Abstract
Plasma cell-free DNA (cfDNA) is a biomarker of ischemia, systemic inflammation, and mortality in humans with gastrointestinal disease. Cell-free DNA has not been investigated as a biomarker for equine colic, to our knowledge. We hypothesized that cfDNA could be measured accurately in neat equine plasma using a benchtop fluorometer and that plasma cfDNA would be elevated in emergency patients compared to healthy horses. Plasma was obtained from blood collected in Roche DNA stabilizing tubes. We used the Qubit 4 fluorometer and 1× dsDNA HS assay kit to measure cfDNA concentration in neat patient plasma and following DNA extraction of plasma with a commercial kit. Assay precision and linearity of dilution were satisfactory for neat plasma cfDNA, but DNA spike and recovery results were variable. Further, cfDNA concentrations in paired neat plasma and extracted-plasma samples (n = 66) were not correlated. Median extracted-plasma cfDNA was higher in emergency patients (n = 50) and a subgroup of colic patients (n = 36), compared to healthy horses (n = 19). Our results with extracted-plasma samples provide proof of concept for further investigation of plasma cfDNA as a biomarker in horses.
Plasma cell-free DNA (cfDNA) is generated from cellular apoptosis, necrosis, and extrusion of neutrophil extracellular traps (NETosis). In humans, plasma cfDNA is a diagnostic and prognostic biomarker for emergency abdominal pain and acute mesenteric ischemia.1,17 In dogs, plasma cfDNA is elevated with gastric dilation–volvulus. 21 Studies on cfDNA in horses are limited. One study documented elevated serum cfDNA in equine recurrent uveitis patients compared to horses without ophthalmic abnormalities; another study found no significant difference in plasma cfDNA between healthy and septic or sick non-septic foals.4,5 Investigation of cfDNA in horses with gastrointestinal disease has not been reported, to our knowledge. Because cfDNA has been identified as a biomarker for gastrointestinal disease in other species, we believe further investigation of cfDNA as a potential biomarker in equine colic is warranted.
Cell-free DNA has many of the characteristics of an ideal biomarker, given that samples can be obtained with minimally invasive methods and can be measured rapidly. Although traditional methods of cfDNA quantification (e.g., real-time PCR [rtPCR], spectrophotometry, fluorescence-based microplate assays) are not ideal for individual patient samples, recent studies have reported use of the Qubit fluorometer (Invitrogen; Thermo Fisher) for simple and rapid quantification of cfDNA.10,13 Strong correlation has been reported between DNA concentrations measured using Qubit and rtPCR.9,16 Various groups have utilized the Qubit instrument to measure human and canine cfDNA, either in neat plasma and/or in samples of DNA extracted from plasma.3,6,7,9,10,13,14,16,23 The small size of the Qubit 4 fluorometer (13.6 × 25 × 5.5 cm, 743 g), combined with reports of its use for the quantification of cfDNA in neat plasma, suggest that it may serve as a practical point-of-care (POC) instrument.
We hypothesized that the Qubit 4 fluorometer would accurately and precisely quantify cfDNA concentration in neat equine plasma and that equine emergency patients in general, and colic patients specifically, would have higher neat plasma and extracted-plasma cfDNA concentrations than healthy horses. Our study protocol was approved by the North Carolina State University (Raleigh, NC, USA) Institutional Animal Care and Use Committee (protocol 18-168-O); consent was obtained from all clients prior to horse enrollment.
We obtained peripheral blood samples from 50 adult horses (≥2-y-old) that had been presented on an emergency basis to an academic veterinary hospital, including a subgroup of 36 colic patients, as well as 19 healthy adult horses and ponies (Suppl. Fig. 1; Suppl. Table 1). To minimize cell lysis and ex vivo release of cfDNA, blood was collected into specialized K3-EDTA cell-free DNA vacutainer tubes (Roche) containing a stabilizing solution. Tubes were refrigerated (4°C) until separation of plasma by centrifugation at 2,300 × g for 10 min. Plasma was harvested without disturbing the buffy coat, and 1-mL aliquots were frozen at −80°C until analysis or extraction. Protocols for sample processing and storage conditions were based on published research in humans and dogs, as well as our own preliminary data.2,8,19,20
DNA was extracted from plasma using a commercial kit (DNeasy blood and tissue kit; Qiagen) following a modified 18 manufacturer’s protocol. Concentrations of cfDNA in thawed plasma and extracted-plasma samples were measured (Qubit 4 benchtop fluorometer, Qubit 1× dsDNA HS assay kit, Invitrogen; Thermo Fisher), in accordance with the manufacturer’s directions and as reported previously for canine and human plasma3,6,10,21,23 and human extracted-plasma DNA samples.9,13,14,16 With a sample volume of 10 μL, the Qubit manufacturer reports the lower limit of quantification as 10 ng/mL, which is equivalent to a lower limit of 0.8 ng/mL for concentrations calculated for extracted-plasma samples (see Suppl. material).
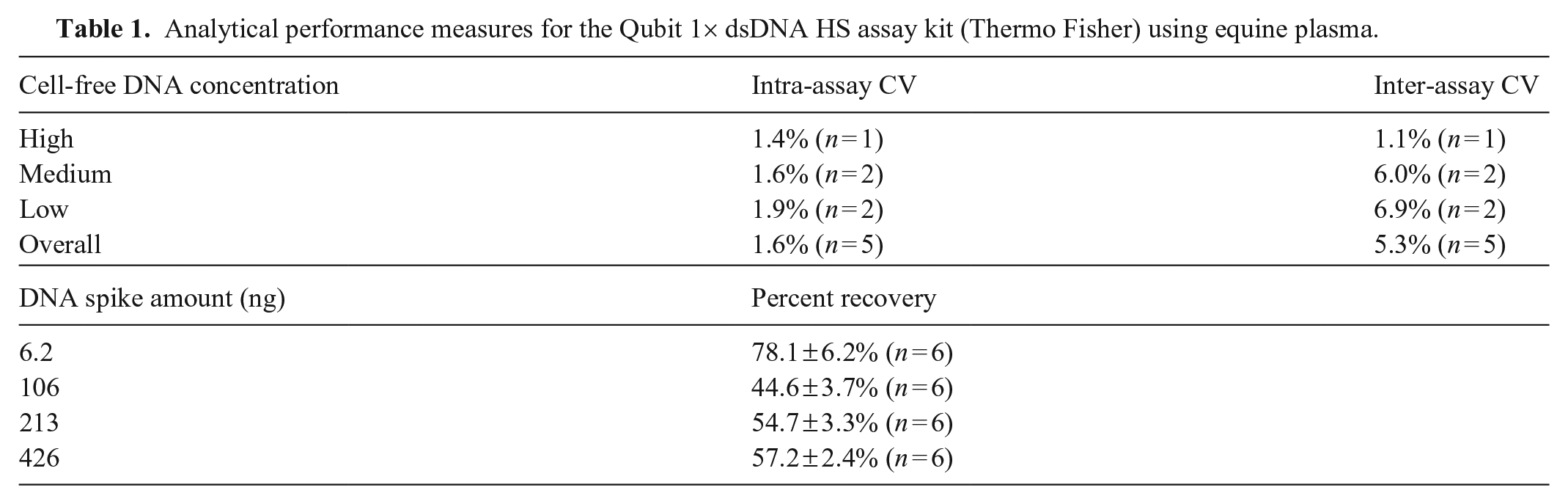
Randomly selected equine emergency patient samples with high, medium, and low neat plasma cfDNA values were used to determine the precision and accuracy of neat plasma cfDNA readings with the Qubit 4 instrument and Qubit 1× dsDNA HS assay kit. Briefly, intra-assay CV was determined by measuring cfDNA concentrations 5 times on the same day in 6 plasma samples. Inter-assay CV was determined by measuring cfDNA concentrations in 5 plasma samples on 4 consecutive days. Linearity was assessed by measuring cfDNA concentrations in dilutions of plasma from 3 horses prepared with DNase-free water (plasma:water ratios of 10:90, 20:80, 30:70, 40:60, 50:50, 60:40, 70:30, 80:20, 90:10, 100:0). Percent recovery was calculated based on aliquots of plasma from 6 horses, each spiked with 4 amounts of DNA standard (D4764; MilliporeSigma). Nonspecific dye fluorescence was measured in 8 plasma samples treated with recombinant DNase I (AM2238; Invitrogen). Autofluorescence was measured in 10 plasma samples using Qubit dilution buffer only.
Normality testing was performed with Shapiro–Wilk tests. Spearman correlation coefficients were calculated for cfDNA concentrations in paired neat plasma and extracted-plasma samples. Mann–Whitney tests were used to compare median extracted-plasma DNA concentrations between horse groups. This comparison was not performed for neat plasma cfDNA because of our demonstrated lack of accuracy in this sample type. Analyses were performed using statistical software (Prism v.9.0; GraphPad), and significance was set at p ≤ 0.05.
Mean intra- and inter-assay CV for measurement of neat plasma cfDNA concentration in emergency patients were 1.6% and 5.3%, respectively (Table 1). Cell-free DNA concentrations in dilutions of plasma samples demonstrated high linearity (R 2 ≥ 0.962; Suppl. Fig. 2). Average percent recoveries of spiked DNA were 44.6–78.1% (Table 1). Treatment of plasma samples with DNase I resulted in a 10–16% decrease in measured cfDNA concentrations (Suppl. Table 2). Qubit measurement of plasma with Qubit dilution buffer only (no reagent) yielded low values consistent with autofluorescence of plasma components (Suppl. Table 2).
Analytical performance measures for the Qubit 1× dsDNA HS assay kit (Thermo Fisher) using equine plasma.

There was no significant correlation between paired neat plasma and extracted-plasma cfDNA concentrations in all emergency patients (Spearman ρ: −0.0133, p = 0.9291, n = 47; Fig. 1A) or in the subset of colic patients (Spearman ρ: 0.07328, p = 0.6757, n = 35). Neat plasma and extracted-plasma cfDNA concentrations from healthy horses also showed no significant correlation (Spearman ρ: 0.4552, p = 0.0502, n = 19; Fig. 1B). The lack of correlation remained when data from the emergency patient and healthy horse groups were pooled (Spearman ρ: 0.2219, p = 0.0733, n = 66; data not shown). Median cfDNA concentrations in extracted-plasma samples were significantly higher in overall equine emergency patients (3.0 ng/mL, range: 0.8–43.7 ng/mL, n = 50, p < 0.0001), or the colic subgroup (3.4 ng/mL, range: 0.8–43.7 ng/mL, n = 36, p < 0.0001), compared with healthy horses (1.2 ng/mL, range: 0.8–2.3 ng/mL, n = 19). Of note, there was overlap between healthy and clinical populations (Fig. 1C).
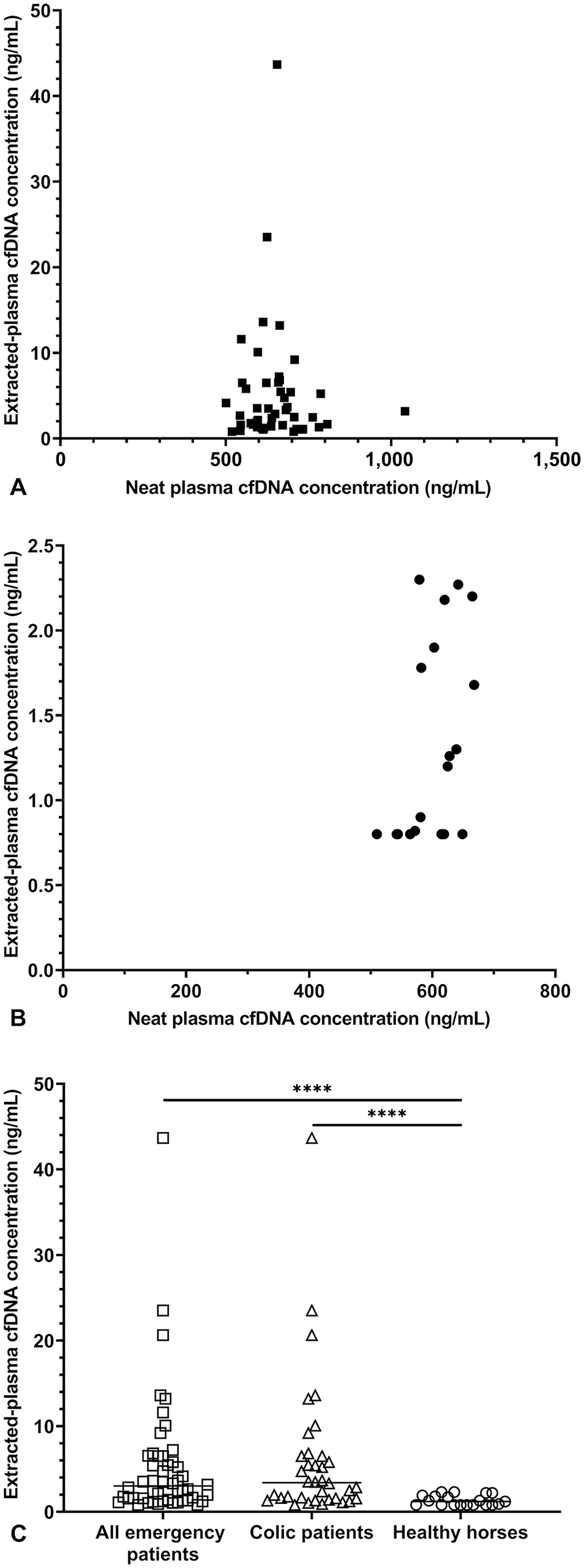
Comparison of cell-free DNA (cfDNA) concentrations in neat plasma and extracted-plasma samples.
In our initial investigation of plasma cfDNA in horses, we hypothesized that measurement of cfDNA in neat equine plasma with the Qubit 4 fluorometer would be precise and accurate. We investigated this hypothesis by determining inter- and intra-assay CV, linearity of dilution, percent recovery following DNA spike-in, and correlation with paired samples of extracted DNA. Our CVs and high degree of linearity of dilution were consistent with satisfactory precision and accuracy, and our intra-assay CVs were comparable to previous reports using canine plasma. 3 However, even though we took care to ensure that our expected DNA concentrations did not exceed the manufacturer-reported range for the assay, percent recovery for spiked DNA revealed assay inaccuracy. Further evidence for inaccuracy came from the lack of correlation between cfDNA concentrations in paired neat plasma and extracted-plasma samples.
We hypothesized that a matrix effect in equine plasma was contributing to our unexpected findings, including 2 potential sources for interference: 1) autofluorescence from compounds in the plasma, and 2) interaction of the fluorescent label with non-DNA molecules in the plasma. To assess the degree of autofluorescence, we measured cfDNA concentration in the absence of the dsDNA reagent. Given that the reagent contains the fluorescent molecule that interacts with dsDNA, omitting this assay component allows estimation of the intrinsic fluorescence of the plasma sample. We detected autofluorescence in all of the equine plasma samples measured without reagent. Interestingly, samples with higher total bilirubin concentration (serum biochemistry analyses at admission) had some of the highest autofluorescence values (Suppl. Fig. 3). This finding is consistent with a previous report of bilirubin increasing autofluorescence in canine plasma. 20
To investigate fluorescent signal arising from non-DNA molecules, we measured the cfDNA concentrations before and after DNA degradation by exogenous DNase I. We confirmed activity of our DNase I by degradation of a stock solution of control genomic DNA. DNase treatment decreased measured cfDNA concentrations in neat equine plasma by 10–16%, which was markedly less than the 98% decrease in the control sample (Suppl. Table 1). This result indicates that a substantial portion of the fluorescent signal detected by the Qubit assay, when measuring cfDNA in neat equine plasma, is not dsDNA. In theory, the difference between cfDNA concentrations measured in DNase I–treated plasma and the autofluorescence values represents binding of the Qubit dsDNA dye to other components in equine plasma besides DNA. The Qubit instrument and assay kit manufacturer has demonstrated excellent specificity of the dsDNA reagent for dsDNA over ssDNA and RNA, and reports that assay performance is satisfactory in the presence of low concentrations of salts and proteins. However, we suspect that proteins and other macromolecules in equine plasma may promote nonspecific interactions between the dsDNA reagent and off-target molecules.
Although our data provide evidence that Qubit measurement of cfDNA in neat equine plasma is inaccurate as a result of a matrix effect, quantification of cfDNA in extracted-plasma samples using the Qubit instrument and dsDNA assay kit is well established in the human medical literature.9,14,15 Indeed, DNA extraction prior to plasma cfDNA measurement is commonplace in human studies, across multiple DNA quantification methods.1,14,16,19 Therefore, we feel that our results with extracted-plasma cfDNA are a significant contribution to our understanding of this biomarker in horses. Our results show that median extracted-plasma cfDNA was significantly higher in emergency cases, and a subset of emergencies with colic, compared to healthy horses. Likely sources of elevated cfDNA in these patients include cell death (apoptosis, necrosis) in patients with tissue damage, and/or NETosis in patients with severe systemic inflammation. We are currently investigating whether cfDNA is a clinically useful biomarker for lesion diagnosis (e.g., intestinal strangulation) in horses with colic, and/or for prognosis in horses with systemic inflammatory response syndrome (SIRS).
We plan to address limitations of this study in our ongoing work. First, we used only one method of DNA quantification. We repeated measurements and analyses, including correlation between paired neat plasma and extracted-plasma cfDNA and indicators of analytical performance, using the Qubit dsDNA BR assay kit with similar findings as reported for the 1× dsDNA HS assay kit (data not shown). We are utilizing additional methods of measuring cfDNA, including spectrophotometry and SYTOX green (Invitrogen) assays, in our continuing investigation of cfDNA as a biomarker in horses.5,12,16 Second, we recognize that DNA extraction does not recover all of the cfDNA present in a plasma sample, and efficiency of extraction can vary. We did not investigate the efficiency of extraction in our study. Regarding the tissue specificity of cfDNA, we did not investigate the sequencing or patterns of methylation or fragmentation of the cfDNA in our samples, which are among methods used to characterize the tissue of origin for cfDNA.11,22 However, these methods are likely more relevant for use of cfDNA in prenatal screening or as a “liquid biopsy” for cancer; the application of this information in the diagnosis of gastrointestinal disease or sepsis is less clear.
A major advantage of the Qubit fluorometer over other DNA quantification techniques is its potential as a POC instrument. Studies in other species have reported measurement of cfDNA in neat plasma using the Qubit instrument, supporting its clinical feasibility. Unfortunately, based on our results, Qubit measurement of cfDNA in neat adult equine plasma is inaccurate. However, our results show that median extracted-plasma cfDNA concentrations are significantly higher in horses presented on emergency, and in horses with colic, compared to healthy horses. These data offer proof of concept for continued investigation of cfDNA as a biomarker in horses, including those with gastrointestinal disease. We are currently conducting studies utilizing DNA extraction, plasma dilution, and measurement of cfDNA in peritoneal fluid to circumvent the error-producing matrix effects of equine plasma and further our investigation of cfDNA as a diagnostic and prognostic biomarker in horses.
Supplemental Material
sj-pdf-1-vdi-10.1177_10406387221078047 – Supplemental material for Investigation of plasma cell-free DNA as a potential biomarker in horses
Supplemental material, sj-pdf-1-vdi-10.1177_10406387221078047 for Investigation of plasma cell-free DNA as a potential biomarker in horses by Rosemary L. Bayless, Bethanie L. Cooper and M. Katie Sheats in Journal of Veterinary Diagnostic Investigation
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Stipend support for Rosemary L. Bayless and partial funding for study supplies were provided by the NIH T32 Ruth Kirschstein Institutional National Research Service Award (NRSA) Training Grant (NIH T32 OD011130). Stipend support for Bethanie L. Cooper was provided by the North Carolina State University College of Veterinary Medicine Veterinary Scholars Program.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
