Abstract
HoBi-like pestivirus (HoBiPeV) is an emerging virus that has been detected in cattle and other ruminants. We diagnosed 2 cases of fatal bovine respiratory disease complex (BRDC) associated with infection with HoBiPeV in a feedlot in Argentina. The main findings in 2 steers autopsied were interstitial bronchopneumonia (case 1) and fibrinous bronchopneumonia (case 2). HoBiPeV was detected by RT-PCR in lungs of both animals and by immunohistochemistry in case 2. Phylogenetic analysis showed that both strains clustered within the “Brazilian-Italian” clade. In case 2, Mannheimia haemolytica was isolated from the lung. There is scant information about the contribution of HoBiPeV to the pathogenesis of BRDC. To our knowledge, HoBiPeV has not been reported previously in association with M. haemolytica pneumonia. Our findings further support the involvement of HoBiPeV in cases of BRDC and contribute to understanding the synergy of this etiologic agent in the pathogenesis of BRD, which is critical for the development of appropriate preventive strategies.
Pestiviruses (Flaviviridae) have been assigned to 11 different species (Pestivirus A–K) based on phylogenetic analysis. 16 Pestivirus A and B (formerly bovine viral diarrhea virus, BVDV1 and BVDV2) are well characterized as important pathogens in cattle, and are distributed in cattle populations worldwide. 26 An emerging pestivirus denoted as Pestivirus H (also known as HoBi-like pestivirus, HoBiPeV) has been described.1,28 HoBiPeV was first isolated from a contaminated batch of fetal bovine serum from Brazil in 2004 1 ; HoBiPeV isolates have since been reported in South America,22,27 Europe,7,9 and Asia.3,28
Infections by pestiviruses in ruminants cause important economic losses to the livestock industry worldwide, 26 and understanding the genetic diversity of pestiviruses within a country is essential for the implementation of regional control programs. 21 HoBiPeV infections in cattle may cause a variety of clinical presentations including diarrhea, mucosal disease–like lesions, respiratory signs, and reproductive failure.1,3,6,7,9,10,30 In experimental infections, HoBiPeV induced leukopenia,8,17 and may cause signs of respiratory disease7,9; therefore, involvement of HoBiPeV in the bovine respiratory disease complex (BRDC) has been proposed. 14 Here we describe 2 fatal cases of BRDC associated with HoBiPeV-infected cattle in a feedlot in Argentina.
In 2017, the owner of a cattle feedlot reported outbreaks of BRDC in 2 pens (1 and 2). The outbreaks occurred on different dates in May, and the pens were 1,025 m apart. The feedlot was located in the province of Cordoba, Argentina, and capacity at the time of the outbreak was 21,895 steers of different ages (5–14-mo-old). No information was available regarding the BVDV persistent infection status of animals entering the system. The feedlot management data indicated low-to-moderate annual mortality (1.14%). The population was composed of weaned beef and dairy calves from northern and central provinces of Argentina, which entered the feedlot when 5–7-mo-old. Feedlot management included prophylactic measures, such as vaccination and metaphylaxis with tilmicosin, to control the BRDC. The calves were vaccinated within 48–72 h after arrival with a commercial multivalent inactivated vaccine (Providean Respiratoria; Tecnovax) containing BVDV1, bovine herpesvirus 1 (BoHV1; Bovine alphaherpesvirus 1), Bovine respirovirus 3 (bovine parainfluenza virus 3, BPIV3), Histophilus somni, Mannheimia haemolytica, and Pasteurella multocida. Animals were revaccinated at 21 d after arrival. In pen 1, BRDC morbidity was 6 of 107 (5.6%) and mortality was 4 of 107 (3.7%) steers. The affected animals of this pen were 7-mo-old mixed steers that had been in the feedlot for 29 d when the first case occurred. In pen 2, morbidity was 4 of 69 (5.8%) and mortality was 3 of 69 (4.3%) steers. The affected animals in this pen were 5-mo-old Aberdeen Angus steers that had been in the feedlot for 38 d when the first case occurred. Affected steers had a cough, excessive nasal discharge, dyspnea, and fever (≥40°C).
Veterinary practitioners performed autopsies (cases 1 and 2, from pens 1 and 2, respectively) immediately after arrival at the premises. In case 1, marked cranioventral consolidation involved 40–50% of the pulmonary parenchyma; affected sections of lung were firm and dark-red. In case 2, severe cranioventral consolidation involved 60% of the pulmonary parenchyma. The pleura was multifocally covered with fibrinous exudate, and there were locally extensive areas of purple-red consolidation and hemorrhage on cut surfaces. An initial gross diagnosis of fibrinous bronchopneumonia was established for both cases. No other significant gross abnormalities were observed in either carcass; in particular, the esophagus, oral cavity, and intestines appeared grossly normal. Lung samples were obtained at autopsy for ancillary bacteriologic, virologic, and histologic testing.
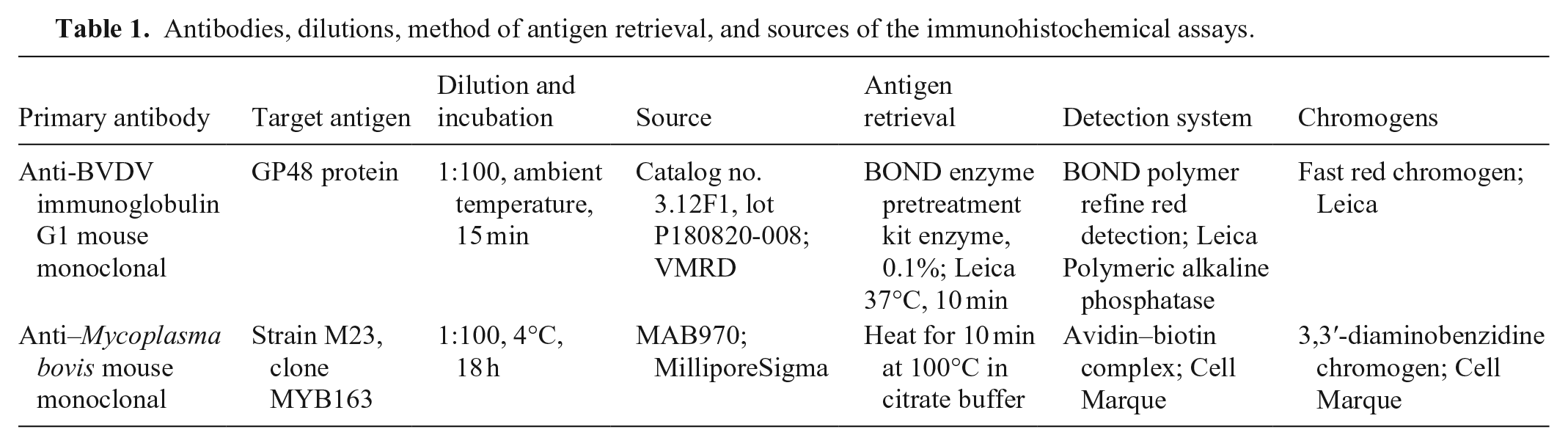
Lung samples were fixed in 10% neutral-buffered formalin for 72 h, processed routinely, and stained with H&E. Immunohistochemistry (IHC) was performed for BVDV (BVDV1 and BVDV2) and Mycoplasma bovis antigens. Tissue sections were mounted on slides, deparaffinized, rehydrated, and then subjected to antigen retrieval and incubated with primary antibodies and detection systems (Table 1). The sections were then counterstained with hematoxylin. Tissues of cattle naturally infected with BVDV (ear notch) or M. bovis (lung) were used as positive controls.
Antibodies, dilutions, method of antigen retrieval, and sources of the immunohistochemical assays.

Portions of the lungs were homogenized in mortars using sterile sand and Earl minimum essential medium supplemented with antibiotics. The tissue suspensions were centrifuged, and the supernatant was stored at −70°C. Viral nucleic acids were extracted from aliquots of samples (High Pure viral nucleic acid kit; Roche). PCR assays were used to detect bovine respiratory syncytial virus (BRSV; Bovine orthopneumovirus), 29 BPIV3, 18 BoHV1, and BoHV5 (Bovine alphaherpesvirus 5). 4 Reverse-transcription PCR (RT-PCR) assays for pestiviruses were performed with 189-389 and N2-R5 primers reported previously, which amplify the partial 5′-untranslated region of the genomes of pestiviruses BVDV1, BVDV2, and HoBiPeV (189-389 primers) 21 or HoBiPeV (N2-R5 primers). 2 Reverse transcription was performed with M-MLV (Moloney murine leukemia virus) enzyme (Promega) and random hexamers (Biodynamics). The PCR was performed with GoTaq enzyme (Promega). The products (200 bp for 189-389 primers, and 150 bp for N2-R5 primers) were purified from agarose gels (Illustra GFX PCR DNA, Cytiva; Gel band purification kit, GE). Amplicons were not cloned but sequenced directly in both directions, and all samples were tested in duplicate. Sequencing was performed by the Genomic Unit of INTA Castelar, Buenos Aires, Argentina. The phylogenetic analysis for pestiviruses was performed as described previously. 23 Briefly, multiple sequence alignment was performed with ClustalW3 v.2.1 (https://www.ebi.ac.uk). The dendrograms were obtained under Kimura distances method and neighbor-joining using MEGA 4 (https://www.megasoftware.net/mega4/mega.html); the bootstrap value was calculated using 1,000 replicates of the alignment. Trees were drawn with Dendroscope 2.7.4. 15 Nucleotide sequences of the strains obtained were submitted to GenBank (MZ189735, MZ189734).
The bacteriologic analysis was performed by inoculating lung tissue on blood agar and MacConkey agar plates. One set of the culture plates was incubated aerobically, and the duplicate under microaerophilic conditions (5% CO2). Plates were incubated at 37°C for 48 h. Identification of isolates cultured was based on Gram stain, colony morphology, biochemical analyses, and using MALDI-TOF mass spectrometry (MS; Microflex LT, Bruker). For MALDI-TOF MS, protein extraction was performed according to the manufacturer’s instructions.
The histologic lesions in the lungs differed in the 2 cases. In case 1, there was moderate-to-severe bronchointerstitial pneumonia characterized by necrotic bronchiolar epithelium and hyperplasia of type II pneumocytes. There were occasional prominent alveolar macrophages in the lumen, and type II pneumocyte hyperplasia, congestion, and expansion of the alveolar walls by a mild infiltrate of mononuclear inflammatory cells. In case 2, there were prominent areas of fibrinosuppurative bronchopneumonia (Fig. 1A), characterized by multifocal areas of necrotizing alveolitis with accumulation of degenerate neutrophils with streaming nuclei (oat cells; Fig. 1B), fewer macrophages, and fibrin within alveolar spaces and bronchiolar lumina. Hemorrhages, thromboses, and edema of the interlobular septa and fibrinous pleuritis were evident in both cases. BVDV IHC was positive in the lung of case 2 in which finely stippled intracellular red staining was present in individual (Fig. 1C) and random clusters of macrophages (Fig. 1D) and type II pneumocytes. In case 1, no immunostaining was observed for BVDV. IHC for M. bovis was negative in both cases.
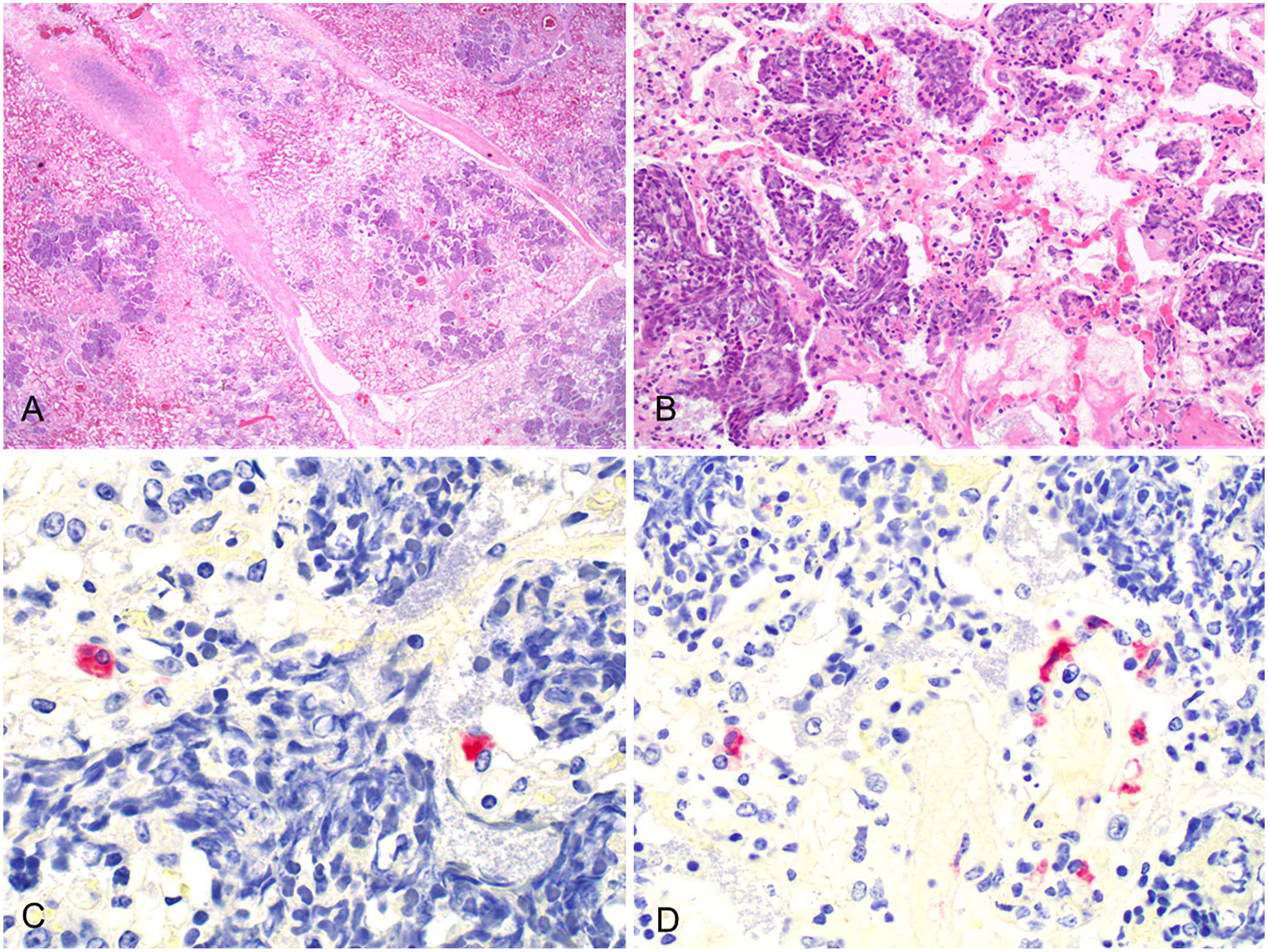
Histologic lesions in a case of bovine respiratory disease with coinfection of HoBi-like pestivirus and Mannheimia haemolytica, case 2.
BRSV, BPIV3, BoHV1, or BoHV5 was not detected in lung samples of either animal. However, PCR assays for pestivirus detection amplified HoBiPeV genetic material in both cases. Phylogenetic analysis showed that both strains clustered in the “Brazilian-Italian” clade (Suppl. Fig. 1). The homology between the strains was 98.5% (Fig. 2). No significant bacterial growth was evident in aerobic and microaerophilic cultures in case 1. Bacterial cultures from case 2 yielded pure growth of M. haemolytica–like bacteria. Based on biochemical tests and MALDI-TOF MS findings, isolates were identified as M. haemolytica.
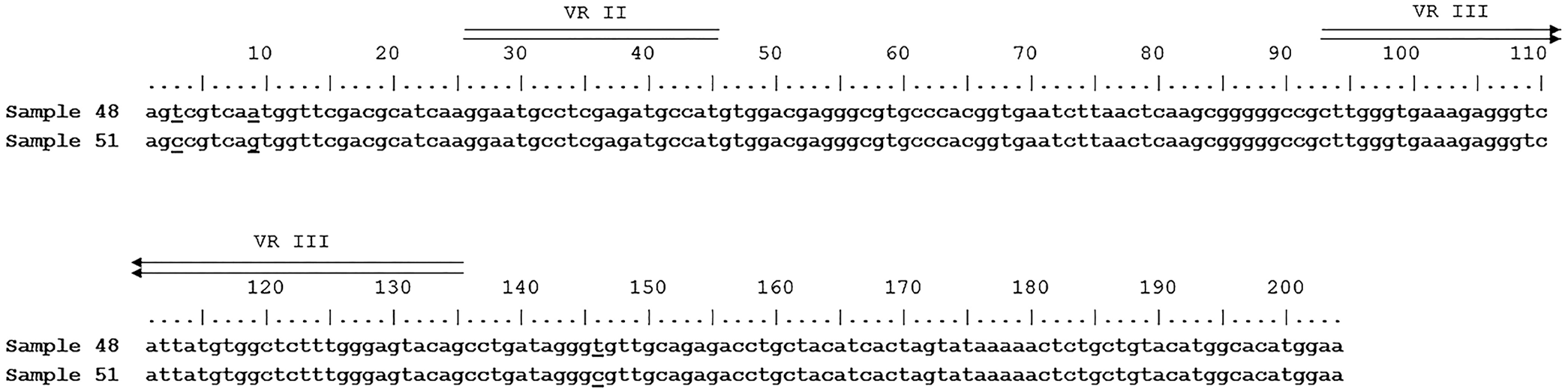
Alignment of nucleotide sequences from the 5′-UTR of HoBi-like strains 48 (MZ189735) and 51 (MZ189734). The nucleotide differences between the samples (positions 3, 9, 146) are underlined. Two variable regions are double overlined (VR II and VR III).
The results of the PCR assay and nucleotide sequencing confirmed the infection by HoBiPeV of the 2 feedlot steers affected with acute BRDC. BRDC is one of the main causes of morbidity and mortality in feedlots, and BVDV1 and BVDV2 are frequently involved in outbreaks of BRDC,11,12 with differences in virulence and pneumopathogenicity described among BVDV genotypes and strains. 26 HoBiPeV and other viruses (BoHV1, BRSV, and BPIV3) have been detected in nasal swabs in an outbreak of BRDC, suggesting that HoBiPeV infection is associated with cases of BRDC, and HoBiPeV should be considered as part of BRDC. 14 Several reports have identified HoBiPeV associated with signs of respiratory disease7,14; however, most of these reports described the detection of HoBiPeV alone, and not in association with another BRDC agent.3,9,20,30 In these reports, there was no evidence of coinfection with other bacterial BRDC agents such as M. haemolytica, P. multocida, M. bovis, or H. somni. In one study, coinfection of HoBiPeV and Streptococcus bovis was identified. 7 To our knowledge, coinfection by HoBiPeV and M. haemolytica, one of the main agents of BRDC,12,13,31 has not been reported previously.
The clinical signs observed in our cases were similar to other cases of respiratory disease.3,7,14 The fact that 2 animals were tested and that both were positive represents strong evidence of the involvement of HoBiPeV in the development of these BRDC cases. In various reports of HoBi-like pestiviral infection,3,7,9 lesions frequently observed included tracheitis, catarrhal bronchopneumonia, or interstitial pneumonia; the lesions in the 2 steers autopsied and reported here were interstitial bronchopneumonia (case 1) and fibrinous bronchopneumonia (case 2). The lesions found in both cases described here are consistent with severe pulmonary infections by M. haemolytica. 12
In IHC slides in case 2, we observed that the monoclonal antibody developed to detect BVDV1 and BVDV2 5 had cross-reactivity with HoBiPeV. This cross-reactivity was expected, given that the GP48 protein is highly conserved in various BVDVs. In our case, immunolabelling of antigenic HoBiPeV was observed more frequently in areas of the lung with moderate-to-severe inflammatory lesions compared to areas with milder lesions. Hence, we believe that HoBiPeV contributed to the development of the lesions. In previous studies,5,19 BVDV antigen was detected by IHC in spleen, mesenteric lymph nodes, thymus, digestive tract, liver, lung, kidney, and CNS of persistently, as well as of acutely, infected animals. We found no reports of the distribution of HoBiPeV detected by IHC in tissues of infected calves. In case 2, we observed that HoBiPeV infected pneumocytes, small vessel endothelial cells, and intra-alveolar macrophages, which is consistent with studies that evaluated animals infected with BVDV1 19 and BVDV2. 5
The detection of HoBiPeV in alveolar macrophages and pneumocytes in case 2 suggests that this virus could have favored bacterial colonization by M. haemolytica, and may also suggest that this virus play an important role, likely altering the innate defense mechanisms of the lung, as other authors have reported in experimental BVDV infections. 26 Various reports have shown synergism between bovine pestiviruses and other BRDC pathogens; infections by BVDV1b are commonly associated with severe secondary infections by M. haemolytica or M. bovis. 12 The absence of specific IHC staining in the lung section in case 1 could indicate lack of sensitivity of the IHC technique to detect this strain, or low viral loads of HoBiPeV in tissue. Unfortunately, we were unable to obtain lymphoid tissue or ear notch samples to determine if the animals were persistently infected with BVDV.
HoBiPeV has been detected in fetal bovine serum in Argentina, although no clinical cases have been described in Argentina before our cases, to our knowledge. 2 The HoBiPeV strains of our outbreak were genetically similar to Brazilian strains.10,14,22 The 2 sequences that we identified were very similar, which would suggest that the same strain was responsible for the outbreaks in the 2 pens. The sequences analyzed in our work were very short; it would be interesting in the future to sequence a more representative region of the genome in order to make a more complete analysis. These outbreaks, involving animals from different origins within a commercial feedlot, in addition to previous detection of the virus in fetal bovine serum from different regions of the country 24 and seropositivity in water buffaloes, 25 suggests that HoBiPeV is distributed widely in Argentina.
In Argentina, several BRDC vaccines include BVDV1, and those available are frequently formulated with inactivated viruses, without further specification of the included viral strains. Vaccination is an efficient method to reduce viral disease; however, the genetic heterogeneity observed in bovine pestiviruses may challenge and compromise the effectiveness of control and eradication programs. 1 Studies involving pestiviral cross-antigenicity have shown limited cross-immunity between different strains, indicating the need to include new genotypes and strains in vaccine formulations.10,20,23 The current practice of sourcing steers from diverse origins for finishing in the central region of Argentina, and the use of current BVDV vaccines, may provide a favorable scenario for the spread of HoBiPeV.
Supplemental Material
sj-pdf-1-jvd-10.1177_10406387221098356 – Supplemental material for HoBi-like pestivirus in 2 cases of fatal respiratory disease of feedlot cattle in Argentina
Supplemental material, sj-pdf-1-jvd-10.1177_10406387221098356 for HoBi-like pestivirus in 2 cases of fatal respiratory disease of feedlot cattle in Argentina by Carlos A. Margineda, Franco Matías Ferreyra, Franco Masnyj, Maximiliano Audrito, Paula Melisa Favaro, Dus Santos María José and Andrea Pecora in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank Dr. Shollie M. Falkenberg for reading this manuscript critically.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our work was funded by project INTA PNSA-1115054, INTA AUDEAS CONADEV-940148, ArgenINTA Foundation, and INTA RIST I111 and I103.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
