Abstract
Porcine circovirus–associated disease, caused primarily by Porcine circovirus 2 (PCV-2), has become endemic in many pig-producing countries and has resulted in significant economic losses to the swine industry worldwide. Tests for PCV-2 infection include PCR, nested PCR, competitive PCR, and real-time PCR (rtPCR). Recombinase polymerase amplification (RPA) has emerged as an isothermal gene amplification technology for the molecular detection of infectious disease agents. RPA is performed at a constant temperature and therefore can be carried out in a water bath. In addition, RPA is completed in ~30 min, much faster than PCR, which usually takes >60 min. We developed a RPA-based method for the detection of PCV-2. The detection limit of RPA was 102 copies of PCV-2 genomic DNA. RPA showed the same sensitivity as rtPCR but was 10 times more sensitive than conventional PCR. Successful amplification of PCV-2 DNA, but not other viral templates, demonstrated high specificity of the RPA assay. This method was also validated using clinical samples. The results showed that the RPA assay had a diagnostic agreement rate of 93.7% with conventional PCR and 100% with rtPCR. These findings suggest that the RPA assay is a simple, rapid, and cost-effective method for PCV-2 detection, which could be potentially applied in clinical diagnosis and field surveillance of PCV-2 infection.
Porcine circovirus 2 (PCV-2; family Circoviridae, genus Circovirus), a nonenveloped virus with an ambisense genome composed of single-stranded circular DNA, is the primary etiologic agent of porcine circovirus–associated diseases, which include postweaning multisystemic wasting syndrome (PMWS), porcine dermatitis and nephropathy syndrome, myocarditis, abortion, and porcine respiratory disease complex. 15 Since PMWS was first reported in western Canada in 1991, 14 the disease has become an endemic syndrome in many pig-producing countries and has caused great economic losses in the swine industry worldwide.
Given that PCV-2 is widely distributed and associated with serious economic damage, it is imperative that rapid and sensitive methods be developed for the detection of PCV-2. Among the methods developed, a substantial number of assays have been developed based on gene amplification analysis, including PCR, 11 real-time PCR (rtPCR), 4 competitive PCR (cPCR), 13 microarray analyses, 9 and loop-mediated isothermal amplification (LAMP). 6
Recombinase polymerase amplification (RPA) is an isothermal gene amplification technology that has been explored for molecular detection. 16 Unlike PCR and many other isothermal technologies, RPA relies on 3 core enzymes. The titular enzyme is a recombinase, which binds to primers to form recombination filaments that can recombine with homologous DNA. The second enzyme is a single-stranded DNA-binding protein, which binds to the strand of DNA displaced by the primer and stabilizes the D-loop that has formed to prevent dissociation of the primer. The third core enzyme is a strand-displacing DNA polymerase that synthesizes DNA by adding bases onto the 3’-end of the primer to force open the DNA double helix as DNA synthesis progresses. 16 Part of the advantage of RPA for molecular detection is that it can tolerate some diversity in the template sequence. 1 RPA can occur with up to 9 changes across the primer and probe binding sites. 3 RPA has been developed for the molecular detection of bacteria, fungi, parasites, and viruses, such as Mycobacterium tuberculosis, 3 Blastomyces dermatitidis, 17 Giardia duodenalis, 7 Plasmodium falciparum, 10 Foot-and-mouth disease virus, 1 and Human immunodeficiency virus 1. 2 However, the application of RPA to detect PCV-2 has not been examined, to our knowledge. Therefore, we conducted this study to explore the applicability of RPA for the detection of PCV-2.
Various porcine respiratory viruses were used in this study, including Classical swine fever virus (CSFV, C strain) a and Foot-and-mouth disease virus (FMDV, type O). b Porcine circovirus 1 (PCV-1), Porcine circovirus 2 (PCV-2), Porcine reproductive and respiratory syndrome virus (PRRSV), porcine parvovirus (PPV), and pseudorabies virus (PRV) were maintained in our laboratory. Forty-eight snap-frozen clinical samples (lymph node, liver, lung, serum, and muscle) were collected from different pig farms in Hebei Province, China and transported to our laboratory for PCV-2 detection. For lymph node, liver, lung, and muscle samples, 10 mg of each sample was homogenized in 1 mL of medium. c After centrifugation at 3,000 × g for 10 min, the supernatant was collected for viral DNA extraction. For serum samples, 1 mL of serum was used directly for viral DNA extraction. The viral DNA of PCV-1, PCV-2, PPV, and PRV were extracted using a commercial kit, d according to the manufacturer’s instructions. The viral RNA of CSFV, PRRSV, and FMDV were extracted using TRIzol, e according to the manufacturer’s instructions. DNA and RNA were quantified using a spectrophotometer. f Viral RNA (100 ng) reverse transcription and purification of synthesized complementary (c)DNA were carried out using commercial kitsg,h following the manufacturer’s instructions. Purified cDNA was quantified using a spectrophotometer. f Viral DNA was extracted from clinical samples using a commercial kit, d and the final viral DNA from each sample was eluted in 20 µL of nuclease-free water. All DNA and cDNA templates were stored at −20°C until assays were performed.
According to the reference sequences of PCV-2 (GenBank accession EF197986), the following primers i specific to the open reading frame 2 of PCV-2 were designed for RPA: forward (5′-ACTCCTCCCGCCATACCATAACCCAGCCCTTC-3′) and reverse (5′-ACATGGTTACACGGATATTGTATTCCTGGTCG-3′). The predicted product size of RPA was ~220 bp. RPA reactions were performed in a 50-µL volume using a commercial kit. j Other components included 29.5 µL of rehydration buffer, 480 nM each RPA primer, 14 mM magnesium acetate, and 1 µL of template. All reagents except the viral DNA and magnesium acetate were prepared in a master mix, which was distributed into each 0.2-mL freeze-dried reaction tube containing a dried enzyme pellet followed by the addition of 1 µL of viral template. Subsequently, magnesium acetate was pipetted into the tube lids, and the lids closed carefully. Magnesium acetate was centrifuged into the rehydrated material using a minispin centrifuge. The tube was vortexed briefly, spun down, and then immediately placed into a water bath at 38°C for 10, 20, 30, and 40 min. The RPA amplified products were purified using a commercial kit, k according to the manufacturer’s instructions. The purified products (5 µL) were electrophoresed on 2.0% (w/v) agarose gel and visualized under ultraviolet light after staining. l Using 105 copies of PCV-2 genomic DNA as template, a weak DNA band with an expected size (~220 bp) was visualized after a 10-min reaction at 38°C (Fig. 1). Semi-quantification by measuring the DNA band density revealed that the DNA yield after a 20-min reaction was ~4 times that generated by a 10-min reaction, whereas no significant difference was observed among products at 20, 30, and 40 min (data not shown). Therefore, 20 min was set for all RPA assays performed in this study. Using 10 ng of viral DNA or cDNA as template for RPA, we showed that only PCV-2 DNA was amplified, whereas PCV-1, PPV, PRV, PRRSV, CSFV, and FMDV templates were not (Fig. 2). Five independent RPA reactions were repeated, and similar results were observed, demonstrating that the RPA assay is specific for the detection of PCV-2. We diluted PCV-2 genomic DNA in 10-fold serial dilutions to achieve DNA concentrations ranging from 106 to 100 copies/µL. One µL of each DNA dilution was used for RPA amplification. The results demonstrated that the detection limit of the RPA assay was 102 copies/reaction (Fig. 3A). The sensitivity was compared among RPA, conventional PCR, and rtPCR. Conventional PCR was conducted following the standard protocol “Detecting porcine circovirus with polymerase chain reaction” published by the Standardization Administration of China (http://shop.standards.ie/nsai/details.aspx?ProductID=1083276). The primer sequences were as follows: forward (5′-CCGCGGGCTGGCTGAACTT-3′) and reverse (5′-ACCCCCGCCACCGCTACC-3′). The predicted PCR product size was ~1,200 bp. Conventional PCR was performed as follows: stage 1, 94°C for 10 min; stage 2, 94°C for 30 s, 62°C for 45 s, and 72°C for 60 s. Stage 2 was repeated for 40 cycles followed by stage 3 of 72°C for 10 min. For rtPCR, the universal master mix was replaced by a commercial rtPCR mix. m The primer sequences for rtPCR were as follows: forward (5′-GGGAGCAGGGCCAGAATT-3′) and reverse (5′-CGCTCTGTGCCCTTTGAATACT-3′). The probe used for rtPCR was 5′-FAM-ACCTTAACCTTTCTTATTCTGMGB-3′. Real-time PCR was performed n as follows: 95°C for 10 min, then 40 cycles of 95°C for 10 s and 60°C for 35 s. The detection limit of conventional PCR was 103 copies/reaction (Fig. 3B); the detection limit of rtPCR was the same as RPA (102 copies/reaction; Fig. 3C). Similar results were obtained from all 5 independent experiments.
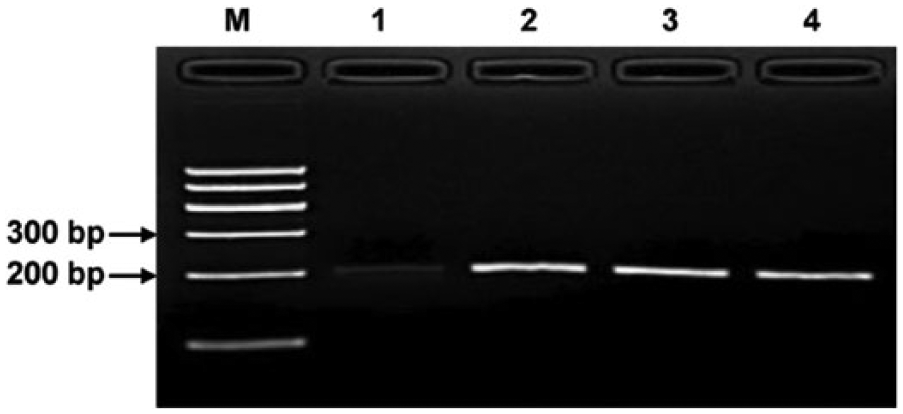
Optimization of recombinase polymerase amplification reaction time for Porcine circovirus-2 (PCV-2) detection. A total of 105 copies of PCV-2 genomic DNA were used as template, and amplified DNA products at different time points were analyzed using agarose gel electrophoresis and staining. l A clear DNA band with an expected size (~220 bp) could be visualized after a 10-min reaction at 38°C. Lane M: DNA marker; lanes 1–4: DNA products from reactions incubated for 10, 20, 30, and 40 min, respectively.
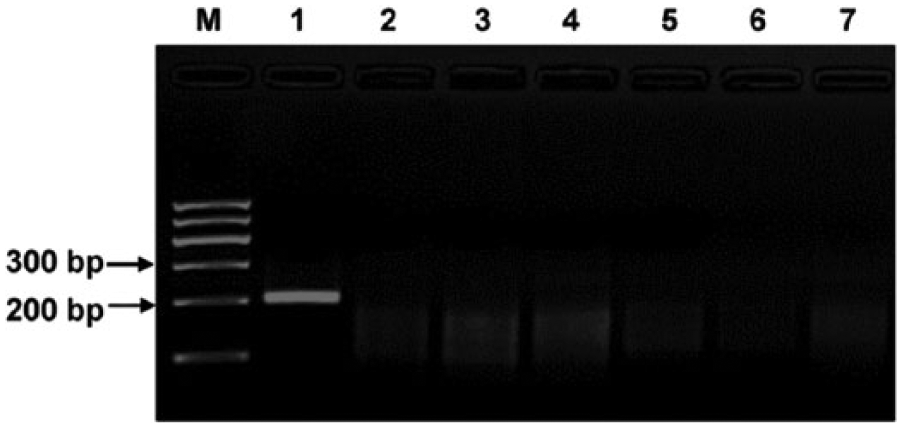
Specificity of recombinase polymerase amplification (RPA) for Porcine circovirus-2 (PCV-2) detection. RPA was carried out at 38°C for 20 min using 10 ng of viral DNA or complementary DNA as template. RPA amplified only PCV-2 DNA; there was no cross-reactivity with other viruses tested. Lane M: DNA marker; lane 1: PCV-2; lane 2: Porcine circovirus-1; lane 3: porcine parvovirus; lane 4: pseudorabies virus; lane 5: Porcine reproductive and respiratory syndrome virus; lane 6: Classical swine fever virus; lane 7: Foot-and-mouth disease virus.
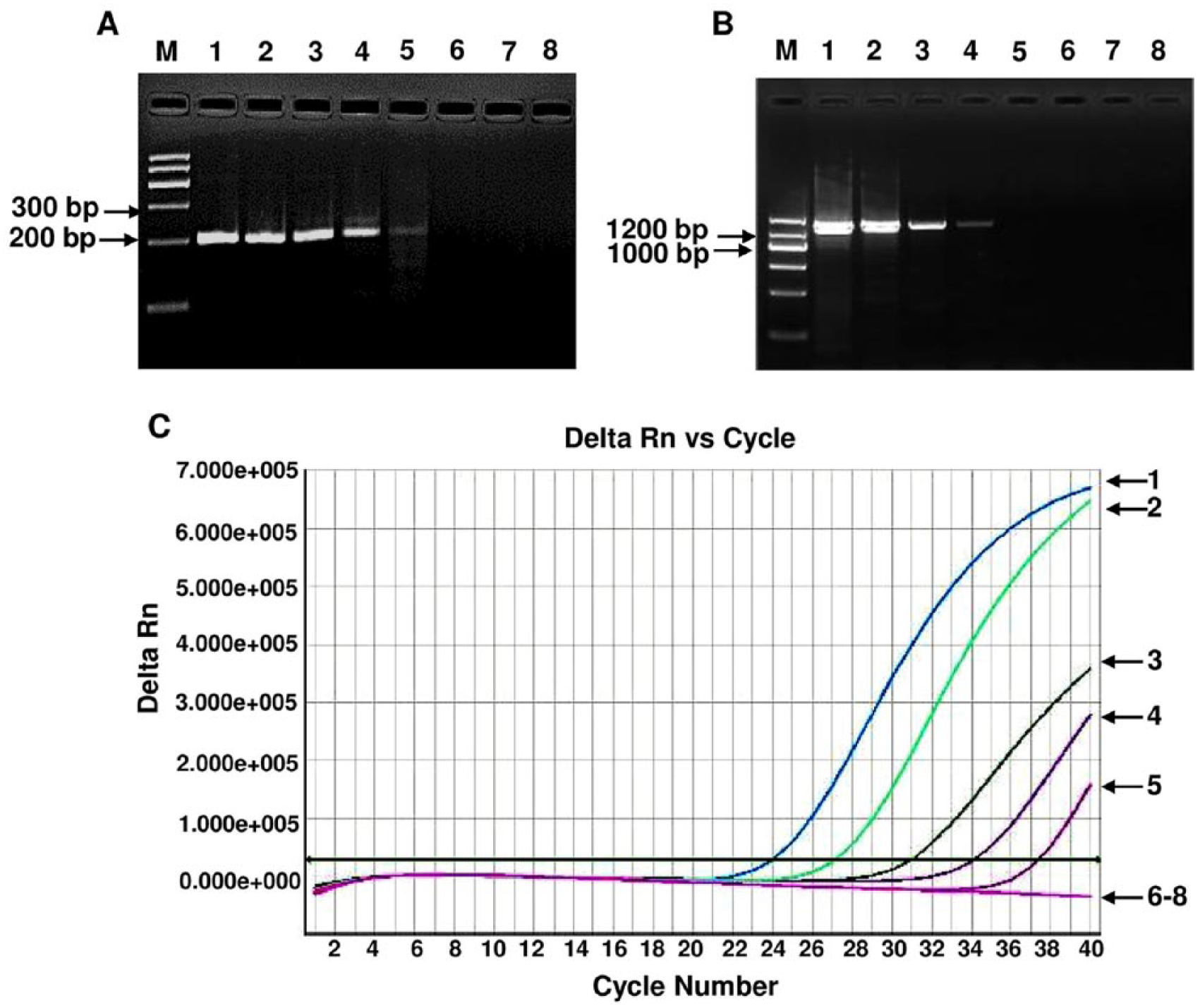
Sensitivity of recombinase polymerase amplification (RPA) for Porcine circovirus-2 (PCV-2) detection. Different copy numbers of PCV-2 DNA were used as template for RPA, conventional PCR, and real-time PCR. The lowest copy number that could be determined by RPA was 102 copies, the same as observed with real-time PCR (
RPA was further validated using clinical samples. Of the total 48 samples, 43 samples tested positive and 5 samples tested negative by both RPA and rtPCR. In contrast, 40 samples tested positive and 8 were negative by conventional PCR. These data showed that RPA had a 100% (48/48) diagnostic agreement rate with rtPCR and a 93.7% (45/48) diagnostic agreement rate with conventional PCR.
Various methods have been developed for the detection of PCV-2, such as rtPCR, 4 cPCR, 13 LAMP, 6 etc. PCR requires an expensive thermocycler and is time-consuming. LAMP is performed at a constant temperature, as is RPA, but the LAMP reaction takes longer (~1 h) to complete than the RPA reaction (10–20 min). In addition, LAMP uses 3 pairs of primers, and the mixing of multiple pairs of primers increases the chance of forming primer–primer interactions, limiting the total amount of primers available for the reaction and reducing amplification efficiency. 5 Additionally, the LAMP products are ladder-like DNA bands. When self-priming of the LAMP primers occurs, it can also produce ladder-like DNA bands, which can create false-positive results. 8 The RPA assay we developed in this study showed that the product, generated by RPA in as little as 10 min, was detectable by agarose gel electrophoresis. The amount of final product in a 20-min reaction was ~4 times that in a 10-min reaction, whereas no significant difference was observed among products generated in 20, 30, and 40 min reactions (data not shown). This is probably because of the complete consumption of dNTPs or other components provided in the commercial kit j after the 20-min reaction. RPA does not require the reaction temperature to be precisely controlled. Previous work has shown successful amplification of methicillin-resistant Staphylococcus aureus by RPA at a temperature range of 25–44°C. 16 Proviral HIV-1 DNA was amplified at 31–43°C. 12 We observed that similar levels of final products were generated at any temperature between 34–42°C (data not shown). RPA, with high sensitivity and high specificity, is a simple, rapid, and cost-effective method for PCV-2 detection, which could be potentially applied in clinical diagnosis, on-the-port quarantine, field surveillance, and molecular epidemiologic investigation of PCV-2 infection.
Footnotes
Authors’ contributions
Jianchang Wang contributed to design of the study; contributed to acquisition, analysis, and interpretation of data; and drafted the manuscript. Jinfeng Wang and L Liu contributed to acquisition and analysis of data. R Li contributed to acquisition and interpretation of data. W Yuan contributed to conception and design of the study; contributed to analysis and interpretation of data; drafted the manuscript; critically revised the manuscript; gave final approval; and agreed to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
Ringpu Bio-Pharmacy Co. Ltd., Tianjin, China.
b.
Obtained from ELISA kit kindly donated by Lanzhou Veterinary Research Institute, Lanzhou, China.
c.
Minimal essential medium, Invitrogen, Beijing, China.
d.
TIANamp virus DNA kit, Tiangen Biotech Co. Ltd., Beijing, China.
e.
Trizol reagent, Tiangen Biotech Co. Ltd., Beijing, China.
f.
ND-1000 spectrophotometer, NanoDrop, Wilmington, DE.
g.
Primescript II 1st strand cDNA Synthesis kit, Takara Co. Ltd., Dalian, China.
h.
cDNA purification kit, Takara Co. Ltd., Dalian, China.
i.
Sangon Biotech Co., Ltd, Shanghai, China.
j.
TwistAmp basic kit, TwistDX, Cambridge, United Kingdom.
k.
Universal DNA purification kit, Tiangen Biotech Co. Ltd., Beijing, China.
l.
GoldView stain, SBS Genetech Co. Ltd., Beijing, China.
m.
Premix Ex Taq, Takara Co. Ltd., Dalian, China.
n.
7500 Fast real-time PCR system, Applied Biosystems, Foster City, CA.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Science and Technology Project Foundation of Animal Husbandry Bureau, Hebei Province, China (grant number 2014-3-03).
