Abstract
Domestic cat hepadnavirus (DCH) is a novel hepadnavirus, first identified in 2018. DCH is generally detected using conventional PCR assays, which include time-consuming agarose gel electrophoresis. We developed a rapid, sensitive, and specific real-time PCR (rtPCR) assay for the detection of the DCH genome. To streamline the procedure, our rtPCR assay was carried out using blood samples, without DNA extraction. A consensus primers/probe set was designed based on the nucleotide sequences of the surface/polymerase gene of all DCH strains available in GenBank. To exclude the possibility that the PCR reaction was blocked by anticoagulants, we also used a primers/probe set for amplifying the cat beta-actin gene as a reference gene. Our direct duplex rtPCR assay had high sensitivity, with a limit of detection of 10 copies/μL of blood for DCH. Our direct duplex rtPCR assay should be a useful tool for DCH detection and surveillance.
Hepadnaviridae is a family of small DNA viruses that are known to cause liver diseases, such as hepatitis, hepatocellular carcinomas, and cirrhosis. Hepadnaviridae viruses have a broad host range, including mammals, birds, fish, reptiles, and frogs. 12 In 2018, a novel virus similar to the hepatitis B virus (HBV), now known as domestic cat hepadnavirus (DCH; Hepadnaviridae, Orthohepadnavirus, Domestic cat hepatitis B virus), was identified in a domestic cat in Australia. This was the first report of hepadnavirus infection in a companion animal. 1 A 2022 survey of canine serum samples discovered hepadnaviral DNA that was genetically close to DCH. 4 This result suggests the possibility of inter-species transmission of DCH in carnivores. There is a clear need for a rapid assay for DCH, both for detection and to monitor the distribution and prevalence of the virus worldwide.
DCH is a small DNA virus that is estimated to be 23–25 nm in diameter. 16 It has a circular DNA genome that is partially double-stranded and is ~3.2 kb in length. Like other hepadnaviruses, the genome contains 4 overlapping open-reading frames encoding the polymerase (L), surface (S), core (C), and X proteins.1,12 DCH is genetically close to HBV, a prototype hepadnavirus, and like HBV is frequently associated with immunosuppressive infections.1,2,10,16
Given that DCH is a relatively newly discovered virus, little information about its genetic diversity and evolution is available. DCH has been reported in Italy, Thailand, Malaysia, the United Kingdom, and Japan,1,2,4,9,10,16,18 but the distribution and prevalence of DCH in other countries have not been investigated in depth. Most studies have used conventional PCR to detect DCH from clinical specimens, and some groups have used real-time PCR (rtPCR).7,10 DCH can be detected in serum, whole blood, heart, lungs, intestines, kidneys, and spleen.1,2,10,16
Given that DCH can be detected in blood samples, it is necessary to consider whether the anticoagulants present in whole blood or plasma samples interfere with the assays, potentially leading to erroneous negative results.5,14 To facilitate the detection and investigation of DCH, it is particularly important to develop a rapid, sensitive, and accurate method for the detection of this virus. We aimed to develop a probe-based, direct duplex rtPCR assay for the detection of DCH in blood samples that could be used to analyze whole blood samples without time-consuming DNA extraction.
Materials and methods
Design of PCR primers
The complete genome sequence of 18 DCH strains and Felis catus Fca126 chromosome E1 (GenBank NC058381) were used as templates for designing primers and probe sets. Two primers and probe sets were designed based on the consensus nucleotide sequences of the surface/polymerase gene of DCH and the ACTB gene of F. catus as a reference gene, using the PrimerQuest tool (https://www.idtdna.com/primerquest/home/index; Table 1; Fig. 1). The sequences of the DCH primers and probe set were aligned with the sequences of 18 DCH strains available in GenBank using CLC Genomics Workbench 22.0.1 in viewing mode (Qiagen) and Primer-BLAST (https://www.ncbi.nlm.nih.gov/tools/primer-blast/) to evaluate any mismatches.
Primers and probe sets used for direct duplex rtPCR for detection of domestic cat hepadnavirus (DCH).
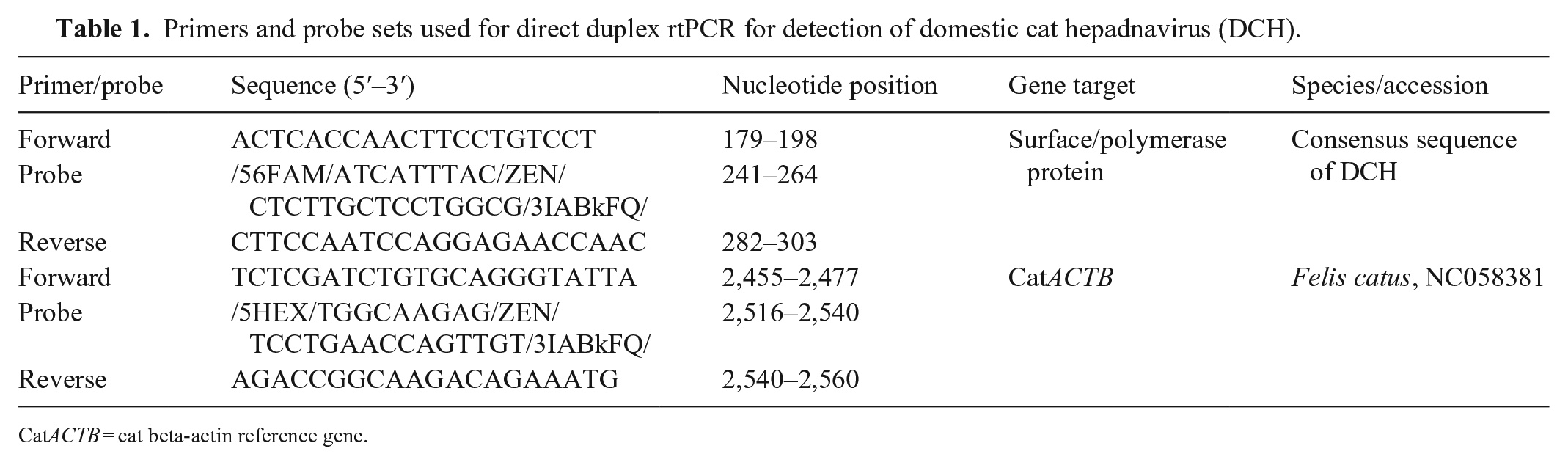
CatACTB = cat beta-actin reference gene.
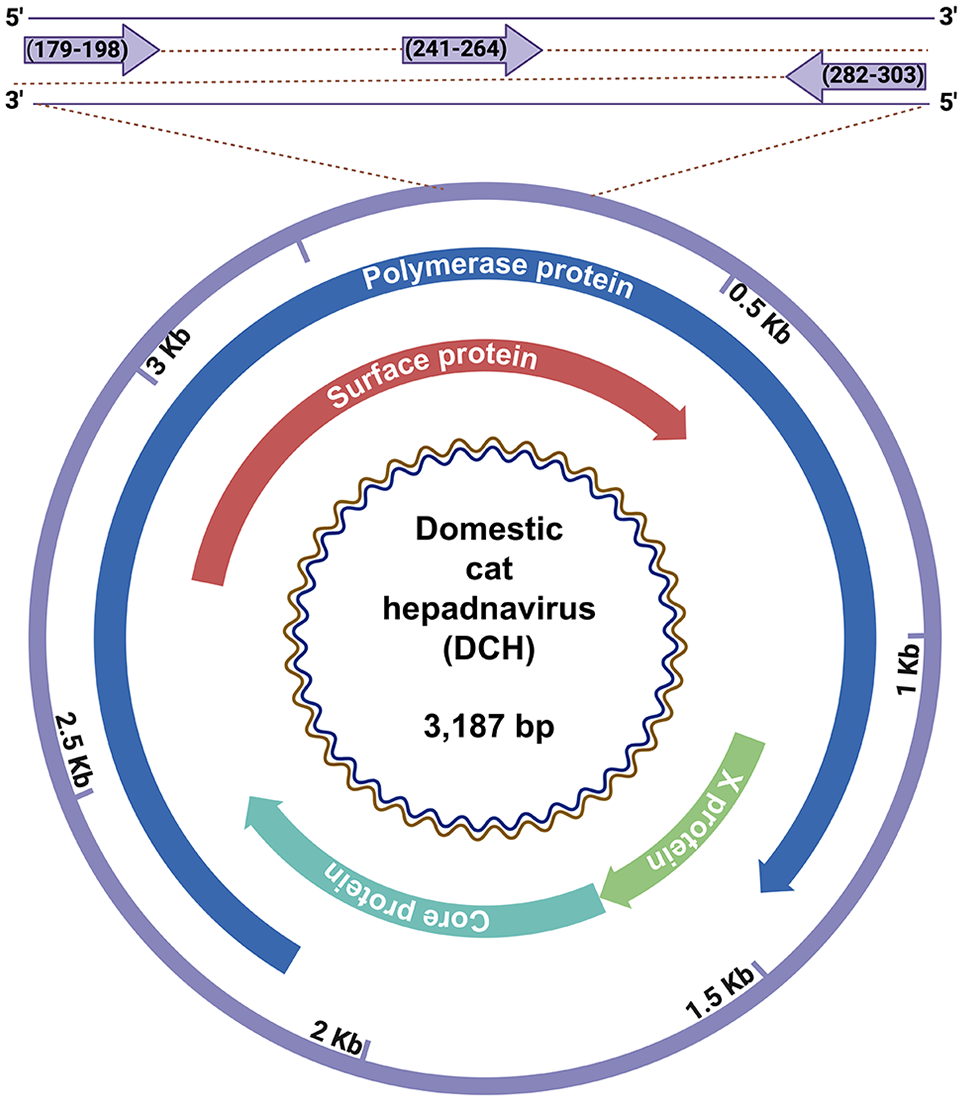
Schematic illustration of the positions of the primers used for detection of domestic cat hepadnavirus (DCH). A primers and probe set for DCH was designed based on the conserved regions of the surface/polymerase gene of DCH.
Real-time PCR assays
The rtPCR reactions conducted (Probe qPCR mix kits; TaKaRa Bio) are relatively tolerant to several inhibitors of PCR, according to the manufacturer. Each 20-μL reaction contained 2 pmol of each primer (forward and reverse), 1 pmol of the probe, 10 μL of Probe qPCR mix, and 0.2 μL of ROX reference dye II, with 1 μL of the blood sample as a template. The reaction was performed (QuantStudio 3 real-time system thermal cycler; Applied Biosystems) at 95°C for 20 s, followed by 40 cycles at 95°C for 3 s, and then 60°C for 30 s. For a positive control for 2 primer and probe sets (DCH and ACTB), we used a DCH-positive blood sample identified previously. 18 A DCH-negative blood sample was used as a negative control for DCH and positive control for ACTB. The Animal Experiment Committee of the University of Miyazaki (authorization 2021-019) approved the use of these blood samples. A reaction without template DNA (distilled water) was used as a negative control in all experiments.
Generation of a DCH DNA standard for rtPCR
The target region of the direct duplex rtPCR (736 bp) was amplified using conventional PCR assays, as described previously. 18 The amplicon was purified (QIAquick PCR purification kits; Qiagen) according to the manufacturer’s instructions. The concentration of DNA was determined (NanoDrop spectrophotometer; Thermo Fisher), and the copy number was calculated using the following equation:
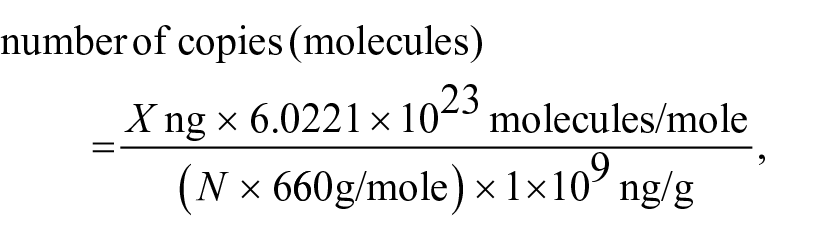
where X is the amount of amplicon (ng), N is the length of the double-stranded DNA (dsDNA) amplicon (736 bp in our study), and 660 g/mole is the average mass per 1 bp dsDNA.
Generation of a standard curve
A 10-fold dilution series of 106–101 DNA copies/μL was prepared using Tris-EDTA buffer (Nippon Gene) as a diluent, and was used to evaluate the standard curve parameters, especially the slope, from which the amplification efficiency and the coefficient of determination (R2) were derived. Each dilution was prepared in triplicate and amplified as described above.
Determination of the limit of detection and sensitivity of assay
The limit of detection (LOD) of DCH was determined using serial dilutions of DCH DNA fragments from 106, 105, 104, 103, 102, 101, 8, 4, 2, 1, and 0.5 copies/μL, with 8 replicates for each concentration and 3 independent PCR assays. The lowest concentration that met the positive detection rate of 95% was considered to be the LOD. 3
To ascertain the performance of the direct duplex rtPCR assay when crude whole blood was used directly as opposed to purified blood DNA, we performed head-to-head comparison assays using spiked blood samples. The 5 μL of DCH-positive samples were 100-fold diluted in 495 μL of DCH-negative blood. This spiked blood was split in half, and DNA was purified from 100 μL of sample (DNeasy blood & tissue kits; Qiagen). DNA was finally eluted in 100 μL of water. The other half of the crude blood samples was directly used for rtPCR, and 1 μL of each purified DNA and blood sample was used for the rtPCR assay in parallel.
Specificity of the direct duplex rtPCR assay
The specificity of the DCH primers and probe set was evaluated using plasmids containing 2 strains of feline leukemia virus (FeLV) and 2 strains of feline immunodeficiency virus (FIV) because previous studies suggested an association between infection with these viruses and DCH.1,2,10,16 The FIV plasmids used in our study were FIV-34T FIV strains F10 and FIV-PPR15,19; the FeLV plasmids were pEECC-FeLV and p61E-FeLV.6,13 These plasmids were obtained through the National Institute of Health (NIH) HIV Reagent Program, National Institute of Allergy and Infectious Diseases (NIAID), Division of AIDS, and used as templates for the direct duplex rtPCR reactions.
Repeatability of the direct duplex rtPCR assays
To evaluate the repeatability of the direct duplex rtPCR assays, we used 7 dilutions of cat blood containing DCH DNA fragments ranging from 106 to 101 DNA copies/μL. The SD and CV were calculated based on the cycle threshold (Ct) values obtained from the triplicate assay.
Statistical analysis
The standard curve, LOD 95%, and repeatability were analyzed (Prism v.9.1.1; GraphPad Software).
Results
Design and evaluation of primers and probes
The consensus primers and probe set designed here (Fig. 1) matched perfectly the sequences of 18 DCH strains (100%) available on GenBank (Fig. 2). Before testing a duplex PCR, we performed a singleplex PCR to check the amplification of DCH and ACTB with the primers and probe set. Singleplex rtPCR using a DCH-positive blood sample showed fluorescence signals for both DCH and ACTB; a DCH-negative blood sample only showed a fluorescence signal for ACTB. The direct duplex rtPCR for the detection of both DCH and ACTB in the same tube showed Ct values comparable to those observed in the singleplex rtPCR (Table 2). We visualized the PCR amplicons on an agarose gel after the duplex rtPCR and found no nonspecific bands (Fig. 3). These results suggested that our direct duplex rtPCR did not generate unwanted complexes of oligos, such as primer dimers.
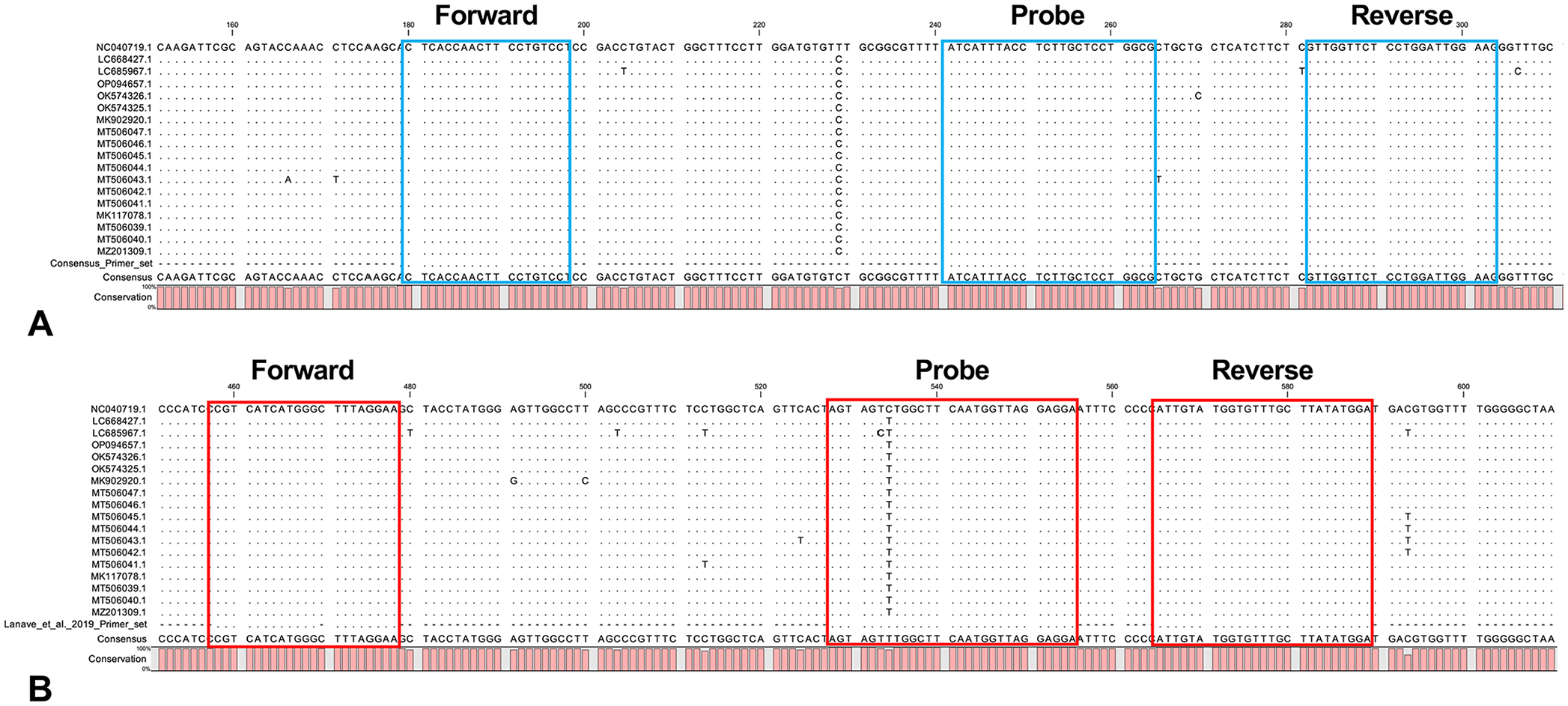
Alignment of the domestic cat hepadnavirus (DCH) primers and probe set with 18 DCH strains. The sequences of the consensus DCH primers and probe sets (
Comparison between singleplex and duplex rtPCR assays for detection of domestic cat hepadnavirus (DCH).
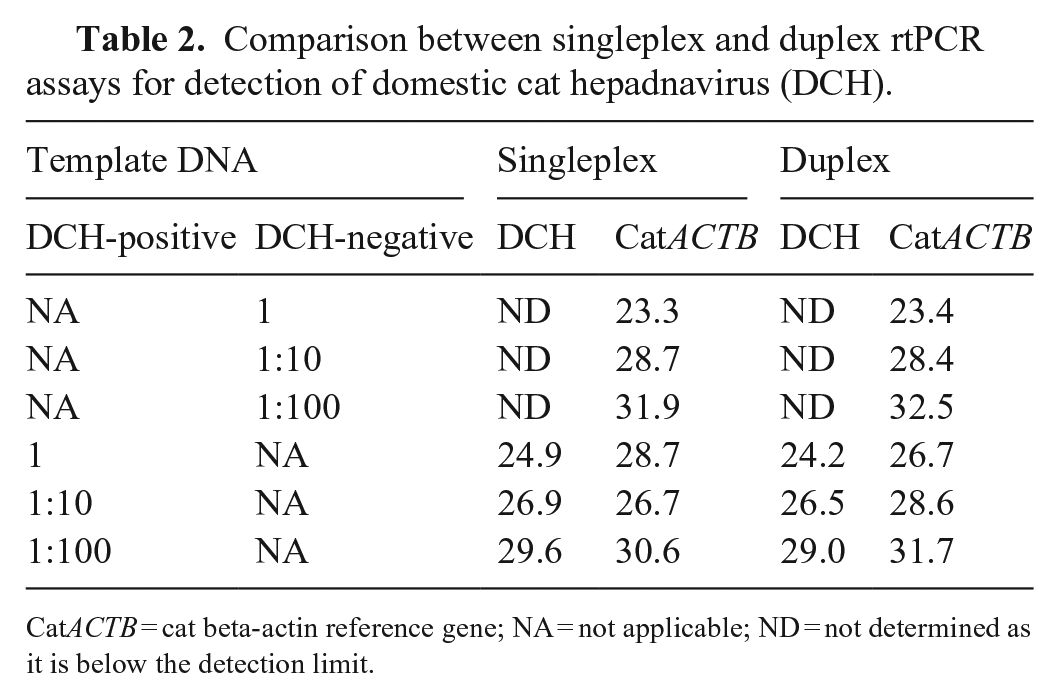
CatACTB = cat beta-actin reference gene; NA = not applicable; ND = not determined as it is below the detection limit.
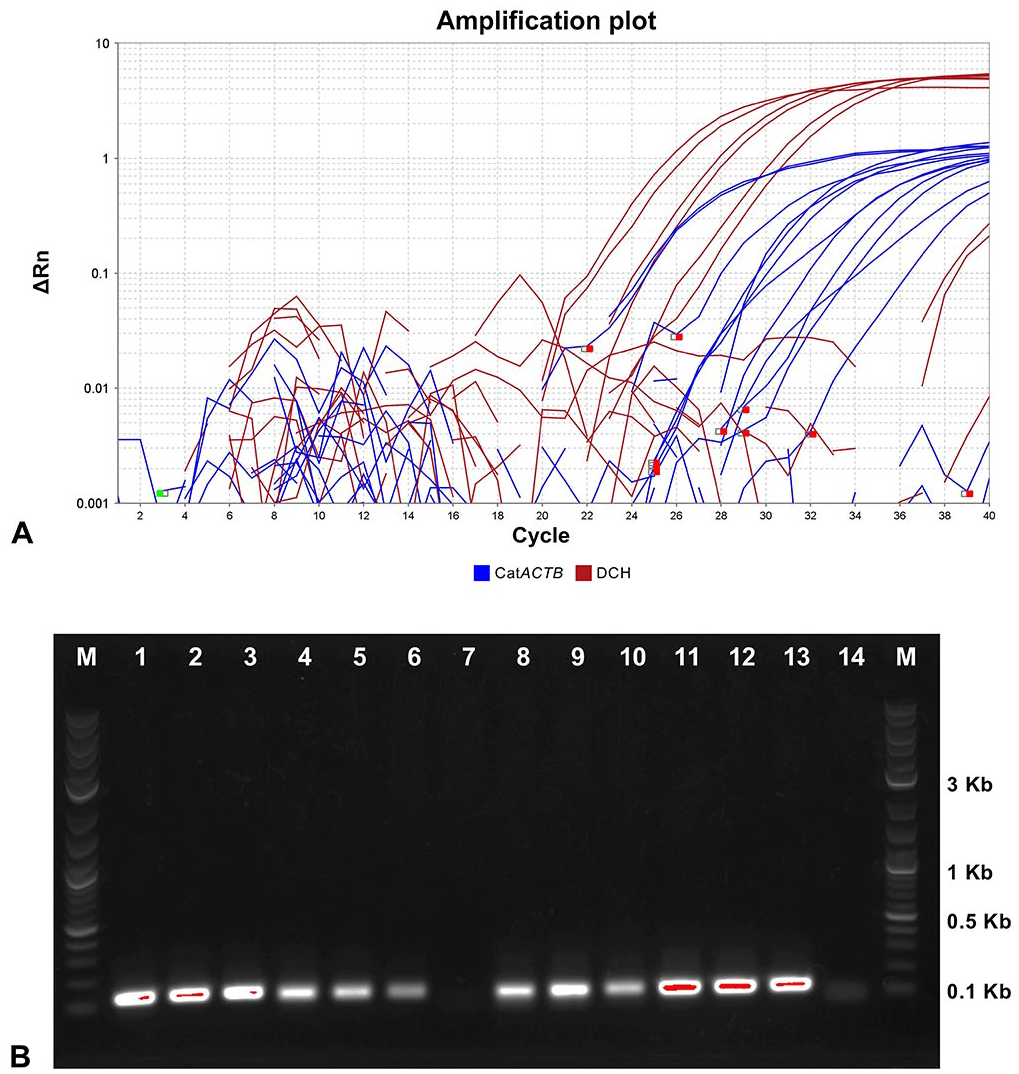
Amplification of the direct duplex real-time PCR (rtPCR) assay and PCR products on an agarose gel.
To compare the linearity and efficiency of the direct duplex rtPCR, we prepared a standard curve for DCH and performed duplex rtPCR. The results showed high linearity (R2 = 0.998) and high efficiency (96%) with a standard curve slope of −3.421, demonstrating a high amplification efficiency and linear equation required for DNA quantification (Suppl. Fig. 1).
Sensitivity, repeatability, and specificity of the direct duplex rtPCR assay
The detection rate of our direct duplex rtPCR of 8 copy/μL for 67% of replicates demonstrates that the LOD was at least 10 copies/μL (Table 3). The direct duplex rtPCR assay had higher Ct values than the rtPCR assay with DNA purification (Suppl. Table 1). Using plasmids encoding 2 FIV strains and 2 FeLV strains, we found that the direct duplex rtPCR assay did not amplify these plasmids encoding non-DCH sequences (Suppl. Table 2).
Sensitivity of the direct duplex rtPCR assay for detection of domestic cat hepadnavirus (DCH).
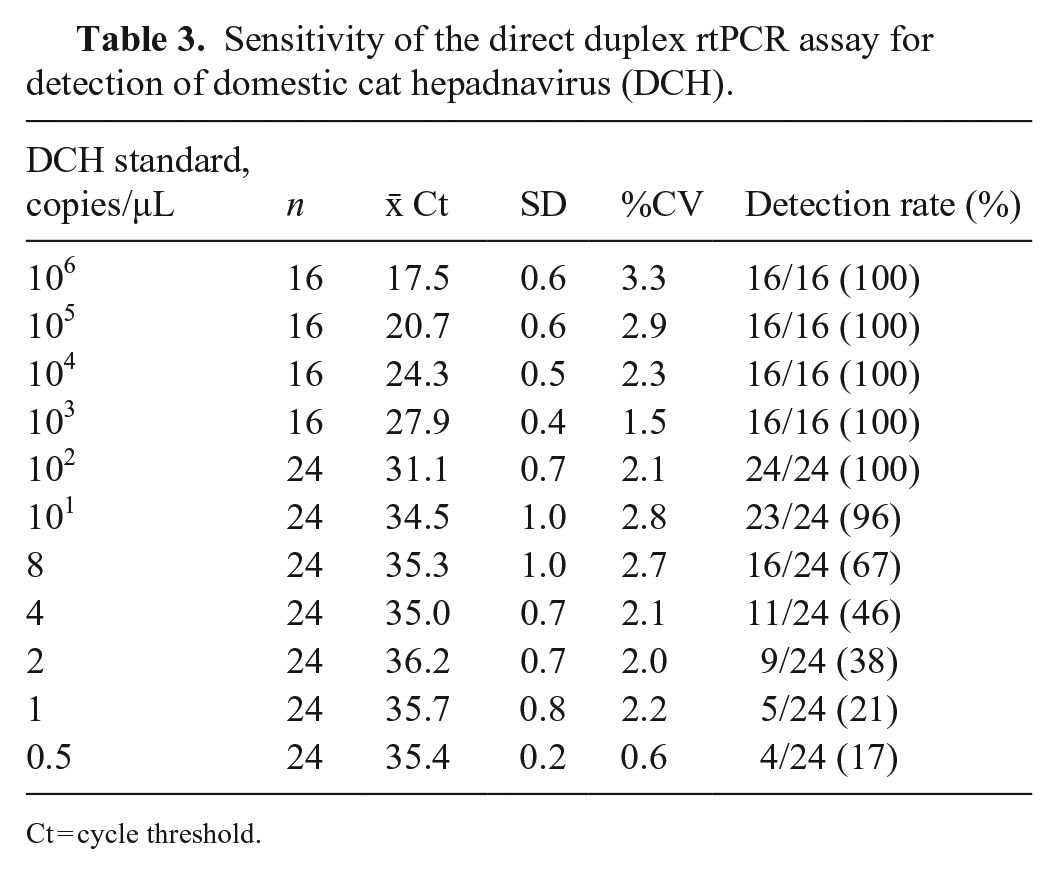
Ct = cycle threshold.
Using serially diluted DNA templates ranging from 106 to 101 copies/μL, we found that our direct duplex rtPCR assay had good repeatability, with a CV of <10% (Table 4). The variance analysis also showed that the p-value of inter-assay variation for each concentration gradient of the template was >0.05, indicating high repeatability of our direct duplex rtPCR assay.
Repeatability of the direct duplex rtPCR assay for detection of domestic cat hepadnavirus (DCH).
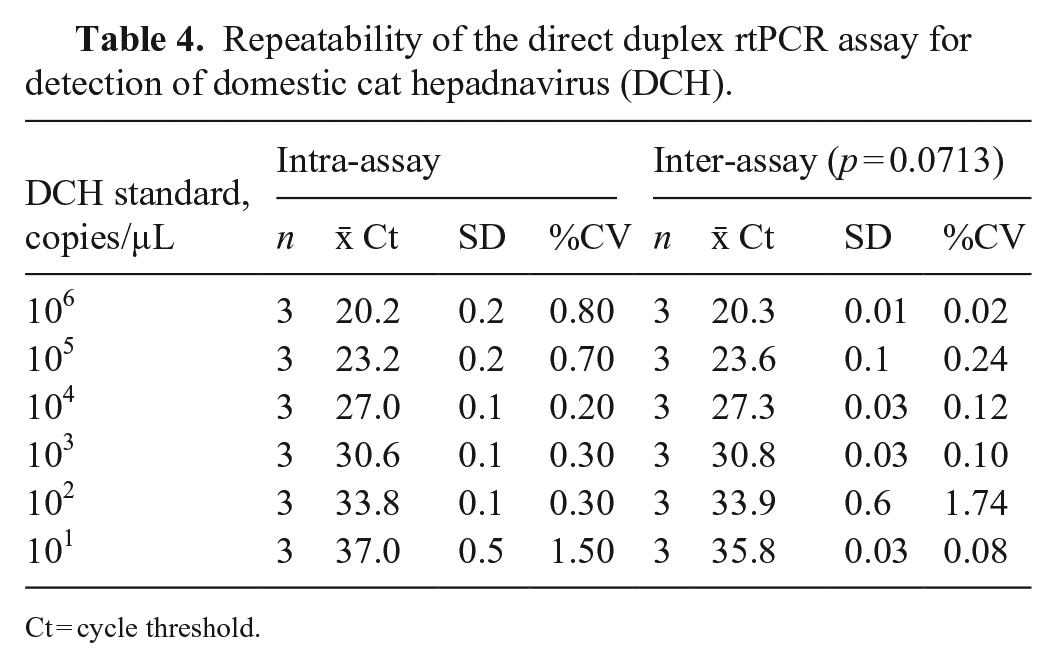
Ct = cycle threshold.
Discussion
Contamination by DNA from other PCR reactions can affect highly sensitive PCR assays, potentially leading to erroneous positive results. 8 Given that our direct duplex rtPCR assay does not require the opening of PCR tubes after the reaction, the risk of contamination is minimized. We had demonstrated previously that a DNA extraction–free method can be used for the detection of DCH in blood samples using a conventional PCR assay. 18 With the elimination of the DNA extraction and electrophoresis steps, our direct duplex rtPCR assay can be performed in a much shorter time than conventional PCR (Suppl. Fig. 2) while retaining high sensitivity, implying that our assay could be used for large-scale screening for DCH.11,20
None of the 4 plasmids encoding 2 strains of FIV or 2 strains of FeLV tested showed any positive signals in our direct duplex rtPCR assay, confirming that our assay was specific for DCH. The CVs of 0.01–1.5% obtained by the repeatability analysis indicated that our assay was highly stable and repeatable.
Our study had some limitations. We included just one DCH-positive sample for the assay. Nevertheless, in silico alignment analysis suggested that our assay would be able to detect 100% of DCH strains available in GenBank. Compared to the previous real-time primer set, 10 alignment-based analysis showed that the previous primers and probe set has a deletion on the forward primer, and one mismatch on the probe with almost all of the DCH strains. These mismatches may affect amplification efficiency and sensitivity; studies have demonstrated that T-T mismatches may result in a 10-to-100-fold underestimation of the initial copy number. 17 Testing more clinical samples is required to verify our assay for large-scale screening.
We believe that the performance of our direct duplex rtPCR assay is better than conventional methods. Considering its high-throughput capacity, our direct duplex rtPCR assay is a time-saving, and cost-effective method for the detection of DCH and cat ACTB. Our assay has considerable potential for routine surveillance, as well as understanding the kinetics of viral loads, which may be associated with disease severity and transmission.
Supplemental Material
sj-pdf-1-vdi-10.1177_10406387231154610 – Supplemental material for Development of a direct duplex real-time PCR assay for rapid detection of domestic cat hepadnavirus
Supplemental material, sj-pdf-1-vdi-10.1177_10406387231154610 for Development of a direct duplex real-time PCR assay for rapid detection of domestic cat hepadnavirus by Maya Shofa, Akiho Ohkawa, Tamaki Okabayashi, Yasuyuki Kaneko and Akatsuki Saito in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
The following reagents were obtained through the NIH HIV Reagent Program, Division of AIDS, NIAID, NIH: feline immunodeficiency virus (FIV) Petaluma infectious molecular clone (FIV-34TF10; ARP-1236) and FIV PPR infectious molecular clone (ARP-1237) contributed by Dr. John H. Elder, and feline leukemia virus (FeLV) EECC infectious molecular clone (pEECC-FeLV; ARP-105) and FeLV 61E infectious molecular clone (p61E-FeLV; ARP-109) contributed by Dr. James I. Mullins. We thank Tomoko Nishiuchi and Yuki Shibatani for their support. We thank Enago (www.enago.com) for the English language review. Fig. 1 and Suppl. Fig. 2 were created with BioRender (![]() ).
).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our work was supported in part by grants to Akatsuki Saito from the Japan Agency for Medical Research and Development (AMED) Research Program on HIV/AIDS (JP21fk0410033; JP22fk0410033; JP22fk0410047), AMED CRDF Global Grant (JP21jk0210039; JP22jk0210039), AMED Japan Program for Infectious Diseases Research and Infrastructure (JP21wm0325009; JP22wm0325009), Japan Society for the Promotion of Science (JSPS) KAKENHI Grant-in-Aid for Scientific Research (C) 19K06382, KAKENHI Grant-in-Aid for Scientific Research (B) 22H02500, Grant for Joint Research Projects of the Research Institute for Microbial Diseases, Osaka University; and to Akatsuki Saito and Tamaki Okabayashi from KAKENHI Grant-in-Aid for Scientific Research (B) 21H02361.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
