Abstract
Newcastle disease is an avian infectious disease caused by avian orthoavulavirus 1, also known as Newcastle disease virus (NDV). This disease has caused significant economic losses to the poultry industry worldwide. The rapid and simple detection of NDV infection is crucial to inform the appropriate control measures. We developed a reverse-transcription recombinase polymerase amplification (RT-RPA) assay combined with a lateral flow assay (LFA) for NDV detection. The RPA assay can be completed at 37°C within 20 min, and the RPA result can be visualized by the LFA within 5 min. The NDV RT-RPA-LFA detected NDV specifically with no cross-reactivity with other pathogens. The detection limit of NDV cDNA with our RT-RPA-LFA was 3.34 × 10−3 ng/μL. Consequently, the RT-RPA-LFA showed good potential for the detection of NDV infection in the field, especially in resource-limited settings.
Newcastle disease, caused by avian orthoavulavirus 1 (Paramyxoviridae, Orthoavulavirus), also known as Newcastle disease virus (NDV), is one of the most serious infectious diseases in the poultry industry. NDV is a nonsegmented, single-stranded, negative-sense RNA virus.3,12 NDV was first discovered in Indonesia and Britain, and spread widely to other regions including Asia, Africa, America, and Europe. 2 NDV has been classified into classes I and II; the virulence of NDV strains ranges from avirulent to highly pathogenic.4,15 Although vaccination and quarantine have been used to control NDV infection, Newcastle disease remains a potential threat to the poultry industry. 5 Many molecular detection methods, such as reverse-transcription PCR (RT-PCR) and real-time RT-PCR,5,17 have been established. However, these methods require high-precision thermocycling for DNA amplification and depend on complex instruments. Thus, rapid and simple methods are needed for the detection of NDV.
Recombinase polymerase amplification (RPA) is a novel technology for isothermal DNA amplification that does not require sophisticated equipment or thermal denaturation. 10 RPA assays have been used successfully for the detection of many pathogens, including foot-and-mouth disease virus, 1 avian influenza A virus (avian IAV), 20 human immunodeficiency virus, 14 and rickettsia. 6 Several enzymes, including recombinase, single-stranded DNA–binding (SSB) protein, and strand-displacing polymerase, are involved in the RPA process. Briefly, recombinases bind to the primers and pair the primers with target homologous DNA. SSB protein binds to the displaced DNA strand and stabilizes the resulting D-loop. Then, DNA amplification is initiated by the polymerase (Suppl. Fig. 1).16,18,19 The RPA assay is performed under isothermal conditions with a short turnaround time and without the requirement for sophisticated equipment. The products of the RPA can be detected by agarose gels 9 or a lateral flow assay (LFA), which is the popular choice given its convenience, rapidity, sensitivity, specificity, and apparatus-independence.8,13,16 The LFA is based on chromatography and immunology technology. During the RPA reaction, the processed probe and the opposing primer generate double-stranded amplification products that co-join 2 antigenic labels (FAM and biotin). In the LFA, the RPA products are complexed with gold-labeled anti-FAM antibodies, then travel in a buffer stream to be trapped at the test line by the interaction between biotin and anti-biotin antibodies, resulting in a red band indicative of a positive result. Non-captured gold particles move through the test line to be fixed at the control line by anti-rabbit antibodies, and produce a red band serving as a negative control (Suppl. Fig. 2). 16
We developed a reverse-transcription RPA (RT-RPA) coupled with a LFA for detection of NDV. NDV (strains F48E9, GM, and Lasota), avian IAV, infectious bursal disease virus (IBDV), infectious bronchitis virus (IBV; Avian coronavirus), Marek disease virus (MDV; Mardivirus), Salmonella (Salmonella enterica ser. Gallinarum biovar Pullorum [Salmonella Pullorum]), and Escherichia coli had been archived in our laboratory. Virulent strain F48E9 was purchased from the China Institute of Veterinary Drug Control (Beijing, China) and propagated in 10-d-old specific pathogen–free (SPF) chicken eggs. The allantoic fluids were harvested and stored at –80°C. Strain F48E9 of NDV was used as the target when developing the NDV RT-RPA-LFA. Common and economically important pathogens in the poultry industry were included for specificity testing purposes. Negative allantoic fluid was obtained from SPF chicken eggs. Blood, cerebrospinal fluid, and lung tissue from SPF chickens were preserved in our laboratory.
Bacterial genomic DNA was extracted (Bacterial DNA kit; Omega Bio-Tek) and viral RNA/DNA was extracted (Viral RNA/DNA extraction kit; Omega Bio-Tek) according to the manufacturer’s instructions. Complementary DNA synthesis was performed (reverse transcriptase M-MLV; TaKaRa Biotechnology). All DNA, cDNA, and RNA were stored at –80°C.
The conserved regions of the NDV F gene were obtained by aligning the sequences from different NDV strains (GenBank FJ480786.1, FJ480785.1, FJ480784.1, FJ480783.1, FJ480782.1, FJ480781.1, FJ480779.1, FJ480778.1, FJ480777.1, FJ480776.1, FJ480775.1, FJ480774.1) using the MegAlign program (DNAstar). RPA primers and probes targeting the conserved regions of the NDV F gene were designed (Primer Premier v.5.0, TwistAmp nfo kit manual; TwistDx; Table 1); only the optimal primer pair and probe are shown. The reverse primer was labeled with a biotin at the 5'-end, and the nfo-probe carried a fluorescein FAM at the 5'-end, an internal tetrahydrofuran residue (THF) at 30 bp from the 5'-end, and a C3 spacer at the 3'-end (Suppl. Fig. 2). 16 All primers and probes were synthesized by Sangon Biotech (Shanghai, China).
Recombinase polymerase amplification (RPA) primers and probe used in our study.
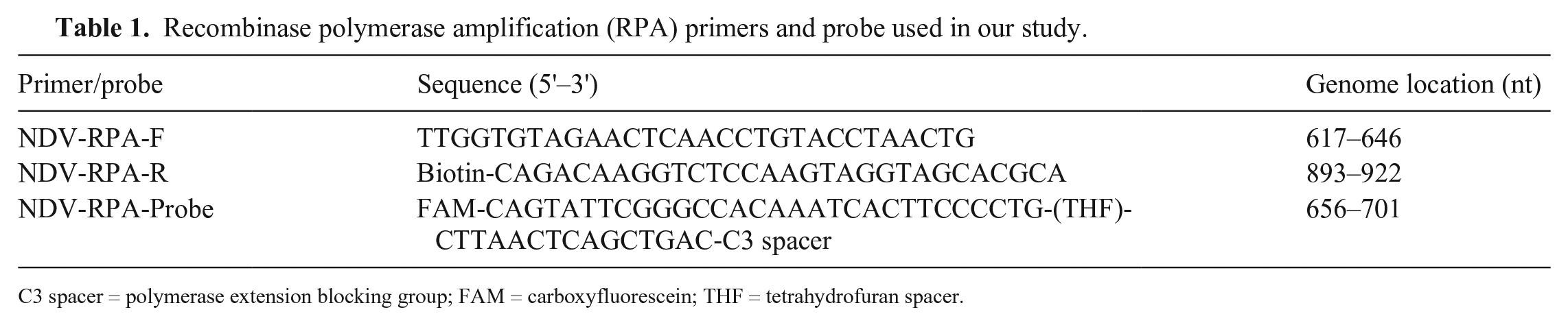

C3 spacer = polymerase extension blocking group; FAM = carboxyfluorescein; THF = tetrahydrofuran spacer.
Complementary DNA of NDV, synthesized as above, was used for our RPA assay. The RPA assay was performed in a 50-μL final reaction volume (TwistAmp RPA nfo kit; TwistDx) according to the manufacturer’s instructions. In brief, the rehydration solution contained 4 μL of cDNA template, 2.1 μL of forward primer (10 μM), 2.1 μL of reverse primer (10 μM), 0.6 μL of the probe (10 μM), 9.2 μL of double-distilled H2O (ddH2O), and 29.5 μL of rehydration buffer (TwistDx). The dried enzyme pellet in a 0.2-mL tube strip provided by the TwistAmp RPA nfo kit was resuspended with 47.5 μL of the rehydration solution, and then the reaction was initiated by adding 2.5 μL of magnesium acetate (280 mM). The tubes were incubated at 37°C for 20 min and then directly determined using a lateral flow dipstick (LFD; Fig. 1A) cassette (Ustar Biotech). First, the reaction tube was placed into the amplicon cartridge. The cartridge was then closed and inserted into the detection chamber; the handle of the detection chamber was closed to lock. The action of locking the detection chamber handle causes a razor blade to cut the tube and the bulb containing the running buffer; the RPA products mix with the running buffer and flow to the fiberglass paper that has been pre-implanted into the detection cassette. The result can be determined 1–2 min after the handle is closed by observing the presence or absence of test (T) and control (C) lines, which indicate the amplification of the target gene and internal control DNA, respectively. A positive result is achieved when both the T line and C line are simultaneously red; a negative result only generates a red C line on the strip (Fig. 1B). If the C line is not observed, the detection result of the sample is deemed invalid.
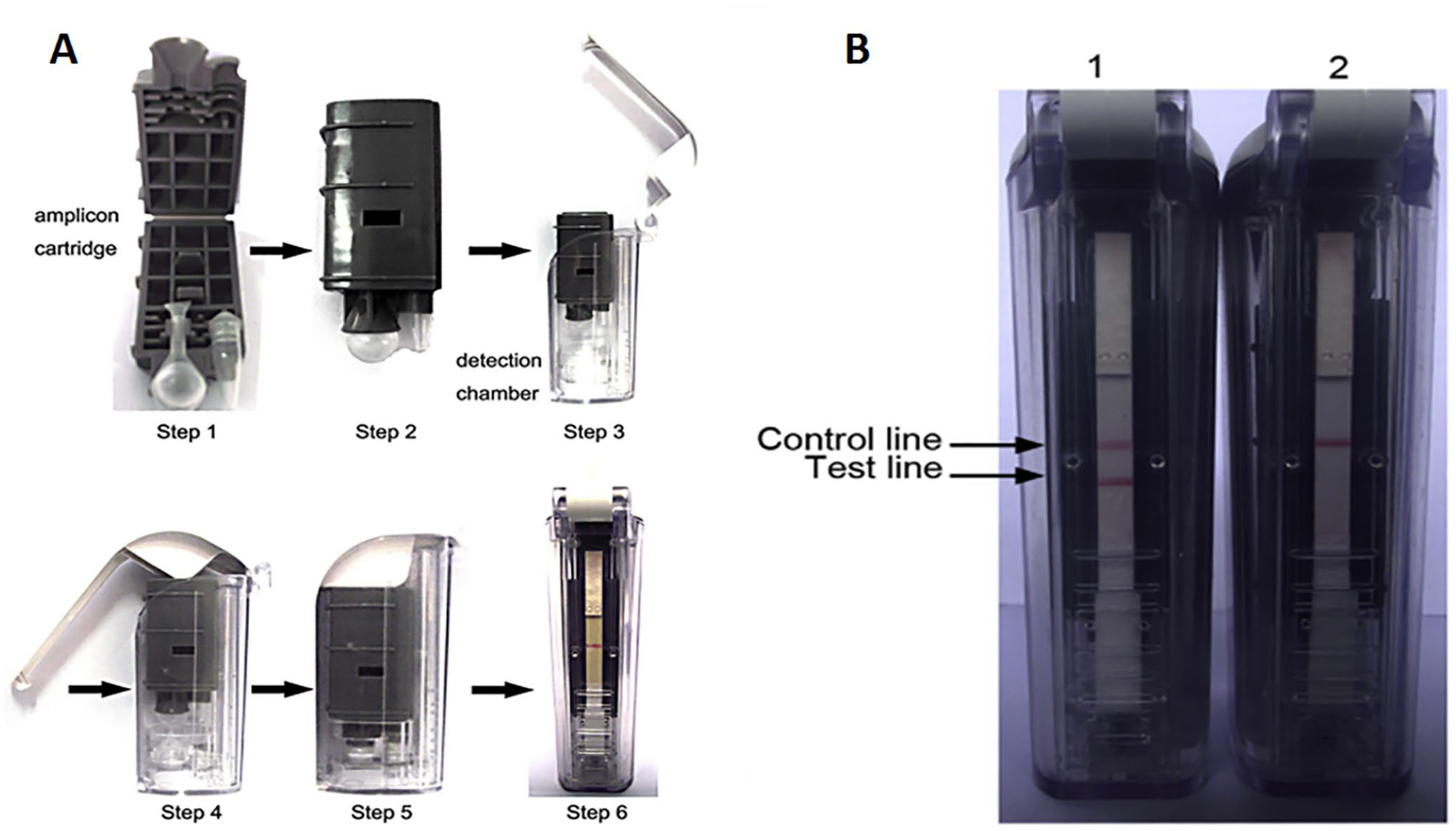

Lateral flow assay (LFA).
When developing the NDV RT-RPA-LFA, we first screened the primers and probes for RPA. After screening different combinations of primers and probe (data not shown), the primer pair of NDV-RPA-F and NDV-RPA-R combined with probe NDV-RPA-Probe were chosen for subsequent evaluation (Table 1), and the RPA amplicon was 267 bp based on theoretical predictions. Then, the RPA reaction, which used NDV-RPA-F and NDV-RPA-R combined with NDV-RPA-Probe, was incubated at 37°C for 20 min, and the result of the RPA assay was determined using the LFD visualization strip cassette. The NDV positive sample produced 2 red bands (T and C line), whereas the negative sample (ddH2O, allantoic fluid, blood, cerebrospinal fluid, and lung tissue) produced only 1 red band (C line; Fig. 2).
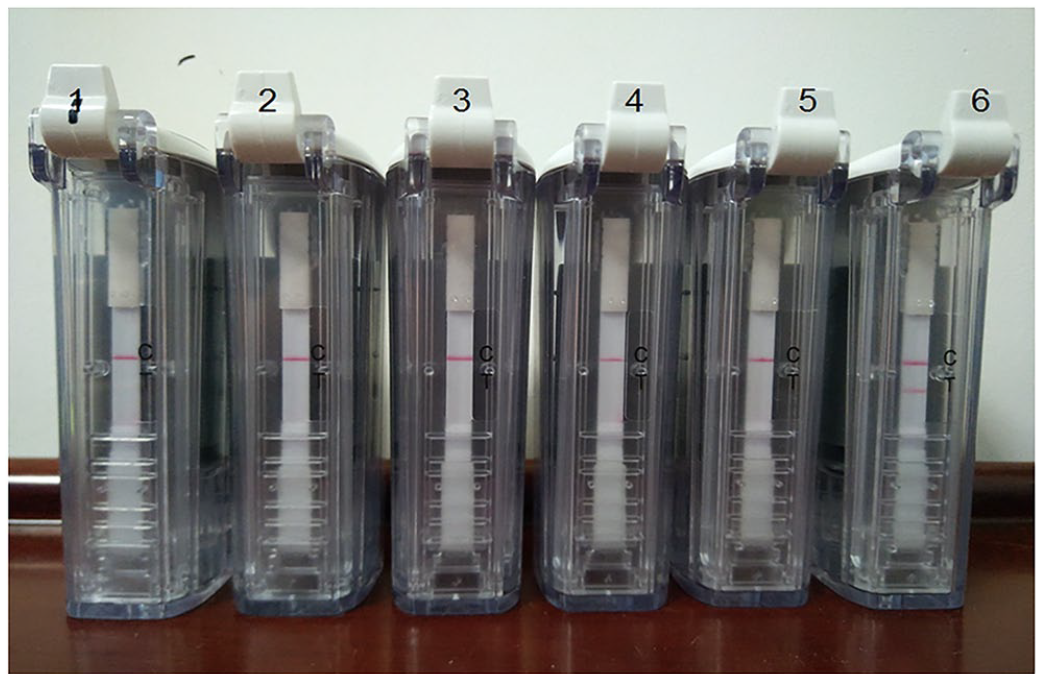

Newcastle disease virus (NDV) reverse-transcription recombinase polymerase amplification assay combined with a lateral flow assay. Lanes: 1 = ddH2O; 2 = negative allantoic fluid; 3 = negative blood; 4 = negative cerebrospinal fluid; 5 = negative lung tissue; 6 = NDV.
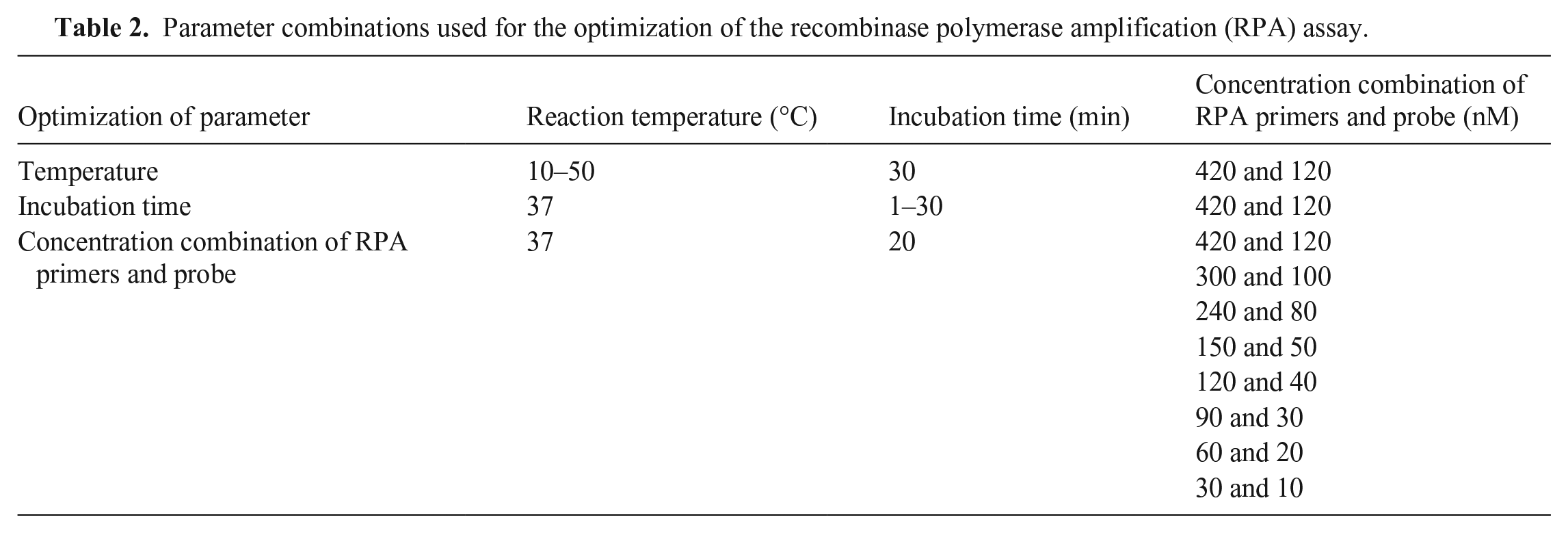
We optimized the RPA assay by testing different reaction temperatures (10, 15, 20, 25, 30, 37, 40, 45, and 50°C), incubation times (1, 5, 10, 15, 20, 25, and 30 min), primer concentrations (420, 300, 240, 150, 120, 90, 60, and 30 nM), and probe concentrations (120, 100, 80, 50, 40, 30, 20, and 10 nM; Table 2). Known positive samples (NDV strain F48E9) and negative controls (ddH2O) were used during the optimization testing. The incubation time was set to 30 min during temperature optimization. When the reaction temperature was ≤25°C or ≥50°C, only the C line was produced or the T line was weak. Both the T line and C line were produced between 30 and 45°C with no observable difference; 37°C was selected for subsequent tests (Suppl. Fig. 3A). The optimal incubation time was determined after temperature optimization. A slight band (T line) appeared when incubating for 5–15 min. When the incubation time was ≥20 min, the T line appeared clearly with no observable difference. Thus, 20 min was selected as the incubation time in subsequent tests (Suppl. Fig. 3B). Then, we optimized the concentrations of primers and probe. Only when the primer concentration was ≥90 nM and probe concentration was ≥30 nM, did the T line appear clearly in the LFD (Suppl. Fig. 3C). The concentrations of primers (90–420 nM) and probe (30–120 nM) were suitable for the RPA assay. According to the optimization results of the RPA assay, the following parameters were used: primer concentration of 420 nM, probe concentration of 120 nM, reaction temperature of 37°C, and incubation time of 20 min.
Parameter combinations used for the optimization of the recombinase polymerase amplification (RPA) assay.
To evaluate the specificity of the developed NDV RT-RPA-LFA, we tested various pathogens, including NDV, avian IAV, IBV, IBDV, MDV, Salmonella Pullorum, and E. coli. The T line was produced only by the NDV-positive sample (Suppl. Fig. 4); cross-reaction was not observed. Hence, the NDV RT-RPA-LFA could be used to detect NDV specifically.
To evaluate the sensitivity of our NDV RT-RPA-LFA, a dilution range of 3.34 × 102 ng/μL to 3.34 × 10–5 ng/μL of NDV cDNA was tested. The detection limit of NDV cDNA with our RT-RPA-LFA was 3.34 × 10–3 ng/μL (Suppl. Fig. 5).
Our NDV RT-RPA-LFA has several advantages compared with other molecular detection methods such as conventional RT-PCR. 5 First, the commercial reagents for the RPA assay are lyophilized particles, which maintains enzyme activity. 11 Second, a simple constant-temperature heating device is sufficient for the RPA assay, whereas PCR requires expensive instruments. Third, the RPA assay takes only 20–30 min to complete DNA amplification, whereas many PCR reactions take >1 h. Finally, the result of the RPA assay can be visualized rapidly by a LFD, whereas the detection of PCR products requires an electrophoresis device. Considering that our RT-RPA-LFA is simple, rapid, and specific for NDV detection and does not need sophisticated equipment, it has the potential to be a valuable tool in the clinical diagnosis of Newcastle disease, especially in resource-limited settings. Additionally, a previous study 7 described a RPA-NALF (nucleic acid lateral flow) immunoassay for the detection of NDV, which could rapidly and efficiently detect NDV infection with a limit of detection of 10 genomic copies/RPA reaction. The specificity of their RPA-NALF assay was evaluated through verifying the primers for their cross-reactivity with other avian respiratory viruses using a BLAST search in NCBI, whereas we carried out the specificity test through testing various common and economically important pathogens in the poultry industry. Compared with previous work, 7 we also used a different kit for the RPA assay and a different device for the detection of RPA products, and then provided an alternative to detect infection by NDV.
Supplemental Material
sj-pdf-1-vdi-10.1177_1040638721990122 – Supplemental material for Development of a reverse-transcription recombinase polymerase amplification assay with a lateral flow assay for rapid detection of avian orthoavulavirus 1
Supplemental material, sj-pdf-1-vdi-10.1177_1040638721990122 for Development of a reverse-transcription recombinase polymerase amplification assay with a lateral flow assay for rapid detection of avian orthoavulavirus 1 by Jindai Fan, Wenxian Chen, Yuanyuan Zhang, Zhixiang Liu, Xiaoming Li, Hongxing Ding, Lin Yi, Jinding Chen and Mingqiu Zhao in Journal of Veterinary Diagnostic Investigation
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by the National Key R&D Projects (2016YFD0500801, 2017YFD0501104) and the Science and Technology Program of Guangzhou, China (201803020005).
Supplementary material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
