Abstract
We report here a transiently culturable oomycete pathogen isolated from a pyogranulomatous tail mass in a cat. The organism was morphologically and genetically distinct from Lagenidium and Pythium species. Following next-generation sequencing (NGS) and assembly of contigs, initial phylogenetic analysis using fragments of the cox1 mitochondrial gene identified this specimen as Paralagenidium sp. after nucleotide alignments with sequences obtained from the Barcode of Life Data System (BOLD). However, further analysis of a concatenation of 13 different mitochondrial genes showed that this organism is unique and different from all known oomycetes. A negative PCR result using primers targeting known oomycete pathogens may not be enough to rule out oomycosis in a suspected case. Additionally, the use of a single gene to classify oomycetes may produce misleading results. The advent of metagenomic sequencing and NGS provides a unique opportunity to further explore the diversity of oomycetes as plant and animal pathogens beyond the current capabilities of global barcoding projects that are based on partial genomic sequences.
Keywords
Oomycetes, like fungi, are osmotrophic eukaryotes that feed by secreting extracellular depolymerizing enzymes into their environment to degrade complex polymers (e.g., cellulose, lignin, proteins), and by transporting the resulting simple, monomeric sugars and amino acids into their own cells. 19 Although similar to fungi in their mycelial growth and potential for dispersal via mitotically formed spores, oomycetes possess cell walls made of glucans and cellulose, rather than chitin, and are not phylogenetically related to fungi.4,16 These filamentous eukaryotes have been found in all parts of the world and in various ecosystems. 23
The most widely known oomycetes of animals are Lagenidium and Pythium species. These pathogens have been associated with disease in dogs, cats, horses, and cattle. 8 Often manifested as chronic progressive subcutaneous skin lesions in these species, oomycosis is microscopically characterized as a mildly eosinophilic pyogranulomatous inflammation containing intralesional septate branching hyphae with non-parallel walls.8,9 Pythium insidiosum infection has also been associated with gastrointestinal and pulmonary nodules in dogs.11,15 In humans, oomycosis occurs as skin and soft tissue, vascular, ocular, and disseminated disease caused by P. insidiosum, P. aphanidermatum, and Lagenidium spp.10,18,21
Over 60% of oomycetes are parasites of plants, causing economically important diseases such as potato blight disease and damping root rot in tobacco plants. 23 With common source infections of plants, animals, and humans in both marine and terrestrial environments, oomycosis presents a unique case in which co-evolution in different host species could lead to the frequent emergence of new pathogens in previously unknown hosts. The use of molecular phylogeny has resulted in the discovery of new genera in the class Oomycota since ~2003, suggesting a greater diversity within this class than previously recognized. 14 However, the use of partial genomic sequences for the classification of newly identified organisms should be done with caution because the term “species identification” could be inappropriate without adequate integration of morphologic, ecologic, and physiologic data with barcode data.6,25
Here we present a case of oomycosis in a cat wherein the assembly of the complete mitochondrial genome showed that the pathogen was neither Pythium nor Lagenidium. Although identified as Paralagenidium using cytochrome c oxidase subunit 1 mitochondrial gene (cox1) phylogeny, genetic analyses of concatenated mitochondrial genes showed that this unique organism is more closely related to plant-associated oomycetes than animal-associated members of this group.
Materials and methods
Case report
A 4-y-old, castrated male, domestic short-haired cat was presented for evaluation of a chronically infected and swollen tail. The client had owned the cat for only a few days. The previous owner of the cat was aware of the infected tail and opted to treat the presumed infection with clindamycin, without effect. The new owner adopted the cat to save him from euthanasia.
On presentation, the swollen area on the tail contained several small fluid pockets or abscesses (Fig. 1). Radiographic examination of the tailbone was unremarkable. Following general anesthesia and intercaudal block with mepivacaine hydrochloride (Carbocaine; Pfizer) to keep the tail flaccid, caudectomy was performed by removing all caudal vertebrae as close to the sacrum as possible, without interrupting pudendal nerve function. The cat recovered fully from surgery, and the surgical site healed without complications. Excised tissue was sent to the Veterinary Medical Diagnostic Laboratory (University of Missouri–Columbia, Columbia, MO, USA) for aerobic and anaerobic culture. The submitted tissue was ground and inoculated onto trypticase soy agar (TSA) with 5% sheep blood, MacConkey agar, Columbia–nalidixic acid agar, Sabouraud dextrose agar, and mycobiotic agar and into thioglycolate broth for aerobic culture at 37°C. Tissue homogenate was also inoculated onto pre-reduced TSA with 5% sheep blood and pre-reduced phenylethyl alcohol agar for anaerobic incubation at 37°C. Additionally, tissue homogenate was inoculated onto chocolate agar for incubation at 37°C under capnophilic conditions (5% CO2) and onto additional TSA + 5% sheep blood, Sabouraud dextrose, and mycobiotic agar plates for incubation at room temperature (23°C). Biopsy specimens were fixed in neutral-buffered formalin; H&E-stained sections were prepared by standard methods. Additional sections were stained with periodic acid–Schiff (PAS) and Gomori methenamine silver–hematoxylin (GMS-H) to better observe fungal organisms.
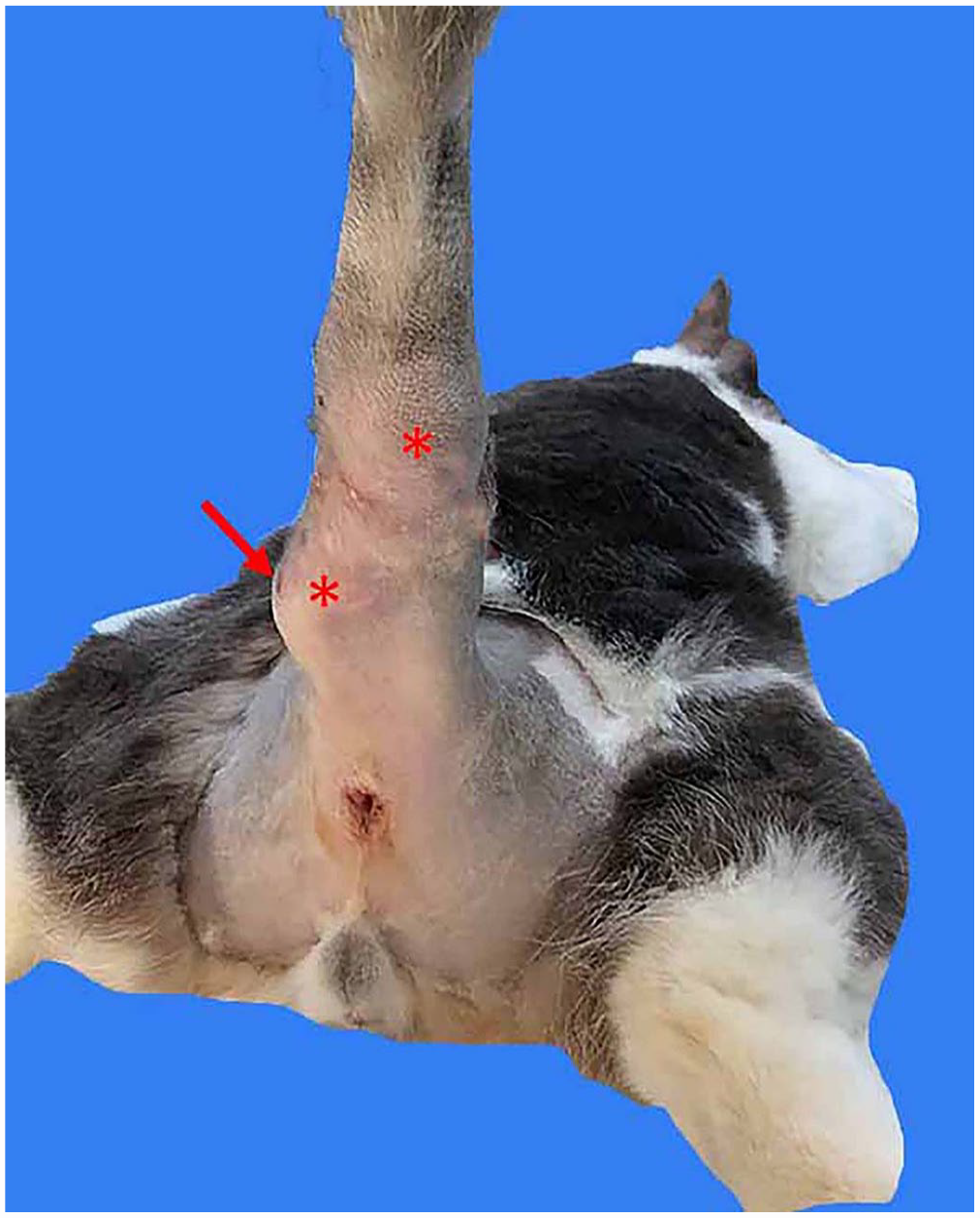
Oomycetic lesions in the tail of cat. Ventral view of the nodular lesions (asterisks), with ulceration (arrow).
Sequencing and assembly
Following centrifugation, the pellet of the thioglycolate broth culture was resuspended in lysis buffer containing proteinase K, vortexed, and incubated overnight on a heating block set to 56°C. A stainless steel bead was added to the sample in a microcentrifuge tube, and tissue disruption was carried out (TissueLyser II; Qiagen). Total nucleic acid was isolated from the disrupted tissue (MagMax magnetic bead-based nucleic acid isolation system; Thermo Fisher). Nucleic acid samples were probed for fungal sequences by a PCR assay targeting the intergenic spacer (ITS) region 2 (ITS86F: GTGAATCATCGAATCTTTGAA; ITS4: TCCTCCGCTTATTGATATGC). 24 For next-generation sequencing (NGS), libraries were constructed (Nextera XT library preparation kit; Illumina) followed by sequencing (MiSeq; Illumina) with paired 250-bp reads.
Paired reads were merged using BBMerge, duplicate reads were removed with Dedupe, and reads were error-corrected and normalized using BBNorm programs executed in the BBTools software suite (v.38.84; https://sourceforge.net/projects/bbmap/). 5 Host genome was removed from reads after alignment to a custom feline genome database created from files downloaded from the genome assembly Felis_Catus_9.0 (National Library of Medicine). MegaBLAST alignment was executed in Geneious Prime (v.2022.0.2, build 2022-01-26) with the following settings: scoring (match mismatch) = 2, maximum e-value = 1e-5, word size = 28, and maximum target seqs = 1 316 134 912.
Contigs were assembled de novo from sequences that returned no hits on the feline genome using Geneious Prime. Annotation of coding regions and prediction of open reading frames (ORFs) was done with the “Find ORFs” function of Geneious Prime using the standard genetic code. The genes identified were confirmed using BLASTX (https://blast.ncbi.nlm.nih.gov/Blast.cgi).
Phylogenetic analyses
Initial phylogenetic analysis consisted of a Bayesian tree (MrBayes v.3.1.2, https://nbisweden.github.io/MrBayes/) constructed in Geneious Prime by comparing 27 sequences of cox1 genes obtained from the Barcode of Life Data System (https://www.boldsystems.org/index.php) and GenBank with 1,100,000 generations, a sampling frequency of 200, and a burn-in of 30%. To generate support for tree topologies, the analysis was bootstrapped with 1,000 replicates using Prototheca zopfii or Achlya hypogyna as outgroups. A concatenated mitochondrial gene set (n = 13) was created using Geneious. The 16-kbp concatenated genome was also used for phylogenetic analysis to generate a Bayesian tree for comparison of tree topologies. Structural comparisons with oomycete pathogens known to infect vertebrates and plants were generated with Circoletto, combining a BLAST search with Circos output. 7 Sequences were submitted to GenBank (OQ557999).
Results
Histopathology
Biopsy specimens had a border of haired skin that covered coalescing inflammatory foci in the dermis and subcutis and extended to all excisional borders (Fig. 2A). Bone was not present. Irregular foci of homogeneous-to-granular debris were comprised of degenerate eosinophils and fewer neutrophils. Cross-sections of degenerate fungal hyphae were present in these foci, outlined by intensely eosinophilic, homogeneous material. Between the eosinophilic foci was coalescing pyogranulomatous inflammation. Fungal hyphae averaged 9 µm in width and had non-parallel walls (Fig. 2B, 2C). Hyphal septa were present but poorly defined. Branching was rare. The remaining tissue had an infiltrate of macrophages, epithelioid cells, and plasma cells. Hyphae stained with PAS and GMS-H. The lesion was characteristic of pyogranulomatous and eosinophilic dermatofasciitis with intra-lesional hyphae consistent with an oomycete, with hyphae surrounded by Splendore–Hoeppli material.
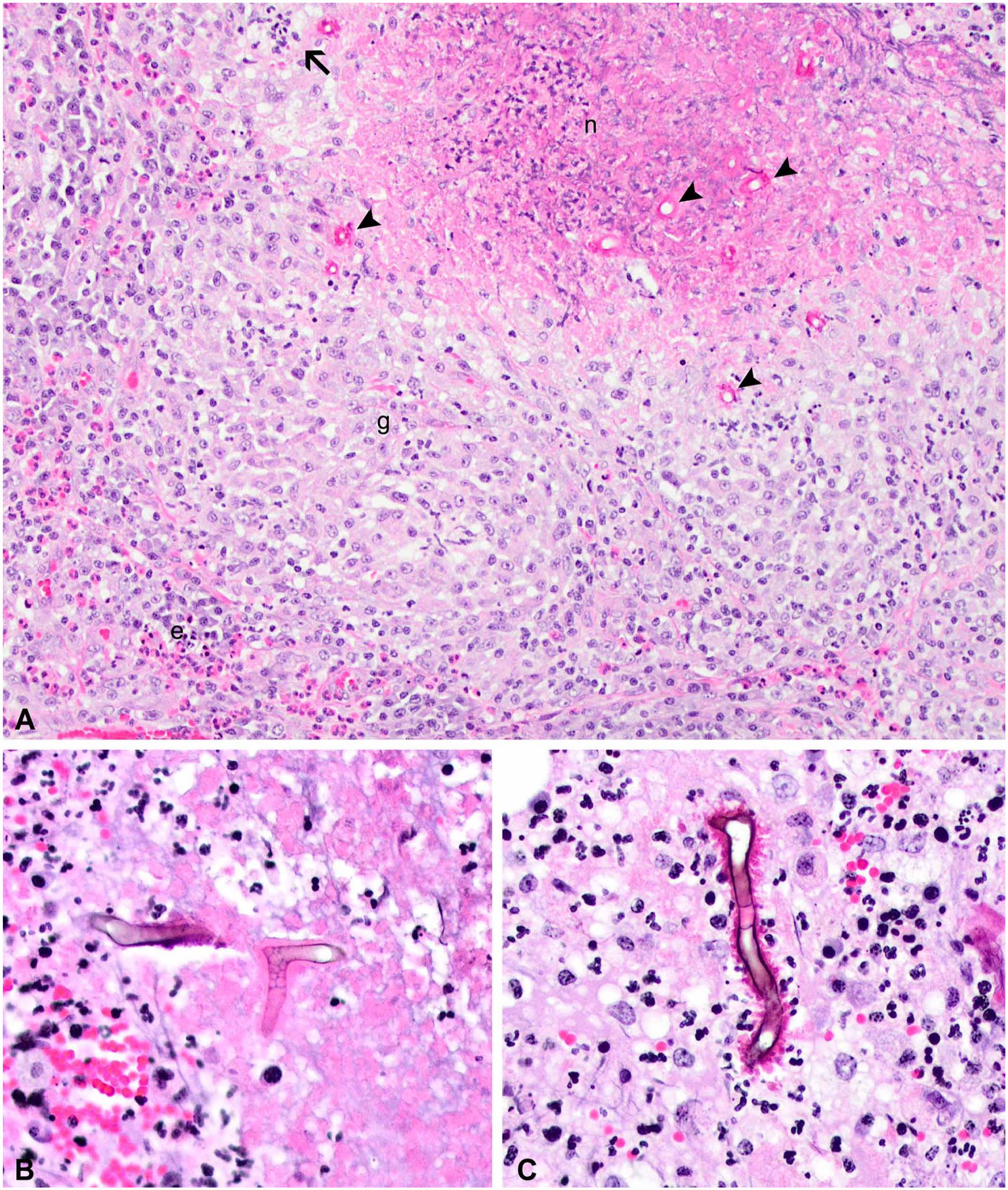
Histologic lesions caused by infection by a novel oomycete in the tail of a cat.
Culture results
Aerobic and anaerobic cultures of excised tissue did not yield known pathogenic bacteria. The thioglycolate enrichment broth yielded a single organism. On agar, the organism grew as firmly embedded clear-to-gray, strongly hemolytic colonies with erose-to-irregular edges. The organism grew on blood agar at 23°C and 35°C. It did not grow on Sabouraud dextrose agar. Microscopically, the isolate was a large (20–50 × 100 µm) ovoid clear-to-basophilic organism that appeared to form pseudohyphae. Staining characteristics did not change with lactophenol cotton blue, Gram, or Wright–Giemsa stains. This organism could not be identified by MALDI-TOF MS using standard libraries (Bruker; Fig. 3).
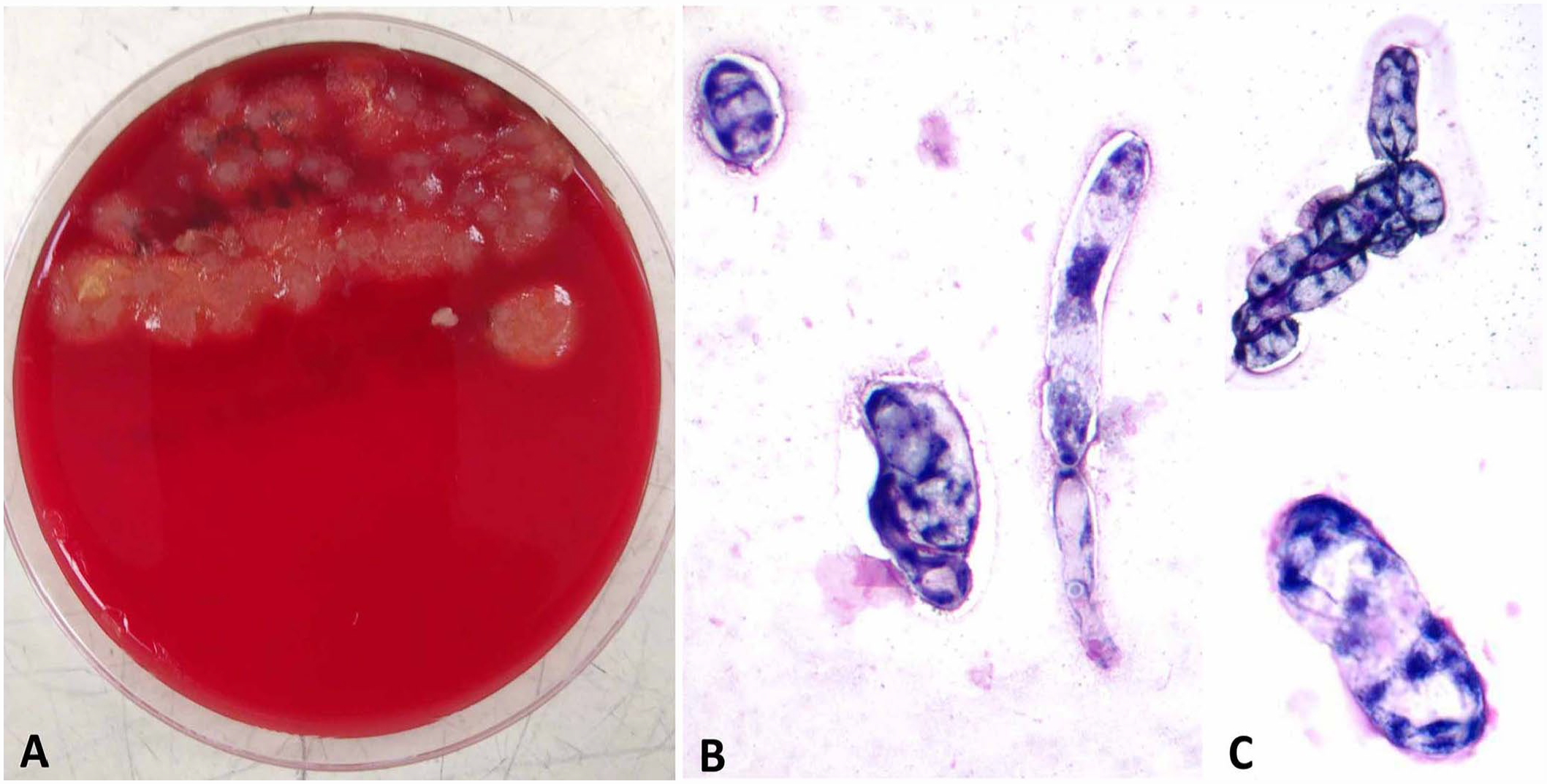
Results of culture of excised tail tissue.
Next-generation sequencing
Our PCR assay using primers specific for fungal ITS2 returned a negative result. However, 2 large contigs totaling 46 kbp were assembled from NGS data. An initial BLASTX search showed that most of the 221 ORFs from our isolate (19-09020) were similar to mitochondrial proteins of known genera of oomycete pathogens associated with plant diseases: Phytophthora, Aphanomyces, Saprolegnia, and Thraustotheca. Similarities with Pythium ultimum and P. insidiosum, oomycete pathogens of animals, were also recorded for some mitochondrial proteins. 1 Phylogenetic analysis using the cox1 gene of the isolate identified it as Paralagenidium sp. (Fig. 4A). Only partial sequences of cox1 from Paralagenidium sp. are available in GenBank. The complete genome of any Paralagenidium sp. is unavailable in GenBank.
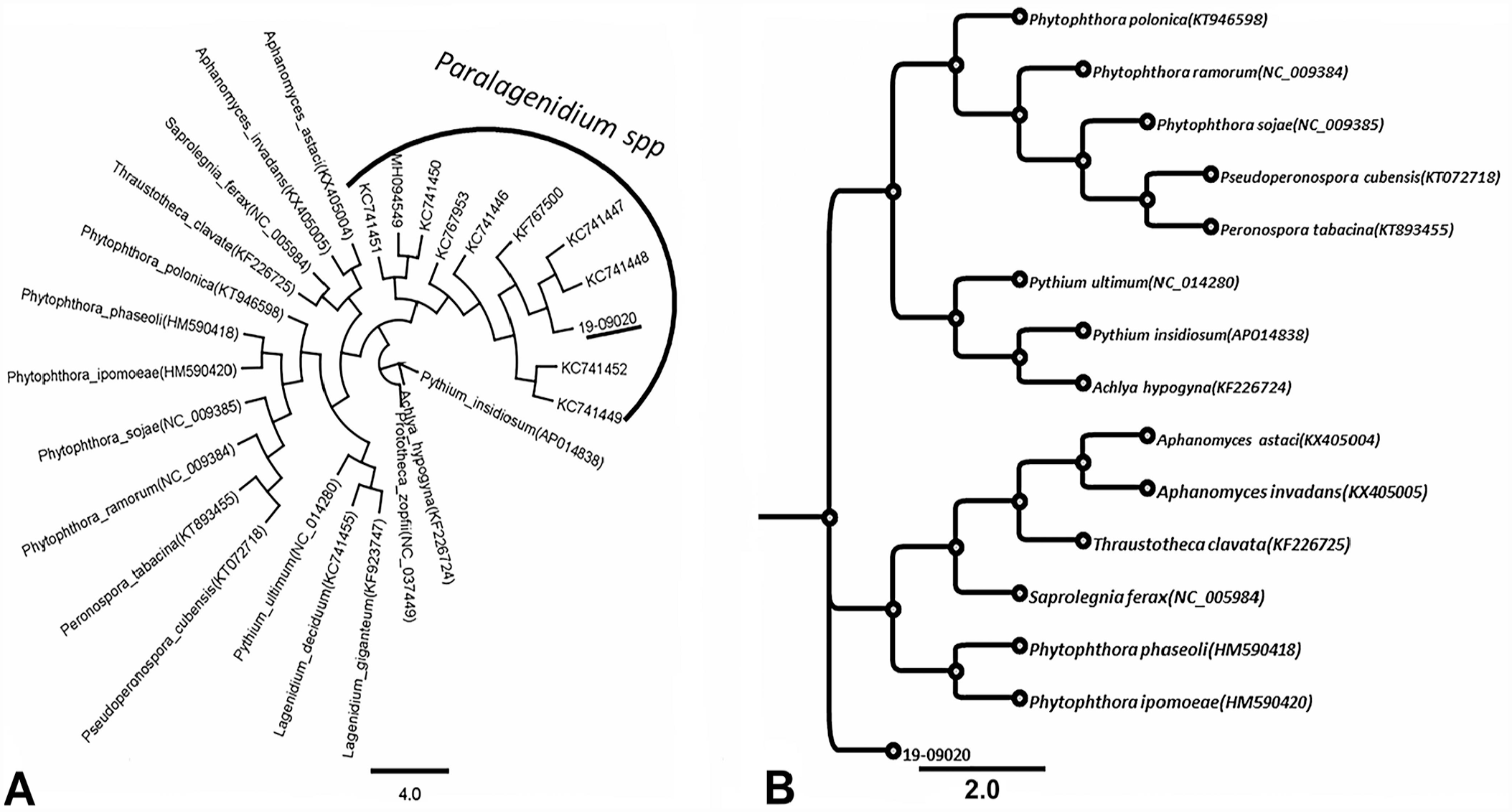
Phylogenetic analysis of the oomycete recovered from lesions in the tail of a cat.
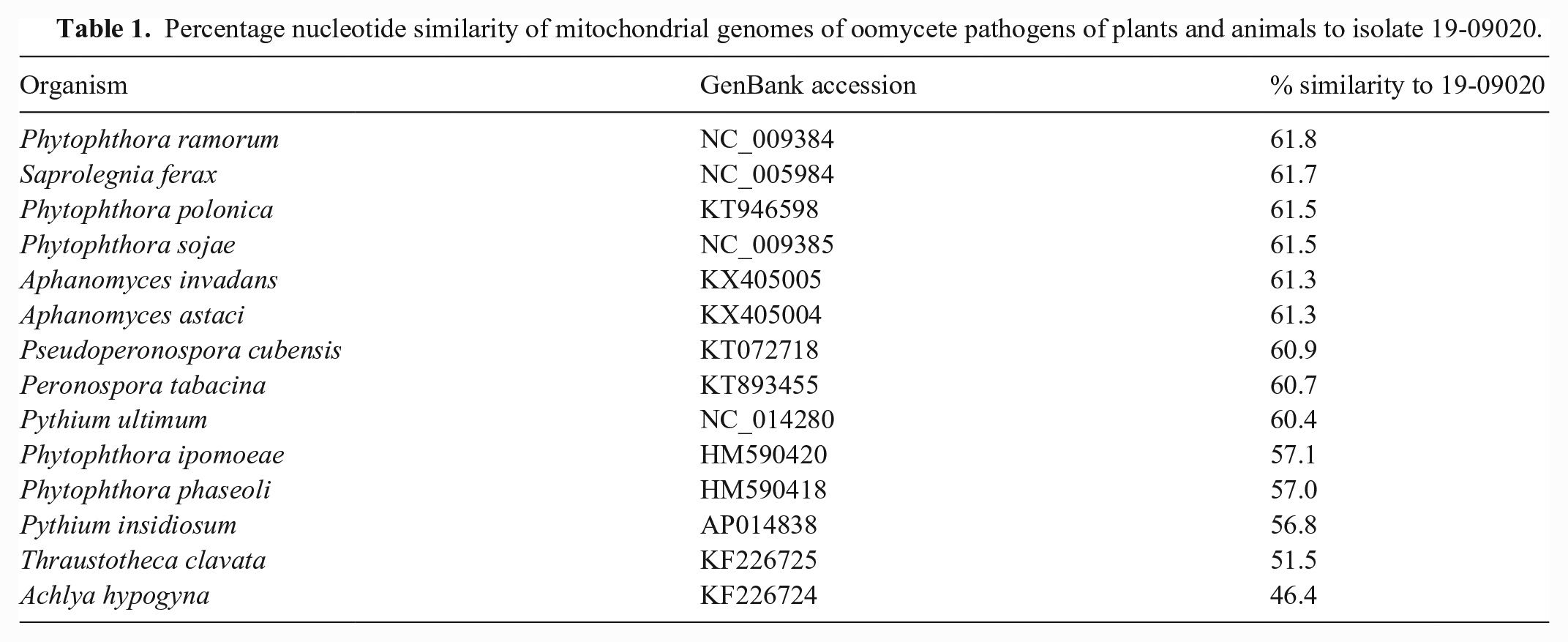
Unlike results obtained using the cox1 gene, phylogenetic analyses that employed genes coding for each of NADH dehydrogenase subunit 5, ATP synthase F1 subunit alpha, and NADH subunit11 had the closest relationship (86–89% similarity) in the aligned regions with Pythium, Phytophthora, Globisporangium, and Plasmopara species (data not shown). Organisms in these genera are oomycetes. This finding suggested that identification based solely on the cox1 gene may not be reliable. Subsequent phylogenetic analysis using concatenated mitochondrial gene sequences (atp1, ATP5F1A, atp6, cob, cox1, cox2, cox3, NAD2, NAD4, NAD5, NAD7, NAD9, NAD11) of members of the class Oomycota showed that our isolate forms a distinct clade in the group (Fig. 4B). On the basis of concatenated mitochondrial gene sequences, isolate 19-09020 had nucleotide similarities of 60.4–61.7% to oomycete pathogens of plants: Saprolegnia ferax, Aphanomyces invadans, Aphanomyces astaci, Peronospora tabacina, Pseudoperonospora cubensis, Phytophthora polonica, Phytophthora sojae, Phytophthora ramorum, and Pythium ultimum. The isolate was least related to P. insidiosum, an oomycete widely associated with disease in animals (Table 1).
Percentage nucleotide similarity of mitochondrial genomes of oomycete pathogens of plants and animals to isolate 19-09020.
To better clarify the relationship between isolate 19-09020 and known oomycete pathogens of plants and animals, a Circoletto plot of 6 selected species of oomycetes was carried out. Isolate 19-09020 was least related to P. ultimum and P. insidiosum (Fig. 5), 7 suggesting that this organism is distinct from oomycete species known to cause disease in cats.
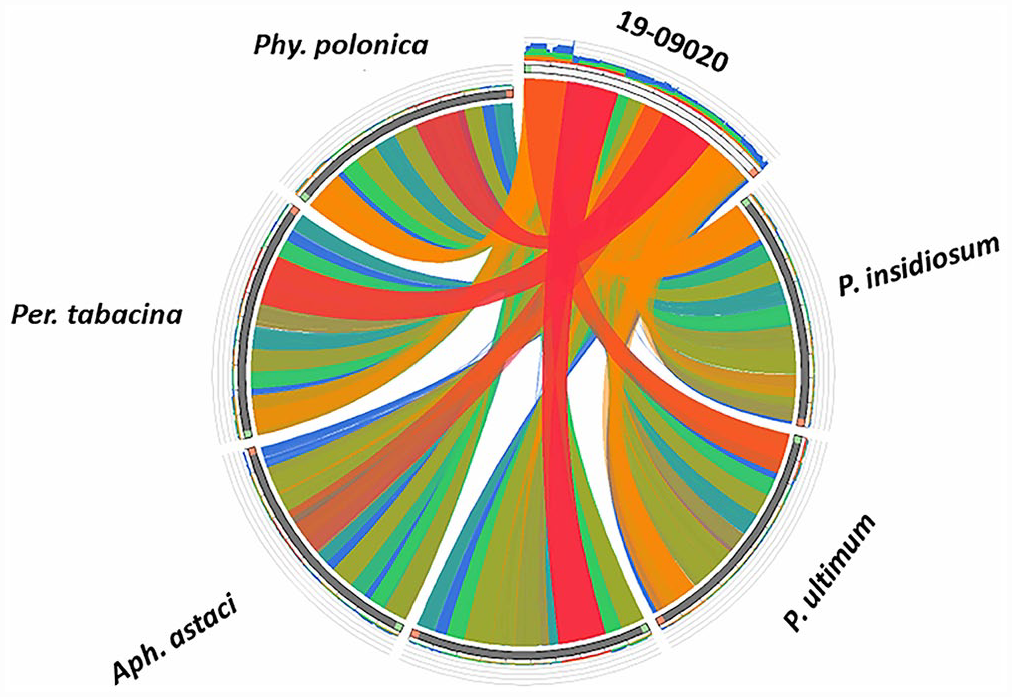
A Circoletto plot of the relationship of the mitochondrial genomes of 19-09020 and oomycete pathogens of plants (Phytophthora polonica, Phy. ipomoeae, Aphanomyces astaci, Peronospora tabacina, Pythium ultimum) and animals (Pythium insidiosum). The width and number of bands joining 2 species indicate genetic relatedness. Isolate 19-09020 is more closely related to oomycete pathogens of plants than those that are known to cause disease in animals.
Discussion
We succeeded in isolating an organism in primary culture in our case. However, we could not maintain the isolate in secondary culture despite several attempts. We surmised that growth in primary culture was probably promoted by growth factors present in feline tissue that formed part of the initial inoculum but were absent in agar and broth media used in secondary culture attempts. Although the tail lesions found in the cat were similar to a previous case of paralagenidiosis in a dog, the organism transiently isolated from the sample did not show cultural and morphologic characteristics described for isolates of Pythium, Lagenidium, or Paralagenidium. 9 Thus, it was impossible to classify our isolate as an oomycete on the basis of morphology. The class Oomycota derived its name from the presence of egg-like structures when organisms in this class were observed under the microscope. 16 However, a major constraint to the use of morphology in the classification of oomycetes is the difficulty in growing and maintaining many of these organisms in culture. 17 Oomycetes are more phylogenetically and morphologically related to diatoms and brown algae in the kingdom Straminipila than they are to fungi. 3
Focusing on the sequence of cox1 and cox2 genes, we showed that our cat isolate did not cluster in the same clade as Lagenidium and Pythium. However, phylogenetic analysis based on comparison with partial cox1 gene sequences available in the database showed a 100% match with Paralagenidium. Because the isolate is not morphologically similar to known isolates of Paralagenidium, we explored phylogenetic classification based on ORFs of sequences coding for mitochondrial enzymes of this isolate. Our multigene concatenation approach recognized a novel member of class Oomycota as the cause of clinical disease in our case.
Although the advent of Sanger sequencing allowed better phylogenetic characterization of members of class Oomycota in the twentieth century, the use of concatenation in phylogenetic analyses has been reported to be a more reliable method to define new species in the class Oomycota.4,22 Partial Sanger sequencing of the gene coding for cox2 has confirmed that oomycetes were indeed phylogenetically distinct from fungi. 16 Beyond the genus level, the global genome barcode project uses the sequence of cox1 as a simple standardized single molecular classifier for delineating species. 13 Indeed, studies using cox1 and sequences of the ITS regions of > 1,000 isolates of oomycetes that covered 23 different genera confirmed that Lagenidium and Pythium, species known to cause disease in mammals, belonged to the same clade and were distinct from Paralagenidium. 20
Following a negative PCR result for Lagenidium and Pythium in a clinical case suggestive of oomycosis in cats, we investigated potential novel members of the class Oomycota. Similarly, a positive PCR result using primers targeting the cox1 gene of Paralagenidium karlingii may not be a reliable marker of paralagenidiosis in domestic animals. Our findings highlight the limitations of Sanger sequencing in delineating species while underscoring the potential contributions of metagenomic sequencing in generating more data to facilitate clear and accurate definition of new species as microbial systematics attempts to exit the era of Carl Linnaeus, in which species classification is based on morphology and physiology, and move to metasystematics,2,12 in which species delineation will be based on metagenomes.
Footnotes
Acknowledgements
We thank the veterinary technicians at the Veterinary Clinics of West Plains (West Plains, MO, USA) for their help in the clinical care of this patient. We appreciate the work of the laboratory technicians in the bacteriology, molecular diagnostics, and histopathology sections of the VMDL (University of Missouri, Columbia, MO, USA) for help in processing the samples.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
