogenesis of AI, and to obtain valuable information on viral biodistribution, tissue tropism, and interactions with host cell factors, in both experimentally and naturally infected birds.13,20,25
Immunohistochemistry (IHC) is a safe, economical, and effective method, widely used for the identification of AIAV antigens in formalin-fixed, paraffin-embedded (FFPE) tissues using specific antibodies. The most reliable and commonly used target, thanks to its highly conserved sequence and abundance within the virion, is the IAV nucleoprotein (NP). 20 Other targets include IAV hemagglutinin, matrix protein, and nonstructural protein 1.14,26,29
RNAscope in situ hybridization (ISH) is a novel and commercially available technique that allows the visualization of single RNA molecules in cytologic and histologic samples, at a single-cell level. 27 RNAscope ISH differs from conventional ISH by the use of a combination of RNA-specific oligonucleotide probes (Z) specifically designed to hybridize to the target RNA in pairs (ZZ). 27 Because a 1-kb region is commonly targeted by 20 probe pairs, sequential hybridizations with the preamplifier, amplifier, and label probes can theoretically yield up to 8,000 labels for each target RNA. Overall, RNAscope ISH has been reported to be more sensitive than conventional ISH and/or IHC, with a substantially similar specificity to both in situ detection tools, and it appeared to be fairly concordant with RT-PCR.2,7,23,27
RNAscope ISH has been widely used in humans, non-human primates, pigs, horses, cattle, cats, dogs, and mice for the identification of a variety of viral pathogens, including severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), porcine epidemic diarrhea virus, equine arteritis virus (Alphaarterivirus equid), bovine viral diarrhea virus (Pestivirus bovis), and feline and canine papillomaviruses (Lambdapapillomavirus 1 and 2, respectively).4–7,15,22,30 This technique has been applied successfully for the detection of duck hepatitis A virus (Avihepatovirus A) genotype 3 in experimentally infected Pekin ducks, a novel orthoreovirus in American corvids, and AIAV NP in a mouse model comparing A(H7N9) and A(H9N2) infections.12,18,31
We aimed to validate an RNAscope ISH assay for the detection of the AIAV matrix protein gene (M gene) and to compare it to the routinely used anti-IAV NP IHC on selected FFPE tissues obtained from naturally infected birds.
Methods
Case selection
We selected 61 FFPE tissue sections from 20 birds sampled between 2009 and 2022, including 3 AIAV-negative commercial chickens (5 tissue sections) and 17 naturally AIAV-infected birds detected by RT-rtPCR, in compliance with official surveillance procedures (Suppl. Table 1). Positive birds included commercial poultry infected with clade 2.3.4.4b H5N8 HPAIAVs from the 2020–2021 and 2016–2017 epizootics (10 mule ducks, 1 quail, 1 goose; n = 48), commercial poultry infected with H5 HPAIAVs distant from Gs/Gd/96-like group HPAIAV from the 2015–2016 French outbreak (1 chicken, 1 guinea fowl; n = 2), 2 captive Houbara bustards infected with H5N8 AIAV (n = 5), and a commercial turkey infected with H6N1 low pathogenicity AIAV (LPAIAV; n = 1).
Tissue microarray construction
For computer-assisted quantitative (CAQ) analysis, a tissue microarray (TMA) was constructed using FFPE tissue blocks obtained from 9 commercial mule ducks naturally infected with clade 2.3.4.4b H5N8 HPAIAVs from the 2020–2021 epizootic (3 flocks, 3 birds/flock). To compare the 2 techniques on tissues with different degrees of viral expression, distribution, and lesion patterns, we selected brain, heart, pancreas, lung, and spleen. Upon histologic assessment, representative lesion areas (regions of interest, ROI) were identified and marked on the blocks (3 ROI for each organ). Subsequently, 1-mm tissue cores were harvested from the donor blocks and transferred into a blank recipient paraffin block (MiniCore 3, Alphelys; Toulouse University Hospital, Toulouse, France). The TMA was designed asymmetrically, and tissues were distributed randomly; triplicates were placed successively.
Histopathology
From the TMA and each FFPE tissue block, 3-µm serial sections were produced and used for H&E staining, anti-IAV NP IHC, and RNAscope ISH targeting the AIAV M gene.
Immunohistochemistry
Immunostaining was performed using a monoclonal mouse antibody directed against IAV NP (HB65, FCG013; Kerafast). Specifically, the immunohistochemical protocol included an antigen retrieval step with pronase 0.05% applied for 10 min at 37°C, a peroxidase blocking step of 5 min at room temperature (S2023; Dako) followed by saturation of nonspecific binding sites with normal goat serum (X0907; Dako) applied for 25 min at room temperature, and overnight incubation with anti-AIAV NP antibody (1:800 dilution) at 4°C. For TMA-based quantitative comparison, immunohistochemical staining was visualized (ENVISIO FLEX HRP Magenta system; Dako Omnis) according to the manufacturer’s recommendations; 3,3′-diaminobenzidine (DAB) chromogen was used for qualitative comparison.
RNAscope ISH
The M gene RNAscope probe (V-H5N8-M1M2) was custom-designed (Advanced Cell Diagnostics) and consisted of 20 antisense oligopairs targeting the well-conserved IAV M genes 1 and 2 of clade 2.3.4.4b H5N8 HPAIAV (A/chicken/France/20P016448/2020; Suppl. File 1). The sequence submitted for probe design was selected to provide the highest homology with other AIAVs, H5 HPAIAVs in particular. RNAscope ISH assays were assessed using RNAscope 2.5 high-definition red assay according to the manufacturer’s recommendations, including mild pretreatment conditions (15-min incubation with protease digestion for antigenic retrieval) and hematoxylin counterstaining. A probe targeting the dihydrodipicolinate reductase (dapB) gene from the Bacillus subtilis strain SMY, served as negative control; a probe directed against the peptidyl-prolyl cis-trans isomerase B (PPIB) gene from chicken and AIAV-positive tissues were used as positive controls for the validation steps (Advanced Cell Diagnostics).
Quantitative comparison
Quantitative comparison was assessed on serial sections obtained from the TMA block stained with both M gene RNAscope ISH and anti-IAV NP IHC. Whole slides were scanned at 40× magnification (3DHISTECH Pannoramic scanner; epredia) and analyzed using open-source software for digital image analysis (QuPath, https://qupath.github.io/). Color deconvolution and automatic cell detection were adjusted for each TMA and each type of organ examined, respectively. Subcellular detection was determined for each core through script-based batch analysis. RNAscope ISH and immunohistochemical detection expression was expressed as an H-score according to the manufacturers’ recommendations.
Statistical analysis
For CAQ analysis, Pearson correlation coefficient (r) and Lin concordance coefficient (ρc) were used to determine correlation and concordance between M gene RNAscope ISH and anti-IAV NP IHC H-score measurements. 19 Additionally, Bland–Altman analysis was performed, and limits of agreement were built after assessing the normality of difference scores. If normality was not observed, limits were built using 2.5 and 97.5 centiles. 3 A Wilcoxon–Mann–Whitney test was used to compare H-score measurements for each organ; p ≤ 0.05 was considered to be statistically significant. All measurements were performed using R software (https://www.R-project.org).
Results
All tissues obtained from AIAV-negative birds were negative with both techniques (Fig. 1A–C). All tested AIAV strains, including H5 HPAIAVs distant from Gs/Gd/96-like group (Fig. 1D–F), clade 2.3.4.4b H5N8 HPAIAV (Fig. 1G–I), and H6N1 LPAIAV, were detected by both techniques in all selected tissues and avian species.
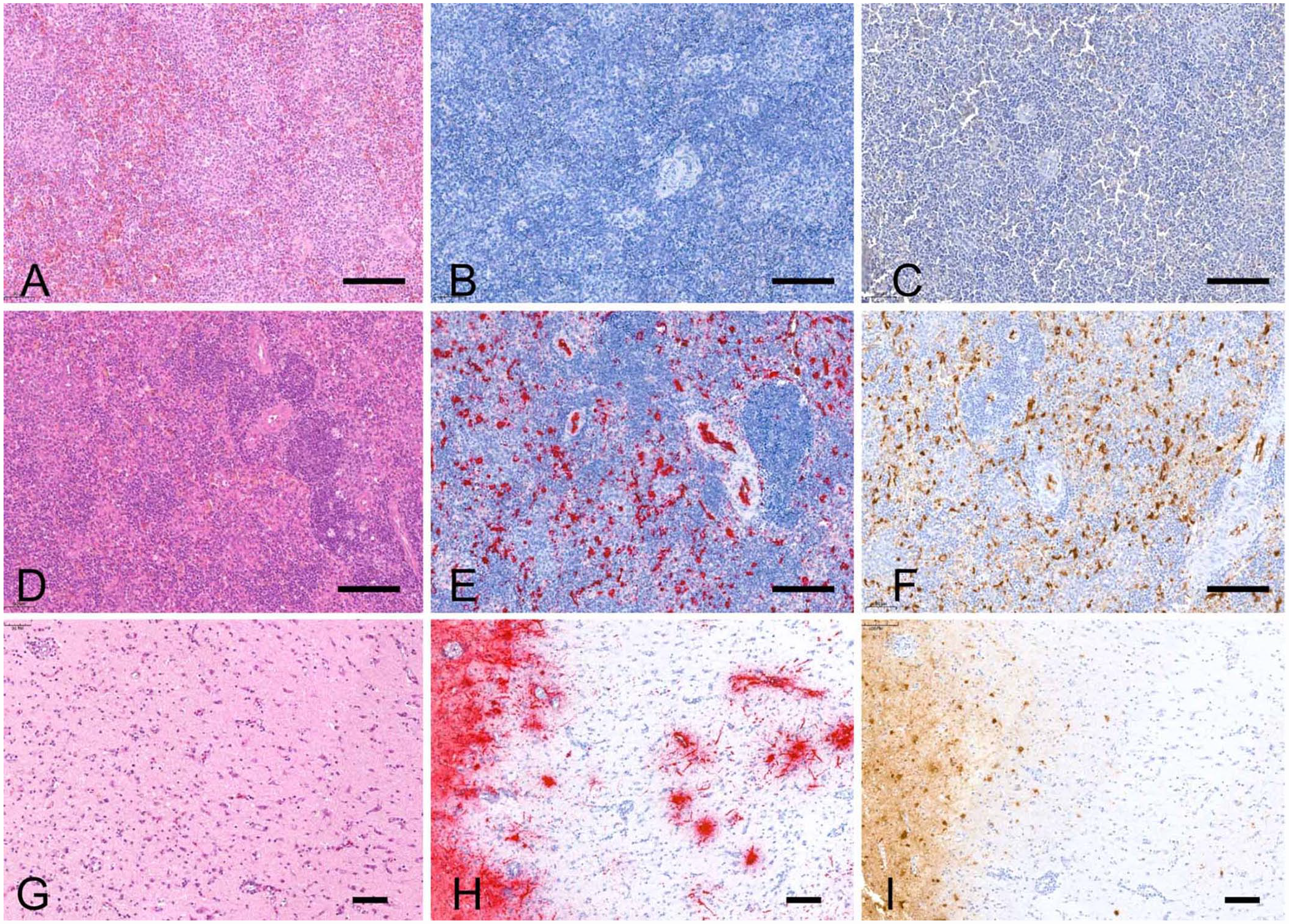
Avian tissues labeled with H&E, anti–influenza A virus (IAV) nucleoprotein (NP) immunohistochemistry (IHC), and RNAscope in situ hybridization (ISH) targeting the matrix (M) gene. Bars = 100 µm.
The distribution pattern of positive cells was similar among techniques. Tissues originating from chickens and guinea fowl naturally infected with H5 HPAIAVs had widespread endothelial expression of both viral NP antigens and M gene RNAs (Fig. 1E, 1F). In ducks, positivity was demonstrated in neurons (Figs. 1H, 1I, 2A), cardiomyocytes (Fig. 2B), pulmonary capillary bed (Fig. 2C), and pancreatic acinar cells (Fig. 2D). In Houbara bustards, mild autolysis of intestinal sections resulted in nonspecific background staining that appeared substantially higher with anti-IAV NP IHC compared to M gene RNAscope ISH. However, positive detection occurred within endothelial cells, myocytes, and mesothelial cells with both techniques. In the turkey naturally infected with H6N1 LPAIAV, both viral antigens and RNAs were detected within tracheal epithelial cells.
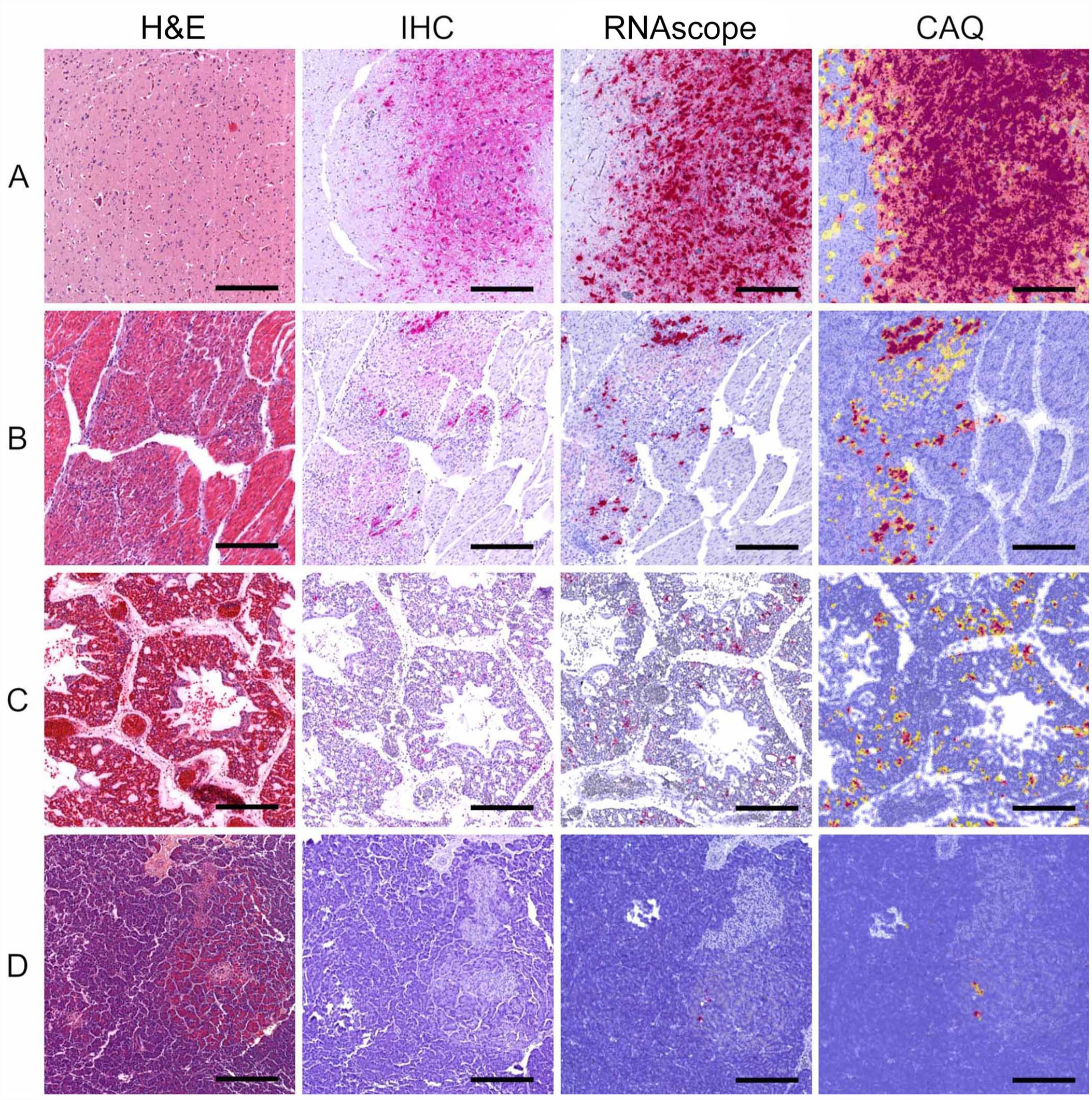
H&E sections and tissue microarray–based computer-assisted quantitative (CAQ) comparison of anti-IAV NP IHC and M gene RNAscope ISH performed on 4 tissue cores of mule ducks naturally infected with H5N8 clade 2.3.4.4b HPAIAV (2020–2021). Bars = 500 µm.
Subcellular distribution appeared slightly different between the 2 techniques. NP antigens could be detected both at a nuclear and a cytoplasmic level by anti-IAV NP IHC. In contrast, M gene RNAscope ISH highlighted viral RNAs mainly as coalescing-to-diffuse cytoplasmic clusters or a mix of cytoplasmic and nuclear dots. On the other hand, M gene RNAscope ISH was able to highlight positive cells, as cytoplasmic and nuclear dots, within IHC-negative areas of brain, heart, and lung tissue sections.
Of 132 tissue cores included in the TMA, 126 were used for CAQ comparison (Fig. 2), and 6 were lost (index core loss: 4.5%). The histologic lesions originally detected in the recipient blocks could also be observed in the tissue cores. Specifically, nonsuppurative inflammation and parenchymal necrosis were invariably present in brain tissue cores, more variably identified in heart tissue cores, and only rarely seen in spleen, lung, and pancreas tissue cores. Spleen tissue cores were unremarkable. The 2 techniques were correlated, with a Pearson correlation coefficient of r = 0.95 (0.94–0.97). The concordance between the 2 techniques was moderate, with a Lin concordance coefficient of ρc = 0.91 (0.88–0.93). 16 Bland–Altman analysis indicated that the 2 techniques had a bias (median difference H-score RNAscope – H-score IHC) of 12.7 (Fig. 3). The limits of agreement appeared wide, ranging from 0 to 102. Disagreement was higher between the 2 techniques for average H-scores of 50–250. Brain tissue cores provided the highest H-score values, with a median of 226 (interquartile range [IQR]: 146–262) and 139 (IQR: 47–202) for M gene RNAscope ISH and anti-IAV NP IHC, respectively (p < 0.05, Fig. 4); heart tissue cores had lower H-scores with median values of 6 (IQR: 3–17) and 5 (IQR: 3–12), respectively. For lung, spleen, and pancreas tissue cores, detection measurements appeared low, with H-scores < 1, and a significant difference between techniques was found for pancreas and lung (p < 0.05; Suppl. Table 2).
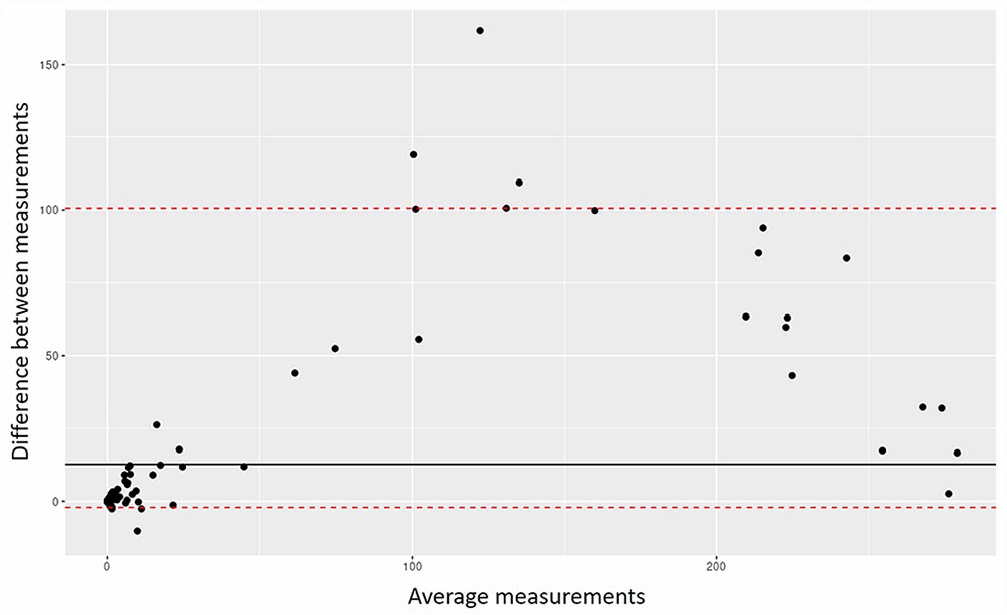
Bland–Altman analysis of M gene RNAscope ISH and anti-IAV NP IHC H-scores. The black line indicates the bias between the 2 techniques, the red dashed lines are the limits of agreement built using the percentiles, and the black dots are the individual tissue core measurements.
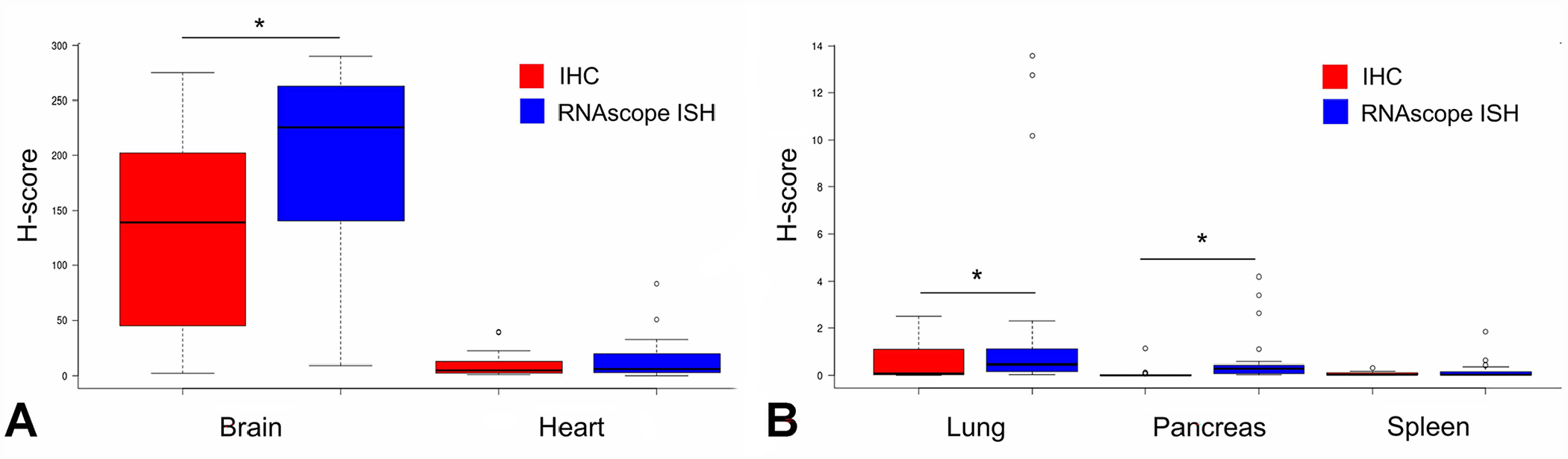
Box plot of M gene RNAscope ISH and anti-AIV NP IHC H-scores.
Discussion
We designed our RNAscope probe to target the AIAV M gene RNAs. Overall, M gene RNAscope ISH appeared to be as specific as anti-IAV NP IHC and was able to successfully detect all selected AIAVs, including clade 2.3.4.4b H5 HPAIAVs, H5 HPAIAV distant from Gs/Gd/96-like H5 HPAIAV, and H6N1 LPAIAV. Quantitative comparison between the 2 techniques indicated high correlation and moderate concordance, explained by higher detection rates with M gene RNAscope ISH, particularly for brain, lung, and pancreatic tissues.
Performance comparison between RNAscope ISH and IHC has been performed in the fields of infectious diseases and oncology. 9 Similar to our results, RNAscope ISH has been reported to be a highly sensitive in situ detection technique that had high concordance with molecular detection (qPCR, RT-PCR, and DNA ISH) and lower concordance with IHC, mostly because of different target molecules (RNAs vs. proteins).2,7,23 In work assessing the use of RNAscope ISH for the detection of SARS-CoV-2 in FFPE human lung tissues, substantial agreement was found between RNAscope ISH and IHC (κ = 0.75), with a specificity of 100% for both techniques 23 ; sensitivity was higher for reverse-transcription droplet digital PCR (96%) compared to RNAscope ISH (36%) and IHC (31%). Another comparative study found that RNAscope ISH was more sensitive than conventional ISH, but not IHC, for the detection of equine arteritis virus. 7
On the one hand, we found that additional positive cells were highlighted by M gene RNAscope ISH compared to anti-IAV NP IHC, suggesting a substantially higher sensitivity of detection within positive tissues. This result can be partially explained by the combination of strong signal amplification and the use of 20 oligopairs specifically designed to hybridize to the targeted RNA. On the other hand, it was occasionally difficult to differentiate nuclear positivity from cytoplasmic positivity with M gene RNAscope ISH, given intense and widespread signal amplification.
Improved viral detection was observed with M gene RNAscope ISH compared to anti-IAV NP IHC in tissues obtained from Houbara bustards, thanks to reduced nonspecific staining. These semi-desert birds reside in geographic areas characterized by high environmental temperatures that can accelerate autolysis, which is a well-recognized cause of background signal. A similar improvement was also observed in tissues originating from wild and captive exotic birds, naturally infected with H5N1 HPAIAVs during the most recent outbreaks registered in France in 2022–2023 (data not shown).
Although the anti-IAV NP IHC can detect a wide range of IAVs, including those affecting humans, equids, and swine, M gene RNAscope ISH is specifically designed to detect AIAV RNAs. A remarkable application of such specificity is the use of M gene RNAscope ISH by our laboratory in lesional tissues obtained from mammals naturally infected with AIAVs, supporting the occurrence of spillover events (data not shown).
Supplemental Material
sj-pdf-1-vdi-10.1177_10406387231182385 – Supplemental material for Validation of an RNAscope assay for the detection of avian influenza A virus
Supplemental material, sj-pdf-1-vdi-10.1177_10406387231182385 for Validation of an RNAscope assay for the detection of avian influenza A virus by Nicolas Gaide, Manuela Crispo, Adam Jbenyeni, Céline Bleuart, Maxence Delverdier, Timothée Vergne, Guillaume Le Loc’h and Jean-Luc Guérin in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank Sophie Peries-Bataille, Christelle Casaroli, and Céline Quintana from the Toulouse University Hospital for their contribution to TMA production, ACDBio for assistance and support with the probe design, the Service of Experimental Histopathology US06/CREFRE-Anexplo for their slide-scanning service, and Dr. Emanuele Turbil from the University Hospital Henri Mondor in Aurillac for his contribution to the statistical analysis. Houbara samples were provided by the International Fund for Houbara Conservation (IFHC). We are grateful to His Highness Sheikh Mohamed bin Zayed Al Nahyan, President of the United Arab Emirates and founder of the IFHC, His Highness Sheikh Theyab bin Mohamed Al Nahyan, Chairman of the IFHC, and His Excellency Mohammed Ahmed Al Bowardi, Deputy Chairman, for their contributions.
Availability of data and materials
Our datasets are available from the corresponding author.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
We performed our study in the framework of the “Chaire de Biosécurité Aviaire”, hosted by the National Veterinary College of Toulouse (ENVT) and funded by the Direction Générale de l’Alimentation, Ministère de l’Agriculture et de l’Alimentation, France. The Chair is also partly funded by the French Comité Interprofessionnel des Palmipèdes à foie gras (CIFOG).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
