Abstract
The local laboratory with a local client-base, that never needs to exchange information with any outside entity, is a dying breed. As marketing channels, animal movement, and reporting requirements become increasingly national and international, the need to communicate about laboratory tests and results grows. Local and proprietary names of laboratory tests often fail to communicate enough detail to distinguish between similar tests. To avoid a lengthy description of each test, laboratories need the ability to assign codes that, although not sufficiently user-friendly for day-to-day use, contain enough information to translate between laboratories and even languages. The Logical Observation Identifiers Names and Codes (LOINC) standard provides such a universal coding system. Each test—each atomic observation—is evaluated on 6 attributes that establish its uniqueness at the level of clinical—or epidemiologic—significance. The analyte detected, analyte property, specimen, and result scale combine with the method of analysis and timing (for challenge and metabolic type tests) to define a unique LOINC code. Equipping laboratory results with such universal identifiers creates a world of opportunity for cross-institutional data exchange, aggregation, and analysis, and presents possibilities for data mining and artificial intelligence on a national and international scale. A few challenges, relatively unique to regulatory veterinary test protocols, require special handling.
Keywords
Background
Prior to the 1990s, most medical laboratories operated independently, serving their local clientele. Even corporate laboratory systems seldom needed to interact with any dissimilar systems. Then 2 forces began to drive a need for interoperability. In human medicine, many patients went from having “their doctor” and “their hospital” to receiving care at many medical facilities throughout their city or state. Coordination of care suffered because medical records did not translate from one facility to the next. In public health, there was a growing recognition that foodborne disease outbreaks were no longer focused on one locality but were connected by food distribution systems. Finding the clusters required aggregating laboratory data from across the country. Public health found a solution in PulseNet, a network of laboratories running identical protocols for fingerprinting bacterial isolates. 5 Integrating clinical care across facilities presented a larger challenge.
One of the early health information networks (HINs) trying to improve care coordination was developed at the Regenstrief Institute in Indianapolis, IN. 2 The first major effort was to exchange laboratory data. As was true everywhere, each laboratory had its own name and code for each test. Trying to map each test at each lab to each hospital’s record system was impractical. Instead, Regenstrief developed a system for assigning a common code and descriptive name to each test. One of the first things identified was that a code was needed for each observation type rather than each test type because some tests result in multiple individual results. This was the beginning of the Laboratory (now Logical) Observation Identifiers Names and Codes (LOINC) standard. 3 In 2002, the American Veterinary Medical Association (AVMA) officially adopted LOINC as the preferred system for encoding laboratory and clinical observations. 1 That same year, the National Animal Health Laboratory Network (NAHLN) was created.
It was becoming clear that the scale of modern production agriculture meant that any significant foreign or emerging disease event would easily overwhelm the U.S. national veterinary laboratories. The NAHLN was created as part of the Public Health Security and Bioterrorism Preparedness and Response Act in 2002. It took cues from both PulseNet and the HIN movement. It established common protocols for testing to ensure inter-laboratory comparability and reliability. It also established a standards-based secure communication protocol. This HL7 messaging process used LOINC codes as its identifiers for orders and observations.
Local test names
The local names and codes used by laboratories are very user-friendly, easy to remember, and short. They draw much of their specificity not from the name itself but from local context. For example, because there are few if any indirect fluorescent antibody (IFA) tests for antigen in veterinary medicine, “disease X IFA” serves as a perfectly good name for “disease X etiologic agent antibody by indirect fluorescent assay.” Local names often lack specificity about the specific antibody type, nucleic acid target, or chemical form measured. In local use, these are made clear in the complete laboratory report and known from then on when ordering tests. In a few cases, this ambiguity may lead to mistakes such as ordering a test for IgG when IgM would be more appropriate for detection of an acute infection. But in most cases, brevity and simplicity win out. Similarly, local codes used within laboratories tend to be short, at least somewhat mnemonic, and easy to type for quickly referencing the correct test. For local use, local names and local codes will never be replaced by fully specified standard names and codes.
When exchanging observation data between dissimilar systems, local codes break down. Geographic differences in disease agent distribution, cultural differences in naming conventions, and spoken language differences all conspire to make it difficult or impossible to reconcile multiple local code systems. Standard code systems, although not as concise or user-friendly, allow results from multiple laboratories to be combined meaningfully. The question is then, when and if the value of such interoperability is worth the effort of standardization.
Application of standardized laboratory data
As the phrase “observation identifier” implies, the most basic use of standard test codes is to determine when 2 tests are the same type of observation. LOINC draws this line at the level of clinical significance. If 2 tests are clinically interchangeable, they will have the same LOINC code regardless of what they are called locally. If they have different LOINC codes, then at least some adjustment will be needed before results can be substituted or combined. Even in a non-electronic world, this can be useful for projects such as merging test catalogs.
The simplest use of standard codes is in the exchange of data between dissimilar information systems. As in the case of a HIN such as Regenstrief, this allows test results from one institution to be viewed in the information system used at another and thus give a more complete picture of the patient’s overall condition. The sending system assigns a standard code to each observation type in its local code system and transmits that standard code to the receiving system. The receiving system maps many of the known standard codes to its internal display names. If it receives a code for which it lacks a mapping, then it can display the standard name associated with the standard code.
Standard codes for observations, as well as standard codes for panels of observations that make up tests, can also be used for the exchange of orders for testing. Such codes are typically published in the compendia of test services to facilitate the identification of codes for tests offered by a laboratory. LOINC codes are labeled to distinguish those that may be assigned to individual observations from those that represent groups of observations and are therefore only appropriate as orders. An order code for a complete blood count, for example, results in various counts of individual cell types, each with a specific observation code.
Things get more interesting when we move from one-to-one exchange of data to data aggregation on a larger scale. In cases such as the NAHLN or PulseNet, a prescribed small set of observations may arrive at a central system to be aggregated and analyzed. This case mirrors one-to-one exchange. For a more loosely constructed network, such as most HINs or the Swine Health Information Center (SHIC) data warehouse, different but related test types may be of interest. Standards such as LOINC include various types of hierarchy. It is possible to identify all tests for a certain analyte or agent and then summarize by specimen type or method of analysis. Because measurement scales are precisely expressed in the code, unit conversion can be performed precisely. Laboratories benefit from these standardized data repositories through a reduction in the number of ad-hoc requests for data by large clients, government regulators, and so on.
A step beyond data aggregation is data mining and artificial intelligence. On one hand, natural language processing and other artificial intelligence advances could make explicit coding of data elements unnecessary. With enough data, these technologies could infer that one test name at one laboratory is the same as another name at another laboratory and that this test detects a specific agent—or at least correlates with detection of that agent. But such inference requires large quantities of labeled data. With consistently coded observation identifiers, the same processing power and same training data quantity can be used for much more clinically important inference, as is discussed elsewhere in this special section. Any information that can be provided in the form of explicit hierarchy, grouping, and machine-processable definitions can increase the power of data mining.
The LOINC code structure and related tools
It is often helpful to break down complex classification into smaller, more manageable parts. Journalists look for the 5 Ws: who, what, where, when, and why. LOINC breaks the identity of a laboratory observation down into 6 parts: component, property, system, scale, timing, and method (Fig. 1).
LOINC names identify the parts that define the code. A real-time PCR for African swine fever virus (ASFV; 73693-4) detects DNA in tissue (Tiss) by nucleic acid amplification and detection with a probe (NAA+probe). Results are reported quantitatively (Qn) as a cycle threshold (Ct). The time (Pt) at which the specimen is collected is the clinically relevant time and is a default value.
The component is the analyte measured with as much precision as possible. In 2020, much effort went into the classification of various tests for Sars-CoV-2 to ensure that the component was correct for tests that detect any betacoronavirus versus those specific for Sars-CoV-2 or any sarbecovirus. 4 When scientists get around to retrospective analysis of COVID-19, these distinctions may be important in filtering out incidental exposure to other sources of RNA.
Property is the hardest part of the LOINC code to keep straight. It represents what about the component is measured. Is it the concentration of an analyte or just the presence or absence? Is presence or absence absolute or based on a cutoff threshold? The property must be coded sufficiently and precisely to ensure that 2 results with the same LOINC can have results safely compared, aggregated, or otherwise analyzed collectively.
The system describes the specimen types that apply to this test. The goal is to separate tests in which differences in specimen type are clinically significant. In many cases, concentrations of various analytes in cerebrospinal fluid are different from the same analyte in serum. When the meaning of the test is essentially the same for 2 different sources or differ only in concentrations, a single LOINC code is often used. In veterinary medicine and public health, there is a growing trend toward providing the specimen type elsewhere and using a system code “XXX” to indicate that the code applies to any specimen type (Fig. 2).
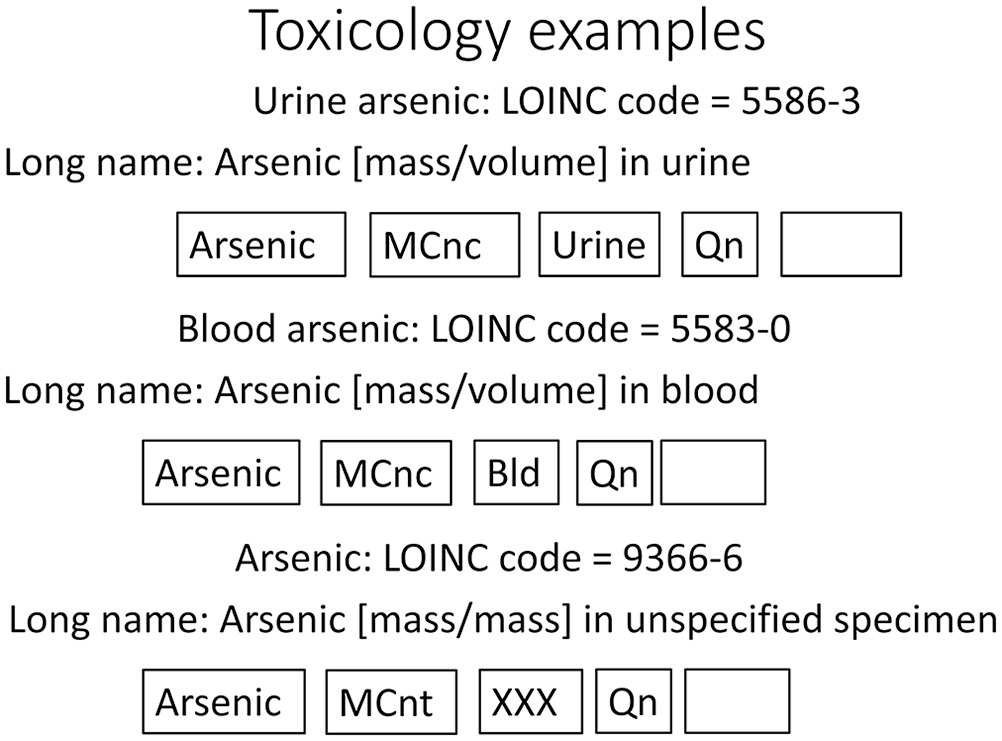

Examples from toxicology illustrate how LOINC differentiates based on clinically relevant differences. Arsenic levels in blood and urine have very different clinical meanings; therefore, the specimen type is part of the LOINC definition. The method used to measure arsenic is seldom of importance to the clinician; therefore, it is left unspecified. On the other hand, any number of solid specimens might have arsenic measured for toxicology; therefore, this LOINC uses XXX, meaning “unspecified specimen, to be included elsewhere in the record.”
Scale is related to property but at a different level. It helps determine how results can be handled. For example, ordinal values, coded classification, and quantitative measures each get their own types of data storage and analysis.
Timing is almost always simply the point in time (abbreviated “Pt”) at which the specimen is collected. This LOINC part becomes important in metabolic studies and other tests that collect data continuously over a fixed time, often 24 h.
Finally, the method is used to distinguish tests by how the property is measured. When the component is something that can be accurately measured against a universal scale, the preference is to omit the method because it is clinically irrelevant. For other tests, such as serology or molecular testing, the method is important (Fig. 3). The level of precision in the method name that has true clinical relevance is sometimes a challenging decision when assigning a LOINC code to a new test.
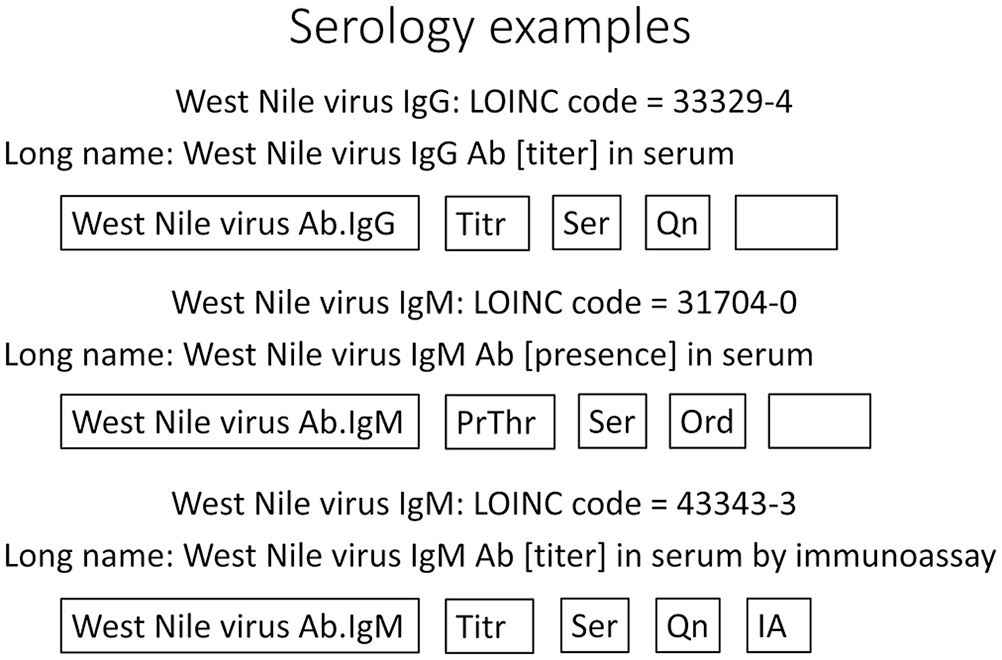

Examples from serology show some clinically important variations that are often overlooked in common test naming. West Nile virus antibody titers are often ordered to evaluate vaccination status but are sometimes confused with IgM antibody presence used to identify a recent infection. In some cases, such as this IgM titer ELISA example, the method is included, but often, such as the first two, it is left to the laboratory to use the best available method. The first and last examples here would have a result such as “1:100”, the second would simply be “Positive” or “Negative.”
Method sometimes creates challenges, especially in serology, where we come to associate the method, which by itself is not clinically important, with specificity and sensitivity of specific test kits, which is important. LOINC tends not to capture these incidental correlations well and often needs suffixes on the method such as “-high sensitivity” to create the distinction. Similarly, commercial kit names are not part of the method but are often associated with performance characteristics and even regulatory requirements. When details of method are important to laboratory and lab network experts but not at the clinical or epidemiologic level, both veterinary medicine and public health laboratories often use a separate data element to carry that detail.
The 6 LOINC parts are combined using standard abbreviations to create short and long LOINC names. These names are somewhat human-readable but are most importantly unambiguous representations of the formal structure. An additional display name has recently been added to make a more appropriate display for user interfaces, reports, or other human-readable displays. The parts and names are available in 20 different language translations.
The good news for users of LOINC is that the 6 parts are becoming increasingly manageable. Test kit vendors are often including LOINC codes in their package inserts. Major medical groups and programs such as the Mayo Clinic, SHIC, and NAHLN, provide LOINC codes for the tests they provide and use.
It is also helpful that more and more laboratory information management system (LIMS) vendors are including provisions for standard code mapping in their software. This eliminates the need to maintain external mapping tables in parallel with the internal test list. Few if any LIMSs use LOINC as the LIMS internal test code system, although the new display names make that a little more reasonable. Mapping to locally chosen test names and local codes is the more common approach. The best systems allow mapping LOINC order codes to panels and tests with multiple results such as multiplex assays and LOINC result codes to specific observation value fields.
LOINC governance
Creating a new code in LOINC—or any other international standard—is a rigorous process. The standard is managed by a professional staff at the Regenstrief Institute in Indianapolis. Submissions are made by LOINC users around the world. Those submitters do need to understand the 6-part structure to create quality code requests. The requests are reviewed by Regenstrief staff who often request supplemental information to ensure proper coding of each part and assignment of appropriate long and short names. The whole process is overseen by 2 volunteer committees, one for Laboratory LOINC and another for Clinical LOINC. To date, veterinary medicine has 2 members on the Laboratory LOINC committee.
The resulting LOINC standard is free for use world-wide. The only significant restriction is that the contents may not be used to create a competing standard. The standard and many supporting documents are available at https://loinc.org.
Conclusion
Local test names and codes work very well in a local context and will always be the way laboratories and their clients will reference specific tests. When there is a need to exchange or aggregate test data across different laboratory and health systems, a common “Esperanto” naming system is needed. LOINC codes provide such a common language and enable a wealth of opportunities to leverage laboratory data more widely.
